-
PDF
- Split View
-
Views
-
Cite
Cite
Luciani R. Carvalho, Michelle L. Brinkmeier, Frederic Castinetti, Buffy S. Ellsworth, Sally A. Camper, Corepressors TLE1 and TLE3 Interact with HESX1 and PROP1, Molecular Endocrinology, Volume 24, Issue 4, 1 April 2010, Pages 754–765, https://doi.org/10.1210/me.2008-0359
Close - Share Icon Share
Abstract
Pituitary hormone deficiency causes short stature in one in 4000 children born and can be caused by mutations in transcription factor genes, including HESX1, PROP1, and POU1F1. HESX1 interacts with a member of the groucho-related gene family, TLE1, through an engrailed homology domain and represses PROP1 activity. Mice with Prop1 deficiency exhibit failed differentiation of the POU1F1 lineage, resulting in lack of TSH, GH, and prolactin. In addition, these mutants exhibit profound pituitary dysmorphology and excess Hesx1 and Tle3 expression. The ability of HESX1 to interact with TLE3 has not been explored previously. We tested the ability of TLE3 to enhance HESX1-mediated repression of PROP1 in cell culture. Both TLE3 and TLE1 repress PROP1 in conjunction with HESX1 with similar efficiencies. TLE1 and TLE3 can each repress PROP1 in the absence of HESX1 via a protein-protein interaction. We tested the functional consequences of ectopic TLE3 and HESX1 expression in transgenic mice by driving constitutive expression in pituitary thyrotrophs and gonadotrophs. Terminal differentiation of these cells was suppressed by HESX1 alone and by TLE3 and HESX1 together but not by TLE3 alone. In summary, we present evidence that HESX1 is a strong repressor that can be augmented by the corepressors TLE1 and TLE3. Our in vitro studies suggest that TLE1 and TLE3 might also play roles independent of HESX1 by interacting with other transcription factors like PROP1.
The pituitary gland is derived from two different layers during embryogenesis. The neuroectoderm, originating from the floor of the diencephalon, will give rise to the posterior pituitary lobe, which stores and releases oxytocin and antidiuretic hormone. The oral ectoderm, derived from the roof of the mouth, will give rise to Rathke’s pouch, the primordium for the anterior and intermediate pituitary lobes (1). The anterior lobe will give rise to five different cell types known as somatotrophs, lactotrophs, gonadotrophs, thyrotrophs, and corticotrophs, which produce and release six different hormones: GH, prolactin, LH and FSH, TSH, and ACTH, respectively. These hormones have important roles in growth, lactation, fertility, metabolism, and stress response in humans and mice.
There are many transcription factors involved in the commitment and differentiation of cell types in the anterior lobe of the pituitary gland. Spontaneous mutants and genetically engineered murine models have demonstrated a role for many of these molecules in the etiology of pituitary hormone deficiency. These include the transcription factors HESX1, PROP1, POU1F1 (PIT1), LHX3, LHX4, OTX2, TBX19 (TPIT), SOX2, and SOX3 (reviewed in Ref. 2). In humans, the phenotypes produced by lesions in these genes range from isolated hormone deficiency to more complex disorders such as septo-optic dysplasia and holoprosencephaly associated with combined pituitary hormone deficiency (CPHD). Mutations in PROP1 are the most prevalent known cause of CPHD in humans. This is likely due to a hot spot in the sequence that is vulnerable to a 2-bp deletion (3), but the etiology of many cases of CPHD remains unknown. Thus, it is clear that other genes remain to be identified, and the characterization of these will elucidate the pathogenesis of these complex conditions and also shed light on normal pituitary development.
One of the first genes to be expressed in murine Rathke’s pouch is Hesx1 at embryonic d 9.0 (e9.0), and its expression wanes significantly after e12.5 (4). The Prop1 gene is expressed in the developing pituitary gland at e10.5, peaks at e12.5, and decreases after e14.5, when Pou1f1 expression is initiated (5). Mouse mutants with lesions in these three genes, Hesx1, Prop1, and Pou1f1, model human CPHD and have contributed to understanding the mechanism of action of each gene (4–11). Mechanistic analysis of known genes is a strategy for identifying genes that could explain unknown genetic etiologies for CPHD. For example, Prop1 is necessary for the inhibition of Hesx1 transcription (11) and activation of Pou1f1 expression, and each of these genes is mutated in some cases of CPHD in humans (4, 12, 13). In mice, Prop1 affects the expression of many other genes, including restriction of Tle3 expression to the dorsal part of the anterior pituitary (14) and the activation of Notch2 (7), but these genes are unlikely to be mutated in humans with CPHD due to their involvement in development of many organs critical for viability.
Tle3 (transducin-like enhancer of split 3), and several other members of the Gro/TLE family of groucho-related genes are expressed during pituitary development (9, 14, 15). The Gro/TLE family is comprised of six different genes Tle1-4, Aes, and Tle6 (16, 17). TLE1-4 each has glutamine repeats (Q region) at the N terminus and a WDR (WD-40 repeat) region for protein-protein interaction at the C terminus. AES has the Q region but lacks the WDR region, whereas TLE6 contains a WDR region but no Q region. The TLE genes act as corepressors by binding to transcription factors such as HESX1 and TCF/LEF family members that have the engrailed homology domain (eh1) and SIX family members that contain short peptide sequences WRPW.
