23 June 2021: Clinical Research
A Comparison of Detection in Sputum by the Conventional Culture and Fluorescent Polymerase Chain Reaction Methods
Xiaoqun Xu12BD, Huoyang Lv3AE, Fengwei Zhang12DF, Houyong Zhu4CDE, Long Cai12AE*DOI: 10.12659/MSM.930293
Med Sci Monit 2021; 27:e930293
Abstract
BACKGROUND: Candida is a pathogenic fungus. In recent years, the increase in immunosuppressive diseases has led to an increase in Candida infections, with the lungs being the most common site. Therefore, the aim of this study was to compare the positive detection rates of Candida in sputum samples by Candida culture and fluorescent polymerase chain reaction (PCR), and to explore a new method for rapid, accurate, and effective detection of Candida in sputum, providing swift evidence of clinical fungal infection.
MATERIAL AND METHODS: From October 2016 to March 2017, 300 sputum samples were collected and detected by the conventional culture method and fluorescent PCR method. The positive rate of Candida detection was compared between the 2 methods.
RESULTS: In the 300 sputum samples, the positive detection rate of Candida was 50% by the culture method and 65.67% by the fluorescent PCR method (P<0.001). Therefore, the positive detection rate of Candida was higher by the fluorescent PCR method.
CONCLUSIONS: The conventional culture method for Candida needs a longer duration (24 h to 48 h) and the positive detection rate is low. However, it takes only 3 h to detect Candida in sputum by the fluorescent PCR method, the positive detection rate is high, and can be used as a screening method for Candida in sputum samples. Additional large-scale clinical trials need to be completed to assess the correlation between fluorescent PCR and pulmonary Candida infection.
Keywords: Candida, Culture, Real-Time Polymerase Chain Reaction, Candidiasis, Culture Techniques, Fluorescence, Polymerase Chain Reaction, Reproducibility of Results, Sensitivity and Specificity, Sputum
Background
Material and Methods
COLLECTION OF SAMPLES:
From October 2016 to March 2017, 300 sputum samples were collected from patients in several wards of our hospital. Basic information about the patients, including the name, sample number, age, sex, and hospitalization number were checked. Sputum samples were divided into 2 parts; 1 part was cultured by the conventional method, and the remaining samples were stored for 3 days in the freezer at −20°C for fluorescent PCR detection. Informed consent from the patients was exempted by the Ethics Committee of Zhejiang Provincial People’s Hospital (Ethics Application Ref: 2020QT351). This study’s protocol complied with the requirements of the Helsinki Declaration of the World Medical Association and the International Ethics Guide for Human Biomedical Research of the Council for International Organizations of Medical Sciences (CIOMS).
:
All the samples were cultured on both bacterial and fungal media. The fungal culture media (Yeast Identification Medium, Mellier [Shanghai] Biological Products Co., Ltd., Shanghai, China) were Sabouraud dextrose agar with chloramphenicol and brain-heart infusion agar with antibacterial agents.
:
DNA was extracted using the centrifugal adsorption column method. DNA is adsorbed by the special silicon matrix adsorption material of the centrifugal tube of the DNA solid extract in an environment of high salt and low pH (Sample treatment solution B of the DNA Extraction Kit [Hangzhou Derlead Biotech Co., Ltd., Hangzhou China]). Then, it is released in a low-salt and high-pH environment (DNA eluent of DL-Sputum [fungus], DNA Extraction Kit, Hangzhou Derlead Biotech Co., Ltd., Hangzhou, China).
PRINCIPLE OF THE FLUORESCENT POLYMERASE CHAIN REACTION (PCR):
Fluorescent PCR [15] is a fluorescent chemiluminescence method for the determination of total products after the PCR. TaqMan probes (Pan-Candida Nucleic Acid Detection Kit Fluorescent PCR, Hangzhou Derlead Biotech Co., Ltd., Hangzhou, China) were used in this study; the TaqMan probe includes fluorescence reporting groups and fluorescence quenching groups. When the 2 groups exist at the same time, the fluorescence emitted by the reporting group is quenched. When the PCR is amplified, the probe is degraded by the probe enzyme and the fluorescence reporting groups and fluorescence quenching groups are separated; then, the fluorescence reporting groups can emit fluorescent signals. The monitoring system can receive each fluorescent signal by amplifying a DNA strand and forming a fluorescent molecule; the accumulation of PCR products and the formation of fluorescent signals are synchronized.
