-
PDF
- Split View
-
Views
-
Cite
Cite
Lin Ding, Susan L Forsburg, Essential Domains of Schizosaccharomyces pombe Rad8 Required for DNA Damage Response, G3 Genes|Genomes|Genetics, Volume 4, Issue 8, 1 August 2014, Pages 1373–1384, https://doi.org/10.1534/g3.114.011346
Close - Share Icon Share
Abstract
Schizosaccharomyces pombe Rad8 is a conserved protein homologous to S. cerevisiaeRad5 and human HLTF that is required for error-free postreplication repair by contributing to polyubiquitylation of PCNA. It has three conserved domains: an E3 ubiquitin ligase motif, a SNF2-family helicase domain, and a family-specific HIRAN domain. Data from humans and budding yeast suggest that helicase activity contributes to replication fork regression and template switching for fork restart. We constructed specific mutations in the three conserved domains and found that both the E3 ligase and HIRAN domains are required for proper response to DNA damage caused by a variety of agents. In contrast, mutations in the helicase domain show no phenotypes in a wild-type background. To determine whether Rad8 functionally overlaps with other helicases, we compared the phenotypes of single and double mutants with a panel of 23 nonessential helicase mutants, which we categorized into five phenotypic groups. Synthetic phenotypes with rad8∆ were observed for mutants affecting recombination, and a rad8 helicase mutation affected the HU response of a subset of recombination mutants. Our data suggest that the S. pombe Rad8 ubiquitin ligase activity is important for response to a variety of damaging agents, while the helicase domain plays only a minor role in modulating recombination-based fork restart during specific forms of replication stress.
Proper response to DNA damage during S phase requires that stalled replication forks are protected and efficiently restarted (Lambert and Carr 2013; Mirkin and Mirkin 2007). Evidence suggests that arrest and restart depend on management of single-stranded DNA and recombination. For example, on release from hydroxyurea (HU), which starves for nucleotides, S. pombe cells show a burst of foci of the single stranded DNA binding protein RPA, followed by foci of homologous recombination (HR) protein Rad52 (Carr and Lambert 2013; Meister et al. 2005; Sabatinos et al. 2012). This is consistent with HR involvement in fork restart (Bailis et al. 2008; Lambert et al. 2010; Meister et al. 2005). HR intermediates are processed and managed by specific helicases (e.g., Rqh1; Fml1) and nucleases (e.g., Mus81) to prevent inappropriate rearrangements (Dehe et al. 2013; Doe et al. 2002; Nandi and Whitby 2012; Sun et al. 2008; Willis and Rhind 2009).
There is good evidence from several systems that one form of fork restart works through a template switching pathway, with recovery through a Holliday junction-like structure (Atkinson and McGlynn 2009; Lambert and Carr 2013). Accumulation of RPA on ssDNA regulates this reaction (Betous et al. 2013; Sirbu et al. 2013). Several lines of evidence link the S. cerevisiaeRad5 to this activity. ScRad5 functions in the error-free branch of postreplication repair (PRR) by promoting the polyubiquitylation of PCNA (Branzei et al. 2004). It also has helicase-dependent fork reversal and restart activity, suggesting that it functions in fork regression downstream of PCNA ubiquitylation (Blastyak et al. 2007; Minca and Kowalski 2010). Cells with mutations that disrupt the ATP binding site in ScRAD5 are sensitive to DNA damage and disrupt the ligation of broken ends in an MRN-dependent pathway (Chen et al. 2005; Minca and Kowalski 2010). Scrad5 mutants are also HU-sensitive, consistent with a role in fork recovery (Kapitzky et al. 2010; Kats et al. 2009). Rad5 has two human homologs, SHPRH and HLTF (Figure 1B) (Unk et al. 2010). SHPRH promotes PCNA polyubiquitylation (Motegi et al. 2006b). HLTF facilitates template switching via its double-stranded DNA translocase activity (Blastyak et al. 2010). It also displaces RPA and PCNA from a modeled replication fork in vitro and contributes to Rad51-independent D-loop formation (Achar et al. 2011; Burkovics et al. 2014). SHPRH and HLTF respond to different forms of damage (Lin et al. 2011). Together, these data suggest that the Rad5/HLTF helicase domain contributes to replication fork stability and restart, possibly in an alternative pathway to Rad51.
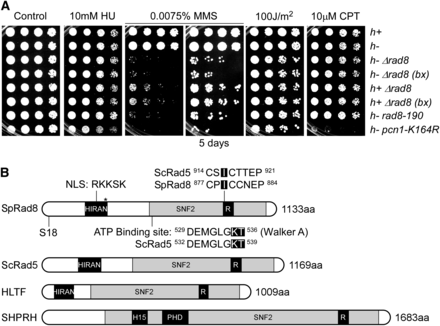
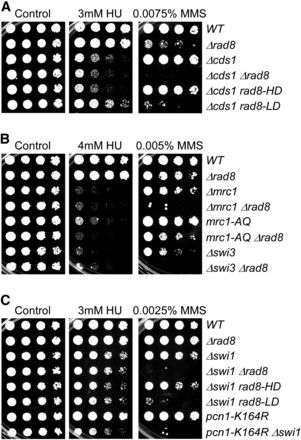
Rad8 is required for response to MMS-induced damage. (A) Representative response to MMS assessed by serial dilution. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 32° for 3 d unless otherwise indicated. bx = FY5627 was backcrossed once to FY528 (wild-type). These assays were repeated five times. (B) Schematic representation of Rad8 functional domains. S18 is a potential phosphorylation site. *Represents the protein product of rad8-190, which has a premature stop codon at amino acid 315. R, RING finger domain; H15, linker histone 1 and histone 5 domains; PHD, PHD-finger.
In S. pombe, the orthologous protein Rad8 (Figure 1B) has been shown to extend the ubiquitin chain on mono-ubiquitinated PCNA on K164 (Frampton et al. 2006), but the role of its ATP-dependent helicase domain has not been assessed. We performed a structure-function analysis of SpRad8. Surprisingly, and in contrast to data from budding yeast, we found no evidence for a role of the Rad8 ATPase domain in the response to replication stress in S. pombe in otherwise wild-type cells. Instead, the ubiquitin ligase domain is required for Rad8 to promote genome stability in response to a variety of stresses, suggesting that PCNA modification is required for multiple facets of genome maintenance. We used a candidate approach to investigate whether the putative Rad8 helicase overlaps with other helicase genes in S. pombe. An analysis of the drug sensitivity of a panel of nonessential helicase mutants allows identification of five distinct phenotypic groups, suggesting specialized helicase functions for specific types of replication stress. There are synthetic interactions between rad8∆ and a subset of genes involved in homologous recombination repair and recombination-mediated fork restart. The only phenotype associated with the helicase domain of Rad8 is in HU, and only in the absence of certain HR activities. This implicates PCNA modification in a wide range of damage response pathways, and suggests that fork regression or D-loop formation mediated by Rad8 helicase is only a minor player in replication fork restart in fission yeast.
