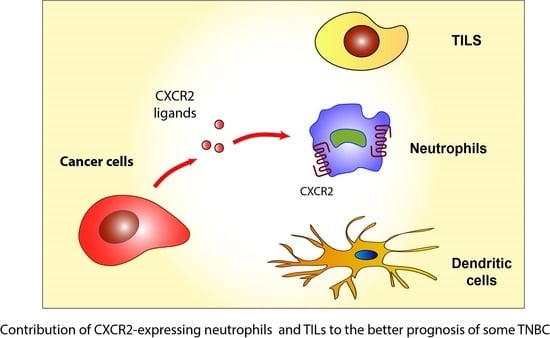
CXCR2 Levels Correlate with Immune Infiltration and a Better Prognosis of Triple-Negative Breast Cancers
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Objectives
2.2. Patients and Tumor Samples
2.3. Immunohistochemistry
2.4. Evaluation of TILs
2.5. Statistical Analysis
3. Results
3.1. Correlations of CD11b, CD66b and CXCR2 Expression with Clinicopathological Features
3.2. Correlations between CD11b, CD66b and CXCR2 in TNBC Samples
3.3. Correlations of CD11b, CD66b and CXCR2 Expression with Immune Tumor Microenvironment
3.4. Survival Analyses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AB | Antibody |
| AR | Androgen receptor |
| CAFs | Cancer associated fibroblasts |
| CI | Confidence interval |
| ECM | Extracellular matrix |
| ER | Estrogen receptor |
| FOXA1 | Forkhead box protein A1 |
| HR | Hazard ratio |
| IHC | Immunohistochemistry |
| NK | Natural killer |
| OS | Overall survival |
| PR | Progesterone receptor |
| ROI | Region of interest |
| RFS | Relapse-free survival |
| TIL | Tumor-infiltrating lymphocytes |
| TMA | Tissue microarray |
| TME | Tumor microenvironment |
| TNBC | Triple-negative breast cancer |
References
- Perou, C.M.; Sorlie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Dent, R.; Trudeau, M.; Pritchard, K.I.; Hanna, W.M.; Kahn, H.K.; Sawka, C.A.; Lickley, L.A.; Rawlinson, E.; Sun, P.; Narod, S.A. Triple-negative breast cancer: Clinical features and patterns of recurrence. Clin. Cancer Res. 2007, 13, 4429–4434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minn, A.J.; Gupta, G.P.; Siegel, P.M.; Bos, P.D.; Shu, W.; Giri, D.D.; Viale, A.; Olshen, A.B.; Gerald, W.L.; Massague, J. Genes that mediate breast cancer metastasis to lung. Nature 2005, 436, 518–524. [Google Scholar] [CrossRef] [PubMed]
- Ahn, S.G.; Kim, S.J.; Kim, C.; Jeong, J. Molecular Classification of Triple-Negative Breast Cancer. J. Breast Cancer 2016, 19, 223–230. [Google Scholar] [CrossRef]
- Lehmann, B.D.; Bauer, J.A.; Chen, X.; Sanders, M.E.; Chakravarthy, A.B.; Shyr, Y.; Pietenpol, J.A. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Investig. 2011, 121, 2750–2767. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burstein, M.D.; Tsimelzon, A.; Poage, G.M.; Covington, K.R.; Contreras, A.; Fuqua, S.A.; Savage, M.I.; Osborne, C.K.; Hilsenbeck, S.G.; Chang, J.C.; et al. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin. Cancer Res. 2015, 21, 1688–1698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guiu, S.; Mollevi, C.; Charon-Barra, C.; Boissiere, F.; Crapez, E.; Chartron, E.; Lamy, P.J.; Gutowski, M.; Bourgier, C.; Romieu, G.; et al. Prognostic value of androgen receptor and FOXA1 co-expression in non-metastatic triple negative breast cancer and correlation with other biomarkers. Br. J. Cancer 2018, 119, 76–79. [Google Scholar] [CrossRef]
- Angius, A.; Cossu-Rocca, P.; Arru, C.; Muroni, M.R.; Rallo, V.; Carru, C.; Uva, P.; Pira, G.; Orru, S.; De Miglio, M.R. Modulatory Role of microRNAs in Triple Negative Breast Cancer with Basal-Like Phenotype. Cancers 2020, 12, 3298. [Google Scholar] [CrossRef]