PROP1 and HESX1 contain paired-type homeodomains, and they bind to DNA as homodimers or heterodimers at a paired homeodomain binding site, P3 (TAATTGAATTA, core sequence is in bold) (5, 9, 18). PROP1 acts as an activator of this promoter in transfection, and its activity is repressed by HESX1. HESX1 recruits TLE1 through an eh1 domain creating a complex that is a stronger repressor of PROP1 activity than HESX1 alone. Mutations in the eh1 domain of HESX1 are associated with impaired PROP1 repression in tissue culture studies and also impaired recruitment of TLE1 in coimmunoprecipitation studies (9). A patient with short stature due to GH deficiency and evolving pituitary hormone deficiency harbored a mutation in the eh1 domain (HESX1I26T), confirming the importance of the eh1 domain in vivo (19).
TLE1 and TLE3 a have similar protein structure, and their patterns of expression during pituitary development are overlapping, but distinct (9, 14, 16). TLE1 is expressed throughout the pituitary at e12.5 and e14.5, whereas TLE3 is initially expressed in the ventral diencephalon at e12.5 and then becomes restricted to the cells dorsal to the lumen of Rathke’s pouch at e14.5. This restricted expression is aberrant in the Prop1 mutant mice in which TLE3 is expressed extensively in the developing pituitary gland (14).
The current study advances our understanding of the activity of TLE1, TLE3, PROP1, and HESX1 on gene expression in cell culture and in pituitary development. Our cell culture results reveal that TLE3 functions similarly to TLE1 in enhancing HESX1 repression of PROP1 activity. Surprisingly, TLE1 and TLE3 can repress PROP1 in the absence of HESX1 through protein interaction. We tested the ability of ectopic TLE3 expression to interfere with pituitary development, as a single transgene or in conjunction with HESX1. The results demonstrate that HESX1 has a strong inhibitory effect on differentiation that is modestly enhanced by TLE3, but TLE3 has no effect on its own in this ectopic context. These results suggest that persistent, ectopic expression of HESX1 alone is sufficient to inhibit cell differentiation in transgenic mice and that HESX1 repression may be mediated by TLE1 and/or TLE3. Taken together, these studies in cells and animals advance our understanding of pituitary development at the molecular level.
Results
TLE3 enhances HESX1 repression of PROP1
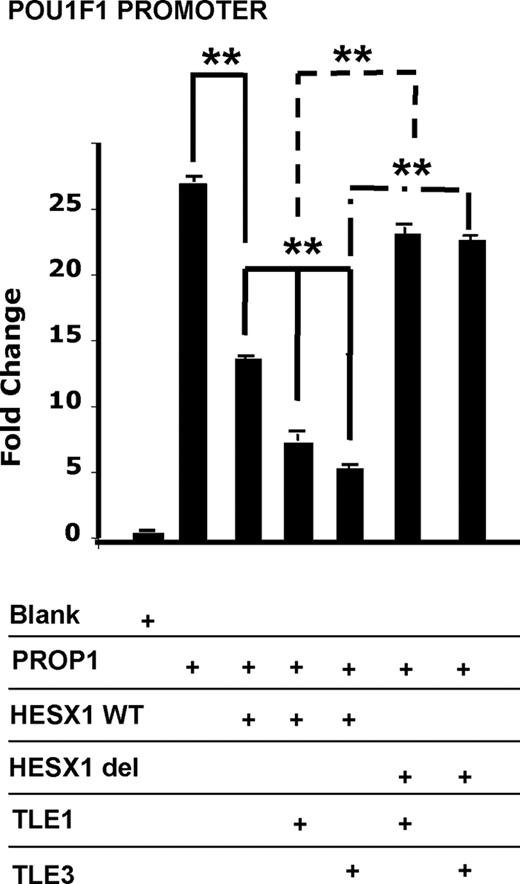
Previous studies have shown that TLE1 and HESX1 interact to repress PROP1 activity (9, 19, 20). To test the potential of TLE1 and TLE3 to act as corepressors with HESX1 in regulating PROP1 activity, transient transfection studies were performed in cell culture using 293T cells (human embryonic kidney). We used expression vectors for human TLE1, TLE3, PROP1, HESX1 wild type (WT), and HESX1(Δ1–50), which is missing the N-terminal 50 amino acids including the eh1 domain. As expected, PROP1 activated the POU1F1 promoter when transfected alone, and HESX1 WT, HESX1(Δ50-185), TLE1, or TLE3 had no activating effect over the reporter alone (data not shown). HESX1 WT repressed PROP1 activation by 46%. HESX1 together with TLE1 or TLE3 enhanced PROP1 repression up to 66 and 79%, respectively (Fig. 1). TLE1 and TLE3 consistently produced significant repressive effects. HESX1(Δ1–50), which harbors a deletion of the eh1 domain, was used to test TLE1 and TLE3 corepression and recruitment through the eh1 domain. Repression was significantly impaired in the absence of the eh1 domain (P < 0.001). The same results were obtained with the multimerized consensus sequence reporter plasmids P36E4 and PRDQ3 (data not shown).