Materials and Methods
Strain construction
All S. pombe strains were constructed and maintained in yeast extract plus supplement (YES) medium or under selection in Edinburgh minimal media (EMM) with appropriate supplements using standard techniques (Sabatinos and Forsburg 2010).
S. pombe strains (listed in Supporting Information, Table S1) were from our collection or purchased from the Bioneer Corporation and the Korea Research Institute of Biotechnology and Bioscience. The ∆rad8::kanMX-Bioneer (FY5132) deletion was isolated from the Bioneer S. pombe Deletion Mutant Library (V2-11-F11). It was backcrossed twice with laboratory wild-type strains. Both mating types were retained (FY5216: h+ or FY5217: h−). ∆rad8::hphMX (FY5625: h+ or FY5627: h−) was generated by replacing the kanamycin-resistant fragment of the Bioneer deletion with hygromycin B–resistant marker (Hentges et al. 2005). ∆rad8::hphMX or backcrossed ∆rad8::kanMX-Bioneer was used to construct double mutants. Strains were generated by tetrad dissection or random spore analysis.
Genomic DNA was extracted using a LioAc-SDS–based method that was derived and optimized from (Looke et al. 2011) for S. pombe. Briefly, in a 1.5-mL eppendorf tube, a single colony was resuspended in 100 μL LiOAc-SDS buffer (200 mM LiOAc, 1% SDS) and incubated at 70° for 15 min. Three volumes (300 μL) of 96% ethanol were added to the sample. After vortexing, the mixture was centrifuged at 15,000g for 3 min. The pellet was completely air-dried, resuspended in 100 μL TE, and centrifuged at 15,000g for 3 min. One microliter of the supernatant was used as template in a 20-μL colony PCR reaction.
Cloning of rad8+ gene and plasmid construction
The rad8+ gene was cloned in two steps. First, two independent fragments, one from ATG to a naturally occurring SpeI site within the ORF (rad8A) and the other from the SpeI site to the TAA codon (rad8B), were amplified by PCR and cloned into a pBlueScript vector. The XhoI site in the rad8B was silenced by site-directed mutagenesis. The complete rad8+ ORF was generated by ligating the rad8A fragment into rad8B-noXhoI containing plasmid at XhoI and SpeI. Mutations were introduced separately in rad8A or rad8B-noXhoI containing plasmids prior to generation of the full gene. Phusion Site-Directed Mutagenesis Kit (Finnzymes) was used for site-directed mutagenesis. Plasmids are listed in Table S2.
Constructing rad8 mutants at its endogenous locus
The rad8 mutants were created as described (Watson et al. 2008). The base strain (FY5622: h− Δrad8::loxP-ura4+-loxM3 ura4-D18 leu1-32 ade6-M210 can1-1) was constructed by replacing the entire rad8+ ORF (from start to stop codon) with the loxP-ura4+-loxM3 cassette that was amplified from the pAW1 plasmid (EUROSCARF-P30537) using primers TATACATGTTATTTTATATTTCTACAGTTTTTGGTAGCTTAAAGTTTGGATAAGCAAACATTACCAAGAAACTCAATAAACGGATCCCCGGGTTAATTAA and GTAGCAATTGCATTTCATATGCATAATATGAAAATACTTTTTTTTTACGATAGCTTTTAATCGGCTTGGTGAAACCGTTGGAATTCGAGCTCGTTTAAAC. Bold sequences indicate the sequences homologous to pAW1. The wild-type (pLD35) and mutated rad8 coding plasmids (pLD36, pLD37, pLD38, and pLD39) were digested with XhoI and SacI and cloned into the pAW8-XhoI plasmid (EUROSCARF-P30585). The resulting plasmids [pLD45: rad8+; pLD46: rad8-∆HIRAN; pLD47: rad8-K535A, T536A (rad8-HD); pLD48: rad8-I879A (rad8-LD); and pLD49: rad8-K535AT536AI879A (rad8-HDLD)] were transformed into the base strain and selected on EMM+Ade+thiamine plates. Transformants were grown in nonselective thiamine-free medium at 32° for 1 d, plated onto 5′-FOA plates, and incubated at 32° for 4 d. The 5-FOA–resistant and LEU− candidates were confirmed by colony PCR and verified by sequencing.
Serial dilution assays
Cells were grown to mid-log phase in YES. Five-fold serial dilutions were prepared in YES and spotted on drug containing rich medium. The plates were incubated 5 d at 25° or 3 d at 32°. Experiments were repeated two to five times.
Protein extracts and immunoblotting
Whole cell extracts (WCE) were prepared using trichloroacetic acid (TCA) extraction as described (Catlett and Forsburg 2003). Eighty micrograms of WCE were separated by SDS-PAGE in a 6% acrylamide gel. FLAG tag was detected with mouse anti-FLAG M2 (1:1000; Sigma) and anti-mouse-IgG-HRP (1:2000; Sigma). Mcm7 was detected with antibody purified from rabbit serum 6184 (1:1000) (Liang and Forsburg 2001) and anti-rabbit-HRP (1:2000; BD Biosciences).
GFP fusions
Two types of GFP fusion were made. For Western blot, the C-terminus of rad8+ was tagged with a GFP fragment at its native locus as described (Bahler et al. 1998). For microscopy (overexpression), linearized plasmids that carry rad8-GFP (under the nmt1 promoter), either wild-type or mutant derivatives, were integrated at leu1-32 locus in rad8∆ background. The nmt1 promoter was repressed in the presence of 15 μM thiamine (Maundrell 1993).
Microscopy
GFP tagged Rad8 strains were grown overnight in EMM-LEU containing 15 μM thiamine. Cells were harvested, washed twice with EMM-LEU, and released in thiamine-free medium to allow overexpression. Pictures were taken 16 hr after induction with a DeltaVision Core epifluorescence wide-field microscope (Applied Precision, WA) using GFP/mCherry Chroma ET C125705 filter (Ex 520/50; Em 630/80; polychroic mirror) and oil-immersion Olympus 60× lens (1.4 NA). Fifteen pictures of z-sections at 0.3 μm were captured, deconvolved, and then projected in softWoRx 5.5. The final figures were cropped and assembled in Canvas 12 (ACD Systems).
