Abstract
Background
The prognostic significance of microsatellite instability (MSI) in colorectal cancers (CRCs) has been addressed in many studies since the initial description of better survival rates in MSI-positive (MSI+) tumors than in MSI-negative (MSI−) tumors. Recent studies have demonstrated that a higher degree of hypomethylation of long interspersed nuclear element-1 (L1) is related to poor prognosis of CRCs and that a wide variation of L1 methylation levels exist within MSI+ CRCs. Our aim was to identify whether L1 and Alu methylation status could predict clinical outcomes within MSI+ CRCs.
Methods
We analyzed 207 MSI+ CRCs for their methylation levels in L1 and Alu repetitive DNA elements using pyrosequencing and correlated them with clinicopathological information including survival data.
Results
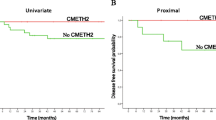
Univariate survival analysis showed that low Alu methylation status (<18.60 %) and low L1 methylation status (<53.00 %) were significantly associated with shorter overall survival time (log-rank test, P = 0.009 and P < 0.001, respectively). Multivariate analysis using nine parameters (Alu methylation status, L1 methylation status, patient’s age, disease stage [tumor, node, metastasis staging system], differentiation, Crohn-like lymphoid reaction, KRAS/BRAF mutation status, CpG island methylator phenotype [CIMP] status, and peritumoral lymphocytic infiltration), which were significantly prognostic in MSI+ CRCs, revealed that low L1 methylation status was an independent prognostic factor of MSI+ CRCs (P = 0.009), whereas low Alu methylation status was not.
Conclusions
Clinical outcomes of MSI+ CRCs depend on L1 methylation status, suggesting that lower L1 methylation status serves as a significant prognostic parameter of adverse prognosis in MSI+ CRCs.


Similar content being viewed by others
References
Sung JJ, Lau JY, Goh KL, Leung WK. Increasing incidence of colorectal cancer in Asia: implications for screening. Lancet Oncol. 2005;6:871–6.
Boland CR, Goel A. Microsatellite instability in colorectal cancer. Gastroenterology. 2010;138:2073–87, e2073.
Poynter JN, Siegmund KD, Weisenberger DJ, et al. Molecular characterization of MSI-H colorectal cancer by MLHI promoter methylation, immunohistochemistry, and mismatch repair germline mutation screening. Cancer Epidemiol Biomarkers Prev. 2008;17:3208–15.
Samowitz WS, Curtin K, Ma KN, et al. Microsatellite instability in sporadic colon cancer is associated with an improved prognosis at the population level. Cancer Epidemiol Biomarkers Prev. 2001;10:917–23.
Bae JM, Kim MJ, Kim JH, et al. Differential clinicopathological features in microsatellite instability–positive colorectal cancers depending on CIMP status. Virchows Arch. 2011;459:55–63.
Kim JH, Shin SH, Kwon HJ, Cho NY, Kang GH. Prognostic implications of CpG island hypermethylator phenotype in colorectal cancers. Virchows Arch. 2009;455:485–94.
Ahn JB, Chung WB, Maeda O, et al. DNA methylation predicts recurrence from resected stage III proximal colon cancer. Cancer. 2011;117:1847–54.
Kang GH. Four molecular subtypes of colorectal cancer and their precursor lesions. Arch Pathol Lab Med. 2011;135:698–703.
Soreide K, Janssen EA, Soiland H, Korner H, Baak JP. Microsatellite instability in colorectal cancer. Br J Surg. 2006;93:395–406.
Thibodeau SN, Bren G, Schaid D. Microsatellite instability in cancer of the proximal colon. Science. 1993;260:816–9.
Benatti P, Gafa R, Barana D, et al. Microsatellite instability and colorectal cancer prognosis. Clin Cancer Res. 2005;11:8332–40.
Guidoboni M, Gafa R, Viel A, et al. Microsatellite instability and high content of activated cytotoxic lymphocytes identify colon cancer patients with a favorable prognosis. Am J Pathol. 2001;159:297–304.
Popat S, Hubner R, Houlston RS. Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol. 2005;23:609–18.
