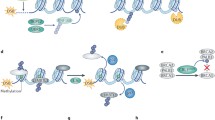
Abstract
DNA is the hereditary material in humans and almost all other organisms. It is essential for maintaining accurate transmission of genetic information. In the life cycle, DNA replication, cell division, or genome damage, including that caused by endogenous and exogenous agents, may cause DNA aberrations. Of all forms of DNA damage, DNA double-strand breaks (DSBs) are the most serious. If the repair function is defective, DNA damage may cause gene mutation, genome instability, and cell chromosome loss, which in turn can even lead to tumorigenesis. DNA damage can be repaired through multiple mechanisms. Homologous recombination (HR) and non-homologous end joining (NHEJ) are the two main repair mechanisms for DNA DSBs. Increasing amounts of evidence reveal that protein modifications play an essential role in DNA damage repair. Protein deubiquitination is a vital post-translational modification which removes ubiquitin molecules or polyubiquitinated chains from substrates in order to reverse the ubiquitination reaction. This review discusses the role of deubiquitinating enzymes (DUBs) in repairing DNA DSBs. Exploring the molecular mechanisms of DUB regulation in DSB repair will provide new insights to combat human diseases and develop novel therapeutic approaches.
摘要
DNA是人类和几乎所有有机体的遗传物质,它对于保持遗传信息的准确传递至关重要。在生命周期中,DNA复制、细胞分裂、基因组损伤,以及由内源性和外源性因素引起的损伤,都可能引起DNA损伤。在所有形式的DNA损伤中,DNA双链断裂(DSB)是最严重的。如果修复功能有缺陷,DNA损伤可能导致基因突变、基因组不稳定、细胞染色体丢失,进而导致肿瘤的发生。DNA损伤可以通过多种机制修复。同源重组(HR)和非同源末端连接(NHEJ)是DSB的两种主要修复机制。另外,大量研究表明,蛋白质修饰在DNA损伤修复中起着至关重要的作用。蛋白质的去泛素化是一种重要的翻译后修饰,它可以从底物中去除泛素分子或多泛素链,从而逆转泛素化降解,稳定底物蛋白。本文综述了去泛素化酶(DUB)在DSB损伤修复中的作用,探讨DUB调控DSB修复的分子机制,为开发人类疾病的新疗法提供了全新思路。
Similar content being viewed by others
Change history
08 April 2021
The typesetting format of the online version of the first issue (2021 22(01)) of Journal of Zhejiang University-SCIENCE B is different from that of the printed version (but all the text, figure and table contents in the article are correct). This is due to the new typesetting company adopted this year.
References
Altun M, Walter TS, Kramer HB, et al., 2015. The human otubain2-ubiquitin structure provides insights into the cleavage specificity of poly-ubiquitin-linkages. PLoS ONE, 10:e0115344. https://doi.org/10.1371/journal.pone.0115344
Britton S, Coates J, Jackson SP, 2013. A new method for highresolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J Cell Biol, 202(3): 579–595. https://doi.org/10.1083/jcb.201303073
Butler LR, Densham RM, Jia JY, et al., 2012. The proteasomal de-ubiquitinating enzyme POH1 promotes the doublestrand DNA break response. EMBO J, 31(19):3918–3934. https://doi.org/10.1038/emboj.2012.232
Cadet J, Berger M, Douki T, et al., 1997. Oxidative damage to DNA: formation, measurement, and biological significance. Rev Physiol Biochem Pharmacol, 131:1–87. https://doi.org/10.1007/3-540-61992-5
Cai JT, Wei JX, Schrott V, et al., 2018. Induction of deubiquitinating enzyme USP50 during erythropoiesis and its potential role in the regulation of Ku70 stability. J Investig Med, 66(1): 1–6. https://doi.org/10.1136/jim-2017-000622
Ceccaldi R, Rondinelli B, D’Andrea AD, 2016. Repair pathway choices and consequences at the double-strand break. Trends Cell Biol, 26(1):52–64. https://doi.org/10.1016/j.tcb.2015.07.009