Results
Rad8 is required for response to a subset of genome damaging agents
To assess the role of Rad8 in damage response, we examined rad8∆ sensitivity to different genotoxins (Figure 1A) and compared this to a commonly used rad8 truncation allele, rad8-190, which we determined encodes a protein product with a premature stop codon at amino acid 315. We compared these to a nonubiquitinable mutant of pcn1 (pcn1-K164R) (Frampton et al. 2006). rad8∆ has no growth defect in the absence of drugs, and no evidence of a meiotic defect (data not shown). As seen previously, it is very sensitive to MMS and slightly sensitive to UV (Doe et al. 1993). rad8∆ MMS sensitivity is not as severe as pcn1-K164R, probably because the error-prone pathway that depends on single ubiquitylation of PCNA is still functional in rad8∆ (Frampton et al. 2006). Surprisingly, and in contrast to S. cerevisiae ScRad5, rad8∆ mutants are not sensitive to HU. We found that rad8∆ is also not sensitive to CPT, a topoisomerase toxin that causes S phase-specific DNA breaks (Wan et al. 1999). However, pcn1-K164R is very CPT-sensitive, possibly due to a sumoylation-mediated pathway involving the same residue (Kai et al. 2007).
Interestingly, h− ∆rad8 has increased sensitivity compared with h+ ∆rad8 when exposed to MMS (Figure 1A). Mutants lacking homologous recombination proteins often are sicker in h− than in h+ configuration, which is presumed to reflect the absence of a template for repair of the mating type imprint and break required for switching in the h− strain (Khasanov et al. 1999). This could suggest a role for Rad8 in aspects of HR repair. We compared h− strains throughout the remainder of this study to avoid differences attributable to mating type.
Domain structure of Rad8
S. pombe Rad8 was first identified as a member of the SNF2 helicase family based on sequence homology (Doe et al. 1993) and has distinct domains matching those in ScRad5 and HLTF (Minca and Kowalski 2010; Unk et al. 2010). There is a SNF2 helicase domain including an ATP binding site. Mutations that change the lysine and threonine to alanine (K538A, T539A) in the ATP binding site abolish the helicase activity in ScRad5 (Chen et al. 2005; Minca and Kowalski 2010). Embedded in the helicase domain, there is a RING-type Zinc finger ubiquitin E3 ligase domain that polyubiquitinates Pcn1 (ScPol30, hPCNA) in concert with the Mms2/Ubc13 E2 heterodimer (Frampton et al. 2006). A point mutation from isoleucine to alanine (I916A) in this motif in ScRad5 abolished the E3 ubiquitin ligase activity by eliminating its interaction with Ubc13 (Ulrich 2003). Near the N-terminus is an uncharacterized HIRAN domain, a motif shared by all the members of this family (HIP116, Rad5p N-terminal domain). It has been suggested that this domain recognizes specific DNA damage or stalled replication forks (Iyer et al. 2006), but it has not been analyzed. The Rad8 HIRAN domain spans amino acid 206 to 319 and contains a potential NLS (nuclear localization signal) RKKSK between amino acids 245 and 251. The rad8-190 truncation allele expresses most of this domain (amino acid 1-314) while deleting the helicase and RING-finger E3 ligase domains. We constructed mutations (Figure 1B) in all three of these conserved domains to assess their function in S. pombe. A large-scale fission yeast phosphorylation analysis (Wilson-Grady et al. 2008) suggested that a serine residue (S18) of Rad8 is a potential phosphorylation site. However, we observed no phenotype of an S18A mutation in DNA damage response, so this was not investigated further (Figure S1).
The HIRAN domain contributes to nuclear localization
To assess protein location, we tagged endogenous Rad8 with GFP at its C-terminus. Under the endogenous promoter, there was not sufficient signal to visualize the protein (data not shown). Therefore, we engineered overproduction strains to increase the GFP signal. We integrated Rad8-GFP at the at leu1+ locus, under the control of the nmt1+ promoter. This promoter allows modest levels of expression in thiamine, and a dramatic overproduction in the absence of thiamine (Forsburg 1993). We observed a distinctive Rad8-GFP signal in the nucleus within 16 hr of removing thiamine (Figure 2A). Prolonged Rad8-GFP overproduction over several days is slightly toxic (Figure S2A).
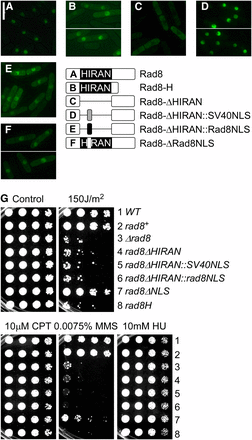
Nuclear localization is necessary but not sufficient for Rad8 function. (A–F) Schematic representation and localization of Rad8-GFP proteins, overproduced from a single copy transgene integrated at leu1-32 in ∆rad8 background. Pictures were taken 16 hr after the removal of thiamine: (A) Rad8-GFP; (B) Rad8-HIRAN-GFP; (C) Rad8-∆HIRAN; (D) Rad8-∆HIRAN::SV40NLS; (E) Rad8-∆HIRAN::rad8NLS; and (F) Rad8-∆NLS (with schematic representations of Rad8 HIRAN–related constructs on the right; not drawn to scale). (G) Drug sensitivity of indicated mutants. These mutants were integrated without a GFP tag at the native locus under the endogenous promoter. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. (1) Wild-type. (2) loxP-rad8+-loxM3. (3) loxP-∆rad8-loxM3. (4) loxP-rad8-∆HIRAN-loxM3. (5) loxP-rad8∆HIRAN::SV40NLS-loxM3. (6) loxP-rad8∆HIRAN::rad8NLS-loxM3. (7) loxP-rad8-∆NLS-loxM3. (8) loxP-rad8-HIRAN-loxM3. Plates were incubated at 32° for 3 d.
We constructed a 3′ truncation allele rad8-H mutant that expresses only the N-terminus of Rad8 up to the end of the HIRAN domain, and a HIRAN deletion mutant (rad8-∆HIRAN) that precisely deletes aa206-319. Rad8-H-GFP (integrated into the leu1+ locus under the nmt1 promoter in the rad8∆ strain) localizes in the nucleus (Figure 2B), whereas Rad8-∆HIRAN-GFP remains in the cytoplasm (Figure 2C). Next, we replaced the entire HIRAN domain with the SV40NLS (PKKKRKV) (Pasion and Forsburg 1999), and this restored the nuclear localization of Rad8 (Figure 2D). However, inserting the putative Rad8NLS (RKKSK) (Doe et al. 1993) in the same configuration only partially restored the nuclear localization (Figure 2E). A smaller deletion (amino acids 246 to 250) removed the RKKSK sequence (Rad8-∆NLS). The majority of this protein localized properly, indicating this is not the primary NLS (Figure 2F).
