Abstract
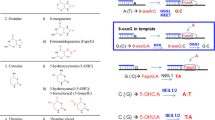
Cellular genomes suffer extensive damage from exogenous agents and reactive oxygen species formed during normal metabolism. The MutT homologs (MutT/MTH) remove oxidized nucleotide precursors so that they cannot be incorporated into DNA during replication. Among many repair pathways, the base excision repair (BER) pathway is the most important cellular protection mechanism responding to oxidative DNA damage. The 8-oxoG glycosylases (Fpg or MutM/OGG) and the MutY homologs (MutY/MYH) glycosylases along with MutT/MTH protect cells from the mutagenic effects of 8-oxoG, the most stable and deleterious product known caused by oxidative damage to DNA. The key enzymes in the BER process are DNA glycosylases, which remove different damaged bases by cleavage of the N-glycosylic bonds between the bases and the deoxyribose moieties of the nucleotide residues. Biochemical and structural studies have demonstrated the substrate recognition and reaction mechanism of BER enzymes. Cocrystal structures of strated the substrate recognition and reaction mechanism of BER enzymes. Cocrystal structures of several glycosylases show that the substrate base flips out of the sharply bent DNA helix and the minor groove is widened to be accessed by the glycosylases. To complete the repair after glycosylase action, the apurinic/apyrimidinic (AP) site is further processed by an incision step, DNA synthesis, an excision step, and DNA ligation through two alternative pathways. The short-patch BER (1-nucleotide patch size) and long-patch BER (2–6-nucleotide patch size) pathways need AP endonuclease to generate a 3′ hydroxyl group but require different sets of enzymes for DNA synthesis and ligation. Protein-protein interactions have been reported among the enzymes involved in BER. It is possible that the successive players in the repair pathway are assembled in a complex to perform concerted actions. The BER pathways are proposed to protect cells and organisms from mutagenesis and carcinogenesis.
Similar content being viewed by others
References
Halliwell, B. and Gutteridge, J. M. C. (1989) Free Radicals in Biology and Medicine. New York, Oxford University Press.
Esterbauer, H., Eckl, P., and Ortner, A. (1990) Possible mutagens derived from lipids and lipid precursors. Mutat. Res. 238, 223–233.
Fraga, C. G., Shigenaga, M. K., Park, J.-W. Degan, P., and Ames, B. N. (1990) Oxidative damage to DNA during aging: 8-hydroxy-2′-deoxyguanosine in rat organ DNA and urine. Proc. Natl. Acad. Sci. USA 87, 4533–4537.
Ames, B. N. and Shigenaga, M. K. (1993) Oxidants are a major contributor to cancer and aging, in DNA and Free Radicals Halliwell, B. and Aruoma, O. (eds.), New York, Ellis Horwood, pp. 1–18.
Wallace, D. C., Schoffner, J. M. Trounce, I., Brown, M. D., Ballinger, S. W., Corral-Debrinski, M., et al. (1995) Mitochondrial DNA mutations in human degenerative diseases and aging. Biochem. Biophys. Acta 1271, 141–151.
Wiseman, H. and Halliwell, B. (1996) Damage to DNA by reactive oxygen and nitrogen species: role in inflammatory disease and progression to cancer. Biochem. J. 313, 17–29.
Shibutani, S., Takeshita, M., and Grollman, A. P. (1991) Insertion of specific bases during DNA synthesis past the oxidation-damaged base 8-oxodG. Nature 349, 431–434.
Cheng, K. C., Cahill, D. S., Kasai, H., Nishimura, S., and Loeb, L. A. (1991) 8-Hydroxyguanine, an abudant form of oxidative DNA damage, causes G-T and A-C substitutions. J. Biol. Chem., 267, 166–172.
Michaels, M. L. and Miller, J. H. (1992) The GO system protects organisms from the mutagenic effect of the spontaneous lesion 8-hydroxyguanine (7,8-dihydro-8-oxo-guanine). J. Bacteriol. 174, 6321–6325.
Moriya, M. (1993) Single-stranded shuttle phagemid for mutagenesis studies in mammalian cells: 8-oxoguanine in DNA induces targeted G:C to T:A transversions in simian kidney cells. Proc. Natl. Acad. Sci. USA 90, 1122–1126.
Wood, M. L., Dizdaroglu, M., Gajewski, E., and Essigmann, J. M. (1990) Mechanistic studies of ionizing radiation and oxidative mutagenesis: genetic effects of single 8-hydroxyguanine (7-hydro-8-oxoguanine) residue inserted at a unique site in a viral genome. Biochemistry 29, 7024–7032.
Swanson, R. L., Morey, N. J., Doetsch, P. W., and Jinks-Robertson, S. (1999) Overlapping specificities of base excision repair, nucleotide excision repair, recombination, and translesion synthesis pathways for DNA base damage in Saccharomyces cerevisiae. Mol. Cell Biol. 19 2929–2935.
McCullough, A. K., Dodson, M. L., and Lloyd, R. S. (1999) Initiation of base excision repair: glycosylase mechanisms and structures. Annu. Rev. Biochem. 68, 255–285.
Akiyama, M., Maki, H., Sekiguchi, M., and Horiuchi, T. (1989) A specific role of MutT protein: to prevent dGdA mispairing in DNA replication. Proc. Natl. Acad. Sci. USA 86, 3949–3952.
Maki, H. and Sekiguchi, M. (1992) MutT protein specifically hydrolyses a potent mutagenic substrate for DNA synthesis. Nature 355, 273–275.
Tchou, J. and Grollman, A. P. (1993) Repair of DNA containing the oxidatively-damaged base 8-hydroxyguanine. Mutat. Res. 299, 277–287.
Chetsanga, C. J. and Lindahl, T. (1979) Release of 7-methylguanine residues whose imidazole rings have been opened from damaged DNA by a DNA glycosylase from Escherichia coli. Nucleic Acids Res. 6, 3673–3683.
Tchou, J., Kasai, H., Shibutani, S., Chung, M.-H., Grollman, A. P., and Nishimura, S. (1991) 8-Oxoguanine (8-hydroxyguanine) DNA glycosylase and its substrate specificity. Proc. Natl. Acad. Sci. USA 88, 4690–4694.