Chauhan D, Tian Z, Nicholson B, et al., 2012. A small molecule inhibitor of ubiquitin-specific protease-7 induces apoptosis in multiple myeloma cells and overcomes bortezomib resistance. Cancer Cell, 22(3):345–358. https://doi.org/10.1016/j.ccr.2012.08.007
Chen XW, Arciero CA, Wang CR, et al., 2006. BRCC36 is essential for ionizing radiation-induced BRCA1 phosphorylation and nuclear foci formation. Cancer Res, 66(10): 5039–5046. https://doi.org/10.1158/0008-5472.CAN-05-4194
Cheng YC, Shieh SY, 2018. Deubiquitinating enzyme USP3 controls CHK1 chromatin association and activation. Proc Natl Acad Sci USA, 115(21):5546–5551. https://doi.org/10.1073/pnas.1719856115
Chiruvella KK, Liang ZB, Wilson TE, 2013. Repair of doublestrand breaks by end joining. Cold Spring Harbor Perspect Biol, 5(5):a012757. https://doi.org/10.1101/cshperspect.a012757
Cohen P, Tcherpakov M, 2010. Will the ubiquitin system furnish as many drug targets as protein kinases? Cell, 143(5):686–693. https://doi.org/10.1016/j.cell.2010.11.016
Coleman KA, Greenberg RA, 2011. The BRCA1-RAP80 complex regulates DNA repair mechanism utilization by restricting end resection. J Biol Chem, 286(15): 13669–13680. https://doi.org/10.1074/jbc.M110.213728
Cooper EM, Cutcliffe C, Kristiansen TZ, et al., 2009. K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1. EMBO J, 28(6):621–631. https://doi.org/10.1038/emboj.2009.27
Cottarel J, Frit P, Bombarde O, et al., 2013. A noncatalytic function of the ligation complex during nonhomologous end joining. J Cell Biol, 200(2):173–186. https://doi.org/10.1083/jcb.201203128
Doil C, Mailand N, Bekker-Jensen S, et al., 2009. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell, 136(3):435–446. https://doi.org/10.1016/j.cell.2008.12.041
Dong YS, Hakimi MA, Chen XW, et al., 2003. Regulation of BRCC, a holoenzyme complex containing BRCA1 and BRCA2, by a signalosome-like subunit and its role in DNA repair. Mol Cell, 12(5):1087–1099. https://doi.org/10.1016/S1097-2765(03)00424-6
Durocher D, Jackson SP, 2001. DNA-PK, ATM and ATR as sensors of DNA damage: variations on a theme? Curr Opin Cell Biol, 13(2):225–231. https://doi.org/10.1016/S0955-0674(00)00201-5
Farshi P, Deshmukh RR, Nwankwo JO, et al., 2015. Deubiquitinases (DUBs) and DUB inhibitors: a patent review. Expert Opin Ther Pat, 25(10):1191–1208. https://doi.org/10.1517/13543776.2015.1056737
Feng L, Wang JD, Chen JJ, 2010. The Lys63-specific deubiquitinating enzyme BRCC36 is regulated by two scaffold proteins localizing in different subcellular compartments. J Biol Chem, 285(40):30982–30988. https://doi.org/10.1074/jbc.M110.135392
Gottlieb TM, Jackson SP, 1993. The DNA-dependent protein kinase: requirement for DNA ends and association with Ku antigen. Cell, 72(1):131–142. https://doi.org/10.1016/0092-8674(93)90057-W
Guervilly JH, Renaud E, Takata M, et al., 2011. USP1 deubiquitinase maintains phosphorylated CHK1 by limiting its DDB1-dependent degradation. Hum Mol Genet, 20(11): 2171–2181. https://doi.org/10.1093/hmg/ddr103
Gupta C, Heinen CD, 2019. The mismatch repair-dependent DNA damage response: mechanisms and implications. DNA Repair, 78:60–69. https://doi.org/10.1016/j.dnarep.2019.03.009
Hanpude P, Bhattacharya S, Dey AK, et al., 2015. Deubiquitinating enzymes in cellular signaling and disease regulation. IUBMB Life, 67(7):544–555. https://doi.org/10.1002/iub.1402
