Abstract
This study aimed to uncover underlying mechanisms and promising intervention targets of heart failure (HF)-related stroke. HF-related dataset GSE42955 and stroke-related dataset GSE58294 were obtained from the Gene Expression Omnibus (GEO) database. Weighted gene co-expression network analysis (WGCNA) was conducted to identify key modules and hub genes. Gene Ontology (GO) and pathway enrichment analyses were performed on genes in the key modules. Genes in HF- and stroke-related key modules were intersected to obtain common genes for HF-related stroke, which were further intersected with hub genes of stroke-related key modules to obtain key genes in HF-related stroke. Key genes were functionally annotated through GO in the Reactome and Cytoscape databases. Finally, key genes were validated in these two datasets and other datasets. HF- and stroke-related datasets each identified two key modules. Functional enrichment analysis indicated that protein ubiquitination, Wnt signaling, and exosomes were involved in both HF- and stroke-related key modules. Additionally, ten hub genes were identified in stroke-related key modules and 155 genes were identified as common genes in HF-related stroke. OTU deubiquitinase with linear linkage specificity (OTULIN) and nuclear factor interleukin 3-regulated (NFIL3) were determined to be the key genes in HF-related stroke. Through functional annotation, OTULIN was involved in protein ubiquitination and Wnt signaling, and NFIL3 was involved in DNA binding and transcription. Importantly, OTULIN and NFIL3 were also validated to be differentially expressed in all HF and stroke groups. Protein ubiquitination, Wnt signaling, and exosomes were involved in HF-related stroke. OTULIN and NFIL3 may play a key role in HF-related stroke through regulating these processes, and thus serve as promising intervention targets.
概要
目的
本研究通过加权基因共表达网络分析(Weighted gene co-expression network analysis, WGCNA)揭示心力衰竭(心衰)相关脑卒中的发病机制, 为临床防治心衰相关脑卒中提供关键靶点.
创新点
本研究基于网络的研究策略, 首次揭示了心衰相关脑卒中的潜在分子相互作用机制以及可能参与的生物学过程, 并筛选出其中的关键基因.
方法
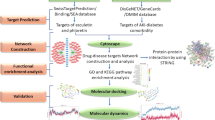
我们从基因表达综合数据库(Gene Expression Omnibus database, GEO)中分别获得心衰和脑卒中相关的芯片数据, 并通过WGCNA鉴定各自的关键模块与关键基因. 然后我们将二者的关键模块与关键基因取交集后, 得到心衰相关脑卒中的关键基因, 并利用Reactome和Cytoscape数据库对关键基因进行功能注释. 最后在多个数据集中对心衰相关脑卒中的关键基因进行验证.
结果
我们在心衰和脑卒中相关芯片数据中分别鉴定出两个关键模块, 随后功能富集分析发现两种疾病的关键模块中的基因均参与了蛋白质泛素化, Wnt信号通路和外泌体三个生物学过程. 取交集后, 我们共得到155个共同基因, 他们可能参与了心衰相关脑卒中的发病机制. 其中, OTULIN和NFIL3被识别为心衰相关脑卒中的关键基因. 功能注释分析显示, OTULIN参与了蛋白质泛素化和Wnt信号通路, NFIL3参与了转录和翻译过程. 最后, 在多个心衰和脑卒中的芯片数据中均证实OTULIN和NFIL3表达显著上调.
结论
蛋白质泛素化, Wnt信号通路和外泌体等生物学过程在心衰相关脑卒中的发病机制中发挥了重要作用. OTULIN和NFIL3通过调节这些通路因而可作为心衰相关脑卒中潜在的干预靶点.