Michaels, M. L., Cruz, C., Grollman, A. P., and Miller, J. H. (1992) Evidence that MutM and MutY combine to prevent mutations by an oxidatively damaged form of guanine in DNA. Proc. Natl. Acad. Sci. USA 89, 7022–7025.
Ohtsubo, T., Nishioka, K., Imaiso, Y., Iwai, S., Shimokawa, H., Oda, H., et al. (2000) Identification of human MutY homolog (hMYH) as a repair enzyme for 2-hydroxyadanine in DNA and detection of multiple forms of hMYH located in nuclei and mitochondria. Nucleic Acids Res. 28, 1355–1364.
Slupska, M. M., Baikalov, C., Luther, W. M., Chiang, J.-H., Wei, Y.-F., and Miller, J. H. (1996) Cloning and sequencing a human homolog (hMYH) of the Escherichia coli mutY gene whose function is required for the repair of oxidative DNA damage. J. Bacteriol. 178, 3885–3892.
Slupska, M. M., Luther, W. M., Chiang, J. H., Yang, H., and Miller, J. H. (1999) Functional expression of hMYH, a human homolog of the Escherichia coli MutY protein. J. Bacteriol. 181, 6210–6213.
Takao, M., Zhang, Q. M., Yonei, S., and Yasui, A. (1999) Differential subcellular localization of human MutY homolog (hMYH) and the functional activity of adenine: 8-oxoguanine DNA glycosylase. Nucleic Acids Res. 27, 3638–3644.
Yeh, Y.-C., Chang, D.-Y., Masin, J., and Lu, A. L. (1991) Two nicking enzymes systems specific for mismatch-containing DNA in nuclear extracts from human cells. J. Biol. Chem. 266, 6480–6484.
Boiteux, S. and Radicella, J. P. (1999) Base excision repair of 8-hydroxyguanine, protects DNA from endogenous oxidative stress. Biochimie 81, 59–67.
Radicella, J. P., Dherin, C., Desmaze, C., Fox, M. S., and Boiteux, S. (1997) Cloning and characterization of hOGG1, a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 94, 8010–8015.
Roldan-Arjona, T., Wei, Y.-F., Carter, K. C., Klungland, A., Anselmino, C., Wang, R.-P. et al. (1997) Molecular cloning and functional expression of a human cDNA encoding the antimutator enzyme 8-hydroxyguanine-DNA glycosylase. Proc. Natl. Acad. Sci. USA 94, 8016–8020.
Shinmura, K., Kasai, H., Sasaki, A., Sugimura, H., and Yokota, J. (1997) 8-hydroxyguanine (7,8-dihydro-8-oxoguanine) DNA glycosylase and AP lyase activities of hOGG1 protein and their substrate specificity. Mutat. Res. 385, 75–82.
McLennan, A. G. (1999) The MutT motif family of nucleotide phosphohydrolases in man and human pathogens. Int. J. Mol. Med. 4, 79–89.
Mo, J.-Y., Maki, H., and Sekiguchi, M. (1992) Hydrolytic elimination of a mutagenic nucleotide, 8-oxodGTP, by human 18-kilodalton protein: sanitization of nucleotide pool. Proc. Natl. Acad. Sci USA 89, 11,021–11,025.
Sakumi, K., Furuichi, M., Tsuzuki, T., Kakuma, T., Kawabata, S.-I., Maki, H., et al. (1993) Cloning and expression of cDNA for a human enzyme that hydrolyzes 8-oxo-dGTP, a mutagenic substrate for DNA synthesis. J. Biol. Chem. 268, 23,524–23,530.
Bhatnagar, S. and Bessman, M. J. (1988) Studies on the mutator gene, mutT of Escherichia coli: molecular cloning of the gene, purification of the gene product, and identification of a novel nucleoside triphosphatase. J. Biol. Chem. 263, 8953–8957.
Tajiri, T., Maki, H., and Sekiguchi, M. (1995) Functional cooperation of MutT, MutM and MutY proteins in preventing mutations caused by spontaneous oxidation of guanine nucleotide in Escherichia coli. Mutat. Res. 336, 257–267.
Fujii, Y., Shimokawa, H., Sekiguchi, M., and Nakabeppu, Y. (1999) Functional significance of the conserved residues for the 23-residue module among MTH1 and MutT family proteins. J. Biol. Chem. 274, 38,251–38,259.
Shimokawa, H., Fujii, Y., Furuichi, M., Sekiguchi, M., and Nakabeppu, Y. (2000) Functional significance of conserved residues in the phosphohydrolase module of Escherichia coli MutT protein. Nucleic Acids Res. 28, 3240–3249.
Kang, D., Nishida, J., Iyama, A., Nakabeppu, Y., Furuichi, M., Fujiwara, T., et al. (1995) Intracellular localization of 8-oxo-dGTPase in human cells, with special reference to the role of the enzyme in mitochondria. J. Biol. Chem. 270, 14,659–14,665.
Fujikawa, K., Kamiya, H., Yakushiji, H., Fuiji, Y., Nakabeppu, Y., and Kasai, H. (1999) The oxidized forms of dATP are substrates for the human MutT homologue, the hMTH1 protein. J. Biol. Chem. 274, 18,201–18,205.
Mildvan, A. S., Weber, D. J., and Abeygunwardana, C. (1999) Solution structure and mechanism of the MutT pyrophos-phohydrolase. Adv. Enzymol. Relat. Areas Mol. Biol. 73, 183–207.
Abeygunawardana, C., Weber, D. J., Gittis, A. G., Frick, D. N., Lin, J., Miller, A. F., et al. (1995) Solution structure of the MutT enzyme, a nucleoside triphosphate pyrophosphohydrolase. Biochemistry 34, 14,997–15,005.
David, S. S. and Williams, S. D. (1998) Chemistry of glycosylase and endonuclease involved in base-excision repair. Chem. Rev. 98, 1221–1261.
Mol, C. D., Parikh, S. S., Putnam, C. D., Lo, T. P., and Tainer, J. A. (1999) DNA repair mechanisms for the recognition and removal of damaged DNA bases. Annu. Rev. Biophys. Biomol. Struct. 28, 101–128.
Wallace, S. S. (1998) Enzymatic processing of radiation-induced free radical damage in DNA. Radiat. Res. 150, S60-S79.