Harper JW, Elledge SJ, 2007. The DNA damage response: ten years after. Mol Cell, 28(5):739–745. https://doi.org/10.1016/j.molcel.2007.11.015
Harrigan JA, Jacq X, Martin NM, et al., 2018. Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat Rev Drug Discov, 17(1):57–78. https://doi.org/10.1038/nrd.2017.152
Harrison JC, Haber JE, 2006. Surviving the breakup: the DNA damage checkpoint. Annu Rev Genet, 40:209–235. https://doi.org/10.1146/annurev.genet.40.051206.105231
Hu X, Kim JA, Castillo A, et al., 2011. NBA1/MERIT40 and BRE interaction is required for the integrity of two distinct deubiquitinating enzyme BRCC36-containing complexes. J Biol Chem, 286(13):11734–11745. https://doi.org/10.1074/jbc.M110.200857
Hu YD, Scully R, Sobhian B, et al., 2011. RAP80-directed tuning of BRCA1 homologous recombination function at ionizing radiation-induced nuclear foci. Genes Dev, 25(7):685–700. https://doi.org/10.1101/gad.2011011
Huang XD, Dixit VM, 2016. Drugging the undruggables: exploring the ubiquitin system for drug development. Cell Res, 26(4):484–498. https://doi.org/10.1038/cr.2016.31
Hurley JH, Lee S, Prag G, 2006. Ubiquitin-binding domains. Biochem J, 399(Pt 3):361–372. https://doi.org/10.1042/BJ20061138
Ismail IH, Davidson R, Gagné JP, et al., 2014. Germline mutations in BAP1 impair its function in DNA doublestrand break repair. Cancer Res, 74(16):4282–4294. https://doi.org/10.1158/0008-5472.CAN-13-3109
Juang YC, Landry MC, Sanches M, et al., 2012. OTUB1 co-opts Lys48-linked ubiquitin recognition to suppress E2 enzyme function. Mol Cell, 45(3):384–397. https://doi.org/10.1016/j.molcel.2012.01.011
Ka HI, Lee S, Han S, et al., 2020. Deubiquitinase USP47-stabilized splicing factor IK regulates the splicing of ATM pre-mRNA. Cell Death Discov, 6:34. https://doi.org/10.1038/s41420-020-0268-1
Kakarougkas A, Jeggo PA, 2014. DNA DSB repair pathway choice: an orchestrated handover mechanism. Br J Radiol, 87(1035):20130685. https://doi.org/10.1259/bjr.20130685
Kakarougkas A, Ismail A, Katsuki Y, et al., 2013. Co-operation of BRCA1 and POH1 relieves the barriers posed by 53BP1 and RAP80 to resection. Nucleic Acids Res, 41(22): 10298–10311. https://doi.org/10.1093/nar/gkt802
Kapuria V, Peterson LF, Fang DX, et al., 2010. Deubiquitinase inhibition by small-molecule WP1130 triggers aggresome formation and tumor cell apoptosis. Cancer Res, 70(22): 9265–9276. https://doi.org/10.1158/0008-5472.CAN-10-1530
Karanam K, Kafri R, Loewer A, et al., 2012. Quantitative live cell imaging reveals a gradual shift between DNA repair mechanisms and a maximal use of HR in mid S phase. Mol Cell, 47(2):320–329. https://doi.org/10.1016/j.molcel.2012.05.052
Kato K, Nakajima K, Ui A, et al., 2014. Fine-tuning of DNA damage-dependent ubiquitination by OTUB2 supports the DNA repair pathway choice. Mol Cell, 53(4): 617–630. https://doi.org/10.1016/j.molcel.2014.01.030
Kawanishi S, Hiraku Y, Pinlaor S, et al., 2006. Oxidative and nitrative DNA damage in animals and patients with inflammatory diseases in relation to inflammation-related carcinogenesis. Biol Chem, 387(4):365–372. https://doi.org/10.1515/BC.2006.049
Kennedy RD, D’Andrea AD, 2005. The Fanconi Anemia/BRCA pathway: new faces in the crowd. Genes Dev, 19(24):2925–2940. https://doi.org/10.1101/gad.1370505
Kerzendorfer C, O’Driscoll M, 2009. Human DNA damage response and repair deficiency syndromes: linking genomic instability and cell cycle checkpoint proficiency. DNA Repair (Amst), 8(9):1139–1152. https://doi.org/10.1016/j.dnarep.2009.04.018
