Abstract
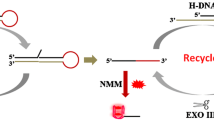
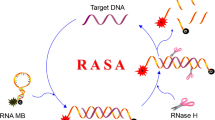
We proposed a dual-template, multi-cycle DNA nanomachine driven by polymerase and a nicking enzyme with high efficiency. The reaction system simply consists of two templates (T1, T2) and two enzymes (KF polymerase, Nb.BbvCI). The two templates are similar in structure (X-X’-Y, Y-Y’-C) with a primer recognition region, a primer analogue generation region, an output region (3′-5′), and two nicking sites. The output strand of T1 is the primer of T2, and the G-rich fragment (G3) is designed as the final product. In the presence of HIV-1, numerous G3 were generated through the multicycle amplification strategy and formed a G-triplex/ThT complex after the addition of thioflavin T (ThT), which greatly enhanced the fluorescence intensity as a signal reporter in the label-free sensing strategy. A dynamic response range of 50 fM - 2 nM for HIV-1 gene detection can be achieved through this multi-cycle G-triplex machine, and benefiting from the high efficiency amplification strategy, the enzymatic reaction can be completed within 45 min and followed by fluorescence measurements. In addition, the analysis of other targets can be achieved by replacing the template sequence. Thus, there is a certain application potential of this strategy for trace biomarker analysis.
Similar content being viewed by others
References
J. Das and S. O. Kelleys, Angew. Chem., Int. Ed. Engl., 2020, 59, 7.
S. R. Shin, Y. S. Zhang, and D. J. Kim, Anal. Chem., 2016, 88, 10019.
Z. Liu, T. Liu, and C. A. Tao, Anal. Sci., 2020, 36, 7.
X. H. Wen, X. W. Xu, and P. F. Huang, Anal. Sci., 2020, 36, 697.
E. Vargas, H. Teymourian, and F. Tehrani, Angew. Chem., Int. Ed. Engl, 2019, 58, 6376.
C. Liu, X. Zeng, and Z. An, ACS Sens., 2018, 3, 1471.
L. C. Brazaca, J. R. Moreto, A. Martin and F. Tehrani, ACS Nano, 2019, 13, 13325.
N. S. Li, W. L. Lin, and Y. P. Hsu, ACS Appl. Bio Mater., 2019, 2, 4847.
S. Mishra, Y. Lee, and J. W. Parks, Anal. Chem., 2018, 90, 12824.
L. Hartwell, D. Mankoff, and A. Paulovich, Nat. Biotechnol., 2006, 24, 905.
Z. Chen, L. Miao, and Y. Liu, Chem. Commun., 2017, 53, 12922.
D. Yang, H. Zhou, and N. E. Dina, R. Soc. Open Sci., 2018, 5, 180955.
L. Cao, X. Cui, and J. Hu, Biosens. Bioelectron., 2017, 90, 459.
Y. Wang, D. X. Liu, and J. P. Deng, Anal. Chim. Acta, 2017, 996, 74.
N. Tomita, Y. Mori, and H. Kanda, Nat. Protoc., 2008, 3, 877.
D. Wang, C. Vietz, and T. Schroder, Nano Lett., 2017, 17, 5368.
S. G. Harroun, C. Prevost-Tremblay, and D. Lauzon, Nanoscale, 2018, 10, 4607.
H. Gao, X. Zheng, and T. Yang, Chem. Commun., 2020, 56, 53.
F. J. Mo, M. Chen, and H. Meng, Sens. Actuators, B, 2020, 309, 7.
J. Wang, S. Li, and J. Xu, Chem. Commun., 2020, 56, 1681.
Y. Cao, L. Li, and B. Han, Biosens. Bioelectron., 2019, 141, 111397.
P. Li, L. Wang, and J. Zhu, Biosens. Bioelectron., 2015, 72, 107.
P. Qin, L. Yao, and J. Xu, Chem. Commun., 2019, 55, 14367.
E. Xiong and L. Jiangs, Analyst, 2019, 144, 634.
R. Zeng, L. Su, and Z. Luo, Anal. Chim. Acta, 2018, 1038, 21.
M. S. Reid, X. C. Le, and H. Zhangs, Angew. Chem., Int., Ed. Engl., 2018, 57, 11856.
H. Qi, S. Yue, and S. Bi, Biosens. Bioelectron., 2018, 110, 207.
H. Zhang, C. Fang, and S. Zhangs, Chemistry, 2010, 16, 12434.
B. Shlyahovsky, D. Li, and Y. Weizmann, J. Am. Chem. Soc., 2007, 129, 3814.
S. Bi, Y. Cui, and Y. Dong, Biosens. Bioelectron., 2014, 53, 207.
Q. Yan, Q. Duan, and Y. Huang, RSCAdv., 2019, 9, 41305.
X. Xu, W. Mao, and F. Lin, Catal. Commun., 2016, 74, 16.
S. Wang, B. Fu, and S. Peng, Chem. Commun., 2013, 49, 7920.
H. Zhou, Z. F. Wu, and Q. J. Han, Anal. Chem., 2018, 90, 3220.
H. Que, X. Yan, and B. Guo, Sens. Actuators, B, 2019, 291, 394.
Q. Liu, X. Sun, and M. Liu, Sens. Actuators, B, 2020, 310, 127804.
A. Kaushik, P. K. Vabbina, and V. Atluri, Biosens. Bioelectron., 2016, 86, 426.
C. Yuan, J. Fang, and Q. Duan, Biosens. Bioelectron., 2019, 133, 243.
S. Xu, Q. Li, and J. Xiang, Sci. Rep., 2016, 6, 24793.
N. K. Schwalb and F. Tempss, Science, 2008, 322, 243.
R. Wang, L. Wang, and H. Zhao, Biosens. Bioelectron., 2016, 86, 834.
H. Xu, D. Wu, and C. Q. Li, Biosens. Bioelectron., 2017, 90, 314.
Z. F. Shen, F. Li, and Y. F. Jiang, Anal. Chem., 2018, 90, 3335.
X. Liu, M. Zou, and D. Li, Anal. Chim. Acta, 2019, 1076, 138.
H. Xu, Y. Zhang, and S. Zhang, Anal. Chim. Acta, 2019, 1047, 172.
H. Xu, Y. Zhang, and S. Zhang, Sens. Actuators, B, 2019, 281, 1016.
Acknowledgements
Thanks for the support of the Science and Technology Research Program of Chongqing Yuzhong District Science and Technology Commission (Grant No. 20180127).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Yi, G., Duan, Q., Yan, Q. et al. Polymerase/Nicking Enzyme Powered Dual-template Multi-cycle G-Triplex Machine for HIV-1 Determination. ANAL. SCI. 37, 1087–1093 (2021). https://doi.org/10.2116/analsci.20P413
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2116/analsci.20P413