Articles
Article Tools
Supplementary
Stats or Metrics
Article
Short Communication
Exp Neurobiol 2022; 31(5): 277-288
Published online October 31, 2022
https://doi.org/10.5607/en22038
© The Korean Society for Brain and Neural Sciences
Glutamate Permeability of Chicken Best1
Jung Moo Lee1,2†, Changdev Gorakshnath Gadhe3†, Hyunji Kang2,4†, Ae Nim Pae3,5* and C. Justin Lee1,2,4*
1KU-KIST Graduate School of Converging Science and Technology, Korea University, Seoul 02841, 2Center for Cognition and Sociality, Institute for Basic Science, Daejeon 34126, 3Brain Science Institute, Korea Institute of Science and Technology, Seoul 02792, 4IBS School, University of Science and Technology, Daejeon 34113, 5KIST School, University of Science and Technology, Seoul 02792, Korea
Correspondence to: *To whom correspondence should be addressed.
Ae Nim Pae, TEL: 82-2-958-5185, FAX: 82-2-958-6999
e-mail: anpae@kist.re.kr
C. Justin Lee, TEL: 82-42-878-9150, FAX: 82-42-878-9151
e-mail: cjl@ibs.re.kr
†These authors contributed equally to this article.
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
Bestrophin-1 (Best1) is a calcium (Ca2+)-activated chloride (Cl-) channel which has a phylogenetically conserved channel structure with an aperture and neck in the ion-conducting pathway. Mammalian mouse Best1 (mBest1) has been known to have a permeability for large organic anions including gluconate, glutamate, and D-serine, in addition to several small monovalent anions, such as Cl‑, bromine (Br-), iodine (I-), and thiocyanate (SCN-). However, it is still unclear whether non-mammalian Best1 has a glutamate permeability through the ion-conducting pathway. Here, we report that chicken Best1 (cBest1) is permeable to glutamate in a Ca2+-dependent manner. The molecular docking and molecular dynamics simulation showed a glutamate binding at the aperture and neck of cBest1 and a glutamate permeation through the ion-conducting pore, respectively. Moreover, through electrophysiological recordings, we calculated the permeability ratio of glutamate to Cl- (PGlutamate/PCl) as 0.28 based on the reversal potential shift by ion substitution from Cl- to glutamate in the internal solution. Finally, we directly detected the Ca2+-dependent glutamate release through cBest1 using the ultrasensitive two-cell sniffer patch technique. Our results propose that Best1 homologs from non-mammalian (cBest1) to mammalian (mBest1) have a conserved permeability for glutamate.
Graphical Abstract
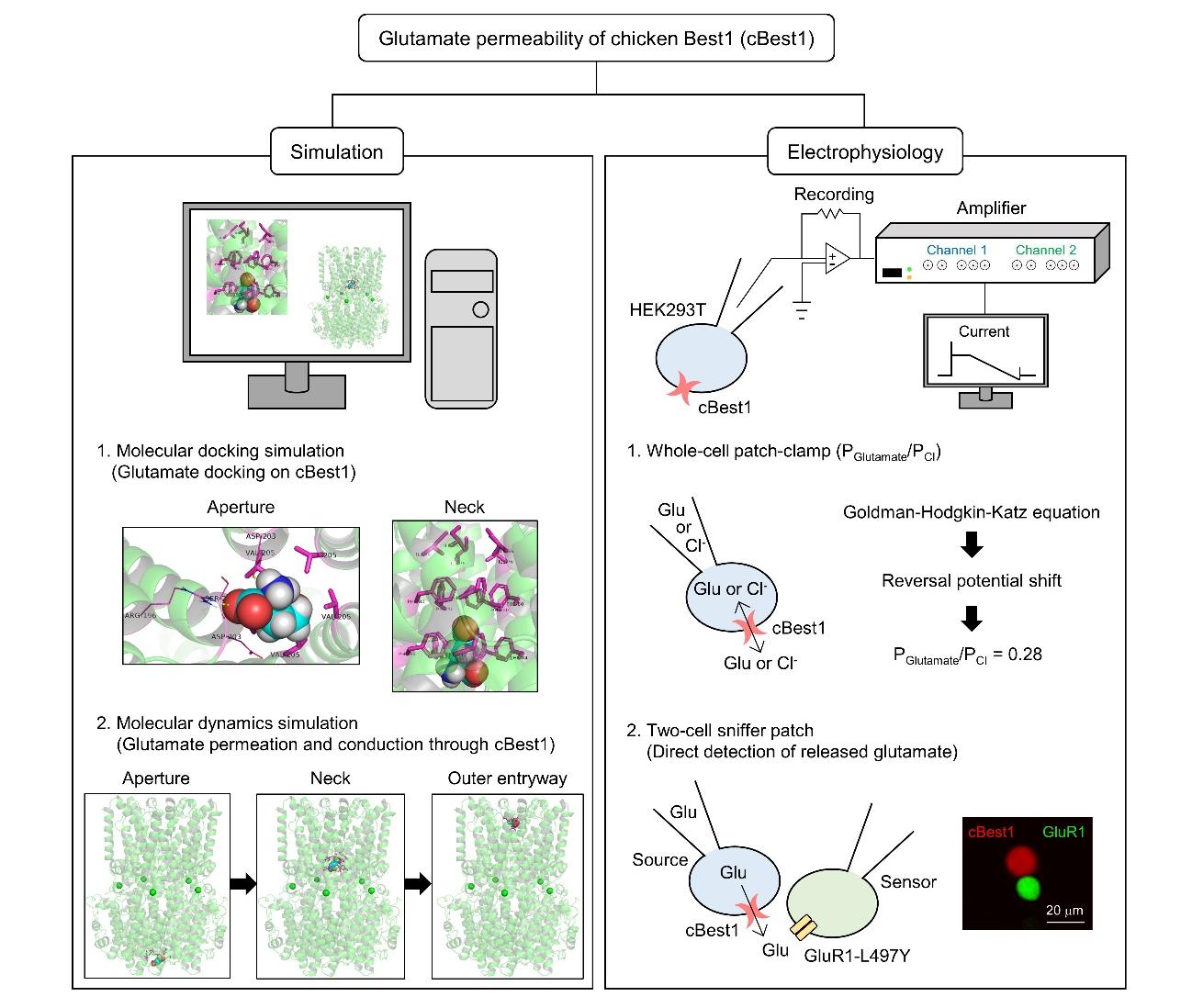
Keywords: Chicken Best1, Glutamate permeability, Molecular docking simulation, Molecular dynamics simulation, Whole-cell patch-clamp recording, Two-cell sniffer patch
INTRODUCTION
Best1, a member of the bestrophin family, is a Ca2+-activated chloride channel found primarily in the retinal pigment epithelium (RPE) of the eye [1]. This channel has been first identified in humans associated with disease-causing mutations in Best vitelliform macular dystrophy [2, 3], which is the genetic form of macular degeneration characterized by progressive loss of visual acuity [4]. Thus, most of the studies on Best1 have been focused on mammalian human (
Non-mammalian Best1 in vertebrates has been exclusively studied at the structural level on the species of