Khanna KK, Jackson SP, 2001. DNA double-strand breaks: signaling, repair and the cancer connection. Nature Genet, 27(3):247–254. https://doi.org/10.1038/85798
Kim H, Huang J, Chen JJ, 2007a. CCDC98 is a BRCA1-BRCT domain-binding protein involved in the DNA damage response. Nat Struct Mol Biol, 14(8):710–715. https://doi.org/10.1038/nsmb1277
Kim H, Chen JJ, Yu XC, 2007b. Ubiquitin-binding protein RAP80 mediates BRCA1-dependent DNA damage response. Science, 316(5828):1202–1205. https://doi.org/10.1126/science.1139621
Komander D, Clague MJ, Urbé S, 2009. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol, 10(8):550–563. https://doi.org/10.1038/nrm2731
Latif C, den Elzen NR, O’Connell MJ, 2004. DNA damage checkpoint maintenance through sustained Chk1 activity. J Cell Sci, 117(Pt 16):3489–3498. https://doi.org/10.1242/jcs.01204
Lee BH, Lee MJ, Park S, et al., 2010. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature, 467(7312):179–184. https://doi.org/10.1038/nature09299
Li FZ, Sun QQ, Liu K, et al., 2019. The deubiquitinase OTUD5 regulates Ku80 stability and non-homologous end joining. Cell Mol Life Sci, 76(19):3861–3873. https://doi.org/10.1007/s00018-019-03094-5
Li YH, Luo KT, Yin YJ, et al., 2017. USP13 regulates the RAP80-BRCA1 complex dependent DNA damage response. Nat Commun, 8:15752. https://doi.org/10.1038/ncomms15752
Lieber MR, 2008. The mechanism of human nonhomologous DNA end joining. J Biol Chem, 283(1):1–5. https://doi.org/10.1074/jbc.R700039200
Lindah T, Barnes DE, 2000. Repair of endogenous DNA damage. Cold Spring Harb Symp Quant Biol, 65:127–133. https://doi.org/10.1101/sqb.2000.65.127
Liu HL, Zhang HX, Wang XH, et al., 2015. The deubiquitylating enzyme USP4 cooperates with CtIP in DNA doublestrand break end resection. Cell Rep, 13(1):93–107. https://doi.org/10.1016/j.celrep.2015.08.056
Liu JL, Xia HG, Kim M, et al., 2011. Beclin1 controls the levels of p53 by regulating the deubiquitination activity of USP10 and USP13. Cell, 147(1):223–234. https://doi.org/10.1016/j.cell.2011.08.037
Liu ZX, Wu JX, Yu XC, 2007. CCDC98 targets BRCA1 to DNA damage sites. Nat Struct Mol Biol, 14(8):716–720. https://doi.org/10.1038/nsmb1279
Lu Q, Zhang FL, Lu DY, et al., 2019. USP9X stabilizes BRCA1 and confers resistance to DNA-damaging agents in human cancer cells. Cancer Med, 8(15):6730–6740. https://doi.org/10.1002/cam4.2528
Luo KT, Li L, Li YH, et al., 2016. A phosphorylation-deubiquitination cascade regulates the BRCA2-RAD51 axis in homologous recombination. Genes Dev, 30(23): 2581–2595. https://doi.org/10.1101/gad.289439.116
Mattiroli F, Vissers JHA, van Dijk WJ, et al., 2012. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell, 150(6):1182–1195. https://doi.org/10.1016/j.cell.2012.08.005
Meuth M, 2010. Chk1 suppressed cell death. Cell Div, 5:21. https://doi.org/10.1186/1747-1028-5-21
Mevissen TET, Hospenthal MK, Geurink PP, et al., 2013. OTU deubiquitinases reveal mechanisms of linkage specificity and enable ubiquitin chain restriction analysis. Cell, 154(1):169–184. https://doi.org/10.1016/j.cell.2013.05.046
Nakada S, Tai I, Panier S, et al., 2010. Non-canonical inhibition of DNA damage-dependent ubiquitination by OTUB1. Nature, 466(7309):941–946. https://doi.org/10.1038/nature09297