- Binnewies, M.; Roberts, E.W.; Kersten, K.; Chan, V.; Fearon, D.F.; Merad, M.; Coussens, L.M.; Gabrilovich, D.I.; Ostrand-Rosenberg, S.; Hedrick, C.C.; et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 2018, 24, 541–550. [Google Scholar] [CrossRef]
- Joyce, J.A.; Pollard, J.W. Microenvironmental regulation of metastasis. Nat. Rev. Cancer 2009, 9, 239–252. [Google Scholar] [CrossRef]
- Zlotnik, A.; Yoshie, O.; Nomiyama, H. The chemokine and chemokine receptor superfamilies and their molecular evolution. Genome Biol. 2006, 7, 243. [Google Scholar] [CrossRef] [PubMed]
- Hughes, C.E.; Nibbs, R.J.B. A guide to chemokines and their receptors. FEBS J. 2018, 285, 2944–2971. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Lazennec, G. Chemokines: Novel targets for breast cancer metastasis. Cancer Metastasis Rev. 2007, 26, 401–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galdiero, M.R.; Marone, G.; Mantovani, A. Cancer Inflammation and Cytokines. Cold Spring Harbor Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lazennec, G.; Lam, P.Y. Recent discoveries concerning the tumor—Mesenchymal stem cell interactions. Biochim. Biophys. Acta 2016, 1866, 290–299. [Google Scholar] [CrossRef] [Green Version]
- Lazennec, G.; Richmond, A. Chemokines and chemokine receptors: New insights into cancer-related inflammation. Trends Mol. Med. 2010, 16, 133–144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bieche, I.; Chavey, C.; Andrieu, C.; Busson, M.; Vacher, S.; Le Corre, L.; Guinebretiere, J.M.; Burlinchon, S.; Lidereau, R.; Lazennec, G. CXC chemokines located in the 4q21 region are up-regulated in breast cancer. Endocr. Relat. Cancer 2007, 14, 1039–1052. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boissiere-Michot, F.; Lazennec, G.; Frugier, H.; Jarlier, M.; Roca, L.; Duffour, J.; Du Paty, E.; Laune, D.; Blanchard, F.; Le Pessot, F.; et al. Characterization of an adaptive immune response in microsatellite-instable colorectal cancer. Oncoimmunology 2014, 3, e29256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chavey, C.; Bibeau, F.; Gourgou-Bourgade, S.; Burlinchon, S.; Boissiere, F.; Laune, D.; Roques, S.; Lazennec, G. Estrogen-receptor negative breast cancers exhibit a high cytokine content. Breast Cancer Res. 2007, 9, R15. [Google Scholar] [CrossRef] [Green Version]
- Escobar, P.; Bouclier, C.; Serret, J.; Bieche, I.; Brigitte, M.; Caicedo, A.; Sanchez, E.; Vacher, S.; Vignais, M.L.; Bourin, P.; et al. IL-1beta produced by aggressive breast cancer cells is one of the factors that dictate their interactions with mesenchymal stem cells through chemokine production. Oncotarget 2015, 6, 29034–29047. [Google Scholar] [CrossRef] [Green Version]
- Freund, A.; Chauveau, C.; Brouillet, J.P.; Lucas, A.; Lacroix, M.; Licznar, A.; Vignon, F.; Lazennec, G. IL-8 expression and its possible relationship with estrogen-receptor-negative status of breast cancer cells. Oncogene 2003, 22, 256–265. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Acharyya, S.; Oskarsson, T.; Vanharanta, S.; Malladi, S.; Kim, J.; Morris, P.G.; Manova-Todorova, K.; Leversha, M.; Hogg, N.; Seshan, V.E.; et al. A CXCL1 paracrine network links cancer chemoresistance and metastasis. Cell 2012, 150, 165–178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strieter, R.M.; Burdick, M.D.; Gomperts, B.N.; Belperio, J.A.; Keane, M.P. CXC chemokines in angiogenesis. Cytokine Growth Factor Rev. 2005, 16, 593–609. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Devalaraja, R.M.; Nanney, L.B.; Du, J.; Qian, Q.; Yu, Y.; Devalaraja, M.N.; Richmond, A. Delayed wound healing in CXCR2 knockout mice. J. Investig. Dermatol. 2000, 115, 234–244. [Google Scholar] [CrossRef] [Green Version]
- Cummings, C.J.; Martin, T.R.; Frevert, C.W.; Quan, J.M.; Wong, V.A.; Mongovin, S.M.; Hagen, T.R.; Steinberg, K.P.; Goodman, R.B. Expression and function of the chemokine receptors CXCR1 and CXCR2 in sepsis. J. Immunol. 1999, 162, 2341–2346. [Google Scholar] [PubMed]
- Timaxian, C.; Raymond-Letron, I.; Bouclier, C.; Gulliver, L.; Le Corre, L.; Chebli, K.; Guillou, A.; Mollard, P.; Balabanian, K.; Lazennec, G. The health status alters the pituitary function and reproduction of mice in a Cxcr2-dependent manner. Life Sci. Alliance 2020, 3. [Google Scholar] [CrossRef]
- Acosta, J.C.; O’Loghlen, A.; Banito, A.; Guijarro, M.V.; Augert, A.; Raguz, S.; Fumagalli, M.; Da Costa, M.; Brown, C.; Popov, N.; et al. Chemokine signaling via the CXCR2 receptor reinforces senescence. Cell 2008, 133, 1006–1018. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Belkadi, A.; Darnall, L.; Hu, T.; Drescher, C.; Cotleur, A.C.; Padovani-Claudio, D.; He, T.; Choi, K.; Lane, T.E.; et al. CXCR2-positive neutrophils are essential for cuprizone-induced demyelination: Relevance to multiple sclerosis. Nat. Neurosci. 2010, 13, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Chavey, C.; Lazennec, G.; Lagarrigue, S.; Clape, C.; Iankova, I.; Teyssier, J.; Annicotte, J.S.; Schmidt, J.; Mataki, C.; Yamamoto, H.; et al. CXC ligand 5 is an adipose-tissue derived factor that links obesity to insulin resistance. Cell Metab. 2009, 9, 339–349. [Google Scholar] [CrossRef] [Green Version]
- Keane, M.P.; Belperio, J.A.; Xue, Y.Y.; Burdick, M.D.; Strieter, R.M. Depletion of CXCR2 inhibits tumor growth and angiogenesis in a murine model of lung cancer. J. Immunol. 2004, 172, 2853–2860. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Yan, C.; Vilgelm, A.E.; Chen, S.C.; Ayers, G.D.; Johnson, C.A.; Richmond, A. Targeted Deletion of CXCR2 in Myeloid Cells Alters the Tumor Immune Environment to Improve Antitumor Immunity. Cancer Immunol. Res. 2020. [Google Scholar] [CrossRef]
- Liu, Y.; O’Leary, C.E.; Wang, L.S.; Bhatti, T.R.; Dai, N.; Kapoor, V.; Liu, P.; Mei, J.; Guo, L.; Oliver, P.M.; et al. CD11b+Ly6G+ cells inhibit tumor growth by suppressing IL-17 production at early stages of tumorigenesis. Oncoimmunology 2016, 5, e1061175. [Google Scholar] [CrossRef] [Green Version]
- Boissiere-Michot, F.; Jacot, W.; Fraisse, J.; Gourgou, S.; Timaxian, C.; Lazennec, G. Prognostic Value of CXCR2 in Breast Cancer. Cancers 2020, 12, 2076. [Google Scholar] [CrossRef]
- Goldhirsch, A.; Glick, J.H.; Gelber, R.D.; Coates, A.S.; Thurlimann, B.; Senn, H.J.; Panel, M. Meeting highlights: International expert consensus on the primary therapy of early breast cancer 2005. Ann. Oncol. 2005, 16, 1569–1583. [Google Scholar] [CrossRef]