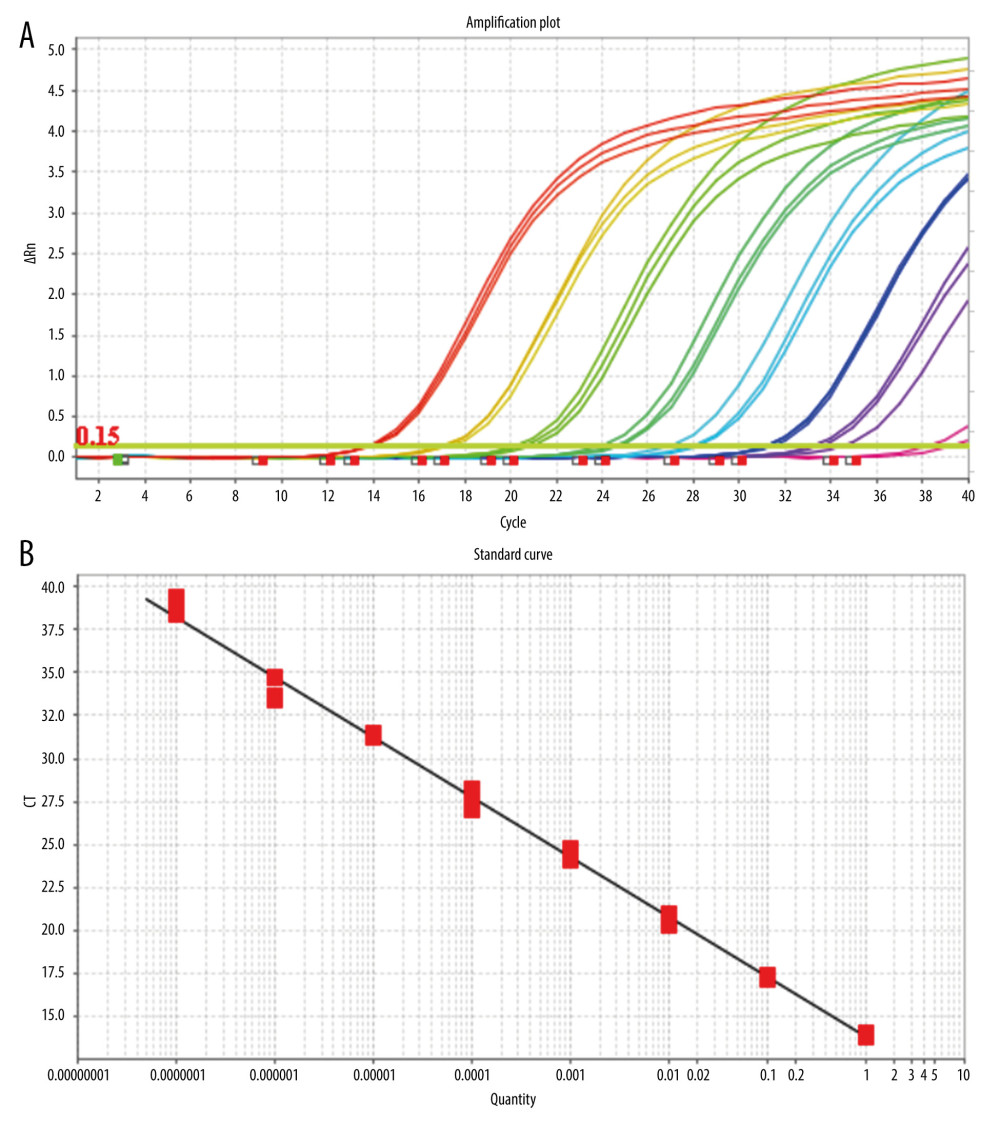
After mixing the fluorescein-labeled TaqMan probes with the template DNA, the TaqMan probe pairs complementary to the template DNA were cut off in the PCR reaction process, and the free fluorescein in the reaction system emitted fluorescence under specific light excitation. With an increase in the number of cycles, the number of target gene fragments increase exponentially, and the copy number of target genes is obtained by real-time detection of the corresponding fluorescent signal intensity using the standard template number of the known concentration (Figure 1A, 1B).