Gro/TLE family members exert effects on HESX1-mediated repression of PROP1 activation. 293T cells were transiently transfected with expression vectors for PROP1, HESX1 WT, HESX1(Δ1-50), TLE1, or TLE3 with the POU1F1 promoter driving the luciferase reporter. The normalized results are expressed as fold change relative to the vector backbone. The experiments were performed in triplicate in at least three independent experiments, and typical results are shown. **, P < 0.001.
PROP1 repression by TLE1 and TLE3 occurs independently of HESX1
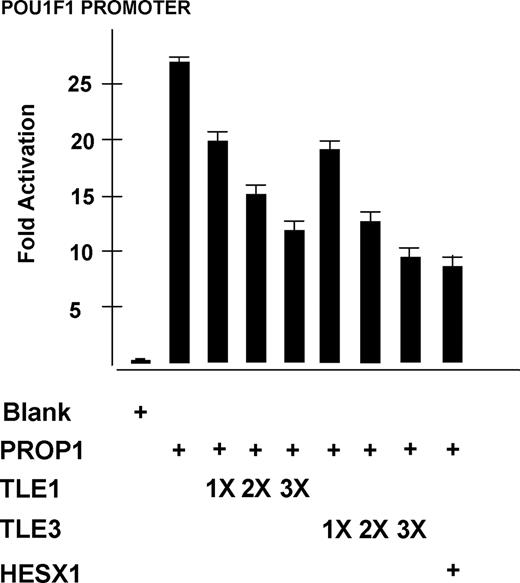
To evaluate the requirement of HESX1 for recruitment of the TLE/Gro family proteins (TLE1 and TLE3), PROP1 activity was analyzed in the absence of HESX1 and the presence of TLE1 or TLE3 individually. PROP1 expression vector was cotransfected with vectors expressing TLE1 or TLE3 in 293T cells with the three previously described reporters coupled to luciferase and varying amounts of empty vector to keep the total amount of transfected DNA constant. With the POU1F1 promoter, the addition of 50 ng TLE1 or TLE3 expression vectors repressed PROP1 activation by 27 and 37%, respectively (Fig. 2). Dose-response curves show that transfecting increasing amounts of TLE1 or TLE3 strengthens the repressive effects, with a maximal effect observed for 200 and 250 ng (plateau phase). The maximal effect of TLE with PROP1 in the absence of HESX1 is the same as that observed with HESX1 and PROP1 independent of TLE. These data suggest that TLE1 and TLE3 can repress PROP1 activation of a minimal promoter in the absence of HESX1. The same results were obtained in αT3-1 cells, a gonadotrope-like cell line, and with the PRDQ3 and P36E4 reporters constructs (data not shown).
Gro/TLE family members repress PROP1 activity in the absence of HESX1. 293T cells were transiently transfected with POU1F1 promoter-luciferase reporter construct with 50 ng PROP1 in the absence of HESX1. Increasing amounts of TLE1 or TLE3 were transfected: 50 ng (1X), 150 ng (2X), and 250 ng (3X). Maximal repressive effects were observed for 200 and 250 ng (plateau phase, data not shown). The luciferase activity produced from PROP1 alone and from PROP1 and TLE1 or TLE3 together was statistically different for the three amounts of DNA transfected (P < 0.001).
TLE repression of PROP1 likely involves protein-protein interaction
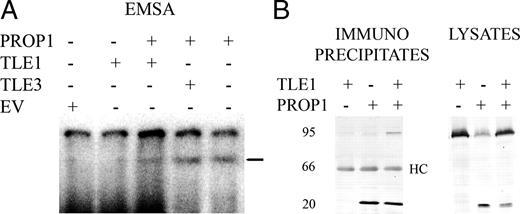
To determine the mechanism by which TLE factors repress PROP1 activity, we performed EMSA to ascertain whether the TLE factors could bind the PROP1 consensus recognition sequence (PRDQ DNA element). We did not observe any interaction between this DNA element and TLE1 or TLE3, whereas PROP1 bound the element as expected (Fig. 3). To determine whether protein interactions might be involved, we transfected expression vectors for PROP1-myc alone, TLE1 alone, and PROP-myc together with TLE1, prepared extracts, immunoprecipitated with anti-myc antibody, and probed Western blots with the anti-myc antibody followed by the anti-TLE antibody (Fig. 3). We observed a band corresponding to TLE1 that coimmunoprecipitated with PROP1, suggesting that there could be an interaction between PROP1 and TLE factors. This interaction may be weak and/or transient given the intensity of the TLE band relative to the amount of DNA transfected.
TLE interaction with PROP1 DNA. A, Radiolabeled PRDQ oligonucleotide was incubated with TLE1, TLE3, and/or PROP1 proteins produced in reticulocyte lysates. Reticulocyte lysates with empty vector (EV) served as a negative control. The samples were separated by gel electrophoresis and autoradiographed. PROP1 bound the probe in this EMSA (bar). B, TLE1 and/or PROP1-myc were transfected in 293T cells (5 μg each in the cotransfection of TLE1 and PROP1, 10 μg for transfection with each individual vector). Cell lysates were prepared and subjected to immunoprecipitation with anti-myc epitope antibodies, followed by SDS-PAGE. Lysates, One eighth of each input lysate, collected before incubation with antibodies, was subjected to gel electrophoresis. After transfer to nitrocellulose, Western blotting (WB) was performed with either anti-myc or anti-Gro/TLE (pan-TLE) antibodies. Immuno-precipitates, Coimmunoprecipitation was performed with the anti-myc antibody. Western blot was performed with myc antibody (left) and pan-TLE antibody. PROP1 migrates at 25 kDa, TLE at 95 kDa, and the heavy chain (HC) at 66 kDa. Endogenous TLE is detectable in lysates from cells not transfected with TLE1. Positions of migration of the IgG heavy chain (HC) are indicated. Positions of size standards are indicated in kilodaltons.
