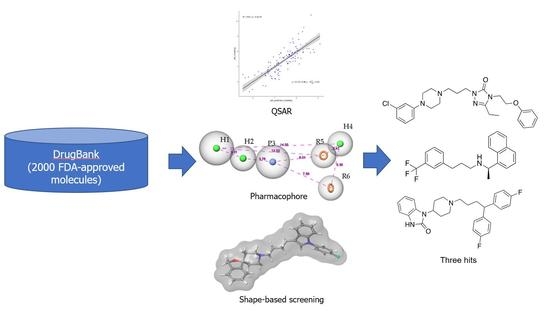
QSAR-Based Computational Approaches to Accelerate the Discovery of Sigma-2 Receptor (S2R) Ligands as Therapeutic Drugs
Abstract
:1. Introduction
2. Results
2.1. 2D-QSAR
2.2. Pharmacophore Model
2.3. Shape-Based Screening
2.4. Virtual Screening Workflow and Experimental Assay
3. Discussion and Conclusions
4. Materials and Methods
4.1. Data Preparation
4.1.1. QSAR Data
4.1.2. Pharmacophore Data Collection
4.2. Data Splitting
4.2.1. Descriptor Selection
4.2.2. Descriptor Screening
4.2.3. Advanced Descriptor Selection: Lasso, Stepwise, and Lars
4.3. Genetic Algorithm (GA)
4.4. GreedGene
4.5. QSAR Model Generation and Validation
4.5.1. Training
4.5.2. Validation
4.5.3. Testing
4.6. Pharmacophore Hypothesis Generation and Evaluation
4.7. Shape-Based Screening
4.8. Virtual Screening Protocol
4.9. Radioligand Binding Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
Appendix A
References
- Martin, W.R.; Eades, C.G.; Thompson, J.A.; Huppler, R.E.; Gilbert, P.E. The effects of morphine- and nalorphine- like drugs in the nondependent and morphine-dependent chronic spinal dog. J. Pharmacol. Exp. Ther. 1976, 197, 517–532. [Google Scholar] [PubMed]
- Su, T.P. Evidence for sigma opioid receptor: Binding of [3H]SKF-10047 to etorphine-inaccessible sites in guinea-pig brain. J. Pharmacol. Exp. Ther. 1982, 223, 284–290. [Google Scholar]
- Hellewell, S.B.; Bowen, W.D. A sigma-like binding site in rat pheochromocytoma (PC12) cells: Decreased affinity for (+)-benzomorphans and lower molecular weight suggest a different sigma receptor form from that of guinea pig brain. Brain Res. 1990, 527, 244–253. [Google Scholar] [CrossRef]
- Kavaliers, M.; Ossenkopp, K.P. Magnetic fields differentially inhibit mu, delta, kappa and sigma opiate-induced analgesia in mice. Peptides 1986, 7, 449–453. [Google Scholar] [CrossRef]
- Chien, C.-C.; Pasternak, G.W. Sigma antagonists potentiate opioid analgesia in rats. Neurosci. Lett. 1995, 190, 137–139. [Google Scholar] [CrossRef]
- Xu, J.; Zeng, C.; Chu, W.; Pan, F.; Rothfuss, J.M.; Zhang, F.; Tu, Z.; Zhou, D.; Zeng, D.; Vangveravong, S.; et al. Identification of the PGRMC1 protein complex as the putative sigma-2 receptor binding site. Nat. Commun. 2011, 2, 1–7. [Google Scholar] [CrossRef]
- Chu, U.B.; Mavlyutov, T.A.; Chu, M.; Yang, H.; Schulman, A.; Mesangeau, C.; McCurdy, C.R.; Guo, L.-W.; Ruoho, A.E. The sigma-2 receptor and progesterone receptor membrane component 1 are different binding sites derived from independent genes. EBioMedicine 2015, 2, 1806–1813. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alon, A.; Schmidt, H.R.; Wood, W.D.; Sahn, J.J.; Martin, S.F.; Kruse, A.C. Identification of the gene that codes for the σ2 receptor. Proc. Natl. Acad. Sci. USA 2017, 114, 7160–7165. [Google Scholar] [CrossRef] [Green Version]