- Jacot, W.; Gutowski, M.; Azria, D.; Romieu, G. Adjuvant early breast cancer systemic therapies according to daily used technologies. Crit. Rev. Oncol. Hematol. 2012, 82, 361–369. [Google Scholar] [CrossRef]
- Mansouri, H.; Alcaraz, L.B.; Mollevi, C.; Mallavialle, A.; Jacot, W.; Boissiere-Michot, F.; Simony-Lafontaine, J.; Laurent-Matha, V.; Roger, P.; Liaudet-Coopman, E.; et al. Co-Expression of Androgen Receptor and Cathepsin D Defines a Triple-Negative Breast Cancer Subgroup with Poorer Overall Survival. Cancers 2020, 12, 1244. [Google Scholar] [CrossRef] [PubMed]
- Jacot, W.; Lopez-Crapez, E.; Mollevi, C.; Boissiere-Michot, F.; Simony-Lafontaine, J.; Ho-Pun-Cheung, A.; Chartron, E.; Theillet, C.; Lemoine, A.; Saffroy, R.; et al. BRCA1 Promoter Hypermethylation is Associated with Good Prognosis and Chemosensitivity in Triple-Negative Breast Cancer. Cancers 2020, 12, 828. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boissiere-Michot, F.; Chabab, G.; Mollevi, C.; Guiu, S.; Lopez-Crapez, E.; Ramos, J.; Bonnefoy, N.; Lafont, V.; Jacot, W. Clinicopathological Correlates of gammadelta T Cell Infiltration in Triple-Negative Breast Cancer. Cancers 2021, 13, 765. [Google Scholar] [CrossRef]
- Salgado, R.; Denkert, C.; Demaria, S.; Sirtaine, N.; Klauschen, F.; Pruneri, G.; Wienert, S.; Van den Eynden, G.; Baehner, F.L.; Penault-Llorca, F.; et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: Recommendations by an International TILs Working Group 2014. Ann. Oncol. 2015, 26, 259–271. [Google Scholar] [CrossRef]
- Salgado, R.; Denkert, C.; Campbell, C.; Savas, P.; Nuciforo, P.; Aura, C.; de Azambuja, E.; Eidtmann, H.; Ellis, C.E.; Baselga, J.; et al. Tumor-Infiltrating Lymphocytes and Associations With Pathological Complete Response and Event-Free Survival in HER2-Positive Early-Stage Breast Cancer Treated With Lapatinib and Trastuzumab: A Secondary Analysis of the NeoALTTO Trial. JAMA Oncol. 2015, 1, 448–454. [Google Scholar] [CrossRef] [PubMed]
- Stanton, S.E.; Adams, S.; Disis, M.L. Variation in the Incidence and Magnitude of Tumor-Infiltrating Lymphocytes in Breast Cancer Subtypes: A Systematic Review. JAMA Oncol. 2016, 2, 1354–1360. [Google Scholar] [CrossRef] [PubMed]
- Adams, S.; Loi, S.; Toppmeyer, D.; Cescon, D.W.; De Laurentiis, M.; Nanda, R.; Winer, E.P.; Mukai, H.; Tamura, K.; Armstrong, A.; et al. Pembrolizumab monotherapy for previously untreated, PD-L1-positive, metastatic triple-negative breast cancer: Cohort B of the phase II KEYNOTE-086 study. Ann. Oncol. 2019, 30, 405–411. [Google Scholar] [CrossRef] [Green Version]
- Safonov, A.; Jiang, T.; Bianchini, G.; Gyorffy, B.; Karn, T.; Hatzis, C.; Pusztai, L. Immune Gene Expression Is Associated with Genomic Aberrations in Breast Cancer. Cancer Res. 2017, 77, 3317–3324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adams, S.; Gray, R.J.; Demaria, S.; Goldstein, L.; Perez, E.A.; Shulman, L.N.; Martino, S.; Wang, M.; Jones, V.E.; Saphner, T.J.; et al. Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J. Clin. Oncol. 2014, 32, 2959–2966. [Google Scholar] [CrossRef]