Cunningham, R. P. (1997) DNA glycosylases. Mutat. Res. 383, 189–196.
Hazra, T. K., Izumi, T., Venkataraman, R., Kow, Y. W., Dizdaroglu, M., and Mitra, S. (2000) Characterization of a novel 8-oxoguanine-DNA glycosylase activity in Escherichia coli and identification of the enzyme as endonuclease VIII. J. Biol. Chem. 275, 27,762–27,767.
Jiang, D., Hatahet, Z., Melamede, R. J., Kow, Y. W., and Wallace, S. S. (1997) Characterization of Escherichia coli endonuclease VIII. J. Biol. Chem. 272, 32,230–32,239.
Zhang, Q. M., Miyabe, I., Matsumoto, Y., Kino, K., Sugiyama, H., and Yonei, S. (2000) Identification of repair enzymes for 5-formyluracil in DNA. Nth, nei, and MutM proteins of Escherichia coli. J. Biol. Chem. 275, 35,471–35,477.
Bruner, S. D., Nash, H. M., Lane, W. S., and Verdine, G. L. (1998) Repair of oxidatively damaged guanine in Saccharomyces cerevisine by an alternative pathway. Curr. Biol. 8, 393–403.
You, H. J., Swanson, R. L., and Doetsch, P. W. (1998) Saccharomyces cerevisiae possesses two functional homologues of Escherichia coli endonuclease III. Biochemistry 37, 6033–6040.
You, H. J., Swanson, R. L., Harrington, C., Corbett, A. H., Jinks-Robertson, S., Senturker, S., et al. (1999) Saccharomyces cerevisiae Ntg1p and Ntg2p: broad specificity N-glycosylases for the repair of oxidative DNA damage in the nucleus and mitochondria. Biochemistry 38, 11,298–11,306.
Asagoshi, K., Yamada, T., Okada, Y., Terato, H., Ohyama, Y., Seki S., et al. (2000) Recognition of formamidopyrimidine by Escherichia coli and mammalian thymine glycol glycosylases. Distinctive paired base effects and biological and mechanistic implications. J. Biol. Chem. 275, 24,781–24,786.
Asagoshi, K., Odawara, H., Nakano, H., Miyano, T., Terato, H., Ohyama, Y., et al. (2000) Comparison of substrate specificities of Escherichia coli endonuclease III and its mouse homologue (mNTH1) using defined oligonucleotide substrates. Biochemistry 39, 11,389–11,398.
Hilbert, T. P., Boorstein, R. J., Kung, H. C., Bolton, P. H., Xing, D., Cunningham, R. P., et al. (1996) Purification of a mammalian homologue of Escherichia coli endonyclease III: identification of a bovine pyrimidine hydrate-thymine glycol DNA-glycosylase/AP lyase by irreversible cross linking to a thymine glycol-containing oligonucleotide. Biochemistry 35, 2505–2511.
Zhang, Q. M., Sugiyama, H., Miyabe, I., Matsuda, S., Saito, I., and Yonei, S. (1997) Replication of DNA templates containing 5-formyluracil, a major oxidative lesion of thymine in DNA. Nucleic Acids Res. 25, 3969–3973.
Bjelland, S., Birkeland, N. K., Benneche, T., Volden, G., and Seeberg, E. (1994) DNA glycosylase activities for thymine residues oxidized in the methyl group are functions of the AlkA enzyme in Escherichia coli. J. Biol. Chem. 269, 30,489–30,495.
Masaoka, A., Terato, H., Kobayashi,M., Honsho, A., Ohyama, Y., and Ide, H. (1999) Enzymatic repair of 5-formyluracil. I. Excision of 5-formyluracil site-specifically incorporated into oligonucleotide substrates by AIKA protein (Escherichia coli 3-methyladenine DNA glycosylase II). J. Biol. Chem. 274, 25,136–25,143.
Hatahet, Z., Kow, Y. W., Purmal, A. A., Cunningham, R. P., and Wallace, S. S. (1994) New substrates for old enzymes: 5-hydroxy-2′-deoxycytidine and 5-hydroxy-2′deoxyuridine are substrates for Escherichia coli endinuclease III and formamidopyrimidine DNA N-glycosylase, while 5-hydroxy-2′-deoxyuidine is a substrate for uracil DNA N-glycosylase. J. Biol. Chem. 269, 18,814–18,820.
Purmal, A. A., Lampman, G. W., Bond, J. P., Hatahet, Z., and Wallace, S. S. (1998) Enzymatic processing of uracil glycol, a major oxidative product of DNA cytosine. J. Biol. Chem. 273, 10,026–10,035.
Castaing, B., Boiteux, S., and Zelwer, C. (1992) DNA containing a chemically reduced apurinic site is a high affinity ligand for the E. coli formamidopyrimidine-DNA glycosylase. Nucleic Acids Res. 20, 389–394.
Blaisdell, J. O., Hatahet, Z., and Wallace, S. S. (1999) A novel role for Escherichia coli endonuclease VIII in prevention of spontaneous G→T transversions. J. Bacteriol. 181, 6396–6402.
Dherin, C., Radicella, J. P., Dizdaroglu, M., and Boiteux, S. (1999) Excision of oxidatively damaged DNA bases by the human alpha-hOgg1 protein and the polymorphic alpha-hOgg1 (Ser326Cys) protein which is frequently found in human populations. Nucleic Acids Res. 27, 4001–4007.
Aburatani, H., Hippo, Y., Ishida, T., Takashima, R., Matsuba, C., Kodama, T., et al. (1997) Cloning and characterization of mammalian 8-hydroxy guanine-specific DNA glycosylase/apurinic, apyrimidinic lyase, a functional MutM homologue. Cancer Res. 57, 2151–2156.
Lu, R., Nash, H. M., and Verdine, G. L. (1997) A mammalian DNA repair enzyme that excises oxidatively damaged guanines maps to a locus frequently lost in lung cancer. Curr. Biol. 7, 397–407.
Takao, M., Aburatani, H., Kobayashi, K., and Yasui, A. (1998) Mitochondrial targeting of human DNA glycosylases for repair of exidative DNA damage. Nucleic Acids Res. 26, 2917–2922.