Ectopic TLE3 and HESX1 expression represses differentiation in transgenic mice
The functional consequences of ectopic expression of the repressor Hesx1 and a corepressor Tle3 in pituitary development were determined through temporal and spatial misexpression in transient transgenic mice. Transgene constructs were designed by placing either mouse TLE3 or HESX1 coding sequences in frame downstream of 4.6 kb of the mouse pituitary glycoprotein hormone α-subunit promoter (αGSU), officially named Cga, followed by the mouse protamine 1 (Prm1) 3′-untranslated region, splice, polyadenylylation, and termination sequences. The well-characterized Cga promoter drives transient expression in Rathke’s pouch and persistent, strong coexpression with Cga in thyrotrophs and gonadotrophs during development and throughout adulthood (21). Tg(Cga-Tle3) and Tg(Cga-Hesx1) transgene constructs were both coinjected and singly injected into fertilized eggs, and fetuses were harvested at e18.5 and e14.5. The presence of each transgene was identified in genomic DNA generated from either embryonic tail or limb biopsies using PCR amplification with transgene-specific primers.
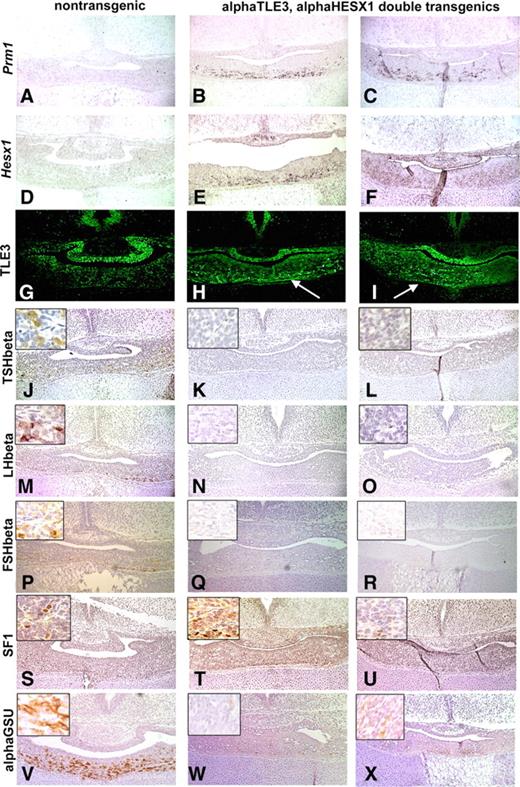
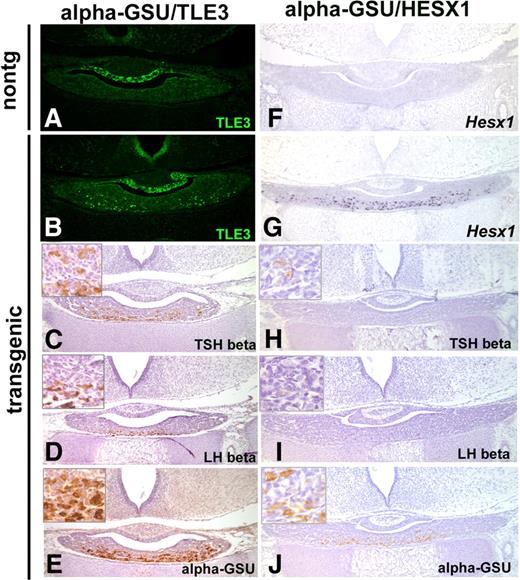
Coinjection of the transgene constructs produced a total of 481 embryos with 8.5% containing both transgenes, 2% containing only Tg(Cga-Hesx1), and 1% containing only Tg(Cga-Tle3). The level of transgene expression in Tg(Cga-Tle3), Tg(Cga-Hesx1) double-transgenic embryos at e18.5 was analyzed by in situ hybridization using a transgene-specific Prm1 probe (Fig. 4, A–C). Strong Prm1 expression was detected in the ventral cells of double-transgenic embryos, whereas no expression was present in nontransgenic controls. HESX1 and TLE3 both contributed substantially to the overall level of transgene expression. Transgenics have detectable Hesx1 mRNA and TLE3 protein in the ventral cells of the anterior pituitary gland, whereas nontransgenic littermate controls lack Hesx1 mRNA and restrict TLE3 protein to the dorsal region of the pituitary gland (Fig. 4, D–I). Six fetuses expressed the transgenes of 24 double transgenics.