In this study, we validated the glutamate permeability of cBest1 using three important experimental methods, molecular docking, molecular dynamics simulation, and electrophysiological recordings. In the simulation, we found that glutamate is docked on the aperture and neck of cBest1 and permeates through the ion-conducting pore. Based on the simulation results, we verified the permeation of glutamate using a whole-cell patch-clamp for measuring the permeability ratio of glutamate to Cl- (PGlutamate/PCl) and an ultrasensitive two-cell sniffer patch for detecting the released glutamate directly. We determined the PGlutamate/PCl of cBest1 to be 0.28, which is a substantial permeability but slightly lower than the mammalian homologs.
MATERIALS AND METHODS
Modeling of cBest1 protein
The X-ray crystal structures of Ca2+-bound, closed-state cBest1 (amino acids 1-405; PDB ID code: 4RDQ) [12] and the cryo-electron microscopy (cryo-EM) structures of Ca2+-bound, open-state cBest1 (amino acids 1-345; PDB ID code: 6N28) [15] were downloaded from the protein data bank server (https://www.rcsb.org/). We deleted the Fab antibody fragments before processing the Ca2+-bound, closed-state cBest1 structure. Using the Discovery Studio (DS) Client 2017 (Dassault Systèmes BIOVIA, USA), both cBest1 model structures were prepared at pH 7.4 with the default parameters, added with hydrogen atoms, and minimized with the CHARMM force field for 200 steps to remove clashes and bad contacts. The resultant model structures were further utilized to set up the docking simulation or molecular dynamics simulation.
Molecular docking simulation
The 2-dimensional (2D) structure of glutamate was downloaded from the PubChem website (https://pubchem.ncbi.nlm.nih.gov/) and prepared as the 3D structure at pH 7.4 using a small molecule preparation wizard in the DS client at the default setting. At pH 7.4, the glutamate with a net charge of -1 was used. We utilized the Ca2+-bound, closed-state cBest1 (amino acids 1-405) model structure, as described above. For the docking simulation of the glutamate at the aperture for the initial positioning of glutamate, the CDOCKER, a molecular dynamics simulated-annealing-based algorithm [16], was used. The pentameric V205 at the aperture region was considered to generate the grid sphere for the binding site. The default option of CDOCKER algorithm was used to generate random conformations at 1000 Kelvin (K), and the top 10 conformations were docked and refined at the binding site. The ranking of docked poses was performed by calculating the binding energy (BE). Before calculating BE, the poses were minimized
Molecular dynamics simulation
The molecular dynamics simulation was performed using the GROMACS-2016.4 [17]. Ligand-protein complex was used for the explicit membrane-embedded molecular dynamics simulation setup. A relative membrane position of glutamate-cBest1 (ligand-protein) was obtained from the Orientations of Proteins in Membranes (OPM) database (https://opm.phar.umich.edu/ppm_server). The output file from this database was fed to CHARMM-GUI website (https://charmm-gui.org/), and we further utilized this 3D structure to set up the initial molecular dynamics simulation files. The lipid bilayer containing 450 dipalmitoylphosphatidylcholine (DPPC) lipids was constructed at the OPM-identified membrane position. A cubic box is extended by 28 Å to XYZ-direction from the cBest1 and explicitly hydrated by the 52,235 TIP3P water molecules. The overall charge of simulation systems was neutralized by the addition of 0.15 M concentration of potassium chloride (KCl) solution (184 K+ and 158 Cl-). The glutamate parameter was taken from the CHARMM General Force Fields (CGenFF), and the protein and lipids were described using CHARMM36 force fields [18].