In this experiment, the primers and probes for fluorescent PCR were designed based on the highly conserved sequences of the 18S, 5.8S, and 28S subunits of fungal ribosomal DNA and the specific sequences of the internal transcribed spacer, as well as the specific primers and probes.
DNA EXTRACTION: We referred to the study by Ramos et al for DNA extraction [16]. The kit used for DNA extraction was the DL-Sputum (fungus) DNA Extraction Kit (Hangzhou Derlead Biotech Co., Ltd, Hangzhou, China); its composition is shown in Table 1. For DNA extraction, 1.2 mL of sputum digestion solution was added into a 2-mL Eppendorf (EP) tube, then 0.6 mL of sputum solution was added and incubated at room temperature (optimum temperature 37°C) for 15 min to 30 min, and oscillated every 3 min for 15 s, until the mixture became clear and had uniform viscosity. Then, it was centrifuged at 10 000 rpm for 3 min, the supernatant liquid was discarded, and the cells were retained. Then, 1 mL of sample cleaning solution was added, oscillated, and suspended, and it was then centrifuged for 3 min at 10 000 rpm, and the supernatant liquid was discarded. The precipitation was suspended by adding 300 μL of sample treatment solution A. All the suspended samples were transferred to the centrifuge tubes containing solid DNA extracts (1 h before use, the centrifuge tubes were centrifuged to collect solids at the bottom of the tube), oscillated for 5 min, and then instantaneously centrifuged. Then, 200 μL of sample treatment solution B was added, mixed well, and centrifuged at 13 000 rpm for 10 min. The supernatant liquid was transferred to a new 1.5-mL EP tube (taking care not to inhale the oil in the lower layer) and mixed with 0.5 times absolute ethanol by volume. All the liquid in the centrifuge tubes was transferred to the DNA purification column with casing, centrifuged for 1 min at 13 000 rpm, and the centrifugal fluid in the casing was discarded. Then, the purification tube was repositioned in the casing. We added 600 μL of washing fluid to the centrifuge tubes. They were centrifuged at 13 000 rpm for 1 min, and the centrifugal fluid in the casing was discarded. The purification column was replaced in the casing and centrifuged for 3 min at 13 000 rpm. After centrifugation, the cap of the purification column was opened and aired for 1 min to 2 min to volatilize the ethanol on the filter membrane. The casing was discarded and the purification column was placed in a new 1.5-ml EP tube. Next, 40 μL of DNA eluent preheated at 70°C was dropped carefully over the filter membrane at the bottom of the purification column and allowed to stand for 3 min. Finally, these EP tubes were centrifuged at 13 000 rpm for 2 min. The cap of the EP tube was cut during centrifugation, and the centrifuged DNA recovery solution was transferred to another sterile EP tube for storage. The DNA recovery solution obtained by centrifugation can be used for subsequent experiments. The extracted fungal DNA should not be stored at room temperature; it should immediately be stored at below −20°C.
FLUORESCENT PCR DETECTION: The Pan-Candida Nucleic Acid Detection Kit (Fluorescent PCR) (Hangzhou Derlead Biotech Co., Ltd, Hangzhou, China) was used for the fluorescent quantitative PCR, and its composition is shown in Table 2. All the reagents need instantaneous centrifugation before use. Each reaction system is shown in Table 3. The dosage of the above reagents (except the sample/reference) was calculated according to the number of reaction tubes, added to the appropriate centrifuge tubes, and mixed well. The liquid in the centrifuge pipette was blown slowly and repeatedly, taking care to avoid splashing of the liquid or producing a large number of bubbles. After instantaneous centrifugation, 16 μL of PCR reaction solution was separately packed into each PCR reaction tube. The prepared PCR reaction tube, negative control, and positive control were transferred to the sample processing area (sampling area). Then, 4 μL of DNA or negative or positive control samples were added to the prepared PCR reaction tube, and then centrifuged immediately after capping the tube (or fixing the sealing film), and transferred to the sample detection area. Finally, the prepared reaction tube was placed in the PCR instrument, ([StepOne Plus ZJSRM Plus] YY-FZ-YQ-03 Hangzhou Boke Co., Ltd., Hangzhou, China) and the reaction and detection was amplified according to the edited sample information, following the conditions shown in Table 4.
