A Review and Database of Snake Venom Proteomes
Abstract
:1. Introduction
2. Results
3. Discussion
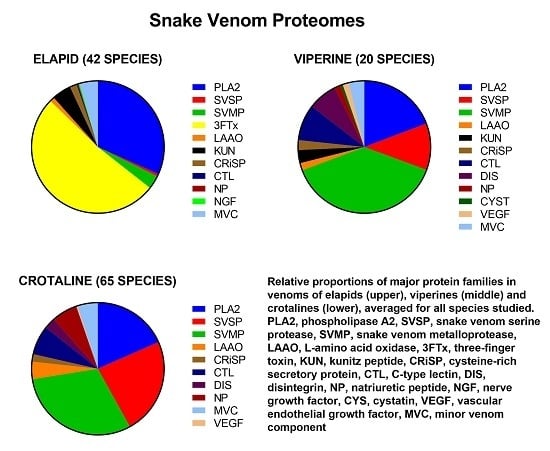
- Dominant protein families (four families): PLA2, SVMP, SVSP and 3FTx.
- Secondary protein families (six families that were commonly present, but in much smaller amounts than the dominant families): KUN, CRiSP, LAAO, CTL, DIS, and NP.
- Minor protein families (nine families): acetylcholinesterase, hyaluronidase, 5′ nucleotidase, phosphodiesterase, phospholipase B, nerve growth factor, vascular endothelial growth factor, vespryn/ohanin and snake venom metalloprotease inhibitor.
- Rare protein families: 36 families listed in Table S3.
- Unique protein families (four families): defensins, waglerin, maticotoxin and cystatins. These families make up to 38% of the whole venom of a single species, but are classified separately as each is present in only one genus.
3.1. Dominant Protein Families
3.2. Secondary Protein Families
3.3. Major Inter-Family Differences
3.3.1. Elapids
3.3.2. Vipers
3.4. Medical Implications
3.5. Evolutionary Biology
4. Methods
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jackson, T.N.W.; Young, B.; Underwood, G.; McCarthy, C.; Kochva, E.; Vidal, N.; van der Weerd, L.; Nabuurs, R.; Dobson, J.; Whitehead, D.; et al. Endless forms most beautiful: The evolution of ophidian oral glands, including the venom system, and the use of appropriate terminology for homologous structures. Zoomorphology 2017, 136, 107–130. [Google Scholar] [CrossRef]
- Vonk, F.J.; Admiraal, J.F.; Jackson, K.; Reshef, R.; de Bakker, M.A.; Vanderschoot, K.; van den Berge, I.; van Atten, M.; Burgerhout, E.; Beck, A. Evolutionary origin and development of snake fangs. Nature 2008, 454, 630. [Google Scholar] [CrossRef] [PubMed]
- Doley, R.; Nguyen Ngoc Bao, T.; Reza, M.A.; Kini, R.M. Unusual accelerated rate of deletions and insertions in toxin genes in the venom glands of the pygmy copperhead (Austrelaps labialis) from kangaroo island. BMC Evol. Biol. 2008, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Chatrath, S.T.; Chapeaurouge, A.; Lin, Q.; Lim, T.K.; Dunstan, N.; Mirtschin, P.; Kumar, P.P.; Kini, R.M. Identification of Novel Proteins from the Venom of a Cryptic Snake Drysdalia coronoides by a Combined Transcriptomics and Proteomics Approach. J. Proteome Res. 2011, 10, 739–750. [Google Scholar] [CrossRef] [PubMed]
- Paiva, O.; Pla, D.; Wright, C.E.; Beutler, M.; Sanz, L.; Gutiérrez, J.M.; Williams, D.J.; Calvete, J.J. Combined venom gland cDNA sequencing and venomics of the New Guinea small-eyed snake, Micropechis ikaheka. J. Proteom. 2014, 110, 209–229. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.H.; Tan, K.Y.; Tan, N.H. Revisiting Notechis scutatus venom: On shotgun proteomics and neutralization by the “bivalent” Sea Snake Antivenom. J. Proteom. 2016, 144, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Herrera, M.; Fernández, J.; Vargas, M.; Villalta, M.; Segura, Á.; León, G.; Angulo, Y.; Paiva, O.; Matainaho, T.; Jensen, S.D.; et al. Comparative proteomic analysis of the venom of the taipan snake, Oxyuranus scutellatus, from Papua New Guinea and Australia: Role of neurotoxic and procoagulant effects in venom toxicity. J. Proteom. 2012, 75, 2128–2140. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Bande, B.W.; Welton, R.E.; Paiva, O.K.; Sanz, L.; Segura, A.; Wright, C.E.; Calvete, J.J.; Gutierrez, J.M.; Williams, D.J. Proteomics and antivenomics of Papuan black snake (Pseudechis papuanus) venom with analysis of its toxicological profile and the preclinical efficacy of Australian antivenoms. J. Proteom. 2017, 150, 201–215. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Ghezellou, P.; Paiva, O.; Matainaho, T.; Ghassempour, A.; Goudarzi, H.; Kraus, F.; Sanz, L.; Williams, D.J. Snake venomics of two poorly known Hydrophiinae: Comparative proteomics of the venoms of terrestrial Toxicocalamus longissimus and marine Hydrophis cyanocinctus. J. Proteom. 2012, 75, 4091–4101. [Google Scholar] [CrossRef] [PubMed]
- Laustsen, A.H.; Gutiérrez, J.M.; Rasmussen, A.R.; Engmark, M.; Gravlund, P.; Sanders, K.L.; Lohse, B.; Lomonte, B. Danger in the reef: Proteome, toxicity, and neutralization of the venom of the olive sea snake, Aipysurus laevis. Toxicon 2015, 107, 187–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomonte, B.; Pla, D.; Sasa, M.; Tsai, W.-C.; Solórzano, A.; Ureña-Díaz, J.M.; Fernández-Montes, M.L.; Mora-Obando, D.; Sanz, L.; Gutiérrez, J.M.; et al. Two color morphs of the pelagic yellow-bellied sea snake, Pelamis platura, from different locations of Costa Rica: Snake venomics, toxicity, and neutralization by antivenom. J. Proteom. 2014, 103, 137–152. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.H.; Tan, K.Y.; Lim, S.E.; Tan, N.H. Venomics of the beaked sea snake, Hydrophis schistosus: A minimalist toxin arsenal and its cross-neutralization by heterologous antivenoms. J. Proteom. 2015, 126, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.H.; Wong, K.Y.; Tan, K.Y.; Tan, N.H. Venom proteome of the yellow-lipped sea krait, Laticauda colubrina from Bali: Insights into subvenomic diversity, venom antigenicity and cross-neutralization by antivenom. J. Proteom. 2017, 166, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Oh, A.M.F.; Tan, C.H.; Ariaranee, G.C.; Quraishi, N.; Tan, N.H. Venomics of Bungarus caeruleus (Indian krait): Comparable venom profiles, variable immunoreactivities among specimens from Sri Lanka, India and Pakistan. J. Proteom. 2017, 164, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Rusmili, M.R.A.; Yee, T.T.; Mustafa, M.R.; Hodgson, W.C.; Othman, I. Proteomic characterization and comparison of Malaysian Bungarus candidus and Bungarus fasciatus venoms. J. Proteom. 2014, 110, 129–144. [Google Scholar] [CrossRef] [PubMed]
- Ziganshin, R.H.; Kovalchuk, S.I.; Arapidi, G.P.; Starkov, V.G.; Hoang, A.N.; Thi Nguyen, T.T.; Nguyen, K.C.; Shoibonov, B.B.; Tsetlin, V.I.; Utkin, Y.N. Quantitative proteomic analysis of Vietnamese krait venoms: Neurotoxins are the major components in Bungarus multicinctus and phospholipases A2 in Bungarus fasciatus. Toxicon 2015, 107, 197–209. [Google Scholar] [CrossRef] [PubMed]
- Lauridsen, L.P.; Laustsen, A.H.; Lomonte, B.; Gutiérrez, J.M. Toxicovenomics and antivenom profiling of the Eastern green mamba snake (Dendroaspis angusticeps). J. Proteom. 2016, 136, 248–261. [Google Scholar] [CrossRef] [PubMed]
- Laustsen, A.H.; Lomonte, B.; Lohse, B.; Fernández, J.; Gutiérrez, J.M. Unveiling the nature of black mamba (Dendroaspis polylepis) venom through venomics and antivenom immunoprofiling: Identification of key toxin targets for antivenom development. J. Proteom. 2015, 119, 126–142. [Google Scholar] [CrossRef] [PubMed]
- Malih, I.; Ahmad rusmili, M.R.; Tee, T.Y.; Saile, R.; Ghalim, N.; Othman, I. Proteomic analysis of Moroccan cobra Naja haje legionis venom using tandem mass spectrometry. J. Proteom. 2014, 96, 240–252. [Google Scholar] [CrossRef] [PubMed]