Similar content being viewed by others
Change history
27 January 2024
An Erratum to this paper has been published: https://doi.org/10.1631/jzus.B20e0544
References
Adams HP, Bendixen BH, Kappelle LJ, et al., 1993. Classification of subtype of acute ischemic stroke. Definitions for use in a multicenter clinical trial. TOAST. Trial of Org 10172 in Acute Stroke Treatment. Stroke, 24(1):35–41. https://doi.org/10.1161/01.str.24.1.35
Aguilar MI, Hart R, Pearce LA, 2007. Oral anticoagulants versus antiplatelet therapy for preventing stroke in patients with non-valvular atrial fibrillation and no history of stroke or transient ischemic attacks. Cochrane Database Syst Rev, 3:CD006186. https://doi.org/10.1002/14651858.CD006186.pub2
Ashburner M, Ball CA, Blake JA, et al., 2000. Gene ontology: tool for the unification of biology. Nat Genet, 25(1):25–29. https://doi.org/10.1038/75556
Bang C, Batkai S, Dangwal S, et al., 2014. Cardiac fibroblast-derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. J Clin Invest, 124(5): 2136–2146. https://doi.org/10.1172/jci70577
Barac YD, Emrich F, Krutzwakd-Josefson E, et al., 2017. The ubiquitin-proteasome system: a potential therapeutic target for heart failure. J Heart Lung Transplant, 36(7): 708–714. https://doi.org/10.1016/j.healun.2017.02.012
Blankesteijn WM, van de Schans VAM, ter Horst P, et al., 2008. The Wnt/frizzled/GSK-3β pathway: a novel therapeutic target for cardiac hypertrophy. Trends Pharmacol Sci, 29(4):175–180. https://doi.org/10.1016/j.tips.2008.01.003
Bogiatzi C, Hackam DG, McLeod AI, et al., 2014. Secular trends in ischemic stroke subtypes and stroke risk factors. Stroke, 45(11):3208–3213. https://doi.org/10.1161/strokeaha.114.006536
Chen JL, Chopp M, 2018. Exosome therapy for stroke. Stroke, 49(5):1083–1090. https://doi.org/10.1161/strokeaha.117.018292
Chung S, Kim TH, Uhm JS, et al., 2020. Stroke and systemic embolism and other adverse outcomes of heart failure with preserved and reduced ejection fraction in patients with atrial fibrillation (from the COmparison Study of Drugs for symptom control and complication prEvention of Atrial Fibrillation [CODE-AF]). Am J Cardiol, 125(1):68–75. https://doi.org/10.1016/j.amjcard.2019.09.035
Cleland JGF, Findlay I, Jafri S, et al., 2004. The Warfarin/Aspirin study in heart failure (WASH): a randomized trial comparing antithrombotic strategies for patients with heart failure. Am Heart J, 148(1):157–164. https://doi.org/10.1016/j.ahj.2004.03.010
Cokkinos DV, Haralabopoulos GC, Kostis JB, et al., 2006. Efficacy of antithrombotic therapy in chronic heart failure: the HELAS study. Eur J Heart Fail, 8(4):428–432. https://doi.org/10.1016/j.ejheart.2006.02.012
di Tullio MR, Qian M, Thompson JLP, et al., 2016. Left ventricular ejection fraction and risk of stroke and cardiac events in heart failure: data from the warfarin versus aspirin in reduced ejection fraction trial. Stroke, 47(8):2031–2037. https://doi.org/10.1161/strokeaha.116.013679
Dichgans M, Pulit SL, Rosand J, 2019. Stroke genetics: discovery, biology, and clinical applications. Lancet Neurol, 18(6):587–599. https://doi.org/10.1016/s1474-4422(19)30043-2
Ding HY, Xie YN, Dong Q, et al., 2019. Roles of hyaluronan in cardiovascular and nervous system disorders. J Zhejiang Univ-SciB (Biomed & Biotechnol), 20(5):428–436. https://doi.org/10.1631/jzus.B1900155
Gao Y, Gong YL, Xia L, et al., 2019. Simulation of inter atrial block based on a human atrial model. J Zhejiang Univ-SciB (Biomed & Biotechnol), 20(4):300–309. https://doi.org/10.1631/jzus.B1800420
García-Berrocoso T, Palà E, Consegal M, et al., 2020. Cardioembolic ischemic stroke gene expression fingerprint in blood: a systematic review and verification analysis. Transl Stroke Res, 11(3):326–336. https://doi.org/10.1007/s12975-019-00730-x