The molecular dynamics simulation system was minimized for the maximum number of 50,000 steps or the threshold force value of 1000 kJ/mol/nm. This minimization was performed to remove the steric clashes introduced during the preparation of molecular dynamics simulation files. Next, system equilibration was performed in six steps with the reduced force constant (FC) at the backbone (FC=4000, 2000, 1000, 500, 200, 50), side chains (FC=2000, 1000, 500, 200, 50, 0), lipids (FC=1000, 400, 400, 200, 40, 0), and dihedrals (FC=1000, 400, 200, 200, 100, 0). Step 1~3 equilibrated for 25 picoseconds (ps) each and outputs were saved at each ps. System heating was performed using the Berendsen temperature coupling method with the temperature of 323.15 K and the coupling constant of 1.0. Further equilibration steps were performed for 100 ps (step 4, 5) and 1000 ps (step 6), where pressure was coupled semi-isotropically with the Berendsen coupling. The reference pressure of 1 bar and compressibility of 4.5×10-5 bar-1 were used with coupling constant 5.0. The linear constraint solver (LINCS) algorithm [19] and particle mesh Ewald (PME) method [20] were used to constrain the covalent interaction and to process electrostatic interactions, respectively. For production simulation, temperature coupling with tau-t of 1 ps was performed using the Nose-Hoover algorithm [21, 22]. The pressure coupling with tau-p of 5 ps and compressibility of 4.5×10-5 bar-1 was performed using the Parrinello-Rahman method [23].
The DPPC lipid has a high phase transition temperature of 314.15 K and exists in the 62.9~64 Å2 of area per lipid. The temperature used in the simulation should be based on the physical properties of the lipid, most notably phase transition temperature where the lipid exists in the liquid-condensed phase. Therefore, we performed the molecular dynamics simulation at 323.15 K and did not find any deformities, such as melting of protein and breaking of peptide bonds in protein, during the simulation.
Umbrella sampling simulation
After six steps of equilibration, we performed the pulling simulation of glutamate from the aperture to the extracellular space using the GROMACS-2016.4. The umbrella sampling simulation was performed for the 1,300 ps and the external pulling force outputs were saved at each 0.1 ps. All the covalent bonds were constrained during umbrella simulation using the LINCS algorithm. Temperature coupling was performed using the Nose-Hoover algorithm with the reference temperature of 323.15 K and the coupling constant of 0.5. Four different groups (protein, ligand, DPPC, and solvent with ions) were used for the temperature coupling. The pulling of glutamate towards the Z-direction was performed using the semi-isotropic pressure coupling (independent of the Z-direction), where uniform pressure scaling at the XY-direction was maintained. Periodic boundary conditions (PBC) were applied in the XYZ-direction. Glutamate was pulled towards the Z-direction under the external pulling force of 1000 kJ/mol/nm at the rate of 0.01 nm/ps. Two groups (protein and ligand) were used for the pulling. For calculating the potential energy and temperature during the umbrella sampling simulation, we used the g_energy utility. For calculating the root mean square deviation (RMSD), radius of gyration, and the number of hydrogen bonds during the umbrella sampling simulation, we used the g_rms, g_gyrate, and g_hbond utility, respectively. For the intermolecular hydrogen bonds in the simulation system, we considered only hydrogen bonds between glutamate and cBest1.
Cloning of cBest1
The
HEK293T cell line culture
Human embryonic kidney 293T (HEK293T) cells were purchased from ATCC (RRID: CVCL_0063). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, 10-013, Corning) supplemented with 25 glucose, 1 sodium pyruvate (in mM), 10% heat-inactivated fetal bovine serum (HI-FBS, 10082-147, Gibco) and 100 units/ml penicillin-streptomycin (15140-122, Gibco). Cells were maintained at 37°C in a humidified 5% CO2 incubator.