SETTING ANALYSIS CONDITIONS FOR THE RESULTS OF QUANTITATIVE FLUORESCENT PCR: When analyzing the results of the PCR amplification curve, the mapping type can be set as Rn (the fluorescence intensity measured at each point minus the fluorescence baseline intensity) vs cycle. Baseline setting: The baseline value was set between 2 and 3 cycles of PCR with the parameters are shown in Table 4. The main purpose of setting the baseline is to calibrate the test results of the samples, so that the experimental results can better reflect the authenticity of the data. Threshold setting: The threshold line of analysis software was set manually. The threshold line is located in the linear part of the exponential growth period of the amplification curve above the baseline. The point where the amplification curve intersects the threshold line is the CT value, which represents the variable value Rn (Rn=Rn [post-PCR reading]-Rn [pre-PCR reading]) before and after the fluorescence intensity amplification of the standardized reporting group. CT stands for the number of cycles that the fluorescence signal in each sample goes through when it reaches the set field value. The CT value was negatively correlated with the logarithmic value of the initial target DNA fragment (reference value). The standard curve is shown in Figure 1A and 1B.
STANDARD OF QUALITY CONTROL:
Unless the negative and positive controls of the kit (Pan-Candida Nucleic Acid Detection Kit Fluorescent PCR, Hangzhou Derlead Biotech Co., Ltd, Hangzhou, China) satisfy the following conditions at the same time, the experiment is considered invalid and needs to be repeated. Negative control: The CT value for negative quality control is ≥34, or Undetermined. Positive control: The CT value for positive quality control is <34. A sample CT value of <34 means a positive result and a sample CT value of >34 or Undetermined means a negative result.
STATISTICAL ANALYSIS:
The data were analyzed using SPSS 25.0 (SPSS, Inc., Chicago, IL). The McNemar test was used to analyze the rate according to the amount of data. A bilateral
Results
All 300 samples collected were used for
Discussion
The diagnosis and treatment of pulmonary candidiasis is complex. Clinicians need to find microbiological evidence to confirm the diagnosis before administering antifungal treatment. Confirmed pulmonary candidiasis according to the 2008 European Organization for Research and Treatment of Cancer/Mycosis Study Group’s (EORTC/MSG) revised definition [19] is to test for the microorganisms in pathological tissue or blood samples from sterile material, and suspected pulmonary candidiasis was defined as a positive sample with at least 1 sputum culture. If a patient presents with clinical symptoms or risk factors associated with pulmonary candidiasis and a conventional fungal culture is shown to be negative after 24 h to 48 h, then clinicians need to wait for a second culture result to obtain a positive culture result before using antifungal therapy for suspected infectious patients.