- Loi, S.; Drubay, D.; Adams, S.; Pruneri, G.; Francis, P.A.; Lacroix-Triki, M.; Joensuu, H.; Dieci, M.V.; Badve, S.; Demaria, S.; et al. Tumor-Infiltrating Lymphocytes and Prognosis: A Pooled Individual Patient Analysis of Early-Stage Triple-Negative Breast Cancers. J. Clin. Oncol. 2019, 37, 559–569. [Google Scholar] [CrossRef]
- Leon-Ferre, R.A.; Polley, M.Y.; Liu, H.; Gilbert, J.A.; Cafourek, V.; Hillman, D.W.; Elkhanany, A.; Akinhanmi, M.; Lilyquist, J.; Thomas, A.; et al. Impact of histopathology, tumor-infiltrating lymphocytes, and adjuvant chemotherapy on prognosis of triple-negative breast cancer. Breast Cancer Res. Treat. 2018, 167, 89–99. [Google Scholar] [CrossRef]
- Gao, Z.H.; Li, C.X.; Liu, M.; Jiang, J.Y. Predictive and prognostic role of tumour-infiltrating lymphocytes in breast cancer patients with different molecular subtypes: A meta-analysis. BMC Cancer 2020, 20, 1150. [Google Scholar] [CrossRef]
- Mahmoud, S.M.; Paish, E.C.; Powe, D.G.; Macmillan, R.D.; Grainge, M.J.; Lee, A.H.; Ellis, I.O.; Green, A.R. Tumor-infiltrating CD8+ lymphocytes predict clinical outcome in breast cancer. J. Clin. Oncol. 2011, 29, 1949–1955. [Google Scholar] [CrossRef]
- Lee, H.J.; Seo, J.Y.; Ahn, J.H.; Ahn, S.H.; Gong, G. Tumor-associated lymphocytes predict response to neoadjuvant chemotherapy in breast cancer patients. J. Breast Cancer 2013, 16, 32–39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brown, J.R.; Wimberly, H.; Lannin, D.R.; Nixon, C.; Rimm, D.L.; Bossuyt, V. Multiplexed quantitative analysis of CD3, CD8, and CD20 predicts response to neoadjuvant chemotherapy in breast cancer. Clin. Cancer Res. 2014, 20, 5995–6005. [Google Scholar] [CrossRef] [Green Version]
- Szekely, B.; Bossuyt, V.; Li, X.; Wali, V.B.; Patwardhan, G.A.; Frederick, C.; Silber, A.; Park, T.; Harigopal, M.; Pelekanou, V.; et al. Immunological differences between primary and metastatic breast cancer. Ann. Oncol. 2018, 29, 2232–2239. [Google Scholar] [CrossRef] [PubMed]
- Pardoll, D.M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 2012, 12, 252–264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quezada, S.A.; Peggs, K.S. Exploiting CTLA-4, PD-1 and PD-L1 to reactivate the host immune response against cancer. Br. J. Cancer 2013, 108, 1560–1565. [Google Scholar] [CrossRef]
- Ahmed, F.S.; Gaule, P.; McGuire, J.; Patel, K.; Blenman, K.; Pusztai, L.; Rimm, D.L. PD-L1 Protein Expression on Both Tumor Cells and Macrophages are Associated with Response to Neoadjuvant Durvalumab with Chemotherapy in Triple-negative Breast Cancer. Clin. Cancer Res. 2020, 26, 5456–5461. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, K.; Ishida, M.; Yanai, H.; Tsuta, K.; Sekimoto, M.; Sugie, T. Prognostic significance of PD-L1-positive cancer-associated fibroblasts in patients with triple-negative breast cancer. BMC Cancer 2021, 21, 239. [Google Scholar] [CrossRef] [PubMed]
- Hammond, M.E.; Hayes, D.F.; Wolff, A.C.; Mangu, P.B.; Temin, S. American society of clinical oncology/college of american pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Oncol. Pract. 2010, 6, 195–197. [Google Scholar] [CrossRef] [Green Version]