Whole-cell patch-clamp recording
For measuring Ca2+-activated Cl- current from cBest1, cBest1 (amino acids 1-405)-IRES2-EGFP clone or pIRES2-EGFP (control vector) was transiently transfected into HEK293T cells using the transfection reagent (Effectene; 301425, QIAGEN) on the day before experiment. After 18 hours, cells were replated onto the glass coverslips coated with 0.1 mg/ml poly-D-Lysine (PDL; P6407, Sigma-Aldrich) and used for whole-cell patch-clamp recording within 10 hours. The external solution contained 150 NaCl, 10 HEPES, 3 KCl, 2 CaCl2, 2 MgCl2, and 5.5 glucose (in mM) with pH adjusted to 7.3 by NaOH and osmolality adjusted to 325 mOsmol/kg. For the presence of ~4.5 µM free Ca2+, a patch electrode (6~8 MΩ) was filled with the high Ca2+ internal solution containing 146 CsCl, 5 (Ca2+)-EGTA (Ethylene glycol-bis(2-aminoethylether)-
For measuring the glutamate permeability of cBest1, 146 mM CsCl in the high Ca2+ internal solution was substituted for 146 mM Cs-Glutamate. The external solution was prepared as described above. To calculate the PGlutamate/PCl, we measured the reversal potential shift between two I-V curves obtained from each internal solution (146 mM CsCl and 146 mM Cs-Glutamate) after a correction for measured liquid junction potentials. PGlutamate/PCl was calculated with the following equation derived from the Goldman-Hodgkin-Katz equation:
The ‘o’ indicates external solution. The ‘146 Cl’ and ‘146 Glu’ represent 146 mM CsCl-containing high Ca2+ internal solution and 146 mM Cs-Glutamate-containing high Ca2+ internal solution, respectively. For plotting the normalized current-voltage curve, each current was normalized to the current at +100 mV. All normalized current-voltage curves were obtained from the average of three to five constant and consecutive sweeps within 5 min after the rupture of the cell membrane. All measured currents showed a slight current rundown [14, 24] in our experimental condition.
Two-cell sniffer patch
Two-cell sniffer patch was performed using a modified protocol from the previous report [6]. For the preparation of the source cell or the sensor cell, cBest1 (amino acids 1-405)-IRES2-DsRED (source) or non-desensitizing form of AMPA receptor subunit, GluR1-L497Y-IRES2-EGFP (sensor), was transiently transfected into HEK293T cells using the transfection reagent (Effectene) on the day before experiment. For the sensor cell, 5 μM 6-Cyano-7-nitroquinoxaline-2,3-dione (CNQX; 0190, Tocris) was supplemented in the medium to block the AMPA receptor-mediated cytotoxicity. After 18 hours, both source and sensor cells were replated together onto the PDL-coated glass coverslips with the addition of 5 μM CNQX. All coverslips were used for whole-cell patch-clamp recording within 10 hours. The external solution contained 150 NaCl, 10 HEPES, 3 KCl, 2 CaCl2, 2 MgCl2, and 5.5 glucose (in mM) with pH adjusted to 7.3 by NaOH and osmolality adjusted to 325 mOsmol/kg. For the presence of ~4.5 μM free Ca2+ in the source cells, a patch electrode (6~8 MΩ) was filled with the internal solution containing 146 Cs-Glutamate, 5 (Ca2+)-EGTA-NMDG, 2 MgCl2, 10 HEPES, 10 Sucrose, 4 Mg-ATP, and 0.3 Na2-GTP (in mM) with pH adjusted to 7.3 by CsOH and osmolality adjusted to 320 mOsmol/kg. For the zero Ca2+ in the source cells, a patch electrode (6~8 MΩ) was filled with the zero Ca2+ internal solution containing 146 Cs-Glutamate, 10 BAPTA, 2 MgCl2, 10 HEPES, 4 Mg-ATP, and 0.3 Na2-GTP (in mM) with pH adjusted to 7.3 by CsOH and osmolality adjusted to 310~320 mOsmol/kg. The internal solution for the sensor cell contained 110 Cs-Gluconate, 30 CsCl, 0.5 CaCl2, 10 HEPES, 10 BAPTA, 4 Mg-ATP, and 0.3 Na2-GTP (in mM) with pH adjusted to 7.3 by CsOH and osmolality adjusted to 290~310 mOsmol/kg. We used the holding voltage of -70 mV for both source and sensor cells. After the giga-ohm (GΩ) seal of the source cell and the rupture of the sensor cell, we started to simultaneously record the currents from each cell using MiniDigi 1B two-channel digitizer (Molecular Devices, USA). While the source cell was ruptured, the responsive current from the sensor cell was measured. To normalize different amounts of expression of GluR1-L497Y on the sensor cells, we applied the 1 mM glutamate in the external solution to maximally activate the GluR1-L497Y and measured the full activation currents from the sensor cells. Each sensor cell current was normalized to the 1 mM glutamate-induced full activation current (percentage of full activation).
Statistical analysis
For all experiments, data normality was analyzed using a D’Agostino-Pearson omnibus normality test. For data following normal distribution, difference between groups was evaluated by an unpaired t test with Welch’s correction. For data not following normal distribution, a Kruskal-Wallis test with Dunn’s multiple comparison test was performed. The significance level was represented as asterisks (*p<0.05; **p<0.01; ****p<0.0001). GraphPad Prism 9.4.1 for Windows (GraphPad Software, USA) was used for these analyses and to create the plots.