The positive rate of a single sputum culture is low [20,21], and repeated sputum cultures need 4 additional days (average) before antifungal treatment can be started, resulting in an additional 4 days of hospitalization [22]. However, there remain some patients who do not have a definite diagnosis of fungal infection and do not receive antifungal treatment before hospital discharge. Some reasons are that unmedicated individuals have less severe disease and their fungal infection resolves without treatment. However, for those patients with liver cirrhosis, blood tumors, other infections, and immune disorders, there is a clear correlation between the time antifungal treatment is initiated and the length of hospitalization [23–26]. Venkatesh et al [27,28] demonstrated that early precision treatment of a
There are some limitations to this study. First, the PCR method is more expensive than sputum culture, which can limit its use for mildly ill patients. However, it will be useful for patients with severe infection, when antifungal treatment needs to be considered and the PCR method can quickly obtain evidence, and for patients in whom Candida infection is highly suspected but there is no basis for antifungal treatment. Second, as the aim of this study was to investigate the detection rate of
In this study, 300 sputum samples were collected and tested by the conventional culture method and fluorescent PCR method to compare their positive detection rates. In the 300 sputum samples, the positive rate of
Conclusions
A conventional culture of
References
1. Ren C, Guo Q: Chinese Journal of lung diseases (Electronic Edition), 2016; 9; 471-78
2. Lortholary O, Renaudat C, Sitbon K, Worrisome trends in incidence and mortality of candidemia in intensive care units (Paris area, 2002–2010): Intensive Care Med, 2014; 40(9); 1303-12
3. Tragiannidis A, Tsoulas C, Kerl K, Invasive candidiasis: Update on current pharmacotherapy options and future perspectives: Expert Opin Pharmacother, 2013; 14(11); 1515-28
4. Vazquez JA, Miceli MH, Alangaden G, Invasive fungal infections in transplant recipients: Ther Adv Infect Dis, 2013; 1(3); 85-105
5. Pappas PG, Kauffman CA, Andes DR, Clinical practice guideline for the management of candidiasis: 2016 update by the Infectious Diseases Society of America: Clin Infect Dis, 2016; 62(4); e1-50
6. Prakash A, Sharma C, Singh A: Clin Microbiol Infect, 2016; 22; 271-77
7. Kapoor G, Saigal S, Saigal K, Why candidemia occurs early: A comment on “Incidence, characteristics and outcome of ICU-acquired candidemia in India”: Intensive Care Med, 2015; 41(2); 377
8. Sievert DM, Ricks P, Edwards JR, Antimicrobial-resistant pathogens associated with healthcare-associated infections: summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009–2010: Infect Control Hosp Epidemiol, 2013; 34(1); 1-14
9. Bassetti M, Righi E, Ansaldi F, A multicenter multinational study of abdominal candidiasis: Epidemiology, outcomes and predictors of mortality: Intensive Care Med, 2015; 41(9); 1601-10
10. Bassetti M, Righi E, Ansaldi F, A multicenter study of septic shock due to candidemia: Outcomes and predictors of mortality: Intensive Care Med, 2014; 40(6); 839-45
11. Lamoth F, Lockhart SR, Berkow EL, Changes in the epidemiological landscape of invasive candidiasis: J Antimicrob Chemother, 2018; 73(Suppl 1); i4-13
12. Puig-Asensio M, Padilla B, Garnacho-Montero J: Clin Microbiol Infect, 2014; 20(4); O245-54