- Sanford, R.A.; Song, J.; Gutierrez-Barrera, A.M.; Profato, J.; Woodson, A.; Litton, J.K.; Bedrosian, I.; Albarracin, C.T.; Valero, V.; Arun, B. High incidence of germline BRCA mutation in patients with ER low-positive/PR low-positive/HER-2 neu negative tumors. Cancer 2015, 121, 3422–3427. [Google Scholar] [CrossRef] [PubMed] [Green Version]



| Patient Features | Number of Patients(n = 290) | % | |
|---|---|---|---|
| Age (years), median (min to max) | 57.72 | [28.54–89.10] | |
| <55 | 129 | 44.5 | |
| ≥55 | 161 | 55.5 | |
| Tumor size | |||
| T1 | 134 | 46.2 | |
| T2 | 138 | 47.6 | |
| T3/T4 | 18 | 6.2 | |
| Nodal status | |||
| N− | 189 | 65.2 | |
| N+ | 101 | 34.8 | |
| Histological grade (4 missing values) | |||
| 1–2 | 66 | 23.1 | |
| 3 | 220 | 76.9 | |
| Histology (3 missing values) | |||
| Ductal | 238 | 82.9 | |
| Lobular | 15 | 5.2 | |
| Other (1) | 34 | 11.9 | |
| Adjuvant chemotherapy (1 missing value) | |||
| No | 71 | 24.6 | |
| Yes | 218 | 75.4 | |
| Basal-like phenotype (2 missing value) | |||
| No | 101 | 35.1 | |
| Yes (basal) | 187 | 64.9 | |
| Molecular apocrine phenotype (17 missing values) | |||
| No | 159 | 58.2 | |
| Yes (molecular apocrine) | 114 | 41.8 | |
| TIL %, median (6 missing values) | |||
| <5% | 134 | 47.2 | |
| ≥5% | 150 | 52.8 | |
| CD3+ cell density (2 missing values) | |||
| Low | 144 | 50.0 | |
| High | 144 | 50.0 | |
| CD8+ cell density (6 missing values) | |||
| Low | 142 | 50.0 | |
| High | 142 | 50.0 | |
| PD-L1TC (24 missing values) | |||
| <1% | 119 | 44.7 | |
| ≥1% | 147 | 55.3 | |
| PD-L1SC (27 missing values) | |||
| 0 | 48 | 18.3 | |
| [0–10] | 85 | 32.3 | |
| [10–50] | 72 | 27.4 | |
| ≥50 | 58 | 22.1 | |
| PD-1SC (21 missing values) | |||
| 0 | 69 | 25.7 | |
| [0–10] | 72 | 26.8 | |
| [10–50] | 106 | 39.4 | |
| ≥50 | 22 | 8.2 | |
| CD11b+ cell density (15 missing values) | |||
| Low | 137 | 49.8 | |
| High | 138 | 50.2 | |
| CD66b+ cell density (14 missing values) | |||
| Low | 138 | 50.0 | |
| High | 138 | 50.0 | |
| CXCR2+ cell density (3 missing values) | |||
| Low | 144 | 50.2 | |
| High | 143 | 49.8 | |
| Variables | CD11b | CD66b | CXCR2 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Low | High | p-Value | Low | High | p-Value | Low | High | p-Value | ||||||||
| N | % | N | % | N | % | N | % | N | % | N | % | |||||
| Age (years) | ||||||||||||||||
| <55 | 53 | 38.7 | 73 | 52.9 | 0.018 | 72 | 52.2 | 55 | 39.9 | 0.040 | 61 | 42.4 | 68 | 47.6 | 0.377 | |
| ≥55 | 84 | 61.3 | 65 | 47.1 | 66 | 47.8 | 83 | 60.1 | 83 | 57.6 | 75 | 52.5 | ||||
| Tumor size | ||||||||||||||||
| T1 | 56 | 40.9 | 71 | 51.5 | 0.079 | 70 | 50.7 | 57 | 41.3 | 0.116 | 68 | 47.2 | 64 | 44.8 | 0.675 | |
| T2/T3/T4 | 81 | 59.1 | 67 | 48.6 | 68 | 49.3 | 81 | 58.7 | 76 | 52.8 | 79 | 55.2 | ||||
| Nodal status | ||||||||||||||||
| N− | 84 | 61.3 | 95 | 68.8 | 0.190 | 88 | 63.8 | 92 | 66.7 | 0.613 | 90 | 62.5 | 97 | 67.8 | 0.343 | |
| N+ | 53 | 38.7 | 43 | 31.2 | 50 | 36.2 | 46 | 33.3 | 54 | 37.5 | 46 | 32.2 | ||||
| Histological grade | ||||||||||||||||
| 1–2 | 48 | 35.8 | 14 | 10.2 | <0.001 | 47 | 34.8 | 14 | 10.2 | <0.001 | 49 | 34.8 | 16 | 11.3 | <0.001 | |