- Lauridsen, L.P.; Laustsen, A.H.; Lomonte, B.; Gutiérrez, J.M. Exploring the venom of the forest cobra snake: Toxicovenomics and antivenom profiling of Naja melanoleuca. J. Proteom. 2017, 150, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Petras, D.; Sanz, L.; Segura, Á.; Herrera, M.; Villalta, M.; Solano, D.; Vargas, M.; León, G.; Warrell, D.A.; Theakston, R.D.G.; et al. Snake Venomics of African Spitting Cobras: Toxin Composition and Assessment of Congeneric Cross-Reactivity of the Pan-African EchiTAb-Plus-ICP Antivenom by Antivenomics and Neutralization Approaches. J. Proteome Res. 2011, 10, 1266–1280. [Google Scholar] [CrossRef] [PubMed]
- Shan, L.-L.; Gao, J.-F.; Zhang, Y.-X.; Shen, S.-S.; He, Y.; Wang, J.; Ma, X.-M.; Ji, X. Proteomic characterization and comparison of venoms from two elapid snakes (Bungarus multicinctus and Naja atra) from China. J. Proteom. 2016, 138, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-W.; Liu, B.-S.; Chien, K.-Y.; Chiang, L.-C.; Huang, S.-Y.; Sung, W.-C.; Wu, W.-G. Cobra venom proteome and glycome determined from individual snakes of Naja atra reveal medically important dynamic range and systematic geographic variation. J. Proteom. 2015, 128, 92–104. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Zhao, H.-Y.; Yin, Y.; Shen, S.-S.; Shan, L.-L.; Chen, C.-X.; Zhang, Y.-X.; Gao, J.-F.; Ji, X. Combined venomics, antivenomics and venom gland transcriptome analysis of the monocoled cobra (Naja kaouthia) from China. J. Proteom. 2017, 159, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Tan, K.Y.; Tan, C.H.; Fung, S.Y.; Tan, N.H. Venomics, lethality and neutralization of Naja kaouthia (monocled cobra) venoms from three different geographical regions of Southeast Asia. J. Proteom. 2015, 120, 105–125. [Google Scholar] [CrossRef] [PubMed]
- Dutta, S.; Chanda, A.; Kalita, B.; Islam, T.; Patra, A.; Mukherjee, A.K. Proteomic analysis to unravel the complex venom proteome of eastern India Naja naja: Correlation of venom composition with its biochemical and pharmacological properties. J. Proteom. 2017, 156, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Sintiprungrat, K.; Watcharatanyatip, K.; Senevirathne, W.D.S.T.; Chaisuriya, P.; Chokchaichamnankit, D.; Srisomsap, C.; Ratanabanangkoon, K. A comparative study of venomics of Naja naja from India and Sri Lanka, clinical manifestations and antivenomics of an Indian polyspecific antivenom. J. Proteom. 2016, 132, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Tan, N.H.; Wong, K.Y.; Tan, C.H. Venomics of Naja sputatrix, the Javan spitting cobra: A short neurotoxin-driven venom needing improved antivenom neutralization. J. Proteom. 2017, 157, 18–32. [Google Scholar] [CrossRef] [PubMed]
- Petras, D.; Heiss, P.; Süssmuth, R.D.; Calvete, J.J. Venom Proteomics of Indonesian King Cobra, Ophiophagus hannah: Integrating Top-Down and Bottom-Up Approaches. J. Proteome Res. 2015, 14, 2539–2556. [Google Scholar] [CrossRef] [PubMed]
- Fernández, J.; Vargas-Vargas, N.; Pla, D.; Sasa, M.; Rey-Suárez, P.; Sanz, L.; Gutiérrez, J.M.; Calvete, J.J.; Lomonte, B. Snake venomics of Micrurus alleni and Micrurus mosquitensis from the Caribbean region of Costa Rica reveals two divergent compositional patterns in New World elapids. Toxicon 2015, 107, 217–233. [Google Scholar] [CrossRef] [PubMed]
- Corrêa-Netto, C.; Junqueira-de-Azevedo, I.d.L.M.; Silva, D.A.; Ho, P.L.; Leitão-de-Araújo, M.; Alves, M.L.M.; Sanz, L.; Foguel, D.; Zingali, R.B.; Calvete, J.J. Snake venomics and venom gland transcriptomic analysis of Brazilian coral snakes, Micrurus altirostris and M. corallinus. J. Proteom. 2011, 74, 1795–1809. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Rey-Suárez, P.; Fernández, J.; Sasa, M.; Pla, D.; Vargas, N.; Bénard-Valle, M.; Sanz, L.; Corrêa-Netto, C.; Núñez, V.; et al. Venoms of Micrurus coral snakes: Evolutionary trends in compositional patterns emerging from proteomic analyses. Toxicon 2016, 122, 7–25. [Google Scholar] [CrossRef] [PubMed]
- Rey-Suárez, P.; Núñez, V.; Fernández, J.; Lomonte, B. Integrative characterization of the venom of the coral snake Micrurus dumerilii (Elapidae) from Colombia: Proteome, toxicity, and cross-neutralization by antivenom. J. Proteom. 2016, 136, 262–273. [Google Scholar] [CrossRef] [PubMed]
- Margres, M.J.; Aronow, K.; Loyacano, J.; Rokyta, D.R. The venom-gland transcriptome of the eastern coral snake (Micrurus fulvius) reveals high venom complexity in the intragenomic evolution of venoms. BMC Genom. 2013, 14, 531. [Google Scholar] [CrossRef] [PubMed]
- Rey-Suárez, P.; Núñez, V.; Gutiérrez, J.M.; Lomonte, B. Proteomic and biological characterization of the venom of the redtail coral snake, Micrurus mipartitus (Elapidae), from Colombia and Costa Rica. J. Proteom. 2011, 75, 655–667. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, J.; Alape-Giron, A.; Angulo, Y.; Sanz, L.; Gutierrez, J.; Calvete, J.; Lomonte, B. Venomic and antivenomic analyses of the Central American Coral Snake, Micrurus nigrocinctus (Elapidae). J Proteome Res. 2011, 10, 1816–1827. [Google Scholar] [CrossRef] [PubMed]
- Sanz, L.; Pla, D.; Pérez, A.; Rodríguez, Y.; Zavaleta, A.; Salas, M.; Lomonte, B.; Calvete, J.J. Venomic Analysis of the Poorly Studied Desert Coral Snake, Micrurus tschudii tschudii, Supports the 3FTx/PLA2 Dichotomy across Micrurus Venoms. Toxins 2016, 8, 178. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Escolano, J.; Sanz, L. Snake Venomics of Bitis Species Reveals Large Intragenus Venom Toxin Composition Variation: Application to Taxonomy of Congeneric Taxa. J. Proteome Res. 2007, 6, 2732–2745. [Google Scholar] [CrossRef] [PubMed]
- Fahmi, L.; Makran, B.; Pla, D.; Sanz, L.; Oukkache, N.; Lkhider, M.; Harrison, R.A.; Ghalim, N.; Calvete, J.J. Venomics and antivenomics profiles of North African Cerastes cerastes and C. vipera populations reveals a potentially important therapeutic weakness. J. Proteom. 2012, 75, 2442–2453. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.K.; Kalita, B.; Mackessy, S.P. A proteomic analysis of Pakistan Daboia russelii russelii venom and assessment of potency of Indian polyvalent and monovalent antivenom. J. Proteom. 2016, 144, 73–86. [Google Scholar] [CrossRef] [PubMed]
- Kalita, B.; Patra, A.; Mukherjee, A.K. Unraveling the Proteome Composition and Immuno-profiling of Western India Russell’s Viper Venom for In-Depth Understanding of Its Pharmacological Properties, Clinical Manifestations, and Effective Antivenom Treatment. J. Proteome Res. 2017, 16, 583–598. [Google Scholar] [CrossRef] [PubMed]
- Tan, N.H.; Fung, S.Y.; Tan, K.Y.; Yap, M.K.K.; Gnanathasan, C.A.; Tan, C.H. Functional venomics of the Sri Lankan Russell’s viper (Daboia russelii) and its toxinological correlations. J. Proteom. 2015, 128, 403–423. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Harrison, R.A.; Wüster, W.; Wagstaff, S.C. Comparative venom gland transcriptome surveys of the saw-scaled vipers (Viperidae: Echis) reveal substantial intra-family gene diversity and novel venom transcripts. BMC Genom. 2009, 10, 564. [Google Scholar] [CrossRef] [PubMed]