RESULTS AND DISCUSSION
Glutamate is docked on the aperture and neck of the cBest1
We first investigated the possibility of glutamate permeation through cBest1 at the simulation level using the cBest1 model structure based on the X-ray crystal structures of cBest1 with amino acids 1-405 (PDB ID code: 4RDQ) (Fig. 1A) [12]. Using this model structure, we performed the molecular docking simulation to check whether glutamate can be docked on the aperture and neck of the pore. For all simulations, we used a negatively charged glutamate as a monovalent organic anion at pH 7.4 (Fig. 1B). We found an apparent binding of glutamate at the hydrophobic aperture formed by pentameric V205 (Fig. 1C; VAL-205). The free carboxylate group of glutamate interacted with the side chain of V205, and the main chain carboxylate and the amino group were exposed to solvent (Fig. 1C). The β and γ-carbon atoms of glutamate interacted with A~C chains of cBest1 via hydrophobic interactions, and the docked pose of glutamate near V205 was slightly diagonal (Fig. 1A, C). Moreover, we observed that glutamate was properly inserted inside the neck, which consists of pentameric I76, F80, and F84 (Fig. 1D; ILE-76, PHE-80, and PHE-84). The free carboxylate group of glutamate inside the neck formed π-electronic interaction with the pentameric F80, and the β and γ-carbon atoms of glutamate interacted with the side chains of pentameric F84 via hydrophobic interactions (Fig. 1D). These results suggest that glutamate can bind well to the aperture and neck of the cBest1, which are a prerequisite for permeation through cBest1.
Glutamate permeates through the ion-conducting pore of cBest1
To investigate whether glutamate can pass through the ion-conducting pathway from the intracellular aperture to the extracellular space at the simulation level, we performed the molecular dynamics simulation and umbrella sampling simulation with a simplified simulation system including cBest1 model structure with amino acids 1-405, glutamate, lipid bilayer with DPPC lipids, water, and KCl (Fig. 2A). We first evaluated the stability of the entire simulation system by calculating the potential energy of the system during the umbrella sampling simulation. The system’s potential energy dropped from the average value of -241.0×104 kJ/mol to the average value of -241.5×104 kJ/mol at the end of the simulation (Fig. 2B), indicating that the simulation system was gradually stabilized. Moreover, we kept a sufficiently high temperature of 323.15 K during the simulation (Fig. 2C), because the phase transition temperature for DPPC lipids is around 314.15 K [25]. We then checked the conformational stability of the cBest1 model structure by calculating the root mean square deviation (RMSD) of backbone atoms. We found that the cBest1 model structure was stable with the RMSD values less than 0.1 nm (Fig. 2D). In addition to the RMSD values, we measured the radius of gyration (Rg) of backbone atoms, which provides information about protein compactness and size [26]. Although the Rg values slightly increased as the simulation progressed, there was no drastic change in the Rg values (Fig. 2E). These results indicate that the cBest1 model structure was stable during the whole time of simulation.
Under physiological conditions, the cell membrane experiences the resting membrane potential of -60~-80 mV depending on the cell types in the brain [27]. This resting membrane potential can exert enough force for permeation and conduction for a conducting ion. To see the glutamate permeation and conduction processes through the ion-conducting pathway of cBest1, the external pulling force, to mimic the force exerted by the resting membrane potential, was applied to the glutamate in the Z-direction using the umbrella sampling simulation. The initial position of glutamate was defined near the aperture region (V205). With the external pulling force, glutamate passed the aperture region, and it took around 160 ps (Fig. 2F, G, Supplementary Movie 1). Following the aperture, glutamate traversed to the start point of the neck region (F84) at around 524 ps and repositioned itself from horizontal to vertical (Fig. 2F, G). This glutamate entered the F84 halfway at 600 ps and exited from the whole neck region containing residues (F84, F80, and I76) at 750 ps, which was finally released into the extracellular space (Fig. 2F, G, Supplementary Movie 1). Next, we measured the number of hydrogen bonds between glutamate and residues in the ion-conducting pathway to see if there is a correlation between the orientation of glutamate and the number of hydrogen bonds during the conduction of glutamate. At the aperture region and inner cavity, glutamate formed a maximum of 4~5 and 1~3 hydrogen bonds with the surrounding residues, respectively (Fig. 2H). We observed the hydrogen bonds at the aperture region in detail, showing that the main chain carboxylate of glutamate formed three hydrogen bonds with the main and side chains of S204, a preceding residue of the aperture of B chain, while the protonated amine of glutamate formed a hydrogen bond with D203 of B chain (Fig. 2H, inset; SER-204, ASP-203). In addition to the polar (S204) and acidic (D203) residues, basic (R196) residue might be participating in the hydrogen bonding with glutamate (Fig. 2H, inset; ARG-196). In contrast to the aperture and inner cavity, almost no hydrogen bond (524 ps~750 ps) was present during traversing the neck region (Fig. 2H). This is highly consistent with the majorly horizontal orientation of glutamate at the aperture and inner cavity, and the vertical orientation of glutamate at the neck region (Fig. 2F). These results suggest a critical role of hydrogen bonds between glutamate and surrounding residues during the conduction of glutamate. Furthermore, we measured the external pulling force to permeate glutamate through cBest1 to examine how much force is needed to pass through the specific region of the pore. We found that the external pulling force required to pass the aperture restriction was up to 1068 kJ/mol/nm which is the highest value during permeation (Fig. 2I), indicating that traversing the aperture region is the rate-determining step for glutamate permeation. This highest value of external pulling force could be explained by the function of the aperture as a size-selective filter [13]. In contrast to the aperture, the glutamate only needed the external pulling force of 577 kJ/mol/nm to pass the neck region (Fig. 2I), which is consistent with a previous report showing the neck as a Ca2+-dependent gate, not a size-selective filter [13].