Ectopic, constitutive expression of HESX1 and TLE3 in transgenic embryos affects gonadotroph and thyrotroph differentiation. Levels of transgene expression were examined in coronal sections of normal littermates and double-transgenic embryos at e18.5: Tg(Cga-Tle3), Tg(Cga-Hesx1). The first column (A, D, G, J, M, P, S, and V) display a representative nontransgenic littermate, whereas the second column (B, E, H, K, N, Q, T, and W) and the third column (C, F, I, L, O, R, U, and X) demonstrate two independent double-transgenic embryos designated as αTLE3, αHESX1. Mouse protamine 1 (Prm1) mRNA expression was used to select transgenics with robust expression levels for further analysis (A–C), whereas the expression of the individual transgene constructs was visualized by Hesx1 in situ hybridization (D–F) and TLE3 immunohistochemistry (G–I). The arrows in H and I depict the ectopic expression of TLE3 in the ventral portion of the pituitary gland. The differentiation of thyrotrophs and gonadotrophs was analyzed using immunohistochemistry with antibodies directed against TSHβ (J–L), LHβ (M–O), FSHβ (P–R), SF1 (S–U), and αGSU or CGA (V–X).
The consequence of inappropriate TLE3 and HESX1 expression in the gonadotrophs and thyrotrophs was analyzed in the highest expressing double transgenics. The differentiation of gonadotrophs and thyrotrophs was blocked in Tg(Cga-Tle3), Tg(Cga-Hesx1) double-transgenic embryos. Immunohistochemistry using antibodies against TSH, LH, and FSH readily detected positive cells in nontransgenic controls; however, no immunopositive cells were detected in sections from four of four expressing, double-transgenic embryos (Fig. 4, J–R), suggesting that transgene expression blocked final differentiation of both thyrotrophs and gonadotrophs. Somatotrophs, lactotrophs, and corticotrophs were appropriately represented in double-transgenic embryos (data not shown). The presence of the earliest gonadotroph marker, the nuclear hormone receptor SF1 (NR5A1) in Tg(Cga-Tle3), Tg(Cga-Hesx1) double-transgenic embryos, together with the deficiency in LH and FSH expression, suggests that the gonadotroph cell lineage was determined but not fully differentiated (Fig. 4, S–U). Interestingly, chorionic gonadotropin alpha (CGA, also called αGSU) protein was dramatically reduced in double-transgenic embryos compared with nontransgenic controls, suggesting the repressor/corepressor complex can negatively regulate endogenous αGSU expression (Fig. 4, V–X), but not the transgene itself. The most plausible explanation for this is the presence of an important, negative, regulatory sequence in or around the endogenous gene that is absent in the 4.6 kb of Cga 5′-flanking sequence in the transgene.
Injection of only the Tg(Cga-Tle3) construct produced 22 embryos at e18.5, 14% of which contained the transgene, whereas injection of the Tg(Cga-Hesx1) construct produced 66 embryos at e18.5, 8% of which contained the transgene. Transgenic embryos expressing Tg(Cga-Tle3) alone had appropriate gonadotroph and thyrotroph differentiation (Fig. 5, A–E). In contrast, transgenic embryos expressing Tg(Cga-Hesx1) alone exhibit a dramatic reduction in TSH- and LH-positive cells (Fig. 5, F–J). However, the expression of endogenous Cga was not affected. This suggests that the ectopic expression of HESX1 is sufficient to block terminal differentiation of gonadotrophs and thyrotrophs, although TLE3 may be required for inhibition of endogenous Cga.
Ectopic expression of HESX1 alone, but not TLE3, affects gonadotroph and thyrotroph differentiation. Levels of transgene expression were determined by TLE3 immunohistochemistry for Tg(Cga-Tle3) or αGSU-TLE3 transgenic embryos and nontransgenic littermates at e18.5 (A and B) or Hesx1 in situ hybridization for Tg(Cga-Hesx1) or αGSU-HESX1 transgenic embryos and nontransgenic littermates (nontg) at e18.5 (F and G). TSHβ and LHβ immunohistochemistry is used to screen for differentiated gonadotrophs and thyrotrophs in Tg(Cga-Tle3) embryos (C and D) and Tg(Cga-Hesx1) embryos (H and I). CGA or αGSU protein is present in both Tg(Cga-Tle3) and Tg(Cga-Hesx1) single-transgenic embryos (E and J).
