Abstract
Background
The tissue environment in the region of hepatocellular carcinoma (HCC) influences both vascular invasion and recurrence. Thus, HCC patient prognosis depends on the characteristics not only of the tumor but also those of adjacent surrounding liver tissue.
Materials and Methods
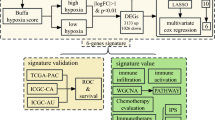
Expression profiles of both tumor and adjacent liver tissue following curative resection were measured to discriminate 56 hepatitis B virus-positive HCC patients into subgroups based on survival risk. This approach was further tested in 40 patients.
Results
Expression profiles of both tumor and adjacent liver tissue successfully discriminated 56 training samples into 2 subgroups, those at low- or high-risk for survival and recurrence. However, the prognostic gene set selected for tumor tissue was quite different from that for adjacent tissues. This variation in prognostic genes resulted in a change in allocation of patients within each low- or high-risk group. Combination of survival subgroups from tumor and adjacent liver tissue significantly improved the prediction of prognostic outcome. This integrative approach was confirmed to be effective in a further 40 test patients. A clinicopathological study showed that survival subgroups divided by tumor and adjacent liver tissue gene expression were also statistically associated with UICC stage and extent of cell differentiation, respectively.
Conclusions
Variation in gene expression during the nontumor stage as well as the tumor stage may affect the prognosis of HCC patients, and integration of the gene expression profiles of HCC and adjacent liver tissue increases discriminatory effectiveness between patient groups, predicting clinical outcomes with enhanced statistical reliability.





Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55:74–108.
Nam SW, Park JY, Ramasamy A, et al. Molecular changes from dysplastic nodule to hepatocellular carcinoma through gene expression profiling. Hepatology. 2005;42:809–18.
Ye QH, Qin LX, Forgues M, et al. Predicting hepatitis B virus-positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat Med. 2003;9:416–23.
Lee JS, Chu IS, Heo J, et al. Classification and prediction of survival in hepatocellular carcinoma by gene expression profiling. Hepatology. 2004;40:667–76.
Kim BY, Lee JG, Park S, et al. Feature genes of hepatitis B virus-positive hepatocellular carcinoma, established by its molecular discrimination approach using prediction analysis of microarray. Biochim Biophys Acta. 2004;1739:50–61.
Neo SY, Leow CK, Vega VB, et al. Identification of discriminators of hepatoma by gene expression profiling using a minimal dataset approach. Hepatology. 2004;39:944–53.
Lee JS, Heo J, Libbrecht L, et al. A novel prognostic subtype of human hepatocellular carcinoma derived from hepatic progenitor cells. Nat Med. 2006;12:410–6.
Boyault S, Rickman DS, de Reyniès A, et al. Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology. 2007;45:42–52.
Chiang DY, Villanueva A, Hoshida Y, et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 2008;68:6779–88.
Pons F, Varela M, Llovet JM. Staging systems in hepatocellular carcinoma. HPB (Oxford). 2005;7:35–41.
Hoshida Y, Villanueva A, Kobayashi M, et al. Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med. 2008;359:1995–2004.
Eng C, Leone G, Orloff MS, Ostrowski MC. Genomic alterations in tumor stroma. Cancer Res. 2009;69:6759–64.
Lee DC, Kang YK, Kim WH, et al. Functional and clinical evidence for NDRG2 as a candidate suppressor of liver cancer metastasis. Cancer Res. 2008;68:4210–20.
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J, et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 2002;30:e15.
Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95:14863–8.
Tibshirani R, Hastie T, Narasimhan B, Chu G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci USA. 2002;99:6567–72.
Weichert W, Röske A, Niesporek S, et al. Class I histone deacetylase expression has independent prognostic impact in human colorectal cancer: specific role of class I histone deacetylases in vitro and in vivo. Clin Cancer Res. 2008;14:1669–77.
Weichert W, Röske A, Gekeler V, et al. Association of patterns of class I histone deacetylase expression with patient prognosis in gastric cancer: a retrospective analysis. Lancet Oncol. 2008;9:139–48.