- Zeng, C.; Weng, C.-C.; Schneider, M.E., Jr.; Puentes, L.; Riad, A.; Xu, K.; Makvandi, M.; Jin, L.; Hawkins, W.G.; Mach, R.H. TMEM97 and PGRMC1 do not mediate sigma-2 ligand-induced cell death. Cell Death Discov. 2019, 5, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Colabufo, N.A.; Abate, C.; Contino, M.; Inglese, C.; Niso, M.; Berardi, F.; Perrone, R. PB183, a sigma receptor ligand, as a potential PET probe for the imaging of prostate adenocarci-noma. Bioorg. Med. Chem. Lett. 2008, 18, 1990–1993. [Google Scholar] [CrossRef] [PubMed]
- Mir, S.U.; Ahmed, I.S.; Arnold, S.; Craven, R.J. Elevated progesterone receptor membrane component 1/sigma-2 receptor levels in lung tumors and plasma from lung cancer patients. Int. J. Cancer 2011, 131, E1–E9. [Google Scholar] [CrossRef]
- Russo, V.; Inglese, C.; Avallone, L.; Roperto, F.; Abate, C.; Zizzo, N.; Munday, J.S.; Berardi, F.; Colabufo, N.A. Sigma 2 receptor expression levels in blood and bladder from healthy and bladder cancer cattle. Vet. Comp. Oncol. 2017, 15, 1503–1512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, S.-R.; Yang, B.; Plössl, K.; Chumpradit, S.; Wey, S.-P.; Acton, P.D.; Wheeler, K.; Mach, R.H.; Kung, H.F. Development of a Tc-99m labeled sigma-2 receptor-specific ligand as a potential breast tumor imaging agent. Nucl. Med. Biol. 2001, 28, 657–666. [Google Scholar] [CrossRef]
- Crawford, K.W.; Bowen, W.D. Sigma-2 receptor agonists activate a novel apoptotic pathway and potentiate antineo-plastic drugs in breast tumor cell lines. Cancer Res. 2002, 62, 313–322. [Google Scholar]
- Craven, R.J. PGRMC1: A new biomarker for the estrogen receptor in breast cancer. Breast Cancer Res. 2008, 10, 113. [Google Scholar] [CrossRef]
- Kashiwagi, H.; McDunn, J.E.; Simon, P.O., Jr.; Goedegebuure, P.S.; Xu, J.; Jones, L.; Chang, K.; Johnston, F.; Trinkaus, K.; Hotchkiss, R.S.; et al. Selective sigma-2 ligands preferentially bind to pancreatic adenocarcinomas: Applications in diag-nostic imaging and therapy. Mol. Cancer 2007, 6, 48. [Google Scholar] [CrossRef] [Green Version]
- Kashiwagi, H.; McDunn, J.E.; Simon, P.O.; Goedegebuure, P.S.; Vangveravong, S.; Chang, K.; Hotchkiss, R.S.; Mach, R.H.; Hawkins, W.G. Sigma-2 receptor ligands potentiate conventional chemotherapies and improve survival in models of pancreatic adenocarcinoma. J. Transl. Med. 2009, 7, 24. [Google Scholar] [CrossRef] [Green Version]
- Hornick, J.R.; Xu, J.; Vangveravong, S.; Tu, Z.; Mitchem, J.B.; Spitzer, D.; Goedegebuure, P.; Mach, R.H.; Hawkins, W.G.; Xu, J. The novel sigma-2 receptor ligand SW43 stabilizes pancreas cancer progression in combination with gemcitabine. Mol. Cancer 2010, 9, 298. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hornick, J.R.; Vangveravong, S.; Spitzer, D.; Abate, C.; Berardi, F.; Goedegebuure, P.; Mach, R.H.; Hawkins, W.G. Lysosomal membrane permeabilization is an early event in sigma-2 receptor ligand mediated cell death in pancreatic cancer. J. Exp. Clin. Cancer Res. 2012, 31, 41. [Google Scholar] [CrossRef] [Green Version]
- Dehdashti, F.; Laforest, R.; Gao, F.; Shoghi, K.; Aft, R.L.; Nussenbaum, B.; Kreisel, F.H.; Bartlett, N.; Cashen, A.; Wagner-Johnson, N.; et al. Assessment of cellular proliferation in tumors by pet using 18F-ISO-1. J. Nucl. Med. 2013, 54, 350–357. [Google Scholar] [CrossRef] [Green Version]