Nash, H. M., Lu, R., Lane, W. S., and Verdine, G. L. (1997) The critical active-site amine of human 8-oxoguanine DNA glycosylase, hOgg1: direct indentification, ablation and chemical reconstitution. Chem. Biol. 4, 693–702.
Bruner, S. D., Norman, D. P., and Verdine, G. L. (2000) Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866.
Nash, H. M., Bruner, S. D., Scharer, O. D., Kawate, T., Addona, T. A., Spooner, E.,et al. (1996) Cloning of a yeast 8-oxoguanine DNA glycosylase reveals the existence of a base-excision DNA-repair protein superfamily. Curr. Biol. 6, 968–980.
Hazra, T. K., Izumi, T., Maidt, L., Floyd, R. A., and Mitra, S. (1998) The presence of two distinct 8-oxoguanine repair enzymes in human cells: their potential complementary roles in preventing mutation. Nucleic Acids Res. 26, 5116–5122.
Au, K. G., Cabrera, M., Miller, J. H., and Modrich, P. (1988) Escherichia coli mutY gene product is required for specific A/G to C: G mismatch correction. Proc. Natl. Acad. Sci. USA 85, 9163–9166.
Lu, A-L. and Chang, D.-Y. (1988) Repair of single base pair transversion mismatches of Escherichia coli in vitro: correction of certain A/G mismatch is independent of dam methylation and host mutHLS gene functions. Genetics 118, 593–600.
Lu, A.-L. and Chang, D.-Y. (1988) A novel nucleotide excision repair for the conversion of an A/G mismatch to C/G base pair in E. coli. Cell 54, 805–812.
Radicella, J.P., Clark, E. A., and Fox, M. S. (1988) Some mismatch repair activities in Escherichia coli. Proc. Natl. Acad. Sci. USA 85, 9674–9678.
Su, S.-S., Lahue, R. S., Au, K. G., and Modrich, P. (1988) Mispair specificity of methyl-directed DNA mismatch correction in vitro. J. Biol. Chem. 263, 6829–6835.
Li, X., Wright, P. M., and Lu, A.-L. (2000). The C-terminal domain of MutY glycosylase determines the 7,8-dihydro-8-oxo-guanine specificity and is crucial for mutation avoidance. J. Biol. Chem. 275, 8448–8455.
Zhang, Q. M., Ishikawa, N., Nakahara, T., and Yonei, S. (1998) Escherichia coli MutY protein has a guanine-DNA glycosylase that acts on 7,8-dihydro-8-oxoguanine: guanine mispair to prevent spontaneous G: C to C: G transversions. Nucleic Acids Res. 26, 4669–4675.
Michaels, M. L., Pham, L., Nghiem, Y., Cruz, C., and Miller, J. H. (1990) MutY, an adenine glycosylase active on G-A mispairs, has homology to endonuclease III. Nucleic Acids Res. 18, 3841–3845.
Tsai-Wu, J.-J., Liu, H.-F., and Lu, A.-L. (1992) Escherichia coli MutY protein has both N-glycosylase and apurinic/apyrimidinic endonuclease activities on A: C and A: G mispairs. Proc. Natl. Acad. Sci. USA 89, 8779–8783.
Gogos, A., Cillo, J., Clarke, N. D., and Lu, A-L., (1996) Specific recognition of A/G and A/8-oxoG mismatches by Escherichia coli MutY: removal of the C-terminal domain preferentially affects A/8-oxoG recognition. Biochemistry 35, 16,665–16,671.
Manuel, R. C., Czerwinski, E. W., and Lloyd, R. S. (1996) Identification of the structural and functional domains of MutY, an Escherichia coli DNA mismatch repair enzyme. J. Biol. Chem. 271, 16,218–16,226.
Manuel, R. C. and Lloyd, R. S. (1997) Cloning, overexpression, and biochemical characterization of the catalytic domain of MutY. Biochemistry 36, 11,140–11,152.
Noll, D. M., Gogos, A., Granek, J. A., and Clarke, N. D. (1999) The C-terminal domain of the adenine-DNA glycosylase MutY confers specificity for 8-oxoguanine adenine mispairs and may have evolved from MutT, an 8-oxodGTPase. Biochemistry 38, 6374–6579.
Guan, Y., Manuel, R. C., Arvai, A. S., Parikh, S. S., Mol, C. D., Miller, J. H., et al. (1998) MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nature Struct. Biol. 5, 1058–1064.
Volk, D. E., House, P. G., Thiviyanathan, V., Luxon, B. A., Zhang, S., Lloyd, R. S. et al. (2000) Structural similarities between MutT and the C-terminal domain of MutY. Biochemistry 39, 7331–7336.
Lu, A-L. and Fawcett, W. P. (1998) Characterization of the recombinant MutY homolog, an adenine DNA glycosylase, from Schizosacchromyces pombe. J. Biol. Chem. 273, 25,098–25,105.
McGoldrick, J. P., Yeh, Y.-C., Solomon, M., Essigmann, J. M., and Lu, A.-L. (1995) Characterization of a mammalian homolog of the Escherichia coli MutY mismatch repair protein. Mol. Cell. Biol. 15, 989–996.
Tsai-Wu, J.-J., Su, H.-T., Wu, Y.-L., Hsu, S.-M., and Wu, C. H. H. (2000) Nuclear localization of the human MutY homologue hMYH. J. Cell. Biochem. 77, 666–677.
Shinmura, K., Yamaguchi, S., Saitoh, T., Takeuchi-Sasaki, M., Kim, S. R., Nohmi, T., et al. (2000) Adenine excisional repair function of MYH protein on the adenine: 8-hydroxyguanine base pair in double-stranded DNA. Nucleic Acids Res. 28, 4912–4918.
Gu, Y. and Lu A-L. (2001) Differential DNA recognition and glycosylase activity of the native human MutY homolog (hMYH) and recombinant hMYH expressed in bacteria. Nucleic Acids Res. 29, 2666–2674.
Parker, A., Gu, Y., and Lu, A-L (2000) Purification and characterization of a mammalian homolog of Escherichia coli MutY mismatch repair protein from calf liver mitochondria. Nucleic Acids Res. 28, 3206–3215.
Lahue, R. S., Au, K. G., and Modrich, P. (1989) DNA mismatch correction in a defined system. Science 245, 160–164.
