Molecular Chaperones: Confirmation for conformational selection
Almost every process in biology relies on proteins in one way or another, and most of these proteins need to have a specific three-dimensional structure to carry out their roles. A network of molecular machines – called chaperones – makes sure that proteins tend to end up folded correctly (Balchin et al., 2016), and a chaperone called Hsp70 is a central hub in this network.
Hsp70 has a domain that binds to substrates at one end (Mayer and Bukau, 2005), and a domain that binds to nucleotides like ATP at the other end. The chaperone’s affinity for substrates decreases when ATP binds to this domain, and it increases again when the ATP is broken down by hydrolysis to yield ADP. Importantly, Hsp70 does not work alone; another protein called Hsp40 both helps to deliver substrates to Hsp70 and also catalyzes the hydrolysis of ATP.
However, despite much progress in recent years (Clerico et al., 2015; Mayer and Kityk, 2015), major questions remain about the interactions between Hsp70 and its substrates. For instance, does Hsp70 passively bind to exposed segments of unfolded proteins, as proposed in the ‘conformational selection’ hypothesis, or does it actively unfold misfolded substrates, as proposed by the ‘induced fit’ hypothesis? Now, in eLife, Ashok Sekhar, Lewis Kay and colleagues report an answer to this long-standing question (Sekhar et al., 2018).
It has proven challenging to decide between these two hypotheses, partly because chaperones are highly dynamic molecular machines, which limits the number of techniques that can be used to study them in action. Fortunately, NMR spectroscopy provided a perfect solution to tackle this problem (Kay, 2016; Huang and Kalodimos, 2017).
According to the conformational selection hypothesis, the substrates switch between being folded and unfolded, and Hsp70 selectively binds to the unfolded state (Figure 1A). The induced fit hypothesis, however, proposes that Hsp70 binds to a folded or misfolded substrate and then unfolds it (Figure 1B). Therefore, if someone could directly detect a substrate switching between an unfolded state and an Hsp70 bound state, that would be evidence for the conformational selection hypothesis. No such exchange should occur if the induced fit model is correct.
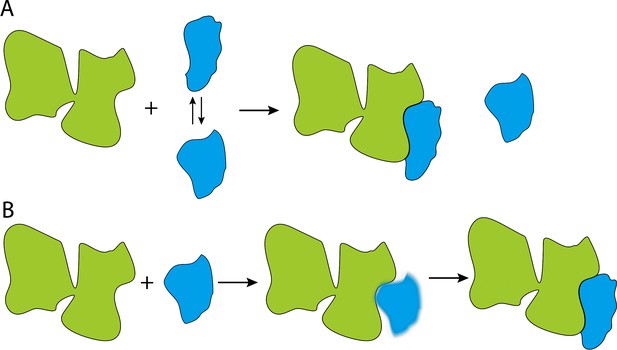
Cartoon illustrations of the conformational selection and induced fit mechanisms.
(A) In the conformational selection mechanism, the substrate (blue) switches between different conformations, and the chaperone (green) selectively interacts with one of these conformations. (B). In the induced fit mechanism, the chaperone directly interacts with the substrate, irrespective of the latter's conformation, and then changes its conformation.
Using advanced NMR methodologies, Sekhar et al. – who are based at the University of Toronto, the Weizmann Institute of Science, the École Normale Supérieure in Paris and the Hospital for Sick Children, also in Toronto – could keep track of two processes – magnetization exchange and chemical exchange – as Hsp70 interacted with a model substrate. Roughly speaking, these two processes can be used to measure conformational changes undergone by the substrate during its interaction with Hsp70.
Sekhar et al. found that Hsp70 from humans and a related bacterial chaperone called DnaK both interact with substrates through the conformational selection mechanism. It was already known that DnaK behaves in a similar way to human Hsp70 (Clerico et al., 2015; Mayer and Kityk, 2015). The results of Sekhar et al. suggest, therefore, that the Hsp70 chaperone machinery prevents the misfolding of proteins by selectively binding to unfolded substrates instead of actively unfolding substrates that have misfolded. The methods reported by Sekhar et al. could also be used to study other chaperone systems such as the Trigger Factor (Saio et al., 2014) and SecB (Huang et al., 2016), both of which capture their substrate proteins in an unfolded state.