13. Doi AM, Pignatari AC, Edmond MB, Epidemiology and microbiologic characterization of nosocomial candidemia from a Brazilian National Surveillance Program: PLoS One, 2016; 11(1); e146909
14. Nucci M, Anaissie E, How we treat invasive fungal diseases in patients with acute leukemia: The importance of an individualized approach: Blood, 2014; 124(26); 3858-69
15. Ramos JT, Villar S, Bouza E, Performance of a quantitative PCR-based assay and beta-d-glucan detection for diagnosis of invasive candidiasis in very-low-birth-weight preterm neonatal patients (CANDINEO Study): J Clin Microbiol, 2017; 55(9); 2752-64
16. Guinea J, Padilla C, Escribano P, Evaluation of MycAssay Aspergillus for diagnosis of invasive pulmonary aspergillosis in patients without hematological cancer: PLoS One, 2013; 8(4); e61545
17. Kullberg BJ, Arendrup MC, Invasive candidiasis: N Engl J Med, 2015; 373(15); 1445-56
18. Schelenz S, Barnes RA, Barton RC, British Society for Medical Mycology best practice recommendations for the diagnosis of serious fungal diseases: Lancet Infect Dis, 2015; 15(4); 461-74
19. De Pauw B, Walsh TJ, Donnelly JP, Revised definitions of invasive fungal disease from the European Organization for Research and Treatment of Cancer/Invasive Fungal Infections Cooperative Group and the National Institute of Allergy and Infectious Diseases Mycoses Study Group (EORTC/MSG) Consensus Group: Clin Infect Dis, 2008; 46(12); 1813-21
20. Borlenghi E, Cattaneo C, Capucci MA, Usefulness of the MSG/IFICG/EORTC diagnostic criteria of invasive pulmonary aspergillosis in the clinical management of patients with acute leukaemia developing pulmonary infiltrates: Ann Hematol, 2007; 86(3); 205-10
21. Kollef M, Micek S, Hampton N: Clin Infect Dis, 2012; 54(12); 1739-46
22. Zhang H, Zhu A, Emerging invasive fungal infections: Clinical features and controversies in diagnosis and treatment processes: Infect Drug Resist, 2020; 13; 607-15
23. Shourian M, Qureshi ST, Resistance and tolerance to cryptococcal infection: An intricate balance that controls the development of disease: Front Immunol, 2019; 10; 66
24. Kobayashi S, Soyama A, Takatsuki M, Relationship between immune function recovery and infectious complications in patients following living donor liver transplantation: Hepatol Res, 2016; 46(9); 908-15
25. Jones TM, Drew RH, Wilson DT, Impact of automatic infectious diseases consultation on the management of fungemia at a large academic medical center: Am J Health Syst Pharm, 2017; 74(23); 1997-2003
26. Aljeboori Z, Gorelik A, Jenkins E, Risk factors for candidaemia and their cumulative effect over time in a cohort of critically ill, non-neutropenic patients: Crit Care Resusc, 2018; 20(4); 313-19
27. Venkatesh MP, Pham D, Fein M: Antimicrob Agents Chemother, 2007; 51(4); 1240-45
28. Roux D, Gaudry S, Khoy-Ear L, Airway fungal colonization compromises the immune system allowing bacterial pneumonia to prevail: Crit Care Med, 2013; 41(9); e191-99
Tables
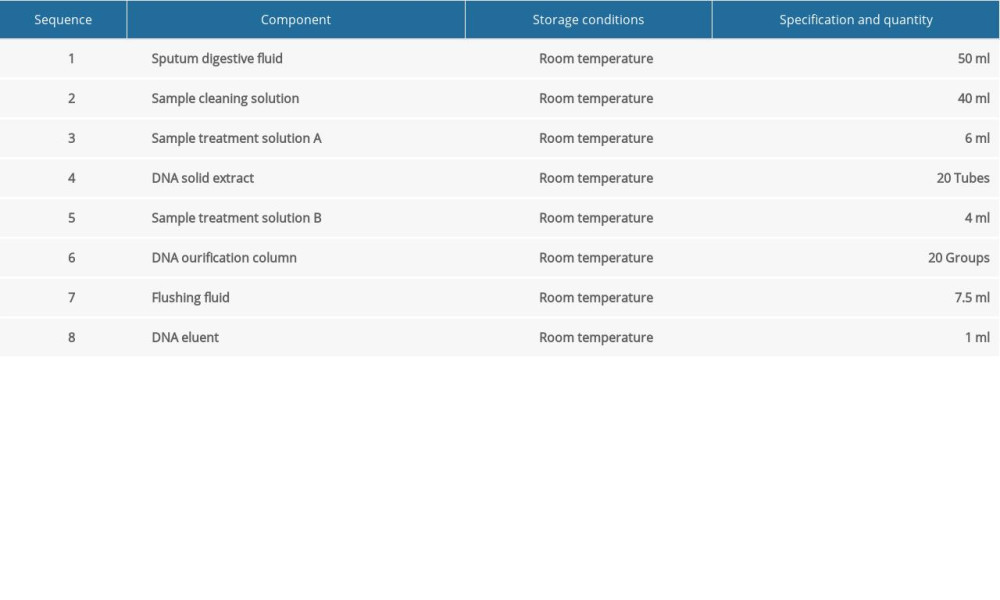 Table 1. The sputum DNA extraction kit.
Table 1. The sputum DNA extraction kit.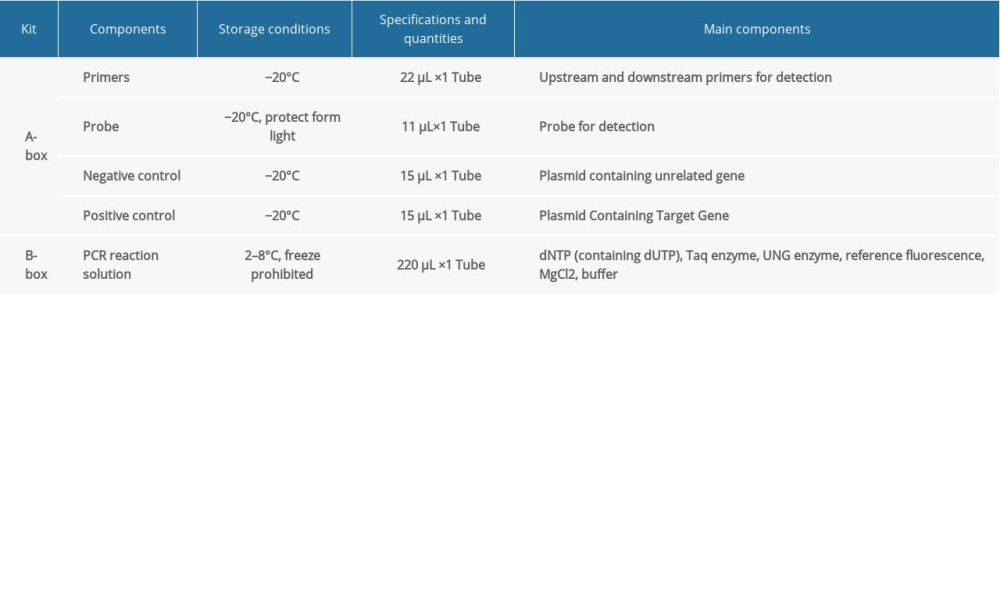 Table 2. The universal nucleic acid detection kit.