- McDonald, E.; Doot, R.K.; Young, A.J.; Schubert, E.K.; Tchou, J.; Pryma, D.A.; Farwell, M.D.; Nayak, A.; Ziober, A.; Feldman, M.D.; et al. Breast cancer 18F-ISO-1 uptake as a marker of proliferation status. J. Nucl. Med. 2019, 61, 665–670. [Google Scholar] [CrossRef]
- Izzo, N.J.; Xu, J.; Zeng, C.; Kirk, M.J.; Mozzoni, K.; Silky, C.; Rehak, C.; Yurko, R.; Look, G.; Rishton, G.; et al. Alzheimer’s therapeutics targeting amyloid beta 1-42 oligomers II: Sigma-2/PGRMC1 receptors mediate Abeta 42 oligomer binding and synaptotoxicity. PLoS ONE 2014, 9, e111899. [Google Scholar] [CrossRef]
- Yi, B.; Sahn, J.J.; Ardestani, P.M.; Evans, A.K.; Scott, L.L.; Chan, J.Z.; Iyer, S.; Crisp, A.; Zuniga, G.; Pierce, J.T.; et al. Small molecule modulator of sigma 2 receptor is neuroprotective and reduces cognitive deficits and neu-roinflammation in experimental models of Alzheimer’s disease. J. Neurochem. 2017, 140, 561–575. [Google Scholar] [CrossRef]
- Grundman, M.; Morgan, R.; Lickliter, J.D.; Schneider, L.S.; DeKosky, S.; Izzo, N.J.; Guttendorf, R.; Higgin, M.; Pribyl, J.; Mozzoni, K.; et al. A phase 1 clinical trial of the sigma-2 receptor complex allosteric antagonist CT1812, a novel ther-apeutic candidate for Alzheimer’s disease. Alzheimer’s Dement. (N. Y.) 2019, 5, 20–26. [Google Scholar] [CrossRef] [PubMed]
- Tropsha, A. Best practices for qsar model development, validation, and exploitation. Mol. Inform. 2010, 29, 476–488. [Google Scholar] [CrossRef] [PubMed]
- Ferorelli, S.; Ferorelli, S.; Abate, C.; Pedone, M.; Colabufo, N.; Contino, M.; Perrone, R.; Berardi, F. Synthesis and binding assays of novel 3,3-dimethylpiperidine derivatives with various lipophilicities as sigma(1) receptor ligands. Bioorg. Med. Chem. 2011, 19, 7612–7622. [Google Scholar] [CrossRef]
- Mach, R.H.; Huang, Y.; Freeman, R.A.; Wu, L.; Vangveravong, S.; Luedtke, R.R. Conformationally-flexible benzamide analogues as dopamine D3 and σ2 receptor ligands. Bioorg. Med. Chem. Lett. 2004, 14, 195–202. [Google Scholar] [CrossRef]
- Huang, Y.; Luedtke, R.R.; Freeman, R.A.; Wu, L.; Mach, R.H. Synthesis of 2-(2,3-dimethoxyphenyl)-4-(aminomethyl)imidazole analogues and their binding affinities for dopamine D2 and D3 receptors. Bioorg. Med. Chem. 2001, 9, 3113–3122. [Google Scholar] [CrossRef]
- Mach, R.H.; Huang, Y.; Freeman, R.A.; Wu, L.; Blair, S.; Luedtke, R.R. Synthesis of 2-(5-bromo-2,3-dimethoxyphenyl)-5-(aminomethyl)-1H-pyrrole analogues and their bind-ing affinities for dopamine D2, D3, and D4 receptors. Bioorg. Med. Chem. 2003, 11, 225–233. [Google Scholar] [CrossRef]
- Yarim, M.; Koksal, M.; Schepmann, D.; Wünsch, B. Synthesis and in vitro evaluation of novel indole-based sigma receptors ligands. Chem. Biol. Drug Des. 2011, 78, 869–875. [Google Scholar] [CrossRef] [PubMed]
- Abate, C.; Ferorelli, S.; Niso, M.; Lovicario, C.; Infantino, V.; Convertini, P.; Perrone, R.; Berardi, F. 2-Aminopyridine derivatives as potential σ(2) receptor antagonists. ChemMedChem 2012, 7, 1847–1857. [Google Scholar] [CrossRef] [PubMed]