Nicholson B, Leach CA, Goldenberg SJ, et al., 2008. Characterization of ubiquitin and ubiquitin-like-protein isopeptidase activities. Protein Sci, 17(6):1035–1043. https://doi.org/10.1110/ps.083450408
Nijman SMB, Huang TT, Dirac AMG, et al., 2005a. The deubiquitinating enzyme USP1 regulates the Fanconi anemia pathway. Mol Cell, 17(3):331–339. https://doi.org/10.1016/j.molcel.2005.01.008
Nijman SMB, Luna-Vargas MPA, Velds A, et al., 2005b. A genomic and functional inventory of deubiquitinating enzymes. Cell, 123(5):773–786. https://doi.org/10.1016/j.cell.2005.11.007
Nishi R, Wijnhoven P, le Sage C, et al., 2014. Systematic characterization of deubiquitylating enzymes for roles in maintaining genome integrity. Nat Cell Biol, 16(10): 1016–1026. https://doi.org/10.1038/ncb3028
Nishi R, Wijnhoven PWG, Kimura Y, et al., 2018. The deubiquitylating enzyme UCHL3 regulates Ku80 retention at sites of DNA damage. Sci Rep, 8:17891. https://doi.org/10.1038/s41598-018-36235-0
Nowsheen S, Deng M, Lou ZK, 2020. Ubiquitin and the DNA double-strand break repair pathway. Genome Instab Dis, 1(2):69–80. https://doi.org/10.1007/s42764-019-00007-5
Olivieri M, Cho T, Álvarez-Quilón A, et al., 2020. A genetic map of the response to DNA damage in human cells. Cell, 182(2):481–496.e21. https://doi.org/10.1016/j.cell.2020.05.040
Orthwein A, Noordermeer SM, Wilson MD, et al., 2015. A mechanism for the suppression of homologous recombination in G1 cells. Nature, 528(7582):422–426. https://doi.org/10.1038/nature16142
Pellegrini L, Yu DS, Lo T, et al., 2002. Insights into DNA recombination from the structure of a RAD51-BRCA2 complex. Nature, 420(6913):287–293. https://doi.org/10.1038/nature01230
Peng YH, Liao QC, Tan W, et al., 2019. The deubiquitylating enzyme USP15 regulates homologous recombination repair and cancer cell response to PARP inhibitors. Nat Commun, 10:1224. https://doi.org/10.1038/s41467-019-09232-8
Pfeiffer A, Luijsterburg MS, Acs K, et al., 2017. Ataxin-3 consolidates the MDC1-dependent DNA double-strand break response by counteracting the SUMO-targeted ubiquitin ligase RNF4. EMBO J, 36(8):1066–1083. https://doi.org/10.15252/embj.201695151
Rehman SAA, Kristariyanto YA, Choi SY, et al., 2016. MINDY-1 is a member of an evolutionarily conserved and structurally distinct new family of deubiquitinating enzymes. Mol Cell, 63(1):146–155. https://doi.org/10.1016/j.molcel.2016.05.009
Riballo E, Kühne M, Rief N, et al., 2004. A pathway of double-strand break rejoining dependent upon ATM, Artemis, and proteins locating to γ-H2AX foci. Mol Cell, 16(5):715–724. https://doi.org/10.1016/j.molcel.2004.10.029
Rodriguez R, Meuth M, 2006. Chk1 and p21 cooperate to prevent apoptosis during DNA replication fork stress. Mol Biol Cell, 17(1):402–412. https://doi.org/10.1091/mbc.e05-07-0594
San Filippo J, Sung P, Klein H, 2008. Mechanism of eukaryotic homologous recombination. Annu Rev Biochem, 77: 229–257. https://doi.org/10.1146/annurev.biochem.77.061306.125255
Sartori AA, Lukas C, Coates J, et al., 2007. Human CtIP promotes DNA end resection. Nature, 450(7169):509–514. https://doi.org/10.1038/nature06337
Schmitt E, Paquet C, Beauchemin, M, et al., 2007. DNA-damage response network at the crossroads of cell-cycle checkpoints, cellular senescence and apoptosis. J Zhejiang Univ-Sci B (Biomed & Biotechnol), 8(6):377–397. https://doi.org/10.1631/jzus.2007.B0377