Lu, A-L (1998) Biochemistry of mammalian DNA mismatch repair, in DNA Repair in Higher Eukaryotes, Hoelm, H., and Nicolaides, N. C. (eds.), Humana Ps, Totowa, NJ, Vol. 2, pp. 95–118.
Kolodner, R. D. and Alani, E. (1994) Mismatch repair and cancer susceptibility. Curr. Opin. Biotech. 5, 585–594.
Modrich, P. and Lahue, R. S. (1996) Mismatch repair in replication fidelity, genetic recombination and cancer biology. Annu. Rev. Biochem. 65, 101–133.
Modrich, P. (1991) Mechanisms and biological effects of mismatch repair. Annu. Rev. Genet. 25, 229–253.
Terato, H., Masaoka, A., Kobayashi, M., Fukushima, S., Ohyama, Y., Yoshida, M., et al. (1999) Enzymatic repair of 5-formyluracil. II. Mismatch formation between 5-formyluracil and guanine during DNA replication and its recognition by two proteins involved in base excision repair (AlkA) and mismatch repair (MutS). J. Biol. Chem. 274, 25,144–25,150.
Masaoka, A., Kobayashi, M., Terato, H., Ohyama, Y., and Ide, H. (1999)Cellular repair mechanism of 5-formyluracil. Nucl. Acids Symp. Ser. 42, 291–292.
Zhao, J. and Winkler, M. E. (2000) Reduction of GC→TA transversion mutation by overexpression of MutS in Escherichia coli K-12. J. Bacteriol. 182, 5025–5028.
Leadon, S. A. and Avrutskaya, A. V. (1997) Differential involvement of the human mismatch repair proteins, hMLH1 and hMSH2, in transcriptio-coupled repair. Cancer Res. 57, 3784–3791.
DeWeese, T. L., Shipman, J. M., Larrier, N. A., Buckley, N. M., Kidd, L. R., Groopman, J. D., et al. (1998) Mouse embryonic stem cells carrying one or two defective Msh2 alleles respond abnormally to oxidative stree inflicted by low-level radiation. Proc. Natl. Acad. Sci. USA 95, 11,915–11,920.
Ni, T. T. M. G. T. and Kolodner, R. (1999) MSH2 and MSH6 are required for removal of adenine misincorporated opposite 8-oxo-guanine in S. cerevisine. Mol. Cell. 4, 439–444.
Hollis, T., Ichikawa, Y., and Ellenberger, T. (2000) DNA bending and a flip-out mechanism for base excision by the helix-hairpin-helix DNA glycosylase, Escherichia coli AlkA. EMBO J. 19, 758–766.
Doherty, A. J., Serpell, L. C., and Ponting, C. P. (1996) The helix-hairpin-helix DNA-binding motif: a structural basis for non-sequence-specific recognition of DNA. Nucleic Acids Res. 24, 2488–2497.
Labahn, J., Scharer, A., Long, A., Ezaz-Nikpay, K., Verdine, G. L., and Ellenberger, T. E. (1996) Structural basis for the excision repair of alkylation-damaged DNA. Cell 86, 321–329.
Yamagata, Y., Kato, M., Odawara, K., Tokuno, Y., Nakashima, Y., Matsushima, N., et al. (1996) Three-dimensional structure of a DNA repair enzyme, 3-methyadenine DNA glycosylase II, from Escherichia coli. Cell 86, 311–319.
Thayer, M. M., Ahern, H., Xing, D., Cunningham, R. P., and Tainer, J. A. (1995) Novel DNA binding motifs in the DNA repair enzyme endonuclease III crystal structure. EMBO J. 14, 4108–4120.
Kuo, C.-F., McRee, D. E., Fisher, C. L., O'Haandley, S. F., Cunningham, R. P., and Tainer, J. A. (1992) Atomic structure of the DNA repair [4Fe-4S] enzyme endonuclease III. Science 258, 434–440.
Volk, D. E., Thiviyanathan, V., House, P. G., Lloyd, R. S., and Gorenstein, D. G. (1999) 1H, 13C and 15N resonance assignments of the C-terminal domain of MutY: an adenine glycosylase active on G: A mismatches. J. Biomol. NMR 14, 385–386.
Li, X. and Lu, A-L. (2000) Intact MutY and its catalytic domain differentially contact with A/8-oxoG-containing DNA. Nucleic Acids Res. 28, 4593–4603.
Sugahara, M., Mikawa, T., Kumasaka, T., Yamamoto, M., Kato, R., Fukuyama, K., et al. (2000) Crystal structure of a repair enzyme of oxidatively damaged DNA, MutM (Fpg), from an extreme thermophile, Thermus thermophilus HB8. EMBO J. 19, 3857–3869.
Zharkov, D. O., Reiger, R. A., Iden, C. R., and Grollman, A. P. (1997) NH2-terminal proline acts as a nucleophile in the glycosylase/APlyase reaction catalyzed by Escherichia coli formamidopyrimidine-DNA glycosylase (Fpg) protein. J. Biol. Chem. 272, 5335–5341.
Sun, B., Latham, K. A., Dodson, M. L., and Lloyd, R. S. (1995), Studies on the catalytic mechanism of five DNA glycosylases: probing for enzyme-DNA imino intermediates. J. Biol. Chem. 270, 19,501–19,508.
O'Connor, T. R. and Laval, J. (1989) Physical association of the 2,6-diamino-4-hydroxy-5N-formamidopyrimidine-DNA glycosylase of Escherichia coli and an activity nicking DNA at apurinic/apyrimidinic sites. Proc. Natl. Acad. Sci. USA 86, 5222–5226.
Williams, S. D. and David, S. S. (1998) Evidence that MutY is a monofunctional glycosylase capable of forming a covalent Schiff base intermediate with substrate DNA. Nucleic Acids Res. 26, 5123–5133.
Zharkov, D. O. and Grollman, A. P. (1998) MutY DNA glycosylase: base release and intermediate complex formation. Biochemistry 37, 12,384–12,394.
Au, K. G., Clark, S., Miller, J. H., and Modrich, P. (1989) Escherichia coli mutY gene encodes an adenine glycosylase active on G/A mispairs. Proc. Natl. Acad. Sci. USA 86, 8877–8881.