- Makran, B.; Fahmi, L.; Pla, D.; Sanz, L.; Oukkache, N.; Lkhider, M.; Ghalim, N.; Calvete, J.J. Snake venomics of Macrovipera mauritanica from Morocco, and assessment of the para-specific immunoreactivity of an experimental monospecific and a commercial antivenoms. J. Proteom. 2012, 75, 2431–2441. [Google Scholar] [CrossRef] [PubMed]
- Sanz, L.; Ayvazyan, N.; Calvete, J.J. Snake venomics of the Armenian mountain vipers Macrovipera lebetina obtusa and Vipera raddei. J. Proteom. 2008, 71, 198–209. [Google Scholar] [CrossRef] [PubMed]
- Göçmen, B.; Heiss, P.; Petras, D.; Nalbantsoy, A.; Süssmuth, R.D. Mass spectrometry guided venom profiling and bioactivity screening of the Anatolian Meadow Viper, Vipera anatolica. Toxicon 2015, 107, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Latinović, Z.; Leonardi, A.; Šribar, J.; Sajevic, T.; Žužek, M.C.; Frangež, R.; Halassy, B.; Trampuš-Bakija, A.; Pungerčar, J.; Križaj, I. Venomics of Vipera berus berus to explain differences in pathology elicited by Vipera ammodytes ammodytes envenomation: Therapeutic implications. J. Proteom. 2016, 146, 34–47. [Google Scholar] [CrossRef] [PubMed]
- Kovalchuk, S.I.; Ziganshin, R.H.; Starkov, V.G.; Tsetlin, V.I.; Utkin, Y.N. Quantitative Proteomic Analysis of Venoms from Russian Vipers of Pelias Group: Phospholipases A2 are the Main Venom Components. Toxins 2016, 8, 105. [Google Scholar] [CrossRef] [PubMed]
- Fenwick, A.M.; Gutberlet, J.R.L.; Evans, J.A.; Parkinson, C.L. Morphological and molecular evidence for phylogeny and classification of South American pitvipers, genera Bothrops, Bothriopsis, and Bothrocophias (Serpentes: Viperidae). Zool. J. Linn. Soc. 2009, 156, 617–640. [Google Scholar] [CrossRef]
- Tang, E.L.H.; Tan, C.H.; Fung, S.Y.; Tan, N.H. Venomics of Calloselasma rhodostoma, the Malayan pit viper: A complex toxin arsenal unraveled. J. Proteom. 2016, 148, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Zainal Abidin, S.A.; Rajadurai, P.; Hoque Chowdhury, M.E.; Ahmad Rusmili, M.R.; Othman, I.; Naidu, R. Proteomic Characterization and Comparison of Malaysian Tropidolaemus wagleri and Cryptelytrops purpureomaculatus Venom Using Shotgun-Proteomics. Toxins 2016, 8, 299. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.-F.; Wang, J.; He, Y.; Qu, Y.-F.; Lin, L.-H.; Ma, X.-M.; Ji, X. Proteomic and biochemical analyses of short-tailed pit viper (Gloydius brevicaudus) venom: Age-related variation and composition–activity correlation. J. Proteom. 2014, 105, 307–322. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.-M.; Yang, Y.-E.; Chen, Y.; Cao, J.; Zhang, C.; Liu, L.-L.; Wang, Z.-Z.; Wang, X.-M.; Wang, Y.-M.; Tsai, I.-H. Transcriptome and proteome of the highly neurotoxic venom of Gloydius intermedius. Toxicon 2015, 107, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Aird, S.D.; Watanabe, Y.; Villar-Briones, A.; Roy, M.C.; Terada, K.; Mikheyev, A.S. Quantitative high-throughput profiling of snake venom gland transcriptomes and proteomes (Ovophis okinavensis and Protobothrops flavoviridis). BMC Genom. 2013, 14, 1–62. [Google Scholar] [CrossRef] [PubMed]
- Aird, S.D. Ophidian envenomation strategies and the role of purines. Toxicon 2002, 40, 335–393. [Google Scholar] [CrossRef]
- Aird, S.D.; Aggarwal, S.; Villar-Briones, A.; Tin, M.M.; Terada, K.; Mikheyev, A.S. Snake venoms are integrated systems, but abundant venom proteins evolve more rapidly. BMC Genom. 2015, 16, 647. [Google Scholar] [CrossRef] [PubMed]
- Villalta, M.; Pla, D.; Yang, S.L.; Sanz, L.; Segura, A.; Vargas, M.; Chen, P.Y.; Herrera, M.; Estrada, R.; Cheng, Y.F.; et al. Snake venomics and antivenomics of Protobothrops mucrosquamatus and Viridovipera stejnegeri from Taiwan: Keys to understand the variable immune response in horses. J. Proteom. 2012, 75, 5628–5645. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Tsai, W.-C.; Ureña-Diaz, J.M.; Sanz, L.; Mora-Obando, D.; Sánchez, E.E.; Fry, B.G.; Gutiérrez, J.M.; Gibbs, H.L.; Sovic, M.G.; et al. Venomics of New World pit vipers: Genus-wide comparisons of venom proteomes across Agkistrodon. J. Proteom. 2014, 96, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Bocian, A.; Urbanik, M.; Hus, K.; Łyskowski, A.; Petrilla, V.; Andrejčáková, Z.; Petrillová, M.; Legáth, J. Proteomic Analyses of Agkistrodon contortrix contortrix Venom Using 2D Electrophoresis and MS Techniques. Toxins 2016, 8, 372. [Google Scholar] [CrossRef] [PubMed]
- Angulo, Y.; Escolano, J.; Lomonte, B.; Gutiérrez, J.M.; Sanz, L.; Calvete, J.J. Snake Venomics of Central American Pitvipers: Clues for Rationalizing the Distinct Envenomation Profiles of Atropoides nummifer and Atropoides picadoi. J. Proteome Res. 2007, 7, 708–719. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Sanz, L.; Sasa, M.; Acevedo, M.E.; Dwyer, Q.; Durban, J.; Pérez, A.; Rodriguez, Y.; Lomonte, B.; Calvete, J.J. Proteomic analysis of venom variability and ontogeny across the arboreal palm-pitvipers (genus Bothriechis). J. Proteom. 2017, 152, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fernández, J.; Lomonte, B.; Sanz, L.; Angulo, Y.; Gutiérrez, J.M.; Calvete, J.J. Snake Venomics of Bothriechis nigroviridis Reveals Extreme Variability among Palm Pitviper Venoms: Different Evolutionary Solutions for the Same Trophic Purpose. J. Proteome Res. 2010, 9, 4234–4241. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Tsai, W.-C.; Bonilla, F.; Solórzano, A.; Solano, G.; Angulo, Y.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics and toxicological profiling of the arboreal pitviper Bothriechis supraciliaris from Costa Rica. Toxicon 2012, 59, 592–599. [Google Scholar] [CrossRef] [PubMed]
- Salazar-Valenzuela, D.; Mora-Obando, D.; Fernández, M.L.; Loaiza-Lange, A.; Gibbs, H.L.; Lomonte, B. Proteomic and toxicological profiling of the venom of Bothrocophias campbelli, a pitviper species from Ecuador and Colombia. Toxicon 2014, 90, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Borges, A.; Segura, Á.; Flores-Díaz, M.; Alape-Girón, A.; Gutiérrez, J.M.; Diez, N.; De Sousa, L.; Kiriakos, D.; Sánchez, E.; et al. Snake venomics and antivenomics of Bothrops colombiensis, a medically important pitviper of the Bothrops atrox-asper complex endemic to Venezuela: Contributing to its taxonomy and snakebite management. J. Proteom. 2009, 72, 227–240. [Google Scholar] [CrossRef] [PubMed]
- Gay, C.; Sanz, L.; Calvete, J.J.; Pla, D. Snake Venomics and Antivenomics of Bothrops diporus, a Medically Important Pitviper in Northeastern Argentina. Toxins 2016, 8, 9. [Google Scholar] [CrossRef] [PubMed]
- Jorge, R.J.B.; Monteiro, H.S.A.; Gonçalves-Machado, L.; Guarnieri, M.C.; Ximenes, R.M.; Borges-Nojosa, D.M.; Luna, K.P.d.O.; Zingali, R.B.; Corrêa-Netto, C.; Gutiérrez, J.M.; et al. Venomics and antivenomics of Bothrops erythromelas from five geographic populations within the Caatinga ecoregion of northeastern Brazil. J. Proteom. 2015, 114, 93–114. [Google Scholar] [CrossRef] [PubMed]