- Niso, M.; Abate, C.; Contino, M.; Ferorelli, S.; Azzariti, A.; Perrone, R.; Colabufo, N.A.; Berardi, F. Sigma-2 receptor agonists as possible antitumor agents in resistant tumors: Hints for collateral sensitivity. ChemMedChem 2013, 8, 2026–2035. [Google Scholar] [CrossRef]
- Abate, C.; Ferorelli, S.; Contino, M.; Marottoli, R.; Colabufo, N.A.; Perrone, R.; Berardi, F. Arylamides hybrids of two high-affinity σ2 receptor ligands as tools for the development of PET radiotracers. Eur. J. Med. Chem. 2011, 46, 4733–4741. [Google Scholar] [CrossRef] [PubMed]
- Berardi, F.; Ferorelli, S.; Abate, C.; Colabufo, N.A.; Contino, M.; Perrone, R.; Tortorella, V. 4-(tetralin-1-yl)- and 4-(naphthalen-1-yl)alkyl derivatives of 1-cyclohexylpiperazine as sigma receptor ligands with agonist sigma2 activity. J. Med. Chem. 2004, 47, 2308–2317. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Li, S.; Xu, J.; Peng, X.; Sai, K.; Chu, W.; Tu, Z.; Zeng, C.; Mach, R.H. Synthesis and structure-activity relationship studies of conformationally flexible tetrahydroisoquinolinyl triazole carboxamide and triazole substituted benzamide analogues as sigma2 receptor ligands. J. Med. Chem. 2014, 57, 4239–4251. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Bergmann, R.; Kniess, T.; Deuther-Conrad, W.; Mamat, C.; Neuber, C.; Liu, B.; Steinbach, J.; Brust, P.; Pietzsch, J. (18)F-Labeled 1,4-Dioxa-8-azaspiro[4.5]decane derivative: Synthesis and biological evaluation of a sigma1 receptor radioligand with low lipophilicity as potent tumor imaging agent. J. Med. Chem. 2015, 58, 5395–5407. [Google Scholar] [CrossRef] [PubMed]
- Berardi, F.; Ferorelli, S.; Abate, C.; Pedone, M.P.; Colabufo, N.A.; Contino, M.; Perrone, R. Methyl substitution on the piperidine ring of N-[omega-(6-methoxynaphthalen-1-yl)alkyl] derivatives as a probe for selective binding and activity at the sigma(1) receptor. J. Med. Chem. 2005, 48, 8237–8244. [Google Scholar] [CrossRef]
- Ferorelli, S.; Abate, C.; Colabufo, N.A.; Niso, M.; Inglese, C.; Berardi, F.; Perrone, R. Design and evaluation of naphthol- and carbazole-containing fluorescent sigma ligands as potential probes for receptor binding studies. J. Med. Chem. 2007, 50, 4648–4655. [Google Scholar] [CrossRef] [PubMed]
- Abate, C.; Niso, M.; Lacivita, E.; Mosier, P.D.; Toscano, A.; Perrone, R. Analogues of σ receptor ligand 1-cyclohexyl-4-[3-(5-methoxy-1,2,3,4-tetrahydronaphthalen-1-yl)propyl]piperazine (PB28) with added polar functionality and reduced lipophilicity for potential use as positron emission tomography radiotracers. J. Med. Chem. 2011, 54, 1022–1032. [Google Scholar] [CrossRef]
- Xie, F.; Kniess, T.; Neuber, C.; Deuther-Conrad, W.; Mamat, C.; Lieberman, B.P.; Liu, B.; Mach, R.H.; Brust, P.; Steinbach, J.; et al. Novel indole-based sigma-2 receptor ligands: Synthesis, structure–affinity relationship and antiproliferative activity. MedChemComm 2015, 6, 1093–1103. [Google Scholar] [CrossRef] [Green Version]