GBD 2015 Mortality and Causes of Death Collaborators, 2016. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the global burden of disease study 2015. Lancet, 388(10053): 1459–1544. https://doi.org/10.1016/s0140-6736(16)31012-1
Gentleman R, Carey V, Huber W, et al., 2021. genefilter: methods for filtering genes from high-throughput experiments. Bioconductor version: Release (3.13). https://doi.org/10.18129/B9.bioc.genefilter
Hahn VS, Knutsdottir H, Luo X, et al., 2021. Myocardial gene expression signatures in human heart failure with preserved ejection fraction. Circulation, 143(2):120–134. https://doi.org/10.1161/circulationaha.120.050498
Horvath S, Dong J, 2008. Geometric interpretation of gene coexpression network analysis. PLoS Comput Biol, 4(8): e1000117. https://doi.org/10.1371/journal.pcbi.1000117
Irizarry RA, Hobbs B, Collin F, et al., 2003. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics, 4(2):249–264. https://doi.org/10.1093/biostatistics/4.2.249
Janky R, Verfaillie A, Imrichová H, et al., 2014. iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLoS Comput Biol, 10(7): e1003731. https://doi.org/10.1371/journal.pcbi.1003731
Jassal B, Matthews L, Viteri G, et al., 2020. The reactome pathway knowledgebase. Nucleic Acids Res, 48(D1): D498–D503. https://doi.org/10.1093/nar/gkz1031
Jickling GC, Xu HC, Stamova B, et al., 2010. Signatures of cardioembolic and large-vessel ischemic stroke. Ann Neurol, 68(5):681–692. https://doi.org/10.1002/ana.22187
Kamel H, Healey JS, 2017. Cardioembolic stroke. Circ Res, 120(3):514–526. https://doi.org/10.1161/circresaha.116.308407
Kanehisa M, Sato Y, 2020. KEGG mapper for inferring cellular functions from protein sequences. Protein Sci, 29(1):28–35. https://doi.org/10.1002/pro.3711
Kang SH, Kim J, Park JJ, et al., 2017. Risk of stroke in congestive heart failure with and without atrial fibrillation. Int J Cardiol, 248:182–187. https://doi.org/10.1016/j.ijcard.2017.07.056
Kotecha D, Banerjee A, Lip GYH, 2015. Increased stroke risk in atrial fibrillation patients with heart failure: does ejection fraction matter? Stroke, 46(3):608–609. https://doi.org/10.1161/strokeaha.114.008421
Leek JT, Johnson WE, Parker HS, et al., 2012. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics, 28(6):882–883. https://doi.org/10.1093/bioinformatics/bts034
Li T Qin JJ, Yang X, et al. 2017. The ubiquitin E3 ligase TRAF6 exacerbates ischemic stroke by ubiquitinating and activating Rac1. J Neurosci, 37(50):12123–12140. https://doi.org/10.1523/jneurosci.1751-17.2017
Liu ZH, Ma CG, Gu JH, et al. 2019. Potential biomarkers of acute myocardial infarction based on weighted gene co-expression network analysis. BioMed Eng OnLine, 18:9. https://doi.org/10.1186/s12938-019-0625-6
Malik R, Dichgans M, 2018. Challenges and opportunities in stroke genetics. Cardiovasc Res, 114(9):1226–1240. https://doi.org/10.1093/cvr/cvy068
Malik R, Chauhan G, Traylor M, et al. 2018. Multiancestry genome-wide association study of 52 0000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat Genet, 50(4):524–537. https://doi.org/10.1038/s41588-018-0058-3
Massie BM, Collins JF, Ammon SE, et al., 2009. Randomized trial of warfarin, aspirin, and clopidogrel in patients with chronic heart failure: the Warfarin and Antiplatelet Therapy in Chronic Heart Failure (WATCH) trial. Circulation, 119(12):1616–1624. https://doi.org/10.1161/circulationaha.108.801753
Mischie AN, Chioncel V, Droc I, et al., 2013. Anticoagulation in patients with dilated cardiomyopathy, low ejection fraction, and sinus rhythm: back to the drawing board. Cardiovasc Ther, 31(5):298–302. https://doi.org/10.1111/1755-5922.12019