- Valente, R.H.; Guimarães, P.R.; Junqueira, M.; Neves-Ferreira, A.G.C.; Soares, M.R.; Chapeaurouge, A.; Trugilho, M.R.O.; León, I.R.; Rocha, S.L.G.; Oliveira-Carvalho, A.L.; et al. Bothrops insularis venomics: A proteomic analysis supported by transcriptomic-generated sequence data. J. Proteom. 2009, 72, 241–255. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves-Machado, L.; Pla, D.; Sanz, L.; Jorge, R.J.B.; Leitão-De-Araújo, M.; Alves, M.L.M.; Alvares, D.J.; De Miranda, J.; Nowatzki, J.; de Morais-Zani, K.; et al. Combined venomics, venom gland transcriptomics, bioactivities, and antivenomics of two Bothrops jararaca populations from geographic isolated regions within the Brazilian Atlantic rainforest. J. Proteom. 2016, 135, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Sousa, L.F.; Nicolau, C.A.; Peixoto, P.S.; Bernardoni, J.L.; Oliveira, S.S.; Portes-Junior, J.A.; Mourão, R.H.V.; Lima-dos-Santos, I.; Sano-Martins, I.S.; Chalkidis, H.M.; et al. Comparison of Phylogeny, Venom Composition and Neutralization by Antivenom in Diverse Species of Bothrops Complex. PLoS Negl. Trop. Dis. 2013, 7, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, R.S.; Boldrini-França, J.; Fonseca, F.P.P.; de la Torre, P.; Henrique-Silva, F.; Sanz, L.; Calvete, J.J.; Rodriques, V.M. Combined snake venomics and venom gland transcriptome analysis of Bothropoides pauloensis. J. Proteom. 2012, 75, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Alape-Girón, A.; Flores-Díaz, M.; Sanz, L.; Madrigal, M.; Escolano, J.; Sasa, M.; Calvete, J.J. Studies on the venom proteome of Bothrops asper: Perspectives and applications. Toxicon 2009, 54, 938–948. [Google Scholar] [CrossRef] [PubMed]
- Sousa, L.F.; Portes-Junior, J.A.; Nicolau, C.A.; Bernardoni, J.L.; Nishiyama, M.Y., Jr.; Amazonas, D.R.; Freitas-de-Sousa, L.A.; Mourão, R.H.V.; Chalkidis, H.M.; Valente, R.H.; et al. Functional proteomic analyses of Bothrops atrox venom reveals phenotypes associated with habitat variation in the Amazon. J. Proteom. 2017, 159, 32–46. [Google Scholar] [CrossRef] [PubMed]
- Núñez, V.; Cid, P.; Sanz, L.; De La Torre, P.; Angulo, Y.; Lomonte, B.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics and antivenomics of Bothrops atrox venoms from Colombia and the Amazon regions of Brazil, Perú and Ecuador suggest the occurrence of geographic variation of venom phenotype by a trend towards paedomorphism. J. Proteom. 2009, 73, 57–78. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Sanz, L.; Pérez, A.; Borges, A.; Vargas, A.M.; Lomonte, B.; Angulo, Y.; Gutiérrez, J.M.; Chalkidis, H.M.; Mourão, R.H.V.; et al. Snake population venomics and antivenomics of Bothrops atrox: Paedomorphism along its transamazonian dispersal and implications of geographic venom variability on snakebite management. J. Proteom. 2011, 74, 510–527. [Google Scholar] [CrossRef] [PubMed]
- Kohlhoff, M.; Borges, M.H.; Yarleque, A.; Cabezas, C.; Richardson, M.; Sanchez, E.F. Exploring the proteomes of the venoms of the Peruvian pit vipers Bothrops atrox, B. barnetti and B. pictus. J. Proteom. 2012, 75, 2181–2195. [Google Scholar] [CrossRef] [PubMed]
- Mora-Obando, D.; Guerrero-Vargas, J.A.; Prieto-Sánchez, R.; Beltrán, J.; Rucavado, A.; Sasa, M.; Gutiérrez, J.M.; Ayerbe, S.; Lomonte, B. Proteomic and functional profiling of the venom of Bothrops ayerbei from Cauca, Colombia, reveals striking interspecific variation with Bothrops asper venom. J. Proteom. 2014, 96, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.M.; Sanz, L.; Escolano, J.; Fernández, J.; Lomonte, B.; Angulo, Y.; Rucavado, A.; Warrell, D.A.; Calvete, J.J. Snake Venomics of the Lesser Antillean Pit Vipers Bothrops caribbaeus and Bothrops lanceolatus: Correlation with Toxicological Activities and Immunoreactivity of a Heterologous Antivenom. J. Proteome Res. 2008, 7, 4396–4408. [Google Scholar] [CrossRef] [PubMed]
- Bernardes, C.P.; Menaldo, D.L.; Camacho, E.; Rosa, J.C.; Escalante, T.; Rucavado, A.; Lomonte, B.; Gutiérrez, J.M.; Sampaio, S.V. Proteomic analysis of Bothrops pirajai snake venom and characterization of BpirMP, a new P-I metalloproteinase. J. Proteom. 2013, 80, 250–267. [Google Scholar] [CrossRef] [PubMed]
- Fernández Culma, M.; Andrés Pereañez, J.; Núñez Rangel, V.; Lomonte, B. Snake venomics of Bothrops punctatus, a semiarboreal pitviper species from Antioquia, Colombia. PeerJ 2014, 2, e246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomonte, B.; Rey-Suárez, P.; Tsai, W.-C.; Angulo, Y.; Sasa, M.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics of the pit vipers Porthidium nasutum, Porthidium ophryomegas, and Cerrophidion godmani from Costa Rica: Toxicological and taxonomical insights. J. Proteom. 2012, 75, 1675–1689. [Google Scholar] [CrossRef] [PubMed]
- Lomonte, B.; Fernández, J.; Sanz, L.; Angulo, Y.; Sasa, M.; Gutiérrez, J.M.; Calvete, J.J. Venomous snakes of Costa Rica: Biological and medical implications of their venom proteomic profiles analyzed through the strategy of snake venomics. J. Proteom. 2014, 105, 323–339. [Google Scholar] [CrossRef] [PubMed]
- Rokyta, D.R.; Lemmon, A.R.; Margres, M.J.; Aronow, K. The venom-gland transcriptome of the eastern diamondback rattlesnake (Crotalus adamenteous). BMC Genom. 2012, 13, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Fasoli, E.; Sanz, L.; Boschetti, E.; Righetti, P.G. Exploring the Venom Proteome of the Western Diamondback Rattlesnake, Crotalus atrox, via Snake Venomics and Combinatorial Peptide Ligand Library Approaches. J. Proteome Res. 2009, 8, 3055–3067. [Google Scholar] [CrossRef] [PubMed]
- Segura, Á.; Herrera, M.; Reta Mares, F.; Jaime, C.; Sánchez, A.; Vargas, M.; Villalta, M.; Gómez, A.; Gutiérrez, J.M.; León, G. Proteomic, toxicological and immunogenic characterization of Mexican west-coast rattlesnake (Crotalus basiliscus) venom and its immunological relatedness with the venom of Central American rattlesnake (Crotalus simus). J. Proteom. 2017, 158, 62–72. [Google Scholar] [CrossRef] [PubMed]
- Durban, J.; Sanz, L.; Trevisan-Silva, D.; Neri-Castro, E.; Alagón, A.; Calvete, J.J. Integrated Venomics and Venom Gland Transcriptome Analysis of Juvenile and Adult Mexican Rattlesnakes Crotalus simus, C. tzabcan, and C. culminatus Revealed miRNA-modulated Ontogenetic Shifts. J. Proteome Res. 2017, 16, 3370–3390. [Google Scholar] [CrossRef] [PubMed]
- Boldrini-França, J.; Corrêa-Netto, C.; Silva, M.M.S.; Rodrigues, R.S.; De La Torre, P.; Pérez, A.; Soares, A.M.; Zingali, R.B.; Nogueira, R.A.; Rodrigues, V.M.; et al. Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: Assessment of geographic variation and its implication on snakebite management. J. Proteom. 2010, 73, 1758–1776. [Google Scholar] [CrossRef] [PubMed]
- Boldrini-França, J.; Rodrigues, R.S.; Fonseca, F.P.P.; Menaldo, D.L.; Ferreira, F.B.; Henrique-Silva, F.; Soares, A.M.; Hamaguchi, A.; Rodrigues, V.M.; Otaviano, A.R.; et al. Crotalus durissus collilineatus venom gland transcriptome: Analysis of gene expression profile. Biochimie 2009, 91, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Georgieva, D.; Öhler, M.; Seifert, J.; Bergen, M.V.; Arni, R.K.; Genov, N.; Betzel, C. Snake Venomic of Crotalus durissus terrificus—Correlation with Pharmacological Activities. J. Proteome Res. 2010, 9, 2302–2316. [Google Scholar] [CrossRef] [PubMed]
- Rokyta, D.R.; Wray, K.P.; Margres, M.J. The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics. BMC Genom. 2013, 14, 394. [Google Scholar] [CrossRef] [PubMed]