- Schininà, B.; Martorana, A.; Colabufo, N.A.; Contino, M.; Niso, M.; Perrone, M.G.; De Guidi, G.; Catalfo, A.; Rappazzo, G.; Zuccarello, E.; et al. 4-Nitro-2,1,3-benzoxadiazole derivatives as potential fluorescent sigma receptor probes. RSC Adv. 2015, 5, 47108–47116. [Google Scholar] [CrossRef]
- Dixon, S.L.; Smondyrev, A.M.; Knoll, E.H.; Rao, S.N.; Shaw, D.E.; Friesner, R.A. PHASE: A new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J. Comput. Mol. Des. 2006, 20, 647–671. [Google Scholar] [CrossRef]
- Moltzen, E.K.; Perregaard, J.; Meier, E. Sigma ligands with subnanomolar affinity and preference for the sigma.2 binding site. 2. spiro-joined benzofuran, isobenzofuran, and benzopyran piperidines. J. Med. Chem. 1995, 38, 2009–2017. [Google Scholar] [CrossRef]
- Chu, W.; McElveen, E.; Xu, J.; Taylor, M.; Luedtke, R.R.; Mach, R.H. Synthesis and in vitro binding of N-phenyl piperazine analogs as potential dopamine D3 receptor ligands. Bioorg. Med. Chem. 2005, 13, 77–87. [Google Scholar] [CrossRef]
- Gitto, R.; Luca, L.D.; Ferro, S.; Scala, A.; Ronsisvalle, S.; Parenti, C.; Prezzavento, O.; Buemi, M.R.; Chimirri, A. From NMDA receptor antagonists to discovery of selective sigma(2) receptor ligands. Bioorg. Med. Chem. 2014, 22, 393–397. [Google Scholar] [CrossRef] [PubMed]
- Stavitskaya, L.; Seminerio, M.J.; Matthews-Tsourounis, M.M.; Matsumoto, R.R.; Coop, A. The effect of the pyridyl nitrogen position in pyridylpiperazine sigma ligands. Bioorg. Med. Chem. Lett. 2010, 20, 2564–2565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mésangeau, C.; Narayanan, S.; Green, A.M.; Shaikh, J.; Kaushal, N.; Viard, E.; Xu, Y.-T.; Fishback, J.A.; Poupaert, J.H.; Matsumoto, R.R.; et al. Conversion of a highly selective sigma-1 receptor–ligand to sigma-2 receptor preferring ligands with anticocaine activity. J. Med. Chem. 2008, 51, 1482–1486. [Google Scholar] [CrossRef]
- Ashford, M.E.; Nguyen, V.H.; Greguric, I.; Pham, T.Q.; Keller, P.; Katsifis, A. Synthesis and in vitro evaluation of tetrahydroisoquinolines with pendent aromatics as sigma-2 (σ2) selective ligands. Org. Biomol. Chem. 2013, 12, 783–794. [Google Scholar] [CrossRef] [Green Version]
- Hajipour, A.R.; Guo, L.W.; Pal, A.; Mavlyutov, T.; Ruoho, A.E. Electron-donating para-methoxy converts a benzamide-isoquinoline derivative into a highly Sigma-2 receptor selective ligand. Bioorg. Med. Chem. 2011, 19, 7435–7440. [Google Scholar] [CrossRef] [Green Version]
- Niso, M.; Pati, M.L.; Berardi, F.; Abate, C. Rigid versus flexible anilines or anilides confirm the bicyclic ring as the hydrophobic portion for optimal σ2 receptor binding and provide novel tools for the development of future σ2 receptor PET radiotracers. RSC Adv. 2016, 6, 88508–88518. [Google Scholar] [CrossRef]
- Sun, Y.-T.; Wang, G.-F.; Yang, Y.-Q.; Jin, F.; Wang, Y.; Xie, X.-Y.; Mach, R.H.; Huang, Y.-S. Synthesis and pharmacological evaluation of 6,7-dimethoxy-1,2,3,4-tetrahydroisoquinoline derivatives as sigma-2 receptor ligands. Eur. J. Med. Chem. 2018, 147, 227–237. [Google Scholar] [CrossRef]