Bulychev, N. V., Varaprasad, C. V., Dorman, G., Miller, J. H., Eisenberg, M., and Grollman, A. P. (1996) Substrate specificity of Escherichia coli MutY protein. Biochemistry 35, 13,147–13,156.
Michaels, M. L., Tchou, J., Grollman, A. P., and Miller, J. H. (1992) A repair system for 8-oxo-7,8-dihydrodeoxyguanine (8-hydroxyguanine). Biochemistry 31, 10,964–10,968.
Lu, A-L., Tsai-Wu, J.-J., and Cillo, J. (1995) DNA determinants and substrate specificities of Escherichia coli MutY. J. Biol. Chem. 270, 23,582–23,588.
Lu, A-L., Yuen, D. S., and Cillo, J. (1996) Catalytic mechanisms and DNA substrate recognition of Eschirichia coli MutY protein. J. Biol. Chem. 271, 24,138–24,143.
Wright, P. M., Yu, J., Cillo, J., and Lu, A.-L. (1999) The active site of the Escherichia coli MutY DNA adenine glycosylase. J. Biol. Chem. 274, 29,011–29,018.
Williams, S. D. and David, S. S. (1999) Formation of a Schiff base intermediate is not required for the adenine glycosylase activity of Escherichia coli Muty. Biochemistry 38, 15,417–15,424.
Zharkov, D. O., Gilboa, R., Yagil, I., Kycia, J. H., Gerchman, S. E., Shoham, G., et al. (2000) Role for lysine 142 in the excision of adenine from A: G mispairs by MutY DNA glycosylase of Escherichia coli. Biochemistry 39, 14,768–14,778.
Williams, S. D. and David, S. S. (2000) A single engineered point mutation in the adenine glycosylase MutY confers bifunctional glycosylase/AP lyase activity. Biochemistry 39, 10,098–10,109.
Mol, C. D., Parikh, S. S., Putnam, C. D., Lo, T. P., and Tainer, J. A. (1999) DNA repair mechanisms for the recognition and removal of damaged DNA bases. Annu. Rev. Biophys. Biomol. Struct. 28, 101–128.
Castaing, B., Fourrey, J. L., Hervouet, N., Thomas, M., Boiteux, S., and Zelwer, C. (1999) AP site structural determinants for Fpg specific recognition. Nucleic Acids Res. 27, 608–615.
Porello, S. L., Williams, S. D., Kuhn, H., Michaels, M. L., and David, S. S. (1996) Specific recognition of substrate analogs by the DNA mismatch repair enzyme MutY. J. Am. Chem. Soc. 118, 10,684–10,692.
Chmiel, N. H., Golinelli, M. P., Francis, A. W., and David, S. S. (2001) Efficient recognition of substrates and substrate analogs by the adenine glycosylase MutY requires the C-terminal domain. Nucleic Acids Res. 29, 553–564.
Hosfield, D. J., Guan, Y., Haas, B. J., Cunningham, R. P., and Tainer, J. A. (1999) Structure of the DNA repair enzyme endonuclease IV and its DNA complex: doublenucleotide flipping at abasic sites and three-metal-ion catalysis. Cell 98, 397–408.
Mol, C. D., Kuo, C. F., Thayer, M. M., Cunningham, R. P., and Tainer, J. A. (1999) Structure and function of the multifunctional DNA-repair enzyme exonuclease III. Nature 374, 381–386.
Mol, C. D., Hosfield, D. J., and Tainer, J. A. (2000) Abasic site recognition by two apurinic/apyrimidinic endonuclease families in DNA base excision repair: the 3′ ends justify the means. Mutat. Res. 460, 211–229.
Dianov, G., Sedgwick, B., Daly, G., Olsson, M., Lovett, S., and Lindahl, T. (1994) Release of 5′-terminal deoxyribose-phosphate residues from incised abasic sites in DNA by the Escherichia coli RecJ protein. Nucleic Acids Res 22, 993–998.
Sandigursky, M. and Franklin, W. A. (1992) DNA deoxyribophosphodiesterase of Escherichia coli is associated with exonuclease 1. Nucleic Acids Res. 20, 4699–4703.
Piersen, C. E., McCullough, A. K., and Lloyd, R. S. (2000) AP lyases and dRPases: commonality of mechanism. Mutat. Res. 459, 43–53.
Dianov, G. and Lindahl, T. (1994) Reconstitution of the DNA base excision-repair pathway. Curr. Biol. 4, 1069–1076.
Dianov, G., Price, A., and Lindahl, T. (1992) Generation of single-nucleotide repair patches following excision of uracil residues from DNA. Mol. Cell Biol. 12, 1605–1612.
Radicella, J. P., Clark, E. A., Chen, S., and Fox, M. S. (1993) Patch length of localized repair event: role of DNA polymerase I in mutY-dependent mismatch repair. J. Bacteriol. 175, 7732–7736.
Tsai-Wu, J.-J. and Lu, A-L. (1994) Escherichia coli mutY-dependent mismatch repair involves DNA polymerase I and a short repair tract. Mol. Gen. Genet. 244, 444–450.
Frosina, G., Cappelli, E., Fortini, P., and Dogliotti, E. (1999) In vitro base excision repair assay using mammalian cell extracts. Methods Mol. Biol. 113, 301–315.
Krokan, H. E., Nilsen, H., Skorpen, F., Otterlei, M., and Slupphaug, G. (2000) Base excision repair of DNA in mammalian cells. FEBS Lett. 476, 73–77.
Matsumoto, Y., Kim, K., Hurwitz, J., Gary, R., Levine, D. S., Tomkinson, A. E., et al. (1999) Reconstitution of proliferating cell nuclear antigen-dependent repair of apurinic/apyrimidinic sites with purified human proteins. J. Biol. Chem. 274, 33,703–33,708.
Srivastava, D. K., Berg, B. J., Prasad, R., Molina, J. T., Beard, W. A., Tomkinson, A. E., et al. (1998) Mammalian abasic site base excision repair. Identification of the reaction sequence and rate-determining steps. J. Biol. Chem. 273, 21,203–21,209.
Pascucci, B., Strucki, M., Jonsson, Z. O., Dogliotti, E., and Hubscher, U. (1999) Long patch base excision repair with purified human proteins: DNA ligase I as patch size mediator for DNA polymerases δ and ε. J. Biol. Chem. 274, 33,696–33,702.