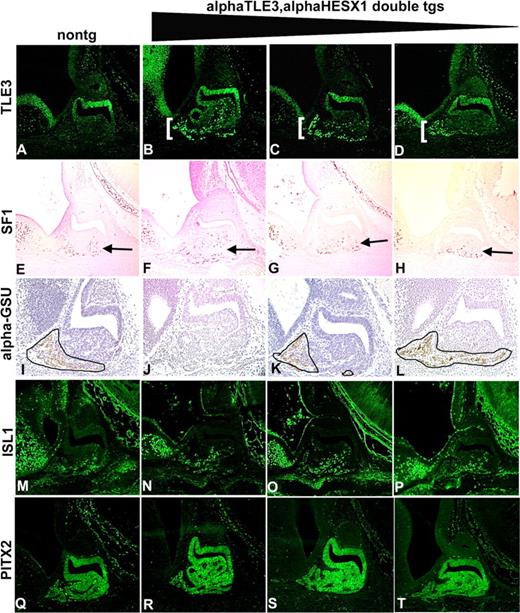
To determine whether the differentiation of gonadotrophs was delayed by expression of HESX1 and/or TLE3 transgenes, transient transgenics were analyzed at the normal timing of onset of SF1 protein expression, e14.5. High levels of ectopic TLE3 protein expression were detected in the ventral cells of the developing anterior pituitary gland in three Tg(Cga-Tle3), Tg(Cga-Hesx1) double transgenics (Fig. 6, A–D). The presence of SF1 in double-transgenic e14.5 embryos demonstrates that an early differentiation step of the gonadotroph cell lineage is not delayed (Fig. 6, E–H). A reduction in endogenous CGA protein levels was visualized in the highest expressing e14.5 double-transgenic embryos, consistent with data obtained from e18.5 double-transgenic embryos (Fig. 6, I–L). Other markers of cell differentiation, ISL1 and PITX2, were not changed between nontransgenic and double-transgenic littermates (Fig. 6, M–T). In vivo data support the role of HESX1 as a strong repressor that can be enhanced by the presence of at least two of the TLE/groucho family members.
Initiation of gonadotroph differentiation in double-transgenic embryos. The three highest expressing Tg(Cga-Tle3), Tg(Cga-Hesx1) double-transgenic embryos (designated αTLE3,αHESX1 double tgs) were selected based on ectopic immunohistochemical staining for TLE3 at e14.5 in the ventral cells of Rathke’s pouch in sagittal sections (columns beginning with B, C, and D; gradation bar indicates decreasing transgene expression levels) compared with a nontransgenic littermate (column beginning with A). SF1 (NR5A1) immunohistochemical staining detected both mature and pre-gonadotrophs (E–H, arrows). Immunohistochemistry using antibodies against αGSU (CGA) (I–L) demonstrated a decrease in expression correlating with level of transgene expression (area of expression outlined). There was no difference in protein levels of ISL1 (M–P) or PITX2 (Q–T) in transgenics and nontransgenic littermates.
Discussion
Transcriptional repressors, activators, and cofactors work together to orchestrate pituitary gland organogenesis. This likely involves interplay between PROP1, HESX1, SIX3, and other transcription factors with coregulators from the TLE/groucho family: TLE1 and TLE3 (22). We demonstrate functional interactions between a subset of these proteins in tissue culture and also through misexpression of HESX1 and TLE3 together and singly in the context of pituitary development.
HESX1 is a strong repressor
HESX1 and TLE1 are coexpressed at e12.5 in the same region of the developing pituitary gland (Fig. 7) (9). TLE1 binds HESX1 and enhances its repressor activity in cell culture and in transgenic mice (9, 19, 20). We confirmed that HESX1 inhibits PROP1-mediated transactivation in cell culture and that TLE1 acts as a corepressor, enhancing this inhibition. The effects we observed are of the same magnitude as other reports. We discovered that ectopic HESX1 alone is a strong repressor of differentiation in transgenic mice, blocking expression of Tshb and Lhb almost completely and modestly repressing Cga expression. This contrasts with the idea that ectopic TLE1 expression is necessary for robust repression of differentiation by ectopic HESX1 transgene expression (9).
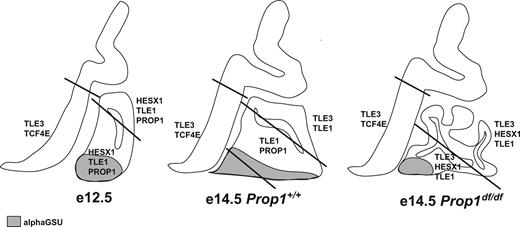
Expression domains of transcription factors, corepressors, and hormone subunits in developing pituitary gland and surrounding tissues. Rathke’s pouch and the overlying neural ectoderm are illustrated diagrammatically in the sagittal plane in normal mice at e12.5 and e14.5 and in Prop1 mutants at e14.5. The diagonal lines mark domains that are defined by differences in gene expression. Tle3 and Tcf7l2 (TCF4E) are expressed in the lower domain of the ventral diencephalon. Rathke’s pouch grows significantly between e12.5 and e14.5 and becomes dysmorphic in Prop1 mutants, probably because of an inability to produce a Notch2-positive transitional zone intermediate between the dorsal proliferating cells and the ventral differentiating cells. Cga, The α-subunit of the glycoprotein hormones LH, FSH, and TSH, is expressed in the differentiated cells of Rathke’s pouch that expand beyond the rostral tip at e12.5 to include more caudomedial cells at e14.5 in normal mice but not Prop1 mutants (gray). The diagram illustrates that 1) HESX1 and TLE1 could repress PROP1 activity at e12.5, 2) ectopic HESX1 and TLE3 expression might contribute to the Prop1 mutant phenotype, and 3) TLE1 and TLE3 likely have interacting partners besides PROP1 and HESX1.
The ability of HESX1 to function as a strong repressor in vitro and in vivo is supported by mutations identified in human patients. Two mutations that lead to hypopituitarism alter HESX1 interactions with TLE factors. A patient with septo-optic dysplasia and hypopituitarism is heterozygous for 1684delG in HESX1 that creates increased DNA binding and more potent repression of PROP1-mediated gene transcription (23). Transgenic mice that misexpress HESX1 have similar features to this patient. A patient with a mutation in the eh1 repression domain of HESX1 (HESX1I26T) exhibited evolving hypopituitarism and no central nervous malformation. This mutation has no effect on DNA binding but impairs HESX1-mediated repression because of failure to recruit TLE1 (19).