- Wu, Z.-W.; Song, S.-Y.; Li, L.; Lu, H.-L.; Lieberman, B.; Huang, Y.-S.; Mach, R.H. Synthesis and evaluation of tetrahydroindazole derivatives as sigma-2 receptor ligands. Bioorg. Med. Chem. 2015, 23, 1463–1471. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Lever, J.R.; Lever, S.Z. Synthesis and in vitro evaluation of tetrahydroisoquinolinyl benzamides as ligands for sigma receptors. Bioorganic Med. Chem. Lett. 2007, 17, 2594–2607. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mach, R.H.; Zeng, C.; Hawkins, W.G. The sigma2 receptor: A novel protein for the imaging and treatment of cancer. J. Med. Chem. 2013, 56, 7137–7160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartman, I.; Gillies, A.R.; Arora, S.; Andaya, C.; Royapet, N.; Welsh, W.J.; Wood, D.W.; Zauhar, R.J. Application of screening methods, shape signatures and engineered biosensors in early drug discovery process. Pharm. Res. 2009, 26, 2247–2258. [Google Scholar] [CrossRef]
- Diller, D.J.; Connell, N.D.; Welsh, W.J. Avalanche for shape and feature-based virtual screening with 3D alignment. J. Comput. Aided Mol. Des. 2015, 29, 1015–1024. [Google Scholar] [CrossRef]
- Nagarajan, K.; Zauhar, R.; Welsh, W.J. Enrichment of ligands for the serotonin receptor using the Shape Signatures approach. J. Chem. Inf. Model. 2005, 45, 49–57. [Google Scholar] [CrossRef]
- Zauhar, R.J.; Gianti, E.; Welsh, W.J. Fragment-based Shape Signatures: A new tool for virtual screening and drug discovery. J. Comput. Aided Mol. Des. 2013, 27, 1009–1036. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.Y.; Ai, N.; Arora, S.; Erenrich, E.; Nagarajan, K.; Zauhar, R.; Young, D.; Welsh, W.J. Identification of previously unrecognized antiestrogenic chemicals using a novel virtual screening approach. Chem. Res. Toxicol. 2006, 19, 1595–1601. [Google Scholar] [CrossRef] [Green Version]
- Zauhar, R.J.; Moyna, G.; Tian, L.; Li, Z.; Welsh, W.J. Shape signatures: A new approach to computer-aided ligand- and receptor-based drug design. J. Med. Chem. 2003, 46, 5674–5690. [Google Scholar] [CrossRef] [PubMed]
- Meek, P.J.; Liu, Z.; Tian, L.; Wang, C.Y.; Welsh, W.J.; Zauhar, R.J. Shape Signatures: Speeding up computer aided drug discovery. Drug Discov. Today 2006, 11, 895–904. [Google Scholar] [CrossRef]
- Hawkins, P.C.; Skillman, A.G.; Nicholls, A. Comparison of shape-matching and docking as virtual screening tools. J. Med. Chem. 2007, 50, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Nicholls, A.; McGaughey, G.B.; Sheridan, R.P.; Good, A.C.; Warren, G.; Mathieu, M.; Muchmore, S.W.; Brown, S.P.; Grant, A.J.; Haigh, J.A.; et al. Molecular shape and medicinal chemistry: A perspective. J. Med. Chem. 2010, 53, 3862–3886. [Google Scholar] [CrossRef] [PubMed]
- Kirchmair, J.; Distinto, S.; Markt, P.; Schuster, D.; Spitzer, G.M.; Liedl, K.R.; Wolber, G. How to optimize shape-based virtual screening: Choosing the right query and including chemical information. J. Chem. Inf. Model. 2009, 49, 678–692. [Google Scholar] [CrossRef]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673–683. [Google Scholar] [CrossRef]