DeMott, M. S., Zigman, S., and Bambara, R. A. (1998) Replication protein A stimulates long patch DNA base excision repair. J. Biol. Chem. 273, 27,492–27,498.
Dianov, G. L., Jensen, B. R., Kenny, M. K., and Bohr, V. A. (1999) Replication protein A stimulates proliferating cell nuclear antigen-dependent repari of abasic sites in DNA by human cell extracts. Biochemistry 38, 11,021–11,025.
Dianov, G. L., Prasad, R., Wilson, S. H., and Bohr, V. A. (1999) Role of DNA polymerase beta in the excision step of long patch mammalian base excision repair. J. Biol. Chem. 274, 13,741–13,743.
Parker, A., Gu, Y., Mahoney, W., Lee, S.-H., Singh, K. K., and Lu, A-L. (2001) Human homolog of the MutY protein (hMYH) physically interacts with protein involved in longpatch DNA base excision repair. J. Biol. Chem. 276, 5547–5555.
Fortini, P., Parlanti, E., Sidorkina, O. M., Laval, J., and Dogliotti, E. (1999) The type of DNA glycosylase determines the base excision repair pathway in mammalian cells. J. Biol. Chem. 274, 15,230–15,236.
Bharati, S., Krokan, H. E., Kristiansen, L., Otterlei, M., and Slupphaug, G. (1998) Human mitochondrial uracil-DNA glycosylase prefrom (UNG1) is processed to two forms one of which is resistant to inhibition by AP sites. Nucleic Acids Res. 26, 4953–4959.
Mol, C. D., Izumi, T., Mitra, S., and Tainer, J. A. (2000) DNA-bound structures and mutants reveal abasic DNA binding by APE1 and DNA repair coordination. Nature 403, 451–456.
Gorman, M. A., Morera, S., Rothwell, D. G., de La, F. E., Mol, C. D., Tainer, J. A., et al. (1997) The crystal structure of the human DNA repair endonuclease HAP1 suggests the recognition of extra-helical deoxyribose at DNA abasic sites. EMBO J. 16, 6548–6558.
Parikh, S. S., Mol, C. D., Slupphaug, G., Bharati, S., Krokan, H. E., and Tainer, J. A. (1998) Base excision repair initiation revealed by crystal structures and binding kinetics of human uracil-DNA glycosylase with DNA. EMBO J. 17, 5214–5226.
Waters, T. R., Gallinari, P., Jiricny, J., and Swann, P. F. (1999) Human thymine DNA glycosylase binds to apurinic sites in DNA but is displaced by human apurinic endonuclease 1. J. Biol. Chem. 274, 67–74.
Yang, H., Clendenin, W. M., Wong, D., Demple, B., Slupska, M. M., Chaing, J. H. and Miller, J. H. (2001) Enhanced activity of adenine-DNA glycosylase (Myh) by apurinic/apyrimidinic endonuclease (Ape1) in mammalian base excision repair of an A/GO mismatch. Nucleic Acids Res. 29, 743–752.
Klungland, A., Hoss, M., Gunz, D., Constantinou, A., Clarkson, S. G., Doetsch, P. W., et al. (1999) Base excision repair of oxidative DNA damage activated by XPG protein. Mol. Cell 3, 33–42.
Bennett, R. A., Wilson, D. M., III, Wong, D., and Demple, B. (1997) Interaction of human apurinic endonuclease and DNA polymerase beta in the base excision repair pathway. Proc. Natl. Acad. Sci. USA 94, 7166–7169.
Krishna, T. S., Kong, X. P., Gary, S., Burgers, P. M., and Kuriyan, J. (1994) Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell 79, 1233–1243.
Kelman, Z. (1997) PCNA: structure, functions and interactions. Oncogene 14, 629–640.
Warbrick, E. (1998) PCNA binding through a conserved motif. BioEssays 20, 195–199.
Zhang, P., Mo, J. Y., Perez, A., Leon, A., Liu, L., Mazloum, N., et al. (1999) Direct interaction of proliferating cell nuclear antigen with the p125 catalytic subunit of mammalian DNA polymerase δ. J. Biol. Chem. 274, 26,647–26,653.
Tsurimoto, T. (1999) PCNA binding proteins. Front. Biosci. 4, D849-D858.
Kelman, Z. and Hurwitz, J. (1998) Protein-PCNA interactions: a DNA-scanning mechanism? Trends Biochem. Sci. 23, 236–238.
Mer, G., Bochkarev, A., Gupta, R., Bochkareva, E., Frappier, L., Ingles, C. J., et al. (2000) Structural basis for the recognition of DNA repair proteins UNG2, XPA, and RAD52 by replication factor RPA. Cell 103, 449–456.
Nagelhus, T. A., Haug, T., Singh, K. K., Keshav, K. F., Skorpen, F., Otterlei, M., et al. (1997) A sequence in the N-terminal region of human uracil-DNA glycosylase with homology to XPA interacts with the C-terminal part of the 34-kDa subunit of replication protein A. J. Biol. Chem. 272, 6561–6566.
Park, M. S., Ludwig, D. L., Stigger, E., and Lee, S. H. (1996) Physical interaction between human RAD52 and RPA is required for homologous recombination in mammalian cells. J. Biol. Chem. 271, 18,996–19,000.
Wold, M. S. (1997) Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 66, 61–92.
Biswas, E. E., Zhu, F. X., and Biswas, S. B. (1997) Stimulation of RTH1 nuclease of the yeast Saccharomyces cerevisiae by replication protein A. Biochemistry 36, 5955–5962.
Prasad, R., Singhal, R. K., Srivastava, D. K., Molina, J. T., Tomkinson, A. E., and Wilson, S. H. (1996) Specific interaction of DNA polymerase beta and DNA ligase I in a multiprotein base excision repair complex from bovine testis. J. Biol. Chem. 271, 16,000–16,007.
Marintchev, A., Robertson, A., Dimitriadis, E. K., Prasad, R., Wilson, S. H., and Mullen, G. P. (2000) Domain specific interaction in the XRCC1-DNA polymerase beta complex. Nucleic Acids Res. 28, 2049–2059.