To test the drug sensitivity of the HIRAN mutants, we integrated them without GFP under the endogenous rad8+ promoter at the native locus using the Cre recombinase–mediated cassette exchange (RMCE) system (see Materials and Methods) (Watson et al. 2008). Both rad8-H and rad8-∆HIRAN phenocopy rad8∆ in damage sensitivity (Figure 2G). Importantly, restoration of nuclear localization with SV40NLS did not rescue the drug sensitivity. The Rad8-∆NLS strain was modestly sensitive to damage, consistent with the slight defect in localization (Figure 2G). Taken together, these results suggest that nuclear localization is necessary but not sufficient for Rad8 function, and that the HIRAN domain provides additional functions beyond nuclear localization.
Mutations in the E3 ligase and helicase domains have different phenotypes
Using the same strategies, we examined localization and drug sensitivity of Rad8 proteins containing point mutations in the putative helicase or ring finger domains. The point mutants correspond to the separation-of-function mutants in S. cerevisiae (Chen et al. 2005; Minca and Kowalski 2010). Similar to rad8+, the helicase-dead mutant rad8-K535A, T536A (rad8-HD), ubiquitin ligase-dead mutant rad8-I879A (rad8-LD), and the mutant with all three mutations rad8-K535AT536AI879A (rad8-HDLD) were all nuclear localized (Figure 3, A–D). The Pnmt1-rad8-GFP fully complemented ∆rad8 and cells mounted the same response to damaging drugs in the presence of thiamine (Figure S2B). However, both wild-type and mutant forms of Rad8 show a slight reduction in growth and increased sensitivity to drugs when strongly overproduced in thiamine-free medium (Figure S2C). This could be due to toxicity or a media effect. Interestingly, because we see a similar effect with both HD and LD mutants, this does not appear to be related to catalytic activity but could reflect a function for the HIRAN domain or a structural role.
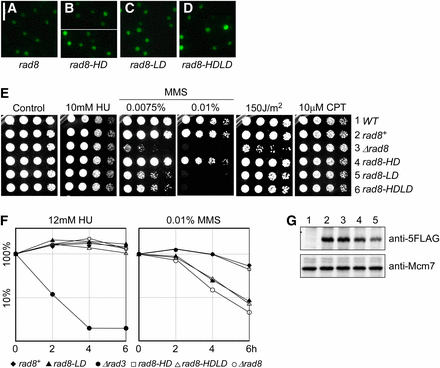
Ring finger is essential for Rad8 damage response. (A–D) Localization of Rad8-GFP proteins, overproduced from a single copy transgene integrated at leu1-32 in ∆rad8 background. Pictures were taken 16 hr after the removal of thiamine: (A) Rad8-GFP; (B) Rad8-HD-GFP; helicase dead; (C) Rad8-LD-GFP, ubiquitin ligase dead; and (D) Rad8-HDLD-GFP, double mutant. (E) Drug sensitivity of indicated mutants. These mutants were integrated without a GFP tag at the native locus under the endogenous promoter. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. (1) Wild-type. (2) loxP-rad8+-loxM3. (3) loxP-∆rad8-loxM3. (4) loxP-rad8-K535AT536A-loxM3. (5) loxP-rad8-I879A-loxM3. (6) loxP-rad8-K535AT536AI879A-loxM3. Plates were incubated at 32° for 3 d. (F) Representative relative survival curves of indicated mutants to acute drug exposure. HU (left), MMS (right). rad3Δ MMS survival curve was not plotted due to complete loss of viability at the 2-, 4-, and 6-hr time points. For each condition, two biological repeats were conducted. (G) Protein level of rad8 mutants. rad8 mutants were tagged with 5FLAG C terminally. Whole cell lysates were prepared using TCA extraction. Mcm7 was used as a loading control. Lane 1: wild-type; lane 2: rad8-5FLAG; lane 3: loxP-rad8-5FLAG; lane 4: loxP-rad8-HD-5FLAG; and lane 5: loxP-rad8-LD-5FLAG.
We integrated the mutants at the endogenous locus without the GFP tag and under the natural promoter. Surprisingly, and different from S. cerevisiae (Minca and Kowalski 2010), rad8-HD mutant did not affect growth in chronic or acute treatment with MMS (Figure 3 E and F). The Rad8 ubiquitin ligase domain proved to be the major contributor to all damage responses (Figure 3 E and F). The amount of Rad8 protein in these mutants is similar (Figure 3G).
ScRad5 channels the PRR to the error-free sub-pathway by polyubiquitinating Pcn1 (ScPol30/PCNA) and promoting fork reversal (Blastyak et al. 2007; Ramasubramanyan et al. 2010; Unk et al. 2010). Previous work with S. pombe showed that SpRad8 is required for PCNA polyubiquitylation (Frampton et al. 2006). We compared the phenotypes of rad8∆ and the rad8 point mutations in combination with different PRR mutants and examined their sensitivity to MMS, HU, and UV. As expected, rad8∆ is in a common epistasis group with rhp18∆ and mms2∆ (Figure S3A), whereas it has increased sensitivity to MMS when combined with mutations in the TLS polymerases in the error-prone arm of the pathway (Figure S3B).
As seen previously (Frampton et al. 2006), pcn1-K164R (which cannot be ubiquitylated) is epistatic to rad8∆. We observed a similar phenotype for rad8-LD, which lacks the ubiquitin ligase domain (Figure 4). Although we observe no obvious phenotype of the helicase mutant alone, the pcn1-K164R rad8-HD double mutant has reduced viability in response to MMS compared with pcn1-K164R rad8∆ (Figure 4). Moreover, pcn1-K164R rad8-HD is slightly sensitive to HU. This suggests that the helicase domain may become important in response to replication stress only if PCNA cannot be modified at K164.
A mutation in Rad8 ligase domain responsible for genetic interactions with pcn1-K164R. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 32° for 3 d unless otherwise indicated.
A damage fingerprint of helicase mutants
Given the observations with the budding yeast and human orthologs, we were surprised at the absence of any phenotypes associated with the rad8-HD mutant. We investigated whether Rad8 is redundant with other helicases in S. pombe, which has 23 annotated nonessential helicases. First, we isolated mutants available in the Bioneer deletion collection (Deshpande et al. 2009) and compared their responses to DNA-damaging drugs or treatment. Based on their patterns of sensitivity to different DNA damaging agents, we identified five distinct groups (summarized in Figure 5; see references in Table S3 and data in Figure 6, Figure 7, Figure S4, and Figure S5).