These calculated force values (Fig. 2I) are based on the Ca2+-bound, closed-state cBest1 with amino acids 1-405 (PDB ID code: 4RDQ) [12]. According to the Ca2+-bound, open-state cBest1 with amino acids 1-345 (PDB ID code: 6N28), the diameter of the neck widens from less than 3.5 Å to 13 Å, whereas the diameter of aperture remains unaltered at less than 4 Å [15]. Therefore, the force value of 577 kJ/mol/nm at the neck could be much less in the Ca2+-bound, open-state cBest1. In fact, at the pore diameter of the neck in the open-state (13 Å), a glutamate molecule with the dimension of 6.5×10.8 Å [28] should freely enter and permeate through the neck by a diffusional force. The minimum diffusional force of glutamate can be estimated to be 1.87 kJ/mol/nm by the simple formula of E (Joule)=V (Volt)×Q (Coulomb) at the resting membrane potential of -70 mV, the net charge of -1 for glutamate, and the thickness of the DPPC lipid bilayer as about 3.6 nm [29], assuming no steric hindrance and frictional force. This estimated value of 1.87 kJ/mol/nm is comparable to 3 kJ/mol/nm, which is based on the predicted external pulling force value in the open-state of the neck region by calculating the average external pulling force value near the end point of the membrane-spanning neck region of the closed-state (42 kJ/mol/nm, averaged over 659~750 ps in Fig. 2I) and correcting for the expanded diameter of the neck in the open-state, 13 Å (42 kJ/mol/nm divided by a factor of 14), under the assumption of negative linear correlation between force and area of the pore. Indeed, we performed the umbrella sampling simulation using the Ca2+-bound, open-state cBest1 model structure with amino acids 1-345, and found that the force required to pass near the end point of the membrane-spanning neck region (averaged over 592~683 ps) was 122 kJ/mol/nm (Fig. 2J, Supplementary Movie 2). This indicates that the actual external pulling force value in the open-state of the neck region could be much higher than the predicted value from the closed-state, possibly due to the steric hindrance and frictional force. This apparently high external pulling force value in the open-state is consistent with the fact that the Bestrophin family of channels show extremely small single-channel conductance [30], possibly due to an energy barrier imposed by the neck [12], despite the relatively high glutamate permeability [6, 31]. Taken together, our molecular dynamics simulation predicts that glutamate can pass through the ion-conducting pore of cBest1.
The cBest1 is a glutamate-permeable channel
Next, we investigated the actual existence of glutamate permeability of cBest1 by performing electrophysiological recordings with the heterologous expression of cBest1 in HEK293T cells. Firstly, we cloned
To examine whether cBest1 is permeable to glutamate, we measured the cBest1-mediated currents using the internal solution with either Cl- or glutamate as the predominant anions and the high Ca2+ (Fig. 3E). We found substantial inward currents, carried by glutamate, with a negatively shifted reversal potential in the presence of internal glutamate (146 mM Cs-Glutamate; -32.77±3.48 mV) compared to that in the presence of internal Cl- (146 mM CsCl; -1.97±0.48 mV) (Fig. 3F, G). Based on the average shift in the reversal potential of 30.80 mV, we calculated the PGlutamate/PCl to be 0.28 (Fig. 3G) using a modified Goldman-Hodgkin-Katz equation as previously described [6, 34, 35]. This calculated PGlutamate/PCl of cBest1 was about 42% of the reported PGlutamate/PCl of mBest1 (0.67) [6]. These results indicate that non-mammalian cBest1 has a considerable glutamate permeability but is slightly lower than the mammalian homologs. Moreover, we directly measured the released glutamate through cBest1 using a two-cell sniffer patch with a pair of source cell expressing cBest1 and a neighboring sensor cell expressing GluR1-L497Y as a biosensor for glutamate (Fig. 3H), as previously described [6]. As a result, upon rupture of the source cell membrane with the internal solution containing 146 mM glutamate and high Ca2+, we found robust glutamate-induced inward currents from the neighboring sensor cell, while there was almost no observable sensor current in the absence of intracellular Ca2+ (Fig. 3I, J). Taken together, these results indicate that cBest1 is also capable of releasing glutamate in a Ca2+-dependent manner like mammalian homologs.
Our study is the first to identify the glutamate permeability of cBest1 using advanced techniques of molecular simulations and electrophysiology. Based on the simulations, we found that glutamate is docked on the aperture and neck of cBest1, and it can pass the pore region of cBest1 with the external pulling force. Based on the electrophysiological recordings, we successfully demonstrate that cBest1 has a substantial permeability for glutamate with the PGlutamate/PCl as 0.28, and the released glutamate can be directly detected by the neighboring cells.