| 3 | 86 | 64.2 | 123 | 89.8 | 88 | 65.2 | 123 | 89.8 | 92 | 65.3 | 126 | 88.7 | ||||
| Histology | ||||||||||||||||
| Ductal | 109 | 79.6 | 118 | 87.4 | 0.082 | 109 | 79.6 | 120 | 88.2 | 0.051 | 111 | 78.2 | 124 | 87.3 | 0.041 | |
| Other | 28 | 20.4 | 17 | 12.6 | 28 | 20.4 | 16 | 11.8 | 31 | 21.8 | 18 | 12.7 | ||||
| Adjuvant chemotherapy | ||||||||||||||||
| No | 40 | 29.4 | 26 | 18.8 | 0.041 | 25 | 18.3 | 42 | 30.4 | 0.019 | 38 | 26.6 | 32 | 22.4 | 0.409 | |
| Yes | 96 | 70.6 | 112 | 81.2 | 112 | 81.8 | 96 | 69.6 | 105 | 73.4 | 111 | 77.6 | ||||
| Basal-like phenotype | ||||||||||||||||
| No | 55 | 40.4 | 41 | 29.9 | 0.069 | 46 | 33.8 | 52 | 37.7 | 0.505 | 56 | 39.4 | 45 | 31.5 | 0.160 | |
| Yes (Basal) | 81 | 59.6 | 96 | 70.1 | 90 | 66.2 | 86 | 62.3 | 86 | 60.6 | 98 | 68.5 | ||||
| Molecular apocrine phenotype | ||||||||||||||||
| No | 61 | 48.4 | 91 | 68.4 | 0.001 | 71 | 55.9 | 83 | 61.5 | 0.359 | 66 | 50.0 | 93 | 66.4 | 0.006 | |
| Yes (Molecular apocrine) | 65 | 51.6 | 42 | 31.6 | 56 | 44.1 | 52 | 38.5 | 66 | 50.0 | 47 | 33.6 | ||||
| Variables | CD11b Density | CD66b Density | CXCR2 Density | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Low | High | p-Value | Low | High | p-Value | Low | High | p-Value | ||||||||
| N | % | N | % | N | % | N | % | N | % | N | % | |||||
| TILs | ||||||||||||||||
| <5% | 88 | 64.7 | 39 | 29.1 | <0.001 | 70 | 51.1 | 56 | 41.8 | 0.125 | 84 | 60.0 | 49 | 34.8 | <0.001 | |
| ≥5% | 48 | 35.3 | 95 | 70.9 | 67 | 48.9 | 78 | 58.2 | 56 | 40.0 | 92 | 65.3 | ||||
| CD3+ density | ||||||||||||||||
| Low | 94 | 69.1 | 42 | 30.4 | <0.001 | 73 | 52.9 | 63 | 45.7 | 0.229 | 91 | 63.6 | 52 | 36.4 | <0.001 | |
| High | 42 | 30.9 | 96 | 69.6 | 65 | 47.1 | 75 | 54.4 | 52 | 36.4 | 91 | 63.6 | ||||
| CD8+ density | ||||||||||||||||
| Low | 78 | 58.2 | 57 | 41.6 | 0.006 | 77 | 56.2 | 57 | 41.9 | 0.018 | 90 | 63.8 | 51 | 36.2 | <0.001 | |
| High | 56 | 41.8 | 80 | 58.4 | 60 | 43.8 | 79 | 58.1 | 51 | 36.2 | 90 | 63.8 | ||||
| PD-L1TC | ||||||||||||||||
| <1% | 68 | 56.7 | 45 | 33.8 | <0.001 | 59 | 48.0 | 52 | 39.4 | 0.168 | 72 | 56.7 | 47 | 34.1 | <0.001 | |
| ≥1% | 52 | 43.3 | 88 | 66.2 | 64 | 52.0 | 80 | 60.6 | 55 | 43.3 | 91 | 65.9 | ||||
| PD-L1SC | ||||||||||||||||
| <10% | 72 | 61.0 | 53 | 39.9 | 0.001 | 77 | 62.6 | 49 | 37.7 | <0.001 | 76 | 60.8 | 57 | 41.6 | 0.002 | |
| ≥10% | 46 | 39.0 | 80 | 60.2 | 46 | 37.4 | 81 | 62.3 | 49 | 39.2 | 80 | 58.4 | ||||
| PD-1SC | ||||||||||||||||
| <10% | 76 | 61.8 | 59 | 44.7 | 0.006 | 84 | 65.6 | 51 | 38.9 | <0.001 | 84 | 63.6 | 57 | 41.9 | <0.001 | |
| ≥10% | 47 | 38.2 | 73 | 55.3 | 44 | 34.4 | 80 | 61.1 | 48 | 36.4 | 79 | 58.1 | ||||
| Variable | OS | RFS | |||||
|---|---|---|---|---|---|---|---|
| HR | 95% CI | p-Value | HR | 95% CI | p-Value | ||
| Age (years) | |||||||
| <55 | 1 | 1 | |||||
| ≥55 | 2.07 | 1.31–3.27 | 0.001 | 1.43 | 0.89–2.31 | 0.137 | |
| Tumor size | |||||||
| T1 | 1 | 1 | |||||
| T2/T3/T4 | 2.82 | 1.75–4.55 | <0.001 | 2.59 | 1.55–4.34 | <0.001 | |
| Nodal status | |||||||
| N− | 1 | 1 | |||||
| N+ | 2.25 | 1.48–3.42 | <0.001 | 4.34 | 2.67–7.05 | <0.001 | |
| Histological grade | |||||||