- Lavecchia, A.; Cerchia, C. In silico methods to address polypharmacology: Current status, applications and future perspectives. Drug Discov. Today 2016, 21, 288–298. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Bellis, L.J.; Bento, A.P.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; et al. ChEMBL: A large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012, 40, D1100–D1107. [Google Scholar] [CrossRef] [Green Version]
- Ganapathy, M.E.; Prasad, P.D.; Huang, W.; Seth, P.; Leibach, F.H.; Ganapathy, V. Molecular and ligand-binding characterization of the sigma-receptor in the Jurkat human T lymphocyte cell line. J. Pharmacol. Exp. Ther. 1999, 289, 251–260. [Google Scholar]
- Mysinger, M.M.; Carchia, M.; Irwin, J.J.; Shoichet, B.K. Directory of useful decoys, enhanced (DUD-E): Better ligands and decoys for better benchmarking. J. Med. Chem. 2012, 55, 6582–6594. [Google Scholar] [CrossRef] [PubMed]
- Willett, P.; Barnard, J.M.; Downs, G.M. Chemical Similarity Searching. J. Chem. Inf. Comp. Sci. 1998, 38, 983–996. [Google Scholar] [CrossRef] [Green Version]
- Harder, E.; Damm, W.; Maple, J.; Wu, C.; Reboul, M.; Xiang, J.Y.; Wang, L.; Lupyan, D.; Dahlgren, M.K.; Knight, J.L.; et al. OPLS3: A force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 2016, 12, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Guvench, O.; Weiser, J.; Shenkin, P.; Kolossváry, I.; Still, W.C. Application of the frozen atom approximation to the GB/SA continuum model for solvation free energy. J. Comput. Chem. 2002, 23, 214–221. [Google Scholar] [CrossRef]
- Team, R.C. R: A Language and Environment for Statistical Computing. Available online: http://r.meteo.uni.wroc.pl/web/packages/dplR/vignettes/intro-dplR.pdf (accessed on 21 August 2021).
- Tibshirani, R. Regression shrinkage and selection via the Lasso. J. R. Stat. Soc. Ser. B-Methodol. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Efron, B.; Hastie, T.; Johnstone, I.; Tibshirani, R. Least angle regression. Ann. Stat. 2004, 32, 407–451. [Google Scholar] [CrossRef] [Green Version]
- Holland, J.H. Genetic Algorithms. Sci. Am. 1992, 267, 66–72. [Google Scholar] [CrossRef]
- Golbraikh, A.; Tropsha, A. Beware of q2! J. Mol. Graph. Model. 2002, 20, 269–276. [Google Scholar] [CrossRef]
- Dixon, S.L.; Smondyrev, A.M.; Rao, S.N. PHASE: A novel approach to pharmacophore modeling and 3D database searching. Chem. Biol. Drug Des. 2006, 67, 370–372. [Google Scholar] [CrossRef]
- Bender, A.; Glen, R.C. A discussion of measures of enrichment in virtual screening: Comparing the information content of descriptors with increasing levels of sophistication. J. Chem. Inf. Model. 2005, 45, 1369–1375. [Google Scholar] [CrossRef]
- Sastry, G.M.; Dixon, S.L.; Sherman, W. Rapid shape-based ligand alignment and virtual screening method based on atom/feature-pair similarities and volume overlap scoring. J. Chem. Inf. Model. 2011, 51, 2455–2466. [Google Scholar] [CrossRef]