- Castro, E.N.; Lomonte, B.; del Carmen Gutiérrez, M.; Alagón, A.; Gutiérrez, J.M. Intraspecies variation in the venom of the rattlesnake Crotalus simus from Mexico: Different expression of crotoxin results in highly variable toxicity in the venoms of three subspecies. J. Proteom. 2013, 87, 103–121. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Pérez, A.; Lomonte, B.; Sánchez, E.E.; Sanz, L. Snake Venomics of Crotalus tigris: The Minimalist Toxin Arsenal of the Deadliest Neartic Rattlesnake Venom. Evolutionary Clues for Generating a Pan-Specific Antivenom against Crotalid Type II Venoms. J. Proteome Res. 2012, 11, 1382–1390. [Google Scholar] [CrossRef] [PubMed]
- Saviola, A.J.; Pla, D.; Sanz, L.; Castoe, T.A.; Calvete, J.J.; Mackessy, S.P. Comparative venomics of the Prairie Rattlesnake (Crotalus viridis viridis) from Colorado: Identification of a novel pattern of ontogenetic changes in venom composition and assessment of the immunoreactivity of the commercial antivenom CroFab®. J. Proteom. 2015, 121, 28–43. [Google Scholar] [CrossRef] [PubMed]
- Sanz, L.; Gibbs, H.L.; Mackessy, S.P.; Calvete, J.J. Venom Proteomes of Closely Related Sistrurus Rattlesnakes with Divergent Diets. J. Proteome Res. 2006, 5, 2098–2112. [Google Scholar] [CrossRef] [PubMed]
- Madrigal, M.; Sanz, L.; Flores-Díaz, M.; Sasa, M.; Núñez, V.; Alape-Girón, A.; Calvete, J.J. Snake venomics across genus Lachesis. Ontogenetic changes in the venom composition of Lachesis stenophrys and comparative proteomics of the venoms of adult Lachesis melanocephala and Lachesis acrochorda. J. Proteom. 2012, 77, 280–297. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Sanz, L.; Molina-Sánchez, P.; Zorita, V.; Madrigal, M.; Flores-Díaz, M.; Alape-Girón, A.; Núñez, V.; Andrés, V.; Gutiérrez, J.M.; et al. Snake venomics of Lachesis muta rhombeata and genus-wide antivenomics assessment of the paraspecific immunoreactivity of two antivenoms evidence the high compositional and immunological conservation across Lachesis. J. Proteom. 2013, 89, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Charris, E.; Montealegre-Sanchez, L.; Solano-Redondo, L.; Mora-Obando, D.; Camacho, E.; Castro-Herrera, F.; Fierro-Pérez, L.; Lomonte, B. Proteomic and functional analyses of the venom of Porthidium lansbergii lansbergii (Lansberg’s hognose viper) from the Atlantic Department of Colombia. J. Proteom. 2015, 114, 287–299. [Google Scholar] [CrossRef] [PubMed]
- Tashima, A.K.; Sanz, L.; Camargo, A.C.M.; Serrano, S.M.T.; Calvete, J.J. Snake venomics of the Brazilian pitvipers Bothrops cotiara and Bothrops fonsecai. Identification of taxonomy markers. J. Proteom. 2008, 71, 473–485. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G. Venomous Reptiles and their Toxins; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- Sanz, L.; Escolano, J.; Ferritti, M.; Biscoglio, M.J.; Rivera, E.; Crescenti, E.J.; Angulo, Y.; Lomonte, B.; Gutiérrez, J.M.; Calvete, J.J. Snake venomics of the South and Central American bushmasters. Comparison of the toxin composition of Lachesis muta gathered from proteomic versus transcriptomic analysis. J. Proteom. 2008, 71, 46–60. [Google Scholar] [CrossRef] [PubMed]
- Doley, R.; Pahari, S.; Mackessy, S.P.; Kini, R.M. Accelerated exchange of exon segments in Viperid three-finger toxin genes (Sistrurus catenatus edwardsii; Desert massasauga). BMC Evol. Biol. 2008, 8, 196. [Google Scholar] [CrossRef] [PubMed]
- Shelke, R.R.J.; Sathish, S.; Gowda, T.V. Isolation and characterization of a novel postsynaptic/cytotoxic neurotoxin from Daboia russelli russelli venom. J. Pept. Res. 2002, 59, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Skejic, J.; Hodgson, W.C. Population divergence in venom bioactivities of elapid snake Pseudonaja textilis: Role of procoagulant proteins in rapid rodent prey incapacitation. PLoS ONE 2013, 8, e63988. [Google Scholar] [CrossRef] [PubMed]
- Isbister, G.K. Procoagulant snake toxins: Laboratory studies, Diagnosis, and understanding snakebite coagulopathy. Semin. Thromb. Hemost. 2009, 35, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Strickland, J.L.; Carter, S.; Kraus, F.; Parkinson, C.L. Snake evolution in Melanesia: Origin of the Hydrophiinae (Serpentes, Elapidae), and the evolutionary history of the enigmatic New Guinean elapid Toxicocalamus. Zool. J. Linn. Soc. 2016, 178, 663–678. [Google Scholar] [CrossRef]
- Sanders, K.L.; Lee, M.S.Y.; Leys, R.; Foster, R.; Scott Keogh, J. Molecular phylogeny and divergence dates for Australasian elapids and sea snakes (hydrophiinae): Evidence from seven genes for rapid evolutionary radiations. J. Evol. Biol. 2008, 21, 682–695. [Google Scholar] [CrossRef] [PubMed]
- Scanlon, J.D.; Lee, M.S.Y. Phylogeny of Australasian venomous snakes (Colubroidea, Elapidae, Hydrophiinae) based on phenotypic and molecular evidence. Zool. Scr. 2004, 33, 335–366. [Google Scholar] [CrossRef]
- Lukoschek, V.; Keogh, J.S. Molecular phylogeny of sea snakes reveals a rapidly diverged adaptive radiation. Biol. J. Linn. Soc. 2006, 89, 523–539. [Google Scholar] [CrossRef]
- Sanders, K.L.; Lee, M.S.Y.; Mumpuni; Bertozzi, T.; Rasmussen, A.R. Multilocus phylogeny and recent rapid radiation of the viviparous sea snakes (Elapidae: Hydrophiinae). Mol. Phylogenet. Evol. 2013, 66, 575–591. [Google Scholar] [CrossRef] [PubMed]
- Maduwage, K.; O’Leary, M.A.; Isbister, G.K. Diagnosis of snake envenomation using a simple phospholipase A2 assay. Sci. Rep. 2014, 4, 4827. [Google Scholar] [CrossRef] [PubMed]
- Wagstaff, S.C.; Laing, G.D.; David, R.; Theakston, G.; Papaspyridis, C.; Harrison, R.A.; Theakston, R.D.G. Bioinformatics and multiepitope DNA immunization to design rational snake antivenom. PLoS Med. 2006, 3, e184. [Google Scholar] [CrossRef] [PubMed]
- Silva, A.; Hodgson, W.C.; Isbister, G.K. Cross-Neutralisation of In Vitro Neurotoxicity of Asian and Australian Snake Neurotoxins and Venoms by Different Antivenoms. Toxins 2016, 8, 302. [Google Scholar] [CrossRef] [PubMed]
- Isbister, G.K.; Maduwage, K.; Page, C.B. Antivenom cross neutralisation in a suspected Asian pit viper envenoming causing severe coagulopathy. Toxicon 2014, 90, 286–290. [Google Scholar] [CrossRef] [PubMed]
- Torres, A.M.; Wong, H.Y.; Desai, M.; Moochhala, S.; Kuchel, P.W.; Kini, R.M. Identification of a Novel Family of Proteins in Snake Venoms. Purification And Structural Characterization Of Nawaprin From Naja Nigricollis Snake Venom. J. Biol. Chem. 2003, 278, 40097–40104. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G. From genome to “venome”: Molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res. 2005, 15, 403–420. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Huang, S.; Guo, P.; Colli, G.R.; Nieto Montes de Oca, A.; Vitt, L.J.; Pyron, R.A.; Burbrink, F.T. Understanding the formation of ancient intertropical disjunct distributions using Asian and Neotropical hinged-teeth snakes (Sibynophis and Scaphiodontophis: Serpentes: Colubridae). Mol. Phylogenet. Evol. 2013, 66, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Esselstyn, J.A.; Brown, R.M. The role of repeated sea-level fluctuations in the generation of shrew (Soricidae: Crocidura) diversity in the Philippine Archipelago. Mol. Phylogenet. Evol. 2009, 53, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Catullo, R.A.; Scott Keogh, J. Aridification drove repeated episodes of diversification between Australian biomes: Evidence from a multi-locus phylogeny of Australian toadlets (Uperoleia: Myobatrachidae). Mol. Phylogenet. Evol. 2014, 79, 106–117. [Google Scholar] [CrossRef] [PubMed]