Murtagh B, Smalling RW, 2006. Cardioembolic stroke. Curr Atheroscler Rep, 8(4):310–316. https://doi.org/10.1007/s11883-006-0009-9
Niu XW, Zhang JJ, Zhang LL, et al., 2019. Weighted gene co-expression network analysis identifies critical genes in the development of heart failure after acute myocardial infarction. Front Genet, 10:1214. https://doi.org/10.3389/fgene.2019.01214
Pei G, Chen L, Zhang W, 2017. WGCNA application to proteomic and metabolomic data analysis. Methods Enzymol, 585:135–158. https://doi.org/10.1016/bs.mie.2016.09.016
Pu LY, Wang M, Li KX, et al., 2020. Identification micro-RNAs functional modules and genes of ischemic stroke based on weighted gene co-expression network analysis (WGCNA). Genomics, 112(4):2748–2754. https://doi.org/10.1016/j.ygeno.2020.03.011
Pullicino P, Homma S, 2010. Stroke in heart failure: atrial fibrillation revisited? J Stroke Cerebrovasc Dis, 19(1): 1–2. https://doi.org/10.1016/j.jstrokecerebrovasdis.2009.09.002
Raghow R, 2016. An ‘omics’ perspective on cardiomyopathies and heart failure. Trends Mol Med, 22(9):813–827. https://doi.org/10.1016/j.molmed.2016.07.007
Ranjan P, Kumari R, Verma SK, 2019. Cardiac fibroblasts and cardiac fibrosis: precise role of exosomes. Front Cell Dev Biol, 7:318. https://doi.org/10.3389/fcell.2019.00318
Ritchie ME, Phipson B, Wu D, et al., 2015. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res, 43(7):e47. https://doi.org/10.1093/nar/gkv007
Rivkin E, Almeida SM, Ceccarelli DF, et al., 2013. The linear ubiquitin-specific deubiquitinase gumby regulates angiogenesis. Nature, 498(7454):318–324. https://doi.org/10.1038/nature12296
Schoner A, Tyrrell C, Wu M, et al., 2015. Endocardial endothelial dysfunction progressively disrupts initially anti then pro-thrombotic pathways in heart failure mice. PLoS ONE, 10(11):e0142940. https://doi.org/10.1371/journal.pone.0142940
Shannon P, Markiel A, Ozier O, et al., 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res, 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Simmonds SJ, Cuijpers I, Heymans S, et al., 2020. Cellular and molecular differences between HFpEF and HFrEF: a step ahead in an improved pathological understanding. Cells, 9(1):242. https://doi.org/10.3390/cells9010242
Stangl A, Elliott PR, Pinto-Fernandez A, et al., 2019. Regulation of the endosomal SNX27-retromer by OTULIN. Nat Commun, 10:4320. https://doi.org/10.1038/s41467-019-12309-z
Struijs JN, van Genugten MLL, Evers SMAA, et al., 2006. Future costs of stroke in the Netherlands: the impact of stroke services. Int J Technol Assess Health Care, 22(4): 518–524. https://doi.org/10.1017/s0266462306051464
Tamai SI, Imaizumi K, Kurabayashi N, et al., 2014. Neuroprotective role of the basic leucine zipper transcription factor NFIL3 in models of amyotrophic lateral sclerosis. J Biol Chem, 289(3):1629–1638. https://doi.org/10.1074/jbc.M113.524389
The Gene Ontology Consortium, 2019. The Gene Ontology Resource: 20 years and still going strong. Nucleic Acids Res, 47(D1):D330–D338. https://doi.org/10.1093/nar/gky1055
Thomas I, EncisoSilva J, Schlueter M, et al., 2016. Anticoagulation therapy and NOACs in heart failure. In: Bauersachs J, Butler J, Sandner P (Eds.), Heart Failure. Handbook of Experimental Pharmacology, Vol. 243. Springer, Cham, p.515–535. https://doi.org/10.1007/164_2016_126
van de Schans VAM, van den Borne SWM, Strzelecka AE, et al., 2007. Interruption of Wnt signaling attenuates the onset of pressure overload-induced cardiac hypertrophy. Hypertension, 49(3):473–480. https://doi.org/10.1161/01.Hyp.0000255946.55091.24
Velmurugan BK, Chang RL, Marthandam Asokan S, et al., 2018. A minireview of E4BP4/NFIL3 in heart failure. J Cell Physiol, 233(11):8458–8466. https://doi.org/10.1002/jcp.26790