Guastadisegni C, Colafranceschi M, Ottini L, Dogliotti E. Microsatellite instability as a marker of prognosis and response to therapy: a meta-analysis of colorectal cancer survival data. Eur J Cancer. 2010;46:2788–98.
Boland CR. Clinical uses of microsatellite instability testing in colorectal cancer: an ongoing challenge. J Clin Oncol. 2007;25:754–6.
Kim GP, Colangelo LH, Wieand HS, et al. Prognostic and predictive roles of high-degree microsatellite instability in colon cancer: a National Cancer Institute–National Surgical Adjuvant Breast and Bowel Project Collaborative Study. J Clin Oncol. 2007;25:767–72.
Lander ES, Linton LM, Birren B, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921.
Cordaux R, Batzer MA. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 2009;10:691–703.
Yoder JA, Walsh CP, Bestor TH. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 1997;13:335–40.
Ehrlich M. DNA methylation in cancer: too much, but also too little. Oncogene. 2002;21:5400–13.
Weisenberger DJ, Campan M, Long TI, et al. Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res. 2005;33:6823–36.
Baba Y, Huttenhower C, Nosho K, et al. Epigenomic diversity of colorectal cancer indicated by LINE-1 methylation in a database of 869 tumors. Mol Cancer. 2010;9:125.
Ogino S, Nosho K, Kirkner GJ, et al. A cohort study of tumoral LINE-1 hypomethylation and prognosis in colon cancer. J Natl Cancer Inst. 2008;100:1734–8.
Ogino S, Kawasaki T, Nosho K, et al. LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer. 2008;122:2767–73.
Estecio MR, Gharibyan V, Shen L, et al. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS One. 2007;2:e399.
Lee S, Cho NY, Choi M, Yoo EJ, Kim JH, Kang GH. Clinicopathological features of CpG island methylator phenotype–positive colorectal cancer and its adverse prognosis in relation to KRAS/BRAF mutation. Pathol Int. 2008;58:104–13.
Lee S, Cho NY, Yoo EJ, Kim JH, Kang GH. CpG island methylator phenotype in colorectal cancers: comparison of the new and classic CpG island methylator phenotype marker panels. Arch Pathol Lab Med. 2008;132:1657–65.
Tellez CS, Shen L, Estecio MR, Jelinek J, Gershenwald JE, Issa JP. CpG island methylation profiling in human melanoma cell lines. Melanoma Res. 2009;19:146–55.
Pobsook T, Subbalekha K, Sannikorn P, Mutirangura A. Improved measurement of LINE-1 sequence methylation for cancer detection. Clin Chim Acta. 2011;412:314–21.
Kitkumthorn N, Mutirangura A. Long interspersed nuclear element-1 hypomethylation in cancer: biology and clinical applications. Clin Epigenet. 2011;1–16.
Brena RM, Auer H, Kornacker K, et al. Accurate quantification of DNA methylation using combined bisulfite restriction analysis coupled with the Agilent 2100 Bioanalyzer platform. Nucleic Acids Res. 2006;34:e17.
Irahara N, Nosho K, Baba Y, et al. Precision of pyrosequencing assay to measure LINE-1 methylation in colon cancer, normal colonic mucosa, and peripheral blood cells. J Mol Diagn. 2010;12:177–83.
Sunami E, de Maat M, Vu A, Turner RR, Hoon DS. LINE-1 hypomethylation during primary colon cancer progression. PLoS One. 2011;6:e18884.
Acknowledgment
This study was supported by a grant from the National R&D Program for Cancer Control, Ministry of Health and Welfare, Republic of Korea (0720540), by a grant from the Seoul National University Hospital Research Fund (0320110220), by A3 Foresight Program from National Research Foundation of Korea (NRF), and by a Mid-career Researcher Program through NRF grant funded by the Ministry of Education, Science and Technology (MEST) (2011-0015646).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rhee, YY., Kim, M.J., Bae, J.M. et al. Clinical Outcomes of Patients with Microsatellite-Unstable Colorectal Carcinomas Depend on L1 Methylation Level. Ann Surg Oncol 19, 3441–3448 (2012). https://doi.org/10.1245/s10434-012-2410-7
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-012-2410-7