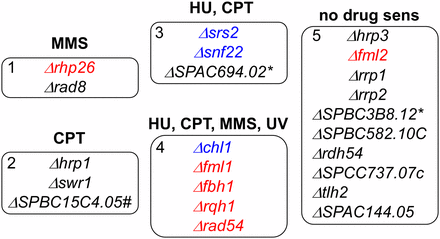
Nonessential helicases cluster in five phenotypic groups. Based on sensitivity to different genotoxins as seen in the serial dilution experiments (see Figure 6, Figure 7, Figure S3, and Figure S4 for data). Red font indicates strains that, when combined with Δrad8, show increased sensitivity to all damaging agents. Blue font indicates strains that, when combined with Δrad8, show mixed phenotypes of decreased and increased sensitivity to damaging agents. Black font indicates strains combined with Δrad8 show no sensitivity to damaging agents. SPBC3B8.12 = SPBC11C11.11C. #RNA/DNA helicase. *RNA helicase.
rad8 genetically interacts with genes involved in homologous recombination. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 32° for 3 d.
Rad8 ubiquitin ligase may contribute to resection. (A–C) Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 32° for 3 d. (D) and (E) Strains were grown overnight at 25°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 25° for 5 d.
The first group is specifically sensitive to MMS, consisting only of rad8∆ itself and rhp26∆, the ortholog of ScRad26 associated with transcription-coupled repair (Kanamitsu and Ikeda 2011). Group 2 mutants were only sensitive to CPT treatment, but not the other agents. This group included two chromatin remodelers and an uncharacterized RNA/DNA helicase. Group 3 contained snf22∆, another member of the SNF2 family, srs2∆, and an uncharacterized putative RNA helicase ∆SPAC694.02. They were sensitive to HU and CPT, which are specific to stress during S phase. The group 4 strains, sensitive to all agents tested, have been linked to aspects of replication fork stabilization and recombination. The last group of mutants showed no sensitivity to any of the agents we tested.
We examined the sensitivity of these mutations in combination with rad8∆ (summarized in Figure 5; see references in Table S4 and data in Figure 6, Figure 7, Figure S4, and Figure S5). Not surprisingly, the rhp26∆ rad8∆ double mutant was profoundly sensitive to MMS. There were no significant synthetic phenotypes observed in combination with group 2 mutants or the group 3 mutant SPAC694.02∆. Synthetic phenotypes were observed with most of the group 4 strains, generally showing increased sensitivity to all damaging agents (indicated in red). In a few cases, the double mutant improved growth on one agent (shown in blue): srs2∆ rad8∆ and snf22∆ rad8∆ show improved growth on HU (Figure 6E and Figure S4), whereas fml1∆ fml2∆ rad8∆ (Figure S5) and chl1∆ rad8∆ (Figure S4) both show improve growth on CPT.
We also examined conditional alleles of several essential helicases, including dna2 (discussed below) and pfh1, which is required to replicate though particular structures in the genome (Pinter et al. 2008; Sabouri et al. 2012; Steinacher et al. 2012; Tanaka et al. 2002). pfh1-R20 is a cold-sensitive allele (Boule and Zakian 2006). The pfh1-R20 rad8∆ mutant showed increased sensitivity to MMS and UV, no effect on CPT, and a partial rescue on HU (Table S4 and Figure S4).
Based on the results from this analysis, we took a candidate approach to investigate whether the rad8 helicase domain is responsible for any of the synthetic phenotypes we identified.
Identifying an effect of the Rad8-helicase mutant in recombination-deficient backgrounds
Homologous recombination is a central component of the response to replication stress (Bailis et al. 2008; Lambert et al. 2010; Meister et al. 2005). ScRad5 and human HLTF are implicated in formation of recombination structures between sister chromatids that can promote replication fork restart, independent of Rad51 (Burkovics et al. 2014; Hu et al. 2013; Minca and Kowalski 2010; Mott and Symington 2011). We examined mutations affecting Rad51, which coats an invading ssDNA filament (Heyer et al. 2010). Filament formation is promoted by mediator complexes including Rad55/Rad57 and Sfr1/Swi5, and is opposed by the helicase Srs2 (Haruta et al. 2008; Heyer et al. 2010)). Invasion and D loop formation are promoted by recruitment of the Rad54 helicase (Heyer et al. 2010).
Consistent with previous observations (Frampton et al. 2006), we find that rad51∆ rad8∆ is growth-impaired even without external genotoxins (Figure 6A). This growth defect was not as dramatic in the rad51∆ rad8-LD double mutant lacking the ubiquitin ligase domain and not apparent at all in the rad51∆ rad8-HD double mutant lacking the helicase motif, thus implicating the E3 ligase domain as an important contributor to genome stability in the absence of Rad51.
Both rad51∆ rad8∆ and in rad51∆ rad8-LD show a dramatic increase in MMS sensitivity, which is consistent with loss of two pathways of repair (template switching and recombination). Increased HU sensitivity was also observed for both rad51∆ rad8∆ and rad51∆ rad8-LD compared with rad51∆ (note the relatively low dose used, because rad51∆ is extremely HU-sensitive). Although the effect was modest, we observed a slight increase in MMS sensitivity in the rad51∆ rad8-HD double mutant. However, there was no synthetic phenotype on HU of rad51∆ rad8-HD.
We examined other members of the HR pathway: rad54∆, rad55∆, and rad57∆. On MMS, we saw similar phenotypes to those for rad51∆: a significantly increased sensitivity in double mutants with rad8∆ or rad8-LD lacking the ligase domain and a very slightly increased sensitivity with the rad8-HD mutation in the helicase domain (Figure 6, B–D). This suggests that they all act in a common pathway with Rad51 for MMS response, and this implicates the helicase domain for a minor contribution in the MMS response when the other pathways are compromised.
However, phenotypes of these additional rad mutants were all different on HU. First, rad54∆ is not quite as HU-sensitive as rad51∆, whereas rad55∆ and rad57∆ are not sensitive at all (Figure S6). Second, there was a strikingly increased HU sensitivity of rad54∆ in all the rad8 double mutants, suggesting that both ligase and helicase activity are important in the absence of Rad54. In contrast, rad55∆ showed increased HU sensitivity combined with rad8∆ or especially rad8-LD, but no effect of rad8-HD, whereas rad57∆ showed only a slight increase in sensitivity to the same extent in all the rad8 double mutants. These data suggest that Rad8 helicase activity may be specifically important for the response in HU if Rad54 is missing, and suggest separable roles of the Rad55/57 mediator components.