Schoenfeld AR, Apgar S, Dolios G, et al., 2004. BRCA2 is ubiquitinated in vivo and interacts with USP11, a deubiquitinating enzyme that exhibits prosurvival function in the cellular response to DNA damage. Mol Cell Biol, 24(17):7444–7455. https://doi.org/10.1128/MCB.24.17.7444-7455.2004
Scully R, Panday A, Elango R, et al., 2019. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nat Rev Mol Cell Biol, 20(11):698–714. https://doi.org/10.1038/s41580-019-0152-0
Shanbhag NM, Rafalska-Metcalf IU, Balane-Bolivar C, et al., 2010. ATM-dependent chromatin changes silence transcription in cis to DNA double-strand breaks. Cell, 141(6):970–981. https://doi.org/10.1016/j.cell.2010.04.038
Shao G, Lilli DR, Patterson-Fortin J, et al., 2009. The Rap80-BRCC36 de-ubiquitinating enzyme complex antagonizes RNF8-Ubc13-dependent ubiquitination events at DNA double strand breaks. Proc Natl Acad Sci USA, 106(9): 3166–3171. https://doi.org/10.1073/pnas.0807485106
Sharma A, Alswillah T, Kapoor I, et al., 2020. USP14 is a deubiquitinase for Ku70 and critical determinant of non-homologous end joining repair in autophagy and PTEN-deficient cells. Nucleic Acids Res, 48(2):736–747. https://doi.org/10.1093/nar/gkz1103
Sobhian B, Shao G, Lilli DR, et al., 2007. RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. Science, 316(5828):1198–1202. https://doi.org/10.1126/science.1139516
Su DX, Ma S, Shan L, et al., 2018. Ubiquitin-specific protease 7 sustains DNA damage response and promotes cervical carcinogenesis. J Clin Invest, 128(10):4280–4296. https://doi.org/10.1172/JCI120518
Sun YL, Jiang XF, Chen SJ, et al., 2005. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc Natl Acad Sci USA, 102(37):13182–13187. https://doi.org/10.1073/pnas.0504211102
Symington LS, Gautier J, 2011. Double-strand break end resection and repair pathway choice. Annu Rev Genet, 45:247–271. https://doi.org/10.1146/annurev-genet-110410-132435
Takeda S, Nakamura K, Taniguchi Y, et al., 2007. Ctp1/CtIP and the MRN complex collaborate in the initial steps of homologous recombination. Mol Cell, 28(3):351–352. https://doi.org/10.1016/j.molcel.2007.10.016
Typas D, Luijsterburg MS, Wiegant WW, et al., 2015. The de-ubiquitylating enzymes USP26 and USP37 regulate homologous recombination by counteracting RAP80. Nucleic Acids Res, 43(14):6919–6933. https://doi.org/10.1093/nar/gkv613
Uckelmann M, Densham RM, Baas R, et al., 2018. USP48 restrains resection by site-specific cleavage of the BRCA1 ubiquitin mark from H2A. Nat Commun, 9:229. https://doi.org/10.1038/s41467-017-02653-3
Wang B, Elledge SJ, 2007. Ubc13/Rnf8 ubiquitin ligases control foci formation of the Rap80/Abraxas/Brca1/Brcc36 complex in response to DNA damage. Proc Natl Acad Sci USA, 104(52):20759–20763. https://doi.org/10.1073/pnas.0710061104
Wang B, Matsuoka S, Ballif BA, et al., 2007. Abraxas and RAP80 form a BRCA1 protein complex required for the DNA damage response. Science, 316(5828):1194–1198. https://doi.org/10.1126/science.1139476
Wang XF, Liu ZY, Zhang L, et al., 2018. Targeting deubiquitinase USP28 for cancer therapy. Cell Death Dis, 9:186. https://doi.org/10.1038/s41419-017-0208-z
Wang ZQ, Zhang HL, Liu J, et al., 2016. USP51 deubiquitylates H2AK13, 15ub and regulates DNA damage response. Genes Dev, 30(8):946–959. https://doi.org/10.1101/gad.271841.115