| 1–2 | 1 | 1 | |||||
| 3 | 0.79 | 0.50–1.26 | 0.328 | 1.02 | 0.59–1.76 | 0.931 | |
| Histology | |||||||
| Ductal | 1 | 1 | |||||
| Other | 0.61 | 0.33–1.15 | 0.108 | 0.91 | 0.49–1.69 | 0.764 | |
| Adjuvant chemotherapy | |||||||
| No | 1 | 1 | |||||
| Yes | 0.33 | 0.21–0.50 | <0.001 | 0.5 | 0.31–0.81 | 0.007 | |
| Basal-like phenotype | |||||||
| No | 1 | 1 | |||||
| Yes (basal) | 1.06 | 0.68–1.66 | 0.787 | 0.85 | 0.53–1.36 | 0.495 | |
| Molecular apocrine phenotype | |||||||
| No | 1 | 1 | |||||
| Yes (molecular apocrine) | 1.6 | 1.04–2.46 | 0.033 | 1.65 | 1.03–2.63 | 0.038 | |
| TILs | |||||||
| <5% | 1 | 1 | |||||
| ≥5% | 0.52 | 0.33–0.80 | 0.003 | 0.47 | 0.29–0.76 | 0.002 | |
| CD3+ density | |||||||
| Low | 1 | 1 | |||||
| High | 0.72 | 0.47–1.10 | 0.126 | 0.64 | 0.40–1.02 | 0.059 | |
| CD8+ density | |||||||
| Low | 1 | 1 | |||||
| High | 1.11 | 0.72–1.70 | 0.634 | 0.91 | 0.57–1.45 | 0.696 | |
| PD-L1TC | |||||||
| <1% | 1 | 1 | |||||
| ≥1% | 0.66 | 0.42–1.02 | 0.061 | 0.59 | 0.37–0.96 | 0.034 | |
| PD-L1SC | |||||||
| <10% | 1 | 1 | |||||
| ≥10% | 0.67 | 0.42–1.06 | 0.081 | 0.57 | 0.35–0.95 | 0.028 | |
| PD-1SC | |||||||
| <10% | 1 | 1 | |||||
| ≥10% | 1.09 | 0.71–1.67 | 0.708 | 0.92 | 0.57–1.47 | 0.725 | |
| CD11b density | |||||||
| Low | 1 | 1 | |||||
| High | 0.72 | 0.46–1.12 | 0.141 | 0.66 | 0.40–1.07 | 0.088 | |
| CD66b density | |||||||
| Low | 1 | 1 | |||||
| High | 1.29 | 0.83–2.01 | 0.251 | 1.2 | 0.74–1.93 | 0.456 | |
| CXCR2 density | |||||||
| Low | 1 | 1 | |||||
| High | 0.61 | 0.40–0.95 | 0.026 | 0.52 | 0.32–0.85 | 0.007 | |
| Variables | OS | RFS | |||||
|---|---|---|---|---|---|---|---|
| HR | 95% CI | p-Value | HR | 95% CI | p-Value | ||
| Tumor size | <0.001 | 0.017 | |||||
| T1 | 1 | 1 | |||||
| T2/T3/T4 | 2.48 | 1.49–4.13 | 1.87 | 1.10–3.17 | |||
| Nodal status | <0.001 | <0.001 | |||||
| N− | 1 | 1 | |||||
| N+ | 2.51 | 1.59–3.97 | 4.28 | 2.57–7.12 | |||
| Adjuvant chemotherapy | <0.001 | 0.002 | |||||
| No | 1 | 1 | |||||
| Yes | 0.32 | 0.20–0.50 | 0.43 | 0.26–0.71 | |||
| Histology | 0.002 | ||||||
| Ductal | 1 | ||||||
| Other | 0.38 | 0.19–0.76 | |||||
| TILs | 0.008 | 0.01 | |||||
| <5% | 1 | 1 | |||||
| ≥5% | 0.54 | 0.34–0.86 | 0.52 | 0.31–0.86 | |||
| CXCR2 | 0.05 | 0.058 | |||||
| Low | 1 | 1 | |||||
| High | 0.64 | 0.40–1.01 | 0.61 | 0.37–1.02 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boissière-Michot, F.; Jacot, W.; Massol, O.; Mollevi, C.; Lazennec, G. CXCR2 Levels Correlate with Immune Infiltration and a Better Prognosis of Triple-Negative Breast Cancers. Cancers 2021, 13, 2328. https://doi.org/10.3390/cancers13102328
Boissière-Michot F, Jacot W, Massol O, Mollevi C, Lazennec G. CXCR2 Levels Correlate with Immune Infiltration and a Better Prognosis of Triple-Negative Breast Cancers. Cancers. 2021; 13(10):2328. https://doi.org/10.3390/cancers13102328
Chicago/Turabian StyleBoissière-Michot, Florence, William Jacot, Océane Massol, Caroline Mollevi, and Gwendal Lazennec. 2021. "CXCR2 Levels Correlate with Immune Infiltration and a Better Prognosis of Triple-Negative Breast Cancers" Cancers 13, no. 10: 2328. https://doi.org/10.3390/cancers13102328