Srs2 helicase is an anti-recombinase that opposes Rad51 filament formation (Macris and Sung 2005). However, Srs2 also promotes fork reversal in repetitive sequences (Kerrest et al. 2009) and contributes to fork restart and template switching at stalled forks (Lambert et al. 2010). In MMS, Srs2 is implicated in restraint of the HR response to promote PRR in budding yeast, but not in fission yeast (Doe and Whitby 2004; Kai et al. 2007). We observed that rad8∆, srs2∆ rad8∆, and srs2∆ rad8-LD all showed similar sensitivity to MMS and UV, whereas srs2∆ rad8-HD resembled the more modest phenotype of srs2∆ (Figure 6E and Figure S4). Intriguingly, loss of rad8 partly suppressed the HU sensitivity observed in srs2∆, and this suppression was strongest in the srs2∆ rad8-HD double mutant lacking the helicase. This is opposite the phenotype observed for rad51∆ rad8, rad54∆ rad8, and rad55∆ rad8 double mutants, and suggests that the Rad8 helicase may antagonize Srs2.
Based on this result, we examined two additional proteins implicated in fork restart and HU response: the Mus81 resolvase and the Rqh1 helicase (Doe et al. 2002; Roseaulin et al. 2008). The double mutants with rad8∆ showed increased defects in growth on HU, MMS, and UV (Figure 6F and Figure S4).
Fml1 operates in an independent pathway
Another interacting helicase revealed in our screen is Fml1 (ScMph1/FANCM). This protein family is capable of fork reversal and promotes recombination at stalled replication forks (Blackford et al. 2012; Gari et al. 2008; Nandi and Whitby 2012; Prakash et al. 2005; Schurer et al. 2004; Sun et al. 2008; Zheng et al. 2011). Recent work suggests that budding yeast Mph1/FancM operates downstream of ScRad5 in repair of interstrand crosslinks (Daee et al. 2012). S. pombe has two Fml proteins: Fml1 and its paralogue, Fml2, which plays a minor role (Sun et al. 2008).
We examined the drug sensitivity of double and triple mutants (Figure S5). These showed no obvious growth defect on plate assays in the absence of replication stress, although the triple mutant was slightly elongated and grew more slowly in liquid media (data not shown). fml1∆ rad8∆ shows an increased sensitivity to MMS, UV, CPT, and HU, relative to both parents. The triple mutant fml1∆ fml2∆ rad8∆ is hypersensitive to MMS, UV, and HU, indicating a role for Rad8 in HU response when the Fml proteins are missing. pcn1-K164R fml1∆ fml2∆ phenocopies fml1∆ fml2∆ rad8∆. The rad8-HD mutant rescues the drug sensitivity of the fml1∆ fml2∆ rad8∆ triple mutant, and this suppression is abolished by pcn1-K164R, indicating that it is dependent on PCNA modification. Similar results were observed in HU, suggesting that the drug-sensitive phenotype of rad8∆ fml∆ mutants in all these cases is linked to ubiquitylation of PCNA.
Genetic evidence that Rad8 contributes to resection
Homologous recombination initiates from DNA ends, and broken ends are a target for resection by the MRN complex. There is evidence that ScRad5 binds to ssDNA as part of an end-joining pathway that involves the MRN complex (Chen et al. 2005). In contrast to the phenotype of Rad51 pathway members, we found only slight increase in MMS or UV sensitivity in double mutants between rad8∆ and some MRN components (rad50∆ and nbs1∆), suggesting a possible common function (Figure 7 and Figure S4).
MRN and the associated Ctp1 protein promote short-range resection, whereas long-range resection is promoted by Exo1 (Langerak et al. 2011). We observed that exo1∆ rad8∆ double mutants have a substantial increase in MMS sensitivity, as does the mre11∆ exo1∆ double mutant (Figure 7B). Similar results have been reported for rad50∆ exo1∆ (Tomita et al. 2003). The exo1∆ rad8-LD with a mutation in the E3 ligase domain was also more sensitive, whereas exo1∆ rad8-HD resembles exo1∆ alone. Finally, the exo1∆ rad8∆ and exo1∆ rad8-LD double mutants also showed increased sensitivity to hydroxyurea. These data suggest a role for Rad8 ubiquitin ligase in early stages of fork resection.
Long-range resection in budding yeast has also been linked to the ScSgs1 helicase and the Dna2 helicase/nuclease, in concert with RPA, and in opposition to fork regression (Cejka et al. 2010; H. Chen et al. 2013; Hu et al. 2012; Karanja et al. 2012; Niu et al. 2010). The corresponding pathway involved in fission yeast is reported to play only a minor role in resection (Langerak et al. 2011), but we examined interactions with mutations in dna2 (Figure 7, C and D and Figure S4) and rqh1 (ScSGS1) (Figure 6F and Figure S4). The dna2-K961T mutant lacks helicase activity (Hu et al. 2012), and the single mutant is not sensitive to DNA-damaging drugs at the dosages used here. The dna2-K961T rad8Δ double mutant is sensitive to HU and even more so to MMS (Figure 7C). dna2ts has defective nuclease activity, and dna2-K961T complements its growth (Hu et al. 2012). The dna2ts rad8Δ double mutant has slightly increased sensitivity to MMS (Figure 7D). In S. pombe, rad11+ is the essential gene that encodes the largest subunit of the trimeric RPA (Parker et al. 1997). Even at permissive temperatures, rad11Ats is sensitive to all genotoxins tested. rad8Δ increased rad11Ats MMS and UV sensitivity, but partly rescued its HU sensitivity (Figure 7E ad Figure S4).
Interactions with the fork protection complex
Finally, we examined activities known to be involved with stabilization of the replication fork and resolution of stalled structures, which are required for MMS response (Figure 8 and Figure S4). The checkpoint kinase Cds1 physically interacts with Mus81 and regulates its function by phosphorylation-induced chromatin dissociation (Boddy et al. 2000; Kai et al. 2005). Both Mus81 and Cds1 contribute to slowing replication in the presence of MMS (Willis and Rhind 2009).
Rad8 works in parallel with replication checkpoint and fork protection complex to maintain fork stability. Strains were grown overnight at 32°, 1:5 serially diluted, and spotted to plain YES rich medium (Control) and YES with indicated drugs. Plates were incubated at 32° for 3 d.
We observed that cds1∆ rad8∆ mutants have increased sensitivity to MMS (Figure 8A). The cds1∆ rad8∆ and cds1∆ rad8-LD showed a similar decline in viability, indicating that this sensitivity depends on the E3 ligase. Consistent with our other data, cds1∆ rad8-HD did not change the sensitivity of cds1∆ on MMS. Curiously, rad8-LD, but not rad8∆ or rad8-HD, modestly rescued both cds1∆ CPT and HU sensitivity (Figure 8A and Figure S4).