- Probst, D.; Reymond, J.L. Exploring DrugBank in virtual reality chemical space. J. Chem. Inf. Model. 2018, 58, 1731–1735. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]




| ID | Reference | pKi Range | Number of Compounds | Structure |
|---|---|---|---|---|
| 1 | Ferorelli, Abate [26] | 5.48–7.71 | 9 |  |
| 2 | Mach, Huang [27] | 6.14–8.09 | 8 |  |
| 3 | Huang, Luedtke [28] | 6.39–6.95 | 4 |  |
| 4 | Mach, Huang [29] | 6.29–7.59 | 9 |  |
| 5 | Yarim, Koksal [30] | 6.18–8.00 | 6 |  |
| 6 | Abate, Ferorelli [31] | 6.51–8.79 | 14 |  |
| 7 | Niso, Abate [32] | 7.54–10.40 | 9 |  |
| 8 | Abate, Ferorelli [33] | 7.29–8.58 | 8 |  |
| 9 | Berardi, Ferorelli [34] | 7.52–9.24 | 4 |  |
| 10 | Bai, Li [35] | 5.99–8.82 | 22 |  |
| 11 | Xie, Bergmann [36] | 6.28–7.64 | 16 |  |
| 12 | Berardi, Ferorelli [37] | 6.62–7.75 | 15 |  |
| 13 | Ferorelli, Abate [38] | 6.17–8.08 | 8 |  |
| 14 | Abate, Niso [39] | 7.63–9.31 | 7 |  |
| 15 | Xie, Kniess [40] | 7.17–8.52 | 10 |  |
| 16 | Schininà, Martorana [41] | 5.33–7.25 | 10 |  |
| Lasso | b_Single | Chi0v_C | Chi1v_C | b_max1len | QRPC + |
| Stepwise | b_single | chi1_C | SMR_VSA2 | BCUT_PEOE_3 | SlogP_VSA9 |
| GA | balabanJ | b_max1len | SMR_VSA0 | Q_VSA_FPNEG | SMR_VSA3 |
| GreedGene | balabanJ | b_max1len | Q_VSA_PNEG | vsa_acc | SlogP_VSA1 |
| Statistical Parameters | Lasso | Stepwise | GA | GreedGene |
|---|---|---|---|---|
| Training R2 | 0.43–0.58 | 0.48–0.60 | 0.58–0.68 | 0.62–0.69 |
| Training Q2 | 0.36–0.52 | 0.42–0.56 | 0.52–0.63 | 0.57–0.64 |
| Validation R2 | 0.27–0.68 | 0.37–0.71 | 0.50–0.73 | 0.53–0.78 |
| % met criteria | 38% | 68% | 100% | 100% |
| Modeling R2 | 0.5 | 0.55 | 0.63 | 0.65 |
| Modeling Q2 | 0.45 | 0.50 | 0.59 | 0.62 |
| Testing R2 | 0.51 | 0.51 | 0.51 | 0.56 |
| Criteria met | Yes | Yes | Yes | Yes |
| Hypo 1 | HHHPRR | 15.2 * |
| Hypo 2 | HHPRD | 7.8 |
| Hypo 3 | HDPRR | 6.4 |
| Hypo 4 | HDPRR | 4.5 |
| Hypo 5 | HAPRR | 3.2 |
| Hypo 6 | HHPRDH | 5.2 |
| Hypo 7 | HAPRR | 3.7 |
| Hypo 8 | AHPRR | 2.1 |
| Hypo 9 | AHPRR | 4.1 |
| Hypo 10 | HHPRR | 4.3 |
| Generic Name | Structure | Inh% at 1 μM |
|---|---|---|
| Ranolazine |  | 13 |
| Flibanserin |  | 13 |
| Nefazodone |  | 76 |
| Cinacalcet |  | 50 |
| Pimozide |  | 55 |
| Vilazodone |  | 26 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, Y.; Dong, H.; Peng, Y.; Welsh, W.J. QSAR-Based Computational Approaches to Accelerate the Discovery of Sigma-2 Receptor (S2R) Ligands as Therapeutic Drugs. Molecules 2021, 26, 5270. https://doi.org/10.3390/molecules26175270
Yu Y, Dong H, Peng Y, Welsh WJ. QSAR-Based Computational Approaches to Accelerate the Discovery of Sigma-2 Receptor (S2R) Ligands as Therapeutic Drugs. Molecules. 2021; 26(17):5270. https://doi.org/10.3390/molecules26175270
Chicago/Turabian StyleYu, Yangxi, Hiep Dong, Youyi Peng, and William J. Welsh. 2021. "QSAR-Based Computational Approaches to Accelerate the Discovery of Sigma-2 Receptor (S2R) Ligands as Therapeutic Drugs" Molecules 26, no. 17: 5270. https://doi.org/10.3390/molecules26175270