- Menegon, M.; Loader, S.P.; Marsden, S.J.; Branch, W.R.; Davenport, T.R.B.; Ursenbacher, S. The genus Atheris (Serpentes: Viperidae) in East Africa: Phylogeny and the role of rifting and climate in shaping the current pattern of species diversity. Mol. Phylogenet. Evol. 2014, 79, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Keogh, J.S.; Scott, I.A.W.; Hayes, C. Rapid And Repeated Origin of Insular Gigantism and Dwarfism in Australian Tiger Snakes. Evolution 2005, 59, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Williams, V.; White, J. Variation in venom constituents within a single isolated population of peninsula tiger snake (Notechis ater niger). Toxicon 1987, 25, 1240–1243. [Google Scholar] [CrossRef]
- Li, M.; Fry, B.G.; Kini, R. Eggs-Only Diet: Its Implications for the Toxin Profile Changes and Ecology of the Marbled Sea Snake (Aipysurus eydouxii). J. Mol. Evol. 2005, 60, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Shine, R. Ecology of Highly Venomous Snakes: The Australian Genus Oxyuranus (Elapidae). J. Herpetol. 1983, 17, 60–69. [Google Scholar] [CrossRef]
- Harvey, A.L. Twenty years of dendrotoxins. Toxicon 2001, 39, 15–26. [Google Scholar] [CrossRef]
- Kornhauser, R.; Hart, A.J.; Reeve, S.; Smith, A.I.; Fry, B.G.; Hodgson, W.C. Variations in the pharmacological profile of post-synaptic neurotoxins isolated from the venoms of the Papuan (Oxyuranus scutellatus canni) and coastal (Oxyuranus scutellatus scutellatus) taipans. NeuroToxicol. 2010, 31, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Kuruppu, S.; Reeve, S.; Banerjee, Y.; Kini, R.M.; Smith, A.I.; Hodgson, W.C. Isolation and Pharmacological Characterization of Cannitoxin, a Presynaptic Neurotoxin from the Venom of the Papuan Taipan (Oxyuranus scutellatus canni). J. Pharmacol. Exp. Ther. 2005, 315, 1196–1202. [Google Scholar] [CrossRef] [PubMed]
- Petras, D.; Heiss, P.; Harrison, R.A.; Süssmuth, R.D.; Calvete, J.J. Top-down venomics of the East African green mamba, Dendroaspis angusticeps, and the black mamba, Dendroaspis polylepis, highlight the complexity of their toxin arsenals. J. Proteom. 2016, 146, 148–164. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, H.L.; Sanz, L.; Sovic, M.G.; Calvete, J.J. Phylogeny-Based Comparative Analysis of Venom Proteome Variation in a Clade of Rattlesnakes (Sistrurus sp.). PLoS ONE 2013, 8, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jorge da Silva, N., Jr.; Aird, S.D. Prey specificity, comparative lethality and compositional differences of coral snake venoms. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2001, 128, 425–456. [Google Scholar] [CrossRef]
- Gibbs, H.L.; Mackessy, S.P. Functional basis of a molecular adaptation: Prey-specific toxic effects of venom from Sistrurus rattlesnakes. Toxicon 2009, 53, 672–679. [Google Scholar] [CrossRef] [PubMed]
- Starkov, V.G.; Osipov, A.V.; Utkin, Y.N. Toxicity of venoms from vipers of Pelias group to crickets Gryllus assimilis and its relation to snake entomophagy. Toxicon 2007, 49, 995–1001. [Google Scholar] [CrossRef] [PubMed]
- Barlow, A.; Pook, C.E.; Harrison, R.A.; Wüster, W. Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc. R. Soc. Lond. B Biol. Sci. 2009, 276, 2443–2449. [Google Scholar] [CrossRef] [PubMed]
- Daltry, J.C.; Wuster, W.; Thorpe, R.S. Diet and snake venom evolution. Nature 1996, 379, 537. [Google Scholar] [CrossRef] [PubMed]
- Poran, N.S.; Coss, R.G.; Benjamini, E. Resistance of California ground squirrels (Spermophilus beecheyi) to the venom of the northern Pacific rattlesnake (Crotalus viridis oreganus): A study of adaptive variation. Toxicon 1987, 25, 767–777. [Google Scholar] [CrossRef]
- Heatwole, H.; Poran, N.S. Resistances of Sympatric and Allopatric Eels to Sea Snake Venoms. Copeia 1995, 1995, 136–147. [Google Scholar] [CrossRef]
- Dowell, N.L.; Giorgianni, M.W.; Kassner, V.A.; Selegue, J.E.; Sanchez, E.E.; Carroll, S.B. The Deep Origin and Recent Loss of Venom Toxin Genes in Rattlesnakes. Curr. Biol. 2016, 26, 2434–2445. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, A.D.; Swain, M.T.; Hegarty, M.J.; Logan, D.W.; Mulley, J.F. Restriction and Recruitment—Gene Duplication and the Origin and Evolution of Snake Venom Toxins. Genome Biol. Evol. 2014, 6, 2088–2095. [Google Scholar] [CrossRef] [PubMed]
- Casewell, N.R.; Wagstaff, S.C.; Harrison, R.A.; Renjifo, C.; Wüster, W. Domain Loss Facilitates Accelerated Evolution and Neofunctionalization of Duplicate Snake Venom Metalloproteinase Toxin Genes. Mol. Biol. Evol. 2011, 28, 2637–2649. [Google Scholar] [CrossRef] [PubMed]
- Lynch, V.J. Inventing an arsenal: Adaptive evolution and neofunctionalization of snake venom phospholipase A2 genes. BMC Evol. Biol. 2007, 7, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Doley, R.; Mackessy, S.P.; Kini, R. Role of accelerated segment switch in exons to alter targeting (ASSET) in the molecular evolution of snake venom proteins. BMC Evol. Biol. 2009, 9, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, H.L.; Sanz, L.; Chiucchi, J.E.; Farrell, T.M.; Calvete, J.J. Proteomic analysis of ontogenetic and diet-related changes in venom composition of juvenile and adult Dusky Pigmy rattlesnakes (Sistrurus miliarius barbouri). J. Proteom. 2011, 74, 2169–2179. [Google Scholar] [CrossRef] [PubMed]
- Andrade, D.V.; Abe, A.S. Relationship of Venom Ontogeny and Diet in Bothrops. Herpetologica 1999, 55, 200–204. [Google Scholar]
- Saldarriaga, M.M.; Otero, R.; Núñez, V.; Toro, M.F.; Díaz, A.; Gutiérrez, J.M. Ontogenetic variability of Bothrops atrox, and Bothrops asper snake venoms from Colombia. Toxicon 2003, 42, 405–411. [Google Scholar] [CrossRef]
- Mackessy, S.P. Venom ontogeny in the Pacific rattlesnakes Crotalus viridis helleri and Crotalus viridis oreganus. In Copeia; American Society of Ichthyologists and Herpetologists (ASIH): Lawrence, KS, USA, 1988. [Google Scholar] [CrossRef]
- Zelanis, A.; Tashima, A.K.; Rocha, M.M.T.; Furtado, M.F.; Camargo, A.C.M.; Ho, P.L.; Serrano, S.M.T. Analysis of the Ontogenetic Variation in the Venom Proteome/Peptidome of Bothrops jararaca Reveals Different Strategies to Deal with Prey. J. Proteome Res. 2010, 9, 2278–2291. [Google Scholar] [CrossRef] [PubMed]
- Tan, N.H.; Armugam, A.; Mirtschin, P.J. The biological properties of venoms from juvenile and adult taipan (Oxyuranus scutellatus) snakes. Comp. Biochem. Physiol. B 1992, 103, 585–588. [Google Scholar] [CrossRef]
- Tan, N.H.; Ponnudurai, G.; Mirtschin, P.J. A comparative study of the biological properties of venoms from juvenile and adult inland taipan (Oxyuranus microlepidotus) snake venoms. Toxicon 1993, 31, 363–367. [Google Scholar] [CrossRef]