Table 2. The universal nucleic acid detection kit.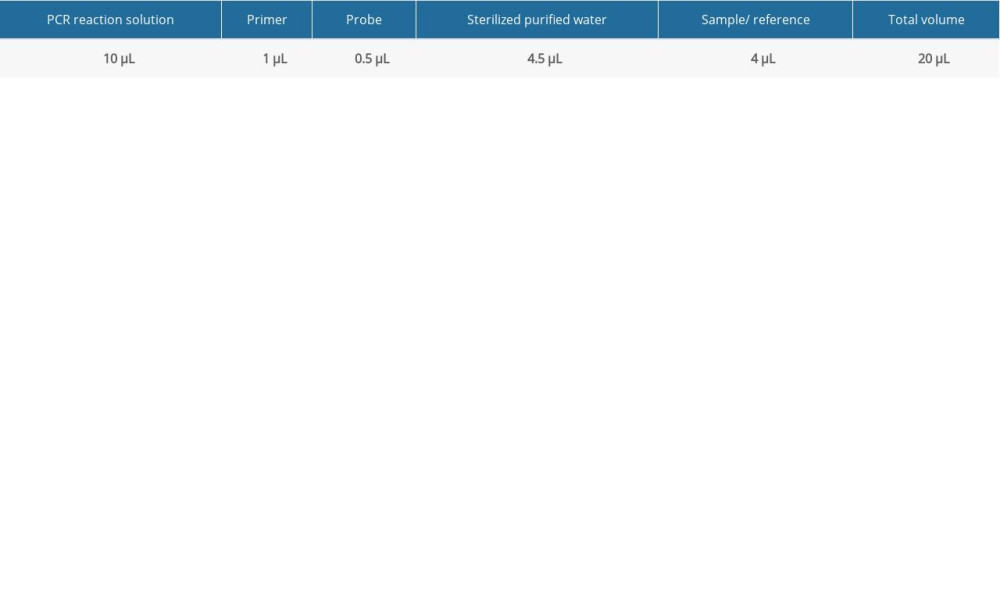 Table 3. Fluorescent PCR detection reaction system.
Table 3. Fluorescent PCR detection reaction system.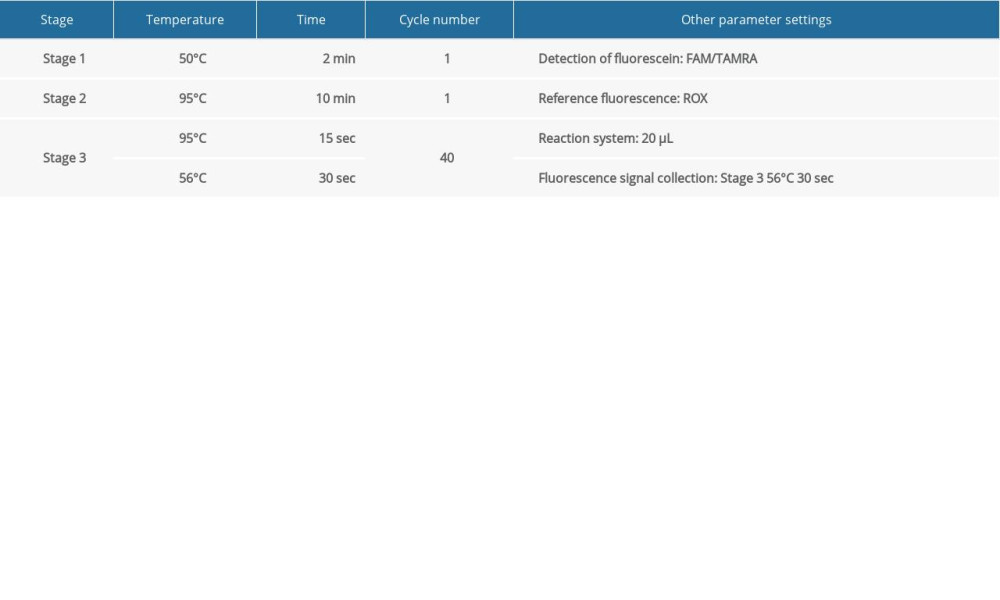 Table 4. Detection conditions of fluorescent PCR.