Mrc1, Swi1, and Swi3 form the fork protection complex (FPC), which stabilizes the replication fork in the presence of stress and is required for MMS response (Noguchi et al. 2004; Sommariva et al. 2005). In addition to the function in fork protection, Mrc1 is also a replication checkpoint adaptor protein that facilitates Cds1 activation mediated by Rad3-Rad26 (Tanaka and Russell 2001; Zhao and Russell 2004; Zhao et al. 2003). The replication checkpoint activity of Mrc1 is abolished in mrc1-AQ mutant (Xu et al. 2006). Deletion of rad8 increased the sensitivity of swi1∆, swi3∆, and mrc1∆ to MMS and UV (Figure 7, B and C and Figure S4). Loss of Rad8 ubiquitin ligase activity was the major contributor to these phenotypes. Consistent with this, there was a strong synthetic phenotype of pcn1-K164R swi1∆, consistent with the primary effect being through the ubiquitin ligase–mediated modification of PCNA.
Discussion
Several lines of evidence implicate the HTLF/Rad5 family of proteins in helicase or translocase function. Human HLTF is capable of displacing RPA and PCNA from a modeled replication fork in vitro and promotes fork regression (Achar et al. 2011; Blastyak et al. 2010). Recent evidence suggests it can promote formation of a D-loop independent of Rad51 or Rad54 (Burkovics et al. 2014). Helicase activity associated with ScRad5 is thought to promote fork regression downstream of PCNA ubiquitylation in the error-free PRR pathway (Blastyak et al. 2007; Minca and Kowalski 2010). ScRad5 helicase is also required to restrain duplication-associated rearrangements (Putnam et al. 2010). Mutations that disrupt the ATP binding site in ScRAD5 result in sensitivity to DNA damage (Chen et al. 2005; Minca and Kowalski 2010). However, in contrast to S. cerevisiae rad5 (Kapitzky et al. 2010; Kats et al. 2009), S. pombe rad8∆ mutants show no HU sensitivity (Figure 1A). This suggests that Rad8 is not required for response to, or recovery from, HU in otherwise wild-type cells; this may reflect the different requirements for HU response in these two fungi (Sabatinos et al. 2012).
We used a structure-function analysis to examine three conserved domains in S. pombe rad8+. Deletion of the HIRAN domain (found specifically in this family of proteins) disrupted nuclear localization and damage response in vivo. We restored localization by adding the nuclear localization sequence from SV40 large T antigen, but this did not restore the normal DNA damage response (Figure 2). The simplest conclusion is that nuclear localization is necessary but not sufficient for Rad8 function and the HIRAN domain makes a unique contribution to the damage response, although we cannot eliminate the possibility that our deletion has an indirect effect that disrupts protein structure.
We mutated the same residues as those shown in ScRAD5 to disrupt the highly conserved ATP-binding site of the SNF2-related helicase domain and the E3 ubiquitin ligase domain. (Chen et al. 2005; Minca and Kowalski 2010; Ulrich 2003). The strongest effects observed were linked to the E3-ligase domain mutation (Figure 3). Although the null was worse in some conditions, the rad8-HDLD mutant generally resembled the rad8-LD mutant, suggesting that there may be a noncatalytic, structural role associated with the physical presence of the protein. We found that the mutation of the E3-ligase domain (rad8-LD) was responsible for all of the damage sensitivity observed in rad8∆ single mutants. We found no evidence for the helicase domain functioning in otherwise wild-type cells.
We used the rad8 point mutations as separation-of-function alleles to see which domains were required for the phenotypes. Epistasis with other components of the PRR pathway gave results as expected for Rad8 playing an essential role in PCNA ubiquitylation. The phenotype of pcn1-K164R rad8∆ and pcn1-K164R rad8-LD on MMS were only slightly worse than the single mutant (Figure 4), suggesting that the predominant role of Rad8 ubiquitin ligase is via the Pcn1 K164 residue. Curiously, however, the pcn1-K164R rad8-HD double mutant was more sensitive to MMS than pcn1-K164R rad8∆ or pcn1-K164R alone, and showed a slight sensitivity to HU. Thus, there is a more complex genetic interaction between rad8 and pcn1 mutants than would be suggested by a simple linear epistasis model, but deciphering this exceeds the generally qualitative nature of serial dilution assays.
We screened candidate helicases to assess any evidence for functional overlap with Rad8. A panel of strains from the Bioneer collection or our collection, which disrupted nonessential helicases, was “fingerprinted” for damage sensitivity (Figure 6, Figure 7, Figure S4, and Figure S5). Some candidates are missing from the version of the Bioneer collection we used and are not represented here. A substantial number of the mutants had no sensitivity to the agents we tested (HU, MMS, CPT, and UV), either alone or in combination with rad8∆, and were not investigated further. Of the four remaining phenotypic groups, the first group is defined by rad8∆, and is sensitive primarily to MMS. It contains one other helicase mutant, rhp26∆, required for transcription-coupled repair (Kanamitsu and Ikeda 2011). Not surprisingly, a double mutant rad8∆ rhp26∆ showed dramatically increased sensitivity. The second group was only sensitive to CPT, an agent that causes S-phase–specific damage due to covalent coupling of topoisomerase to the ends of DNA (Wan et al. 1999). These mutants, which include several chromatin remodeling proteins, showed no synthetic interaction with rad8∆. Group 3 is represented by srs2∆, snf22∆, and a putative RNA helicase, and is only sensitive to CPT and HU, which cause defects specifically during S phase. We did not observe MMS sensitivity under these conditions with this allele of snf22∆, although there are other alleles that are reportedly MMS-sensitive (Dolan et al. 2010).
We observed a mixed interaction between rad8∆ and srs2∆ or snf22∆, in which MMS and CPT sensitivity were increased relative to the single mutants, but HU sensitivity was suppressed (Figure 6E and Figure S4), suggesting different roles for Rad8 in response to HU and MMS in these mutant backgrounds. The fourth group includes recombination-associated activities that are associated with replication fork restart. chl1∆ has genetic interactions with an alternative RFC complex and with the fork protection complex (Ansbach et al. 2008); the double mutant increases MMS sensitivity but decreases CPT sensitivity (Figure S4). The other mutants define genes linked to HR and replication fork stability (Figure 6, B and F, Figure S4, and Figure S5): Fml1 promotes recombination at stalled forks while preventing crossovers (Nandi and Whitby 2012; Sun et al. 2008); Fbh1 directly antagonizes Rad51 (Lorenz et al. 2009) and Rqh1 limits Rad51-mediated fork restart (Lambert et al. 2010). In contrast to these activities limiting recombination, Rad54 promotes Rad51-mediated strand invasion (Muris et al. 1997; Heyer et al. 2010) and stimulates fork regression (Bugreev et al. 2011).