Weake VM, Workman JL, 2008. Histone ubiquitination: triggering gene activity. Mol Cell, 29(6):653–663. https://doi.org/10.1016/j.molcel.2008.02.014
Welchman RL, Gordon C, Mayer RJ, 2005. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat Rev Mol Cell Biol, 6(8):599–609. https://doi.org/10.1038/nrm1700
Wiener R, Zhang XB, Wang T, et al., 2012. The mechanism of OTUB1-mediated inhibition of ubiquitination. Nature, 483(7391):618–622. https://doi.org/10.1038/nature10911
Wijnhoven P, Konietzny R, Blackford AN, et al., 2015. USP4 auto-deubiquitylation promotes homologous recombination. Mol Cell, 60(3):362–373. https://doi.org/10.1016/j.molcel.2015.09.019
Wrigley JD, Gavory G, Simpson I, et al., 2017. Identification and characterization of dual inhibitors of the USP25/28 deubiquitinating enzyme subfamily. ACS Chem Biol, 12(12):3113–3125. https://doi.org/10.1021/acschembio.7b00334
Wu JH, Chen YP, Geng GH, et al., 2019. USP39 regulates DNA damage response and chemo-radiation resistance by deubiquitinating and stabilizing CHK2. Cancer Lett, 449:114–124. https://doi.org/10.1016/j.canlet.2019.02.015
Wu ZQ, Qiu MH, Guo Y, et al., 2019. OTU deubiquitinase 4 is silenced and radiosensitizes non-small cell lung cancer cells via inhibiting DNA repair. Cancer Cell Int, 19:99. https://doi.org/10.1186/s12935-019-0816-z
Yang YF, Yang CZ, Li TT, et al., 2020. The deubiquitinase USP38 promotes NHEJ repair through regulation of HDAC1 activity and regulates cancer cell response to genotoxic insults. Cancer Res, 80(4):719–731. https://doi.org/10.1158/0008-5472.CAN-19-2149
Yu H, Pak H, Hammond-Martel I, et al., 2014. Tumor suppressor and deubiquitinase BAP1 promotes DNA double-strand break repair. Proc Natl Acad Sci USA, 111(1):285–290. https://doi.org/10.1073/pnas.1309085110
Yuan J, Luo KT, Deng M, et al., 2014. HERC2-USP20 axis regulates DNA damage checkpoint through Claspin. Nucleic Acids Res, 42(21):13110–13121. https://doi.org/10.1093/nar/gku1034
Zhang D, Zaugg K, Mak TW, et al., 2006. A role for the deubiquitinating enzyme USP28 in control of the DNA-damage response. Cell, 126(3):529–542. https://doi.org/10.1016/j.cell.2006.06.039
Acknowledgments
The research was supported by the National Natural Science Foundation of China (Nos. 91749115 and 81872298) and the Natural Science Foundation of Jiangxi Province (No. 20181BAB205044), China. The authors thank all Jian YUAN laboratory members for critical comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Yunhui LI participated in searching and summarizing the relevant literature as well as designing and writing the manuscript. Jian YUAN provided the theme and design, and edited the manuscript. Both authors have read and approved the final manuscript.
Corresponding author
Additional information
Compliance with ethics guidelines
Yun-hui LI and Jian YUAN declare that they have no conflicts of interest.
This article does not contain any studies with human or animal subjects performed by either of the authors.
Rights and permissions
About this article
Cite this article
Li, Y., Yuan, J. Role of deubiquitinating enzymes in DNA double-strand break repair. J. Zhejiang Univ. Sci. B 22, 63–72 (2021). https://doi.org/10.1631/jzus.B2000309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B2000309
Key words
- Deubiquitinating enzymes (DUBs)
- DNA double-strand breaks (DSBs)
- DNA repair
- Non-homologous end joining (NHEJ)
- Homologous recombination (HR)