Table 4. Detection conditions of fluorescent PCR.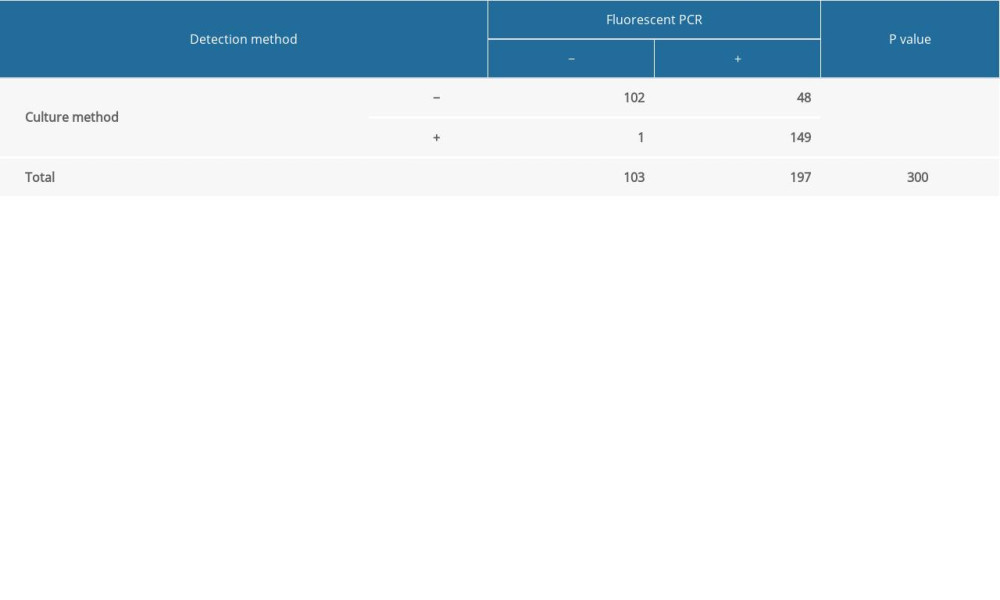 Table 5. Detection results of culture method and fluorescent PCR.
Table 5. Detection results of culture method and fluorescent PCR. Table 1. The sputum DNA extraction kit.
Table 1. The sputum DNA extraction kit. Table 2. The universal nucleic acid detection kit.
Table 2. The universal nucleic acid detection kit. Table 3. Fluorescent PCR detection reaction system.
Table 3. Fluorescent PCR detection reaction system. Table 4. Detection conditions of fluorescent PCR.
Table 4. Detection conditions of fluorescent PCR. Table 5. Detection results of culture method and fluorescent PCR.
Table 5. Detection results of culture method and fluorescent PCR. In Press
12 Mar 2024 : Clinical Research
Preoperative Blood Transfusion Requirements for Hemorrhoidal Severe Anemia: A Retrospective Study of 128 Pa...Med Sci Monit In Press; DOI: 10.12659/MSM.943126
12 Mar 2024 : Clinical Research
Tissue Inhibitors of Metalloproteinase 1 (TIMP-1) and 3 (TIMP-3) as New Markers of Acute Kidney Injury Afte...Med Sci Monit In Press; DOI: 10.12659/MSM.943500
12 Mar 2024 : Review article
Optimizing Behçet Uveitis Management: A Review of Personalized Immunosuppressive StrategiesMed Sci Monit In Press; DOI: 10.12659/MSM.943240
12 Mar 2024 : Clinical Research
Metabolomic Alterations in Methotrexate Treatment of Moderate-to-Severe PsoriasisMed Sci Monit In Press; DOI: 10.12659/MSM.943360
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952