- Castoe, T.A.; Daza, J.M.; Smith, E.N.; Sasa, M.M.; Kuch, U.; Campbell, J.A.; Chippindale, P.T.; Parkinson, C.L. Comparative phylogeography of pitvipers suggests a consensus of ancient Middle American highland biogeography. J. Biogeogr. 2009, 36, 88–103. [Google Scholar] [CrossRef]
- Daza, J.M.; Castoe, T.A.; Parkinson, C.L. Using regional comparative phylogeographic data from snake lineages to infer historical processes in Middle America. Ecography 2010, 33, 343–354. [Google Scholar] [CrossRef]
- Wittenberg, R.D.; Jadin, R.C.; Fenwick, A.M.; Gutberlet, R.L. Recovering the evolutionary history of Africa’s most diverse viper genus: Morphological and molecular phylogeny of Bitis (Reptilia: Squamata: Viperidae). Org. Divers. Evol. 2015, 15, 115–125. [Google Scholar] [CrossRef]
- Pyron, R.A.; Burbrink, F.T.; Colli, G.R.; de Oca, A.N.M.; Vitt, L.J.; Kuczynski, C.A.; Wiens, J.J. The phylogeny of advanced snakes (Colubroidea), with discovery of a new subfamily and comparison of support methods for likelihood trees. Mol. Phylogenet. Evol. 2011, 58, 329–342. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.S.Y.; Sanders, K.L.; King, B.; Palci, A. Diversification rates and phenotypic evolution in venomous snakes (Elapidae). R. Soc. Open Sci. 2016, 3, 150277. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, A.; Thorpe, R.S. A phylogeny of four mitochondrial gene regions suggests a revised taxonomy for Asian pitvipers (Trimeresurus and Ovophis). Mol. Phylogenet. Evol. 2004, 32, 83–100. [Google Scholar] [CrossRef] [PubMed]




| SPECIES | PLA2 | SVSP | SVMP | LAAO | 3FT | KUN | CRiSP | NP | %WV | 3FT + PLA2 | REF |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Austrelaps labialis | 33 | 3 | 45 | 9 | 8 | 98 | 78 | [3] | |||
| Drysdalia coronoides | 86.4 | 9.2 | 2.8 | 98.4 | 86.4 | [4] | |||||
| Micropechis ikaheka | 80 | <0.1 | 7.6 | 0.4 | 9.2 | 0.7 | 1.8 | 99.8 | 89.2 | [5] | |
| Notechis scutatus | 74.5 | 5.9 | 5.6 | 6.9 | 0.3 | 2 | 93.2 | 80.1 | [6] | ||
| Oxyuranus scutellatus | 68–80 | <5 | 5–9 | 0–9 | <10 | <1 | 1 | >90 | 68–89 | [7] | |
| Pseudechis papuanus | 90.2 | 2.8 | 1.6 | 3.1 | 2.3 | 100 | 93.3 | [8] | |||
| Toxicocalamus longissimus | 6.5 | 1.4 | 92.1 | 100 | 98.6 | [9] | |||||
| Aipysurus laevis | 71.2 | 25.3 | 2.5 | 99 | 96.5 | [10] | |||||
| Hydrophis cyanocinctus | 18.9 | 81.1 | 100 | 100 | [9] | ||||||
| H. platurus | 32.9 | 0.9 | 49.9 | 9.1 | 92.8 | 82.8 | [11] | ||||
| H. schistosus | 27.5 | 0.5 | 0.2 | 70.5 | 1.3 | 100 | 98 | [12] | |||
| Laticauda colubrina | 33.3 | 66.1 | 0.05 | 99.45 | 99.4 | [13] | |||||
| Bungarus caeruleus (Sri Lanka) | 64.5 | 1.3 | 19 | 4.4 | 5.5 | 94.7 | 83.5 | [14] | |||
| B. candidus Malaya | 25.2 | 3.9 | 4.9 | 5.8 | 30.1 | 12.6 | 3.9 | 1 | 86.4 | 55.6 | [15] |
| B. fasciatus Vietnam | 66.8 | 3.5 | 7 | 1.3 | 1.8 | 0.4 | 80.8 | 68.1 | [16] | ||
| B.fasciatus Malaya | 44.2 | 5.8 | 4.7 | 5.8 | 17.4 | 9.3 | 1.2 | 88.4 | 61.6 | [15] | |
| Dendroaspis angusticeps | 6.7 | 69.2 | 16.3 | 2 | 94.2 | 69.2 | [17] | ||||
| D. polylepis | 3.2 | 31 | 61.1 | 2.9 | 95.3 | 31 | [18] | ||||
| Naja haje | 4 | 9 | 1 | 60 | 1.9 | 10 | 85.9 | 64 | [19] | ||
| N. melanoleuca | 12.9 | 9.7 | 57.1 | 3.8 | 7.6 | 91.1 | 70 | [20] | |||
| N. katiensis | 29 | 3.3 | 67.1 | 0.2 | 99.6 | 96.1 | [21] | ||||
| N. mossambica | 27.1 | 2.6 | 69.3 | 99 | 96.4 | [21] | |||||
| N. nigricollis | 21.9 | 2.4 | 73.2 | 0.2 | 97.7 | 95.1 | [21] | ||||
| N. nubiae | 26.4 | 2.6 | 70.9 | 99.9 | 97.3 | [21] | |||||
| N. pallida | 30.1 | 1.6 | 67.7 | 99.4 | 97.8 | [21] | |||||
| N. atra China | 12.2 | 1.6 | 84.3 | 1.8 | 99.9 | 96.5 | [22] | ||||
| Naja atra Taiwan | 14–17 | 2–2.6 | 0.2 | 76–80 | 2.2–2.4 | >93 | 90–97 | [23] | |||
| N. kaouthia China | 26.9 | 1.1 | 56.6 | 5.4 | 90 | 83.5 | [24] | ||||
| Naja kaouthia Malaya | 23.5 | 3.3 | 1.1 | 63.7 | 0.5 | 4.3 | 96.4 | 87.2 | [25] | ||
| Naja kaouthia Thailand | 12.2 | 2.6 | 1 | 78.3 | 2.3 | 0.2 | 96.4 | 90.5 | [25] | ||
| Naja kaouthia Vietnam | 17.4 | 1.6 | 0.5 | 76.4 | 0.8 | 96.7 | 93.8 | [25] | |||
| N. naja Eastern India | 11.4 | 0.3 | 1 | 0.8 | 63.8 | 0.4 | 2.1 | 2 | 79.8 | 75.2 | [26] |
| Naja naja North-west India | 21.4 | 0.9 | 74 | 2.5 | 98.8 | 95.4 | [27] | ||||
| Naja naja Sri Lanka | 14 | 0.9 | 80.5 | 3.7 | 99.1 | 94.5 | [27] | ||||
| N. sputatrix | 31.2 | 0.4 | 1.3 | 0.1 | 64.2 | 0.2 | 97 | 95.4 | [28] | ||
| Ophiophagus hannah | 2.8 | 11.9 | 0.5 | 64.5 | 3.3 | 6.5 | 0.2 | 89.5 | 67.3 | [29] | |
| Micrurus alleni | 10.9 | 1.2 | 3 | 77.3 | 92.4 | 88.2 | [30] | ||||
| M. altirostris | 13.7 | 0.9 | 1.2 | 79.5 | 2.1 | 0.1 | 97.5 | 93.2 | [31] | ||
| M. clarki | 36.5 | 1 | 1.6 | 3.8 | 48.2 | 0.9 | 92 | 84.7 | [32] | ||
| M. corallinus | 11.9 | 0.8 | 2.9 | 2.3 | 81.7 | 99.6 | 93.6 | [31] | |||
| M. dumerelii | 52 | 1.9 | 1.8 | 3.1 | 28.1 | 9 | 95.9 | 80.1 | [33] | ||
| M. fulvius | 64.9 | 2.9 | 25.1 | 2.2 | 95.1 | 90 | [34] | ||||
| M. mipartitus | 29 | 1.3 | 1.6 | 4 | 61.1 | 1.9 | 98.9 | 90.1 | [35] | ||
| M. mosquitensis | 55.6 | 0.5 | 2.6 | 2.8 | 22.5 | 9.8 | 93.8 | 78.1 | [30] | ||
| M. multifasciatus | 8.2 | 3.6 | 3.2 | 83 | 1.9 | 99.9 | 91.2 | [35] | |||
| M. nigrocinctus | 48 | 0.7 | 4.3 | 2.3 | 38 | 93.3 | 86 | [36] | |||
| M. tschudii | 4.1 | 0.7 | 95.2 | 1.6 | 100 | 99.3 | [37] |
| SPECIES | PLA2 | SVSP | SVMP | LAAO | CRiSP | CTL/SNACLEC | DIS | NP | KUN | VEGF | CYS | %WV | REF |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bitis arietans | 4.3 | 19.5 | 38.5 | 13.2 | 17.8 | 4.2 | 1.7 | 99.2 | [38] | ||||
| B.caudalis | 59.8 | 15.1 | 11.5 | 1.7 | 1.2 | 4.9 | 2.3 | 3.2 | 99.7 | [38] | |||
| B.gabonica | 11.4 | 26.4 | 22.9 | 1.3 | 2 | 14.3 | 3.4 | 2.8 | 3 | 1 | 9.8 | 98.3 | [38] |
| B.nasicornis | 20.1 | 21.9 | 40.9 | 3.2 | 1.3 | 4.2 | 3.5 | 4.2 | 99.3 | [38] | |||
| B.rhinoceros | 4.8 | 23.9 | 30.8 | 2.2 | 1.2 | 14.1 | 8.5 | 0.3 | 7.5 | 5.3 | 98.6 | [38] | |
| Cerastes cerastes (Morocco) | 19.1 | 6.9 | 63.1 | 0.7 | 1.7 | 8.5 | 100 | [39] | |||||
| C. cerastes (Tunisia) | 16.6 | 13.2 | 55.9 | 6.2 | 3.2 | 4.9 | 100 | [39] | |||||
| Daboia russelii (Pakistan) | 32.8 | 3.2 | 21.8 | 0.6 | 2.6 | 6.4 | 0.4 | 28.4 | 1.5 | 97.7 | [40] | ||
| D. russelii (West India) | 32.5 | 8 | 24.8 | 0.3 | 6.8 | 1.8 | 4.9 | 12.5 | 1.8 | 93.4 | [41] | ||
| D. russelii (Sri Lanka) | 35 | 16 | 6.9 | 5.2 | 2 | 22.4 | 4.6 | 92.1 | [42] | ||||
| Echis carinatus sochureki | 7.97 | 4.58 | 56.57 | 1.19 | 1.99 | 16.53 | 7.7 | 0.4 | 97 | [43] | |||
| E. coloratus | 5.7 | 3.58 | 61.41 | 3.91 | 5.69 | 9.45 | 5.8 | 0.32 | 96 | [43] | |||
| E. ocellatus | 8.5 | 1.71 | 72.43 | 1.36 | 0.34 | 6.46 | 2.72 | 93.5 | [43] | ||||
| E. pyramidium leakeyi | 21.57 | 1.42 | 48.94 | 2.83 | 24.26 | 0.28 | 99.3 | [43] | |||||
| Macrovipera lebetina (Tunisia) | 5 | 5.5 | 63.1 | 3.2 | 15.1 | 3.1 | 3.3 | 98.3 | [44] | ||||
| M. l. obtusa | 14.6 | 14.9 | 32.1 | 1.7 | 2.6 | 14.8 | 11.3 | 5.3 | 97.3 | [45] | |||
| M. mauritanica | 5.5 | 8.3 | 45.4 | 8.1 | 13.8 | 4.5 | 2.5 | 4.9 | 93 | [44] | |||
| Vipera anatolica | 8.1 | 1.6 | 41.5 | 15.9 | 1.1 | 2 | 0.3 | 70.5 * | [46] | ||||
| V. berus | 10 | 31 | 19 | 2 | 8 | 2 | 1 | 11 | 84 * | [47] | |||