There is a striking growth defect in rad51∆ rad8∆ strains that is not observed in double mutants with rad54∆, rad55∆, or rad57∆ (Figure 6, A–D). The sensitivity in rad51∆ can be attributed to the ubiquitin ligase domain, because rad51∆ rad8-LD, but not rad51∆ rad8-HD, showed increased sensitivity on HU. Because rad51∆ mutants show evidence of genome instability and DNA damage in the absence of damaging agents (Nakamura et al. 2008; Sabatinos et al. 2012), these results suggest that Rad8 ubiquitin ligase activity is important to respond to the intrinsic stress in rad51∆ mutants.
Double mutants between rad8∆ and rad54∆, rad55∆, or rad57∆ all had severe synthetic drug sensitivity to MMS treatment, and the same was observed for rad8-LD (Figure 6, A–D), which again was consistent with the central role of PCNA ubiquitylation in response to alkylation damage. However, the rad8-HD double mutants show modestly increased sensitivity relative to the HR single mutants, showing that there is a subtle contribution from the helicase domain when HR is affected.
Curiously, the results in HU were different for each of the separation of function alleles, suggesting that there are distinct differences between them. rad54∆ rad8-LD and rad54∆ rad8-HD had similar levels of sensitivity, which was almost as pronounced as rad54∆ rad8∆ (Figure 6B), suggesting there may be a structural role or function for the HIRAN domain even in the absence of a catalytic activity. We conclude that both catalytic domains of Rad8 are important in the absence of Rad54 in HU, which could implicate Rad8 helicase as an alternative mechanism to Rad54 activity in HU. The rad57∆ mutant was similar to rad54∆, although the degree of synthetic sensitivity was not as dramatic (Figure 6D). Surprisingly, because Rad55 and Rad57 work together (Liu et al. 2011; Sung 1997), rad55∆ rad8-LD and rad55∆ rad8∆ were both HU-sensitive but rad55∆ rad8-HD was not (Figure 6C). Together, these data suggest a role for the Rad8 helicase domain in the absence of the typical Rad51 recombination pathway, which may be related to the capacity of the human HLTF protein to produce D-loops in a Rad51-independent and Rad54-independent pathway (Burkovics et al. 2014). They also suggest that the Rad55 and Rad57 proteins may not contribute equally to replication fork restart in HU.
Given these results in HU, we examined other mutations that disrupt replication fork restart. Srs2 contributes to fork reversal in repetitive sequences (Kerrest et al. 2009), and to fork restart and template switching at stalled forks (Lambert et al. 2010; Lorenz et al. 2009). On MMS, there was no synthetic interaction in srs2∆ rad8∆ double mutants (Figure 6E). However, rad8-HD and rad8∆ strongly rescued srs2∆ when treated with HU. Because Srs2 antagonizes recombination, the opposite effects on HU sensitivity in the absence of rad8 are consistent with Rad8 promoting HR, and opposing Srs2. This would be consistent with the modest increase in sensitivity in the rad54∆ rad8 mutants. We propose that the helicase domain of Rad8 plays a minor role in Rad51-independent replication fork restart, particularly in HU.
The FANCM homolog Fml1 is already characterized as a translocase that regulates recombination at stalled forks (Nandi and Whitby 2012; Sun et al. 2008), and ICL repair by FANCM/ScMph1 depends on ScRad5 ubiquitin ligase activity (Daee et al. 2012). We observed synthetic phenotypes between rad8∆ and fml1∆ and showed these were linked to the E3-ligase domain (Figure S5). Similarly, double mutants between fork protection complex proteins and rad8∆ showed strikingly increased MMS sensitivity that depended on the E3 ligase domain and the PCNA K164 residue that is ubiquitylated. Again, phenotypes on HU suggest that in this condition, both Rad8 domains make a modest contribution to survival.
The requirement for the ubiquitin ligase activity of Rad8 is consistent with evidence that PCNA is required at multiple points of repair (Frampton et al. 2006). The modification of PCNA in the PRR pathway in budding yeast is linked to completion of replication (Branzei et al. 2004). The PRR pathway is also required to suppress gross chromosome rearrangements and repeat associated expansions in budding yeast (Daee et al. 2007; Motegi et al. 2006a; Putnam et al. 2010). PCNA also limits D-loop extension during recombination (Sebesta et al. 2013). During resection, PCNA promotes processivity of Exo1 (X. Chen et al. 2013) and promotes end-joining (Chen et al. 2005).
The initial stages of resection work through an MRN-Ctp1 complex (Limbo et al. 2007). In turn, this initial event promotes Exo1-mediated bulk resection (Mimitou and Symington 2008). We see no obvious phenotype in mre11∆ rad8∆ (Figure 7A and Figure S4), and little additional phenotype of a double mutant combining rad8∆ with the other components of the MRN complex suggest that they function in a common epistasis group. However, a strong MMS-sensitive phenotype was observed when rad8∆ or rad8-LD was combined with exo1∆ (Figure 7B). This implicates PCNA ubiquitylation in resection pathways independent of Exo1, perhaps indicating a contribution to early stages of resection. Recent work shows that the single-stranded DNA binding protein RPA also plays a significant role in promoting resection. We observe a pronounced sensitivity to MMS in a rad11ts rad8∆ double mutant (Figure 7E).
This study suggests that while the general pathways and enzymes for genome stability are similar across eukaryotes, the activities of specific DNA regulators have diverged within those modules. Among the SNF2-specific helicases, for example, metazoans have significantly expanded the family (Hauk and Bowman 2011). Our data suggest that the helicase function of S. pombe Rad8 is potentially required under specialized circumstances, as a minor redundant pathway with the HR proteins most notably in the restart of replication forks in HU, or in the absence of ubiquitin-PCNA. The primary function of this enzyme in fission yeast in the maintenance of genome stability appears to be due to its E3 ubiquitin ligase activity and the HIRAN domain, indicating a fundamental role for PCNA modification in preserving genomic integrity.
Acknowledgments
We thank members of the laboratory for helpful discussions and comments on the manuscript. This work was supported by NIH grants R01 GM059321 and R01 GM081418 (to S.L.F.), training support from NIH Cellular Biochemical, and Molecular Sciences Training Program at the USC Keck School of Medicine and USC-Dornsife College Doctoral fellowship (to L.D.).
Footnotes
Communicating editor: C. S. Hoffman
Literature Cited
Chen, X., Paudyal, S.C., Chin, R.I., and You, Z. 2013 PCNA promotes processive DNA end resection by Exo1. Nucleic Acids Res. 41: 9325–9338
Author notes
Supporting information is available online at http://www.g3journal.org/lookup/suppl/doi:10.1534/g3.114.011346/-/DC1