Deng S, Zhou H, Xiong R, et al. Over-expression of genes and proteins of ubiquitin specific peptidases (USPs) and proteasome subunits (PSs) in breast cancer tissues observed by the methods of RFDD-PCR and proteomics. Breast Cancer Res Treat. 2007;104:21–30.
Agarwala SS, Keilholz U, Gilles E, et al. LDH correlation with survival in advanced melanoma from two large, randomised trials (Oblimersen GM301 and EORTC 18951). Eur J Cancer. 2009;45:1807–14.
Balendiran GK, Martin HJ, El-Hawari Y, Maser E. Cancer biomarker AKR1B10 and carbonyl metabolism. Chem Biol Interact. 2009;178:134–7.
Penning TM. AKR1B10: a new diagnostic marker of non-small cell lung carcinoma in smokers. Clin Cancer Res. 2005;11:1687–90.
Yoshitake H, Takahashi M, Ishikawa H, et al. Aldo-keto reductase family 1, member B10 in uterine carcinomas: a potential risk factor of recurrence after surgical therapy in cervical cancer. Int J Gynecol Cancer. 2007;17:1300–6.
Teramoto R, Minagawa H, Honda M, et al. Protein expression profile characteristic to hepatocellular carcinoma revealed by 2D-DIGE with supervised learning. Biochim Biophys Acta. 2008;1784:764–72.
Macheiner D, Gauglhofer C, Rodgarkia-Dara C, et al. NORE1B is a putative tumor suppressor in hepatocarcinogenesis and may act via RASSF1A. Cancer Res. 2009;69:235–42.
Buckingham L, Penfield Faber L, Kim A, et al. PTEN, RASSF1 and DAPK site-specific hypermethylation and outcome in surgically treated stage I and II nonsmall cell lung cancer patients. Int J Cancer. 2010;126:1630–9.
Misawa A, Tanaka S, Yagyu S, Tsuchiya K, Iehara T, Sugimoto T, et al. RASSF1A hypermethylation in pretreatment serum DNA of neuroblastoma patients: a prognostic marker. Br J Cancer. 2009;100:399–404.
Mousavi SA, Sporstøl M, Fladeby C, Kjeken R, Barois N, Berg T. Receptor-mediated endocytosis of immune complexes in rat liver sinusoidal endothelial cells is mediated by FcgammaRIIb2. Hepatology. 2007;46:871–4.
Thorgeirsson SS, Grisham JW. Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet. 2002;31:339–46.
Lee JS, Thorgeirsson SS. Genome-scale profiling of gene expression in hepatocellular carcinoma: classification, survival prediction, and identification of therapeutic targets. Gastroenterology. 2004;127:S51–5.
Utsunomiya T, Shimada M, Imura S, Morine Y, Ikemoto T, Mori M. Molecular signatures of noncancerous liver tissue can predict the risk for late recurrence of hepatocellular carcinoma. J Gastroenterol. 2010;45:146–52.
Tsuchiya M, Parker JS, Kono H, Matsuda M, Fujii H, Rusyn I. Gene expression in nontumoral liver tissue and recurrence-free survival in hepatitis C virus-positive hepatocellular carcinoma. Mol Cancer. 2010;9:74.
Woo HG, Park ES, Cheon JH, Kim JH, Lee JS, Park BJ, et al. Gene expression-based recurrence prediction of hepatitis B virus-related human hepatocellular carcinoma. Clin Cancer Res. 2008;14:2056–64.
Acknowledgment
This work was supported by grants from the National Nuclear R & D program and Human Genome Project, the Korean Ministry of Science and Technology. M.H.C. was supported by a grant from National Research Foundation of Korea (NRF-2010-0000784).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Bu-Yeo Kim and Kyung-Suk Suh contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, BY., Suh, KS., Lee, JG. et al. Integrated Analysis of Prognostic Gene Expression Profiles from Hepatitis B Virus-Positive Hepatocellular Carcinoma and Adjacent Liver Tissue. Ann Surg Oncol 19 (Suppl 3), 328–338 (2012). https://doi.org/10.1245/s10434-011-1709-0
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-011-1709-0