| V. kaznakovi | 41 | 11 | 16 | 4 | 10 | 12 | 0.53 | 4 | 94.5 | [48] | |||
| V. nikolskii | 65 | 19 | 0.66 | 0.08 | 0.66 | 4 | 8 | 97.4 | [48] | ||||
| V. orlovi | 24 | 24 | 15 | 5 | 12 | 11 | 0.56 | 0.15 | 4 | 91.7 | [48] | ||
| V. raddei | 23.8 | 8.4 | 31.6 | 0.2 | 7.4 | 9.6 | 9.7 | 6 | 0.9 | 2.4 | 100 | [45] | |
| V. renardii | 44 | 8 | 12 | 4 | 8 | 3 | 13 | 0.8 | 3 | 95.8 | [48] |
| SPECIES | PLA2 | SVSP | SVMP | LAAO | CRiSP | CTL/SNACLEC | DIS | NP | DEF | MPi | %WV | REF |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Calloselasma rhodostoma | 4.4 | 14.9 | 41.2 | 7 | 2.5 | 26.3 | 96.3 | [50] | ||||
| Cryptelytrops purpureomaculatus | 8 | 12 | 35 | 10 | 6 | 19 | 2 | 92 | [51] | |||
| Gloydius brevicaudus | 25 | 3.7 | 64.4 | 0.9 | 1.1 | 0.2 | 4.6 | 99.9 | [52] | |||
| G. intermedius | 9.9 | 36.2 | 2.6 | 13.1 | 6.2 | 0.8 | 25.3 | 94.1 | [53] | |||
| Ovophis okinavensis | 0.65 | 93.1 | 4.2 | 0.62 | 0.47 | 99 | [54] | |||||
| Protobothrops elegans | 77.1 | 10.4 | 8 | 0.5 | 0.1 | 0.2 | 96.3 | [55] | ||||
| Protobothrops flavoviridis | 55.5 | 11.8 | 17.3 | 3.1 | 2 | 0.9 | 2.6 | 93.2 | [56] | |||
| P.mucrosquamatus | 22.5 | 10.4 | 43 | 2 | 0.8 | 3.9 | 0.8 | 3.6 | 87 | [57] | ||
| Viridovipera stejnegeri | 24.5 | 11 | 43.1 | 3.3 | 6 | 1.5 | 2.2 | 1.2 | 92.8 | [57] | ||
| Agkistrodon bilineatus (3 subsp) | 34.3–42 | 7.6–16.9 | 24.5–30.8 | 2.6–4.9 | 0–5.6 | 0.4–1.4 | 2.2–3.1 | 4.6–8.7 | 76.7+ | [58] | ||
| A. c. contortrix | 50.7 | 5.85 | 25 | 4 | 2 | 0.8 | 88.35 | [59] | ||||
| A. piscivorus (3 subsp) | 33.6–46 | 10.1–13.9 | 21–33.1 | 0.8–4.5 | 2–3.5 | 0.8–3.2 | 2.2–4.9 | 5.7–5.9 | 76.2 | [58] | ||
| Atropoides nummifer | 36.5 | 22 | 18.2 | 9.1 | 1.9 | 1.3 | 2.5 | 8.6 | 100 | [60] | ||
| A. picadoi | 9.5 | 13.5 | 66.4 | 2.2 | 4.8 | 1.8 | <0.1 | 1.8 | 100 | [60] | ||
| Bothriechis aurifer | 7.3 | 35.1 | 9.5 | 10.7 | 16.4 | 1.4 | 13.4 | 3.2 | 97 | [61] | ||
| B. bicolor | 35.2 | 19.1 | 8.5 | 10.8 | 1 | 4.4 | 7.6 | 3.6 | 4.6 | 94.8 | [61] | |
| B. marchi | 14.3 | 10.1 | 34.2 | 1.1 | 2.8 | 4.2 | 6.5 | 10.6 | 8.5 | 83.8 | [61] | |
| B. lateralis | 8.7 | 11.3 | 55.1 | 6.1 | 6.5 | 11.1 | 98.8 | [61] | ||||
| B. nigroviridis | 38.3 | 18.4 | 0.5 | 2.1 | 37 | 96.3 | [62] | |||||
| B. schlegelii | 43.8 | 5.8 | 17.7 | 8.9 | 2.1 | 13.4 | 91.7 | [61] | ||||
| B. supraciliaris | 13.4 | 15.2 | 6.8 | 5.9 | 4.3 | 1.6 | 21.9 | 69.1 | [63] | |||
| B. thalassinus | 12.1 | 39.6 | 4.3 | 5.1 | 11.5 | 2 | 10.6 | 9.9 | 95.1 | [61] | ||
| Bothrocophias campbelli | 43.1 | 21.3 | 15.8 | 5.7 | 0.9 | 6.4 | 0.3 | 3.9 | 97.4 | [64] | ||
| B. colombiensis | 44.3 | <1 | 42.1 | 5.7 | 0.1 | 5.6 | 0.8 | 99.5 | [65] | |||
| Bothropoides diporus | 24.1 | 7.2 | 34.2 | 7.4 | 2.9 | 1.4 | 15.9 | 2.6 | 95.7 | [66] | ||
| B. erythromelaus (5 populations) | 10.1–15.1 | 4–9.7 | 32.5–59.9 | 0.4 | 8.4–21.6 | 3.4–8.9 | 9.3–14.5 | 68+ | [67] | |||
| B. insularis | 10 | 12.5 | 30 | 1.3 | 1.3 | 31.3 | 11.3 | 97.7 | [68] | |||
| B. jaracara (south-east) | 3.7 | 13.7 | 35.6 | 7.2 | 2.4 | 9.6 | 7 | 16.4 | 95.6 | [69] | ||
| B. jaracara (south) | 20.2 | 28.6 | 10.3 | 8 | 2.6 | 9.4 | 0.2 | 22.6 | 100 | [69] | ||
| B. neuwiedi | 8.4 | 8.8 | 49.9 | 16.7 | 2 | 8.6 | 94.4 | [70] | ||||
| B. pauloensis | 31.9 | 10.5 | 38.1 | 2.8 | 2.2 | 0.6 | 1.3 | 12.4 | 99.8 | [71] | ||
| Bothrops asper (Caribbean coast) | 28.8 | 18.2 | 41 | 9.2 | 0.1 | 0.5 | 2.1 | 99.9 | [72] | |||
| B. asper (Pacific coast) | 45.5 | 4.4 | 44 | 4.6 | 0.1 | 0.5 | 1.4 | 100 | [72] | |||
| B. atrox (Western Para Brazil) | 5.7–7.5 | 9.7–14.1 | 46.5–54 | 8.7–9.4 | 3.7–4.3 | 10.2–13.1 | 84.5+ | [73] | ||||
| B. atrox (Colombia) | 24.1 | 10.9 | 48.5 | 4.7 | 2.6 | 7.1 | 1.7 | 0.3 | 99.9 | [74] | ||
| B. atrox (Venezuela) | 7.7–8.5 | 2.3 | 85 | 1.2–1.5 | 2.8–3.8 | 99+ | [75] | |||||
| B. atrox (Peru) | 11 | 11.1 | 58.2 | 10.5 | 2.4 | 3.6 | 3.2 | 100 | [76] | |||
| B. ayerbi | 0.7 | 9.3 | 53.7 | 3.3 | 1.1 | 10.1 | 2.3 | 8.3 | 88.8 | [77] | ||
| B. barnetti | 6.4 | 6.7 | 74.1 | 0.8 | 3.1 | 3.3 | 5.5 | 99.9 | [76] | |||
| B. caribbaeus | 12.8 | 4.7 | 68.6 | 8.4 | 2.6 | 1.7 | 98.8 | [78] | ||||
| B. jararacussu | 25.7 | 12.3 | 26.2 | 15 | 2.2 | 9.7 | 91.1 | [70] | ||||
| B. lanceolatus | 8.6 | 14.4 | 74.2 | 2.8 | <0.1 | 100 | [78] | |||||
| B. pictus | 14.1 | 7.7 | 68 | 1.1 | 8.9 | 99.8 | [76] | |||||
| B. pirajai | 40.2 | 7.1 | 20.7 | 5.2 | 9.2 | 1.4 | 5.6 | 89.4 | [79] | |||
| B. punctatus | 9.3 | 5.4 | 41.5 | 3.1 | 1.2 | 16.7 | 3.8 | 10.7 | 91.7 | [80] | ||
| Cerrophidion godmani | 23.4 | 19.1 | 32.8 | 5 | 4.2 | 0.5 | 7.5 | 5.7 | 98.2 | [81] | ||
| C. sasai | 23.4 | 19.1 | 32.8 | 5 | 4.2 | 0.5 | 7.5 | 5.7 | 98.2 | [82] | ||
| Crotalus adamanteus | 7.8 | 20 | 24.4 | 5.3 | 1.3 | 22.2 | 16.8 | 97.8 | [83] | |||
| C. atrox | 7.3 | 19.8 | 49.7 | 8 | 4.3 | 3.4 | 6.2 | 3 | 100 | [84] | ||
| C. basiliscus | 14 | 11 | 68 | 2 | 4 | 99 | [85] | |||||
| C. culminatus | 8.3 | 10.1 | 35.5 | 2.7 | 1.9 | 13 | 1.6 | 24.4 | 97.5 | [86] | ||
| C. durissus cascavella | 90.9 | 1.2 | <0.1 | <0.1 | 0.9 | <0.1 | 0.2 | 93.4 | [87] | |||
| C. d. collilineatus | 72 | 1.9 | 0.4 | 0.5 | 1.8 | <0.1 | 0.5 | 20.8 | 98 | [88] | ||
| C. d. terrificus | 48.5 | 25.3 | 3.9 | 77.7 | [89] | |||||||
| C. horridus | 22.8 | 58.2 | 0.1 | 1.1 | 0.8 | 0.22 | 0.2 | 82.3 | [90] | |||
| C. simus simus | 22.4 | 30.4 | 27.4 | 5.7 | 1 | 0.6 | 1.5 | 6.5 | 95.5 | [91] | ||
| C. tigris | 26.8 | 66.2 | 1.9 | 0.2 | 95.1 | [92] | ||||||
| C. tzabacan | 11.1 | 5.4 | 18.5 | 0.5 | 35.2 | 4.2 | 23.5 | 98.4 | [86] | |||
| C. viridis | 7.7–10.2 | 26.8 | 10.9–11.4 | 1.9–2.5 | 2.1–3.9 | 1.8–3.3 | 0.1 | 6.5–8.2 | 35.6–38 | 0.1 | 93.5+ | [93] |
| Sistrurus catenatus (3 subsp.) | 31.3-31.9 | 18.2-24.4 | 40.6-48.6 | 1.6-4.2 | 0.8-10.7 | 0.9-4.2 | 93.4+ | [94] | ||||
| S. miliarius | 32.5 | 17.1 | 36.1 | 2.1 | 2.9 | 7.7 | 98.4 | [94] | ||||
| Lachesis acrochorda | 2.3 | 35.1 | 23.2 | 9.6 | 0.9 | 6.9 | 21.5 | 99.5 | [95] | |||
| L. melanocephala | 13.4 | 21 | 18.9 | 3.6 | 7.5 | 30.2 | 94.6 | [94] | ||||
| L. muta muta | 8.7 | 31.2 | 31.9 | 2.7 | 1.8 | 7.9 | 14.7 | 98.9 | [94] | |||
| L. m. rhombeata | 10.8 | 26.5 | 29.5 | 0.5 | 1.4 | 2.7 | 28 | 99.4 | [96] | |||
| L. stenophrys | 14.1 | 21.2 | 30.6 | 2.7 | 3.6 | 27.1 | 99.3 | [94] | ||||
| Porthidium lansbergii | 16.2 | 4.5 | 35.5 | 3.6 | 1.4 | 6.7 | 12.9 | 12.4 | 93.2 | [97] | ||
| P. nasutum | 11.6 | 9.6 | 52.1 | 3 | 1.3 | 10.4 | 9.9 | 1.9 | 99.8 | [81] | ||
| P. ophryomegus | 13.5 | 7.3 | 45 | 3.3 | 0.6 | 8 | 16.7 | 4.2 | 98.6 | [97] | ||
| Rhinocerophis alternatus | 2 | 5.8 | 52.2 | 14.9 | 2.5 | 14.8 | 92.2 | [70] | ||||
| R. cotiara | 0.6 | 13 | 51 | 19.6 | 2.9 | 4.7 | 91.8 | [70] | ||||
| R. fonescai | 30.1 | 4.1 | 42.5 | 1.9 | 2.4 | 9.8 | 4.4 | 95.2 | [98] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tasoulis, T.; Isbister, G.K. A Review and Database of Snake Venom Proteomes. Toxins 2017, 9, 290. https://doi.org/10.3390/toxins9090290
Tasoulis T, Isbister GK. A Review and Database of Snake Venom Proteomes. Toxins. 2017; 9(9):290. https://doi.org/10.3390/toxins9090290
Chicago/Turabian StyleTasoulis, Theo, and Geoffrey K. Isbister. 2017. "A Review and Database of Snake Venom Proteomes" Toxins 9, no. 9: 290. https://doi.org/10.3390/toxins9090290