Wang M, Wang LJ, Pu LY, et al., 2020. LncRNAs related key pathways and genes in ischemic stroke by weighted gene co-expression network analysis (WGCNA). Genomics, 112(3):2302–2308. https://doi.org/10.1016/j.ygeno.2020.01.001
Wei ZZ, Zhang JY, Taylor TM, et al., 2018. Neuroprotective and regenerative roles of intranasal Wnt-3a administration after focal ischemic stroke in mice. J Cereb Blood Flow Metab, 38(3):404–421. https://doi.org/10.1177/0271678x17702669
Wojcik C, di Napoli M, 2004. Ubiquitin-proteasome system and proteasome inhibition: new strategies in stroke therapy. Stroke, 35(6):1506–1518. https://doi.org/10.1161/01.STR.0000126891.93919.4e
Xu HB, Qin WY, Hu X, et al., 2018. Lentivirus-mediated overexpression of OTULIN ameliorates microglia activation and neuroinflammation by depressing the activation of the NF-κB signaling pathway in cerebral ischemia/reperfusion rats. J Neuroinflammation, 15:83. https://doi.org/10.1186/s12974-018-1117-5
Yang PC, 2018. Induced pluripotent stem cell (IPSC)-derived exosomes for precision medicine in heart failure. Circ Res, 122(5):661–663. https://doi.org/10.1161/circresaha.118.312657
Yip AM, Horvath S, 2007. Gene network interconnectedness and the generalized topological overlap measure. BMC Bioinformatics, 8:22. https://doi.org/10.1186/1471-2105-8-22
Zhang ZG, Chopp M, 2016. Exosomes in stroke pathogenesis and therapy. J Clin Invest, 126(4): 1190–1197. https://doi.org/10.1172/jci81133
Zhou LY, Wang Y, Wang K, et al., 2019. Potential therapeutic drugs for ischemic stroke based on bioinformatics analysis. Int J Neurosci, 129(11):1098–1102. https://doi.org/10.1080/00207454.2019.1634072
Zhou YY, Zhou B, Pache L, et al., 2019. Metascape provides a biologist-oriented resource for the analysis of systemslevel datasets. Nat Commun, 10:1523. https://doi.org/10.1038/s41467-019-09234-6
Zhu WH, Nan YN, Wang SQ, et al., 2019. Bioinformatics analysis of gene expression profiles of sex differences in ischemic stroke. Biomed Res Int, 2019:2478453. https://doi.org/10.1155/2019/2478453
Zou RJ, Zhang DE, Lv L, et al., 2019. Bioinformatic gene analysis for potential biomarkers and therapeutic targets of atrial fibrillation-related stroke. J Transl Med, 17:45. https://doi.org/10.1186/s12967-019-1790-x
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 81900387), the Guangdong Basic and Applied Basic Research Fund (No. 2019A1515011806), the Fundamental Research Funds for the Central Universities (No. 19ykpy97), and the Science and Technology Program of Guangzhou City of China (No. 201803040010).
Author information
Authors and Affiliations
Contributions
Zhaoyu LIU and Jingfeng WANG directed the project. Haifeng ZHANG designed experiments. Chiyu LIU and Sixu CHEN performed experiments and data analysis. Chiyu LIU drafted the manuscript. Yangxin CHEN, Qingyuan GAO, and Zhiteng CHEN perfected the manuscript. All authors have read and approved the final manuscript and, therefore, have full access to all the data in the study and take responsibility for the integrity and security of the data.
Corresponding authors
Ethics declarations
Chiyu LIU, Sixu CHEN, Haifeng ZHANG, Yangxin CHEN, Qingyuan GAO, Zhiteng CHEN, Zhaoyu LIU and Jingfeng WANG declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by any of the authors.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Liu, C., Chen, S., Zhang, H. et al. Bioinformatic analysis for potential biological processes and key targets of heart failure-related stroke. J. Zhejiang Univ. Sci. B 22, 718–732 (2021). https://doi.org/10.1631/jzus.B2000544
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B2000544