Our results are contrary to the previous reports claiming that cBest1 has no permeability for glutamate [12, 13]. Kane Dickson et al. [12] have used a fluorescence-based ion flux assay which can indirectly measure the anion permeability by monitoring fluorescence change by proton uptake to counter the anion influx into the reconstituted proteoliposome. Thus, it is necessary to be cautious in interpreting the results of this assay due to its indirect measurement of glutamate permeability. Moreover, Vaisey et al. [13] have observed no glutamate-mediated anion current in the electrophysiological recordings using the purified cBest1 protein in the artificial planar lipid bilayer system, while we obtained the substantial currents by the efflux of glutamate using heterologously expressed cBest1 in HEK293T cells under the whole-cell patch-clamp configuration. This discrepancy might be due to each distinct experimental condition for current recordings. For example, precise measurement of permeability ratio requires a proper correction for liquid junction potential when exchanging the bath solutions. It is unclear whether Vaisey et al. [13] made a proper correction for liquid junction potential or not, based on the description of the method for exchanging with glutamate-containing bath solution. In addition, 4 mM ATP was added in our internal solution for whole-cell patch-clamp, but not in their
In summary, we have demonstrated that glutamate can permeate and be released through cBest1. It will be of great interest to investigate the possibility of permeation of other large organic solutes, such as GABA [7] and D-serine [8], through cBest1. The ability of cBest1 to release glutamate implies the potential and critical roles of cBest1 in cognition and brain diseases, as it has been demonstrated with the mammalian homologs.
Supplemental Materials
ACKNOWLEDGEMENTS
This work was supported by the Institute for Basic Science (IBS), Center for Cognition and Sociality (IBS-R001-D2); Korea Institute of Science and Technology intramural grant (2E31522) with Korea Institute of Science and Technology Information (KISTI) supercomputing resources (KSC-2019-CRE-0155 and KSC-2021-RND-0059).
Figures



References
- Marmorstein AD, Marmorstein LY, Rayborn M, Wang X, Hollyfield JG, Petrukhin K (2000) Bestrophin, the product of the Best vitelliform macular dystrophy gene (VMD2), localizes to the basolateral plasma membrane of the retinal pigment epithelium. Proc Natl Acad Sci U S A 97:12758-12763
- Petrukhin K, Koisti MJ, Bakall B, Li W, Xie G, Marknell T, Sandgren O, Forsman K, Holmgren G, Andreasson S, Vujic M, Bergen AA, McGarty-Dugan V, Figueroa D, Austin CP, Metzker ML, Caskey CT, Wadelius C (1998) Identification of the gene responsible for Best macular dystrophy. Nat Genet 19:241-247
- Marquardt A, Stöhr H, Passmore LA, Krämer F, Rivera A, Weber BH (1998) Mutations in a novel gene, VMD2, encoding a protein of unknown properties cause juvenile-onset vitelliform macular dystrophy (Best's disease). Hum Mol Genet 7:1517-1525
- Crincoli E, Zhao Z, Querques G, Sacconi R, Carlà MM, Giannuzzi F, Ferrara S, Ribarich N, L'Abbate G, Rizzo S, Souied EH, Miere A (2022) Deep learning to distinguish Best vitelliform macular dystrophy (BVMD) from adult-onset vitelliform macular degeneration (AVMD). Sci Rep 12:12745
- Oh SJ, Lee CJ (2017) Distribution and function of the bestrophin-1 (Best1) channel in the brain. Exp Neurobiol 26:113-121
- Woo DH, Han KS, Shim JW, Yoon BE, Kim E, Bae JY, Oh SJ, Hwang EM, Marmorstein AD, Bae YC, Park JY, Lee CJ (2012) TREK-1 and Best1 channels mediate fast and slow glutamate release in astrocytes upon GPCR activation. Cell 151:25-40
- Lee S, Yoon BE, Berglund K, Oh SJ, Park H, Shin HS, Augustine GJ, Lee CJ (2010) Channel-mediated tonic GABA release from glia. Science 330:790-796
- Koh W, Park M, Chun YE, Lee J, Shim HS, Park MG, Kim S, Sa M, Joo J, Kang H, Oh SJ, Woo J, Chun H, Lee SE, Hong J, Feng J, Li Y, Ryu H, Cho J, Lee CJ (2022) Astrocytes render memory flexible by releasing D-serine and regulating NMDA receptor tone in the hippocampus. Biol Psychiatry 91:740-752
- Han KS, Woo J, Park H, Yoon BJ, Choi S, Lee CJ (2013) Channel-mediated astrocytic glutamate release via Bestrophin-1 targets synaptic NMDARs. Mol Brain 6:4