The HESX1-TLE1 complex is proposed to suppress PROP1-mediated transcription of Pou1f1 until β-catenin displaces TLE1 (9). Are other genes natural targets of the HESX1-TLE1 repressor complex? These genes would be expected to be overexpressed in Hesx1 mutant mice. Foxd3, Pax3, Dmbx1, and Sp5 expression and Wnt signaling are expanded and/or elevated in the anterior neural ridge of Hesx1 mutants (24, 25). Additional investigation of pituitary gene expression in Hesx1 mutants may be necessary to uncover the direct targets of HESX1-mediated repression.
Mechanism of TLE-PROP1 action
TLE1 and TLE3 have highly conserved protein structures. The overall amino acid identity is 80% with 86% identity in the glutamine-rich region (Q) and 95% in the WD repeat region (16). This conservation suggests a similar mechanism of action of TLE1 and TLE3. There are areas of overlapping expression in pituitary development (Fig. 7), and we observe similar effects of TLE1 and TLE3 in transfection studies and transgenic mice. TLE3 and TLE1 repress PROP1 activation of the POU1F1 promoter independent of HESX1. The mechanism of this TLE-mediated repression likely involves protein-protein interaction because TLE does not bind the paired homeodomain recognition sequence in EMSA, but it coimmunoprecipitates with PROP1 when coexpressed in 293T cells. Although another study did not detect coimmunoprecipitation, the protein interaction may be weak or transient (9). However, the interaction of PROP1 and TLE is consistent with chromatin immunoprecipitation studies that show occupancy of the Pou1f1 early enhancer by TLE and PROP1 at e12.5 after HESX1 is released (20). This supports the idea that PROP1 and TLE interaction could have biological significance.
TLE factors interact with transcription factors that contain WRPW or eh1 motifs (17, 22). Analysis of the PROP1 peptide sequence identified only weak alignment with the WRPW and eh1 motifs, suggesting if there is a direct interaction between TLE and PROP1, then other unknown sequences might be involved in mediating the interaction (26). Alternatively, the protein-protein interaction may be indirect. Although the 293T cells do not express endogenous HESX1, we cannot rule out the possibility that another protein is present in the coimmunoprecipitated complex or involved in the TLE-mediated repression of PROP1 activation.
Role for ectopic TLE3 and HESX1 expression in Prop1 mutants?
Prop1 mutants express both Tle3 and Hesx1 ectopically, in overlapping areas, which could contribute to the pituitary developmental abnormalities characteristic of Prop1 mutants (Fig. 7). This idea is supported by our demonstration that TLE3 can enhance the repression mediated by HESX1 in cell culture and transgenic mice. Terminal differentiation of thyrotrophs and gonadotrophs is suppressed by HESX1 in the presence or absence of TLE3, but TLE3 alone has no effect in this context. Endogenous CGA expression is suppressed by TLE3 and HESX1 together but not by HESX1 alone. These results suggest that ectopic HESX1 and TLE3 expression in Prop1 mutants could contribute to failed differentiation. The failed thyrotroph differentiation in Prop1 mutants is attributable to the requirement of PROP1 for Pou1f1 expression, which in turn is essential for caudomedial expression of Tshb. However, the gonadotropin deficiency in humans with PROP1 mutations is poorly understood, and it is one of the clinical hallmarks used to distinguish patients with PROP1 and POU1F1 mutations. Our results suggest that ectopic TLE3 and HESX1 expression may contribute to the gonadotropin deficiency in patients with PROP1 mutations, although other explanations, such as depletion of precursors, and bystander effects causing cell death, could contribute.
What is the normal function of TLE3?
Tle3 is expressed in the prospective intermediate lobe from e14.5–e16.5 around the time that Hesx1 and Prop1 transcripts have normally waned (Fig. 7). The transgenic expression studies were aimed to assess the consequences of ectopic Tle3 and Hesx1 expression in Prop1 mutants. The neutral effect of TLE3 on differentiation in Tg(Cga-Tle3) single transgenics implies that there were no endogenous repressors that could interact with TLE3 in the ventral aspect of Rathke’s pouch to suppress differentiation in this context. The endogenous partners for TLE3, however, are likely coexpressed dorsally in the dividing cells.
The Nkx, Hes, Runx, Lef/Tcf, and Six gene families encode proteins with a WRPW or eh1 motif that can interact with the Gro/TLE family (27, 28). We identified members of some of these families in cDNA libraries of microdissected Rathke’s pouch at e12.5 and e14.5 (29, 30). These include Nkx3.1, Nkx2.4, Runx1, Hes6, and Otx2 and three members of the Six gene family. Six3 is coexpressed with Tle3 and regulates proliferation (22, 31, 32). This suggests that TLE3 may be involved in regulating the differentiation of cells in the dorsal aspect of Rathke’s pouch and/or regulating proliferation. Deficiency of either HES1 or TBX19 (TPIT) permits premature differentiation of these cells into GH- or LH-expressing cells, respectively, so they clearly have the capacity for differentiation into a variety of cell types unless appropriately regulated (33–37). Immunohistochemistry and in situ hybridization studies reveal that Six6 and Nkx3.1 have in a similar temporal and spatial expression pattern to Prop1 (8, 38) (our unpublished observations). In addition, Six3, Tcf3, and Hes1 have also been identified in the developing anterior pituitary gland at e14.5 (20, 34, 35, 39, 40). Any of the genes encoding interactors with TLE1 and TLE3 are candidate genes for screening patients with pituitary abnormalities.