Masson, M., Niedergang, C., Schreiber, V., Muller, S., Menissier-de Murcia, J., and de Murcia, G. (1998) XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol. Cell Biol. 18, 3563–3571.
Levin, D. S., Bai, W., Yao, N., O'Donnell, M., and Tomkinson, A. E. (1997) An interaction between DNA ligase I and proliferating cell nuclear antigen: implications for Okazaki fragment synthesis and joining. Proc. Natl. Acad. Sci. USA 94, 12,863–12,868.
Montecucco, A., Rossi, R., Levin, D. S., Gary, R., Park, M. S., Motycka, T. A., et al. (1998) DNA ligase I is recruited to sites of DNA replication by an interaction with proliferating cell nuclear antigen: identification of a common targeting mechanism for the assembly of replication factories. EMBO J. 17, 3786–3795.
Beard, W. A. and Wilson, S. H. (2000) Structural design of a eukaryotic DNA repair polymerase: DNA polymerase beta. Mutat. Res. 460, 231–244.
Rice, P. A. (1999) Holding damaged DNA together. Nat. Struct. Biol. 6, 805–806.
Wilson, S. H. and Kunkel, T. A. (2000) Passing the baton in base excision repair. Nat. Struct. Biol. 7, 176–178.
Gowen, L. C., Avrutskaya, A. V., Latour, A. M., Koller, B. H., and Leadon, S. A. (1998) BRCA1 required for transcription-coupled repair of oxidative DNA damage. Science 281, 1009–1012.
LePage, F., Kwoh, E. E., Avrutskaya, A., Gentil, A., Leadon, S. A., Sarasin, A., et al. (2000) Transcription-coupled repair of 8-oxoguanine: requirement for XPG, TFIIH, and CSB and implications for Cockayne syndrome. Cell 101, 159–171.
Le Page, F., Klungland, A., Barnes, D. E., Sarasin, A., and Boiteux, S. (2000) Transcription coupled repair of 8-oxoguanine in mirine cells: the OGG1 protein is required for repair in nontranscribed sequences but not in transcribed sequences. Proc. Natl. Acad. Sci. USA 97, 8397–8402.
Thomas, D., Scot, A. D., Barbey, R., Padula, M., and Boiteux, S. (1997) Inactivation of OGG1 increases the incidence of G: C to T: A transversions in Saccharomyces cerevisiae: evidence for endogenous oxidative damage to DNA in eukaryotic cells. Mol. Gen. Genet. 254, 171–178.
van der Kemp P. A., Thomas, D., Barbey, R., de Oliveira, R., and Boiteux, S. (1996) Cloning and expression in Escherichia coli of the Ogg1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc. Natl. Acad. Sci. USA 93, 5197–5202.
Chang, D.-Y., Gu, Y. and Lu, A-L. (2001) Fission yeast (Schizosaccharomyces pombe) cells defective in the MutY-homologous glycosylase activity have a mutator phenotype and are sensitive to hydrogen peroxide. Mol. Genet. Genom., in press.
Ochs, K., Sobol, R. W., Wilson, S. H., and Kaina, B. (1999) Cells deficient in DNA polymerase beta are hypersensitive to alkylating agent-induced apoptosis and chromosomal breakage. Cancer Res. 59, 1544–1551.
Sobol, R. W., Horton, J. K., Kuhn, R., Gu, H., Singhal, R. K., Prasad, R., et al. (1996) Requirement of mammalian DNA polymerasebeta in base-excision repair. Nature 379, 183–186.
Tebbs, R. S., Flannery, M. L., Meneses, J. J., Hartmann, A., Tucker, J. D., Thompson, L. H., et al. (1999) Requirement for the Xrcc1 DNA base excision repair gene during early mouse development. Dev. Biol. 208, 513–529.
Wilson, D. M., III and Thompson, L. H. (1997) Life without DNA repair. Proc. Natl. Acad. Sci. USA 94, 12,754–12,757.
Malins, D. C., Holmes, E. H., Polissar, N. L., and Gunselman, S. J. (1993) The etiology of breast cancer: characteristic alterations in hydroxyl radical-induced DNA base lesions during oncogenesis with potential for evaluating incidence risk. Cancer 71, 3036–3043.
Olinski, R., Zastawny, T., Budzbon, J., Skokowski, J., Zegarski, W., and Dizdaroglu, M. (1992) DNA base modifications in chromatin of human cancerous tissues. FEBS Lett. 309, 193–198.
Szatrowski, T. P. and Nathan, C. F. (1991) Production of large amounts of hydrogen peroxide by human tumor cells. Cancer Res. 51, 794–798.
Hollstein, M., Sidransky, D., Vogelstein, B., and Harris, C. C. (1991) p53 mutations in human cancers. Science 253, 49–53.
Mitsudomi, T., Viallet, J., Mulshine, J. L., Linnoila, R. I., Minna, J. D., and Gazdar, A. F. (1991) Mutations of ras genes distinguish a subset of non-small-cell lung cancer cell lines from small-cell lung cancer lines. Oncogene 6, 1353–1362.
Takama, F., Kanuma, T., Wang D., Nishida, J. I., Nakabeppu, Y., Wake, N. et al. (2000) Mutation analysis of the hMTH1 gene in sporadic human ovarian cancer. Int. J. Oncol. 17, 467–471.
Audebert, M., Chevillard, S., Levalois, C., Gyapay, G., Vieillefond, A., Klijanienko, J., et al. (2000) Alterations of the DNA repair gene OGG1 in human clear cell carcinomas of the kidney. Cancer Res. 60, 4740–4744.
Klungland, A., Rosewell, I., Hollenbach, S., Larsen, E., Daly, G., Epe, B., et al. (1999) Accumulation of premutagenic DNA lesions in mice defective in removal of oxidative base damage. Proc. Natl. Acad. Sci. USA 96, 13,300–13,305.
Friedberg, E. C. and Meira, L. B. (2000) Database of mouse strains carrying targeted mutations in genes affecting cellular responses to DNA damage. Version 4. Mutat. Res. 459, 243–274.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lu, AL., Li, X., Gu, Y. et al. Repair of oxidative DNA damage. Cell Biochem Biophys 35, 141–170 (2001). https://doi.org/10.1385/CBB:35:2:141
Issue Date:
DOI: https://doi.org/10.1385/CBB:35:2:141