- Park H, Han KS, Seo J, Lee J, Dravid SM, Woo J, Chun H, Cho S, Bae JY, An H, Koh W, Yoon BE, Berlinguer-Palmini R, Mannaioni G, Traynelis SF, Bae YC, Choi SY, Lee CJ (2015) Channel-mediated astrocytic glutamate modulates hippocampal synaptic plasticity by activating postsynaptic NMDA receptors. Mol Brain 8:7
- Oh SJ, Lee JM, Kim HB, Lee J, Han S, Bae JY, Hong GS, Koh W, Kwon J, Hwang ES, Woo DH, Youn I, Cho IJ, Bae YC, Lee S, Shim JW, Park JH, Lee CJ (2019) Ultrasonic neuromodulation via astrocytic TRPA1. Curr Biol 29:3386-3401.e8
- Kane Dickson V, Pedi L, Long SB (2014) Structure and insights into the function of a Ca(2+)-activated Cl(-) channel. Nature 516:213-218
- Vaisey G, Miller AN, Long SB (2016) Distinct regions that control ion selectivity and calcium-dependent activation in the bestrophin ion channel. Proc Natl Acad Sci U S A 113:E7399-E7408
- Vaisey G, Long SB (2018) An allosteric mechanism of inactivation in the calcium-dependent chloride channel BEST1. J Gen Physiol 150:1484-1497
- Miller AN, Vaisey G, Long SB (2019) Molecular mechanisms of gating in the calcium-activated chloride channel bestrophin. Elife 8:e43231
- Wu G, Robertson DH, Brooks CL 3rd, Vieth M (2003) Detailed analysis of grid-based molecular docking: a case study of CDOCKER-A CHARMm-based MD docking algorithm. J Comput Chem 24:1549-1562
- Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2:19-25
- Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr (2010) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31:671-690
- Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463-1472
- Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577-8593
- Nosé S (1984) A molecular dynamics method for simulations in the canonical ensemble. Mol Phys 52:255-268
- Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A Gen Phys 31:1695-1697
- Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182-7190
- Xiao Q, Prussia A, Yu K, Cui YY, Hartzell HC (2008) Regulation of bestrophin Cl channels by calcium: role of the C terminus. J Gen Physiol 132:681-692
- Khakbaz P, Klauda JB (2018) Investigation of phase transitions of saturated phosphocholine lipid bilayers via molecular dynamics simulations. Biochim Biophys Acta Biomembr 1860:1489-1501
- Arnittali M, Rissanou AN, Harmandaris V (2019) Structure of biomolecules through molecular dynamics simulations. Procedia Comput Sci 156:69-78
- Hille B (1992) Ionic channels of excitable membranes. 2nd ed. Oxford University Press, Oxford
- Takano T, Kang J, Jaiswal JK, Simon SM, Lin JH, Yu Y, Li Y, Yang J, Dienel G, Zielke HR, Nedergaard M (2005) Receptor-mediated glutamate release from volume sensitive channels in astrocytes. Proc Natl Acad Sci U S A 102:16466-16471
- Curtis EM, Hall CK (2013) Molecular dynamics simulations of DPPC bilayers using "LIME", a new coarse-grained model. J Phys Chem B 117:5019-5030
- Chien LT, Zhang ZR, Hartzell HC (2006) Single Cl- channels activated by Ca2+ in Drosophila S2 cells are mediated by bestrophins. J Gen Physiol 128:247-259
- Park H, Han KS, Oh SJ, Jo S, Woo J, Yoon BE, Lee CJ (2013) High glutamate permeability and distal localization of Best1 channel in CA1 hippocampal astrocyte. Mol Brain 6:54
- Weill U, Krieger G, Avihou Z, Milo R, Schuldiner M, Davidi D (2019) Assessment of GFP tag position on protein localization and growth fitness in yeast. J Mol Biol 431:636-641
- Dave K, Gelman H, Thu CT, Guin D, Gruebele M (2016) The effect of fluorescent protein tags on phosphoglycerate kinase stability is nonadditive. J Phys Chem B 120:2878-2885
- Qu Z, Hartzell HC (2000) Anion permeation in Ca(2+)-activated Cl(-) channels. J Gen Physiol 116:825-844
- Han YE, Kwon J, Won J, An H, Jang MW, Woo J, Lee JS, Park MG, Yoon BE, Lee SE, Hwang EM, Jung JY, Park H, Oh SJ, Lee CJ (2019) Tweety-homolog (Ttyh) family encodes the pore-forming subunits of the swelling-dependent volume-regulated anion channel (VRACswell) in the brain. Exp Neurobiol 28:183-215
- Zhang Y, Kittredge A, Ward N, Ji C, Chen S, Yang T (2018) ATP activates bestrophin ion channels through direct interaction. Nat Commun 9:3126
- Liao Z, Lockhead D, St Clair JR, Larson ED, Wilson CE, Proenza C (2012) Cellular context and multiple channel domains determine cAMP sensitivity of HCN4 channels: ligand-independent relief of autoinhibition in HCN4. J Gen Physiol 140:557-566
- Woo JS, Srikanth S, Gwack Y (2018) Modulation of Orai1 and STIM1 by cellular factors. In: Calcium entry channels in non-excitable cells (Kozak JA, Putney JW Jr eds), pp 73-92. Boca Raton, FL