In vivo, the Hsp70 machinery needs to process long proteins consisting of more than 300 to 400 amino acids (Balchin et al., 2016). Yet Hsp70 has just one relatively small substrate-binding site that can accommodate only seven to eight amino acids. How can Hsp70 fulfill such a challenging task? It is thought that Hsp40 helps deliver substrates to Hsp70, and so it will be important to examine the exact role that Hsp40 plays in recognizing substrates and delivering them to Hsp70. Will the current model, which works for isolated Hsp70, still apply when Hsp40 and the rest of the Hsp70 machinery are present? Many key questions remain unanswered, yet this latest study gives hope that NMR spectroscopy is well suited to address these questions.
References
-
How Hsp70 molecular machines interact with their substrates to mediate diverse physiological functionsJournal of Molecular Biology 427:1575–1588.https://doi.org/10.1016/j.jmb.2015.02.004
-
Structures of large protein complexes determined by nuclear magnetic resonance spectroscopyAnnual Review of Biophysics 46:317–336.https://doi.org/10.1146/annurev-biophys-070816-033701
-
New views of functionally dynamic proteins by solution NMR spectroscopyJournal of Molecular Biology 428:323–331.https://doi.org/10.1016/j.jmb.2015.11.028
-
Hsp70 chaperones: cellular functions and molecular mechanismCellular and Molecular Life Sciences 62:670–684.https://doi.org/10.1007/s00018-004-4464-6
-
Insights into the molecular mechanism of allostery in Hsp70sFrontiers in Molecular Biosciences 2:58.https://doi.org/10.3389/fmolb.2015.00058
Article and author information
Author details
Publication history
- Version of Record published: February 20, 2018 (version 1)
Copyright
© 2018, Jiang et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 1,578
- views
-
- 190
- downloads
-
- 5
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Structural Biology and Molecular Biophysics
Regulated hydrolysis of the phosphoinositide phosphatidylinositol(4,5)-bis-phosphate to diacylglycerol and inositol-1,4,5-P3 defines a major eukaryotic pathway for translation of extracellular cues to intracellular signaling circuits. Members of the lipid-activated protein kinase C isoenzyme family (PKCs) play central roles in this signaling circuit. One of the regulatory mechanisms employed to downregulate stimulated PKC activity is via a proteasome-dependent degradation pathway that is potentiated by peptidyl-prolyl isomerase Pin1. Here, we show that contrary to prevailing models, Pin1 does not regulate conventional PKC isoforms α and βII via a canonical cis-trans isomerization of the peptidyl-prolyl bond. Rather, Pin1 acts as a PKC binding partner that controls PKC activity via sequestration of the C-terminal tail of the kinase. The high-resolution structure of full-length Pin1 complexed to the C-terminal tail of PKCβII reveals that a novel bivalent interaction mode underlies the non-catalytic mode of Pin1 action. Specifically, Pin1 adopts a conformation in which it uses the WW and PPIase domains to engage two conserved phosphorylated PKC motifs, the turn motif and hydrophobic motif, respectively. Hydrophobic motif is a non-canonical Pin1-interacting element. The structural information combined with the results of extensive binding studies and experiments in cultured cells suggest that non-catalytic mechanisms represent unappreciated modes of Pin1-mediated regulation of AGC kinases and other key enzymes/substrates.
-
- Structural Biology and Molecular Biophysics
Roco proteins entered the limelight after mutations in human LRRK2 were identified as a major cause of familial Parkinson’s disease. LRRK2 is a large and complex protein combining a GTPase and protein kinase activity, and disease mutations increase the kinase activity, while presumably decreasing the GTPase activity. Although a cross-communication between both catalytic activities has been suggested, the underlying mechanisms and the regulatory role of the GTPase domain remain unknown. Several structures of LRRK2 have been reported, but structures of Roco proteins in their activated GTP-bound state are lacking. Here, we use single-particle cryo-electron microscopy to solve the structure of a bacterial Roco protein (CtRoco) in its GTP-bound state, aided by two conformation-specific nanobodies: NbRoco1 and NbRoco2. This structure presents CtRoco in an active monomeric state, featuring a very large GTP-induced conformational change using the LRR-Roc linker as a hinge. Furthermore, this structure shows how NbRoco1 and NbRoco2 collaborate to activate CtRoco in an allosteric way. Altogether, our data provide important new insights into the activation mechanism of Roco proteins, with relevance to LRRK2 regulation, and suggest new routes for the allosteric modulation of their GTPase activity.