Materials and Methods
Generation of transient transgenics
The open reading frame of mouse Tle3 was amplified with forward primer 5′-gccaccatggactacaaggacgacgatgacaagatgtatccgcaaggcag-3′ and reverse primer 5′-tcagtagatgacctcgtaaactgtggccttctt-3′ using an e14.5 Rathke’s pouch cDNA library as the template (29, 30). The Tle3 open reading frame was cloned into the TOPO-TA cloning vector (Invitrogen, Carlsbad, CA) using the manufacturer’s protocol, and its identity was confirmed by DNA sequence analysis. Tle3 was placed downstream of the previously characterized mouse αGSU promoter (Cga) and upstream of a mouse Protamine 1 (Prm1) 3′-untranslated and 3′-flanking sequences, which contain an intron and polyadenylation sequences, using the HindIII and EcoRV sites (21). The identity of the Cga-Tle3 transgene construct was confirmed by DNA sequence analysis. Before microinjection, the transgene insert was purified from the vector sequences after digestion with the restriction enzymes Asp718 and NotI.
The open reading frame of mouse Hesx1 was amplified with forward primer 5′-gccaccatggactacaaggacgacgatgacaaggtctcccagccttcgggaaggtgctcag-3′ and reverse primer 5′-ctatttcagaagatctgggttgaagggttttttcgccattagaaa-3′ using cDNA from an e14.5 Prop1df/df Rathke’s pouch library as a template (29, 30). The Hesx1 open reading frame was cloned in the pGEM T-easy cloning vector (Promega, Madison, WI), and the integrity was confirmed by sequence analysis. The Hesx1 open reading frame was excised with EcoRI and subcloned into the pBluescript SK+ (Stratagene, La Jolla, CA). The completed transgene construct was generated by cloning the −4.6-kb mouse αGSU promoter and enhancer sequences into the KpnI and HindIII site upstream of Hesx1, and the mouse Prm1 sequences described above were cloned downstream into the BamHI/XbaI sites. The identity of the mouse Cga-Hesx1 transgene construct was confirmed by DNA sequence analysis. The transgene construct was released from the vector backbone using BssHII. Purified DNA from both transgene constructs was either comicroinjected or singly microinjected into fertilized eggs by The University of Michigan Transgenic Animal Model Core. Fertilized eggs were obtained from mating (C57BL/6 × SJL) F1 female and male mice. Pronuclear microinjection was performed, and injected eggs were implanted into CD-1 surrogate mothers at the two-cell stage.
Embryo genotyping
Embryos were collected at d 14.5 and 18.5 of gestation (e14.5 and e18.5). Embryonic d 0.5 was designated as the day the microinjected fertilized eggs were surgically transferred to pseudo-pregnant foster females. Genomic DNA extracted from either tail or limb biopsies was used to genotype the embryos by PCR amplification with transgene-specific primers. The transgenes were amplified using a forward primer within the mouse Cga promoter, 5′-tcaactttcaggatgttttgtgtaa-3′, and reverse primers specific to either Tle3, 5′-cccgatgatggcgttcaa-3′, or Hesx1, 5′-taggggtgggttgccacc-3′. Products of 740 bp for the Cga-Tle3 transgene and 558 bp for the Cga-Hesx1 transgene were amplified at 94 C for 4 min followed by 30 cycles of 94 C for 30 sec, 55 C for 30 sec, and 72 C for 30 sec and ending in 72 C for 10 min and visualized with ethidium bromide on a 1% agarose gel.
In situ hybridization
Embryos were fixed in buffered 4% paraformaldehyde for 90 min (e14.5) or overnight (e18.5). After fixation, the embryos were washed in PBS and dehydrated through a series of ethanol solutions. The embryos were embedded in paraffin, sectioned to 6 μm thickness, and processed as described below. In situ hybridization was performed using riboprobes labeled with digoxigenin (Roche, Indianapolis, IN) (15). Tge 394-bp Hesx1 cDNA clone was a generous gift from Dr. Paul Thomas (Brisbane, Australia). The BamHI linearized fragment was transcribed with T3 polymerase and hybridized at a 1:100 dilution overnight at 53 C. A 511-bp Prm1 probe was generated from the 3′ end of the transgene construct. The BamHI linearized fragment was transcribed with T3 polymerase, diluted 1:100, and hybridized to tissue sections overnight at 53 C.
Immunohistochemistry
Sagittal (e14.5) and coronal (e18.5) sections of paraffin-embedded embryos were used in the immunohistochemistry experiments. Slides were boiled for 10 min in 10 mm citric acid if destined for analysis with PITX2, ISL1, TLE3, or NR5A1 antibodies. All slides were blocked with a 1:1 mixture of methanol/3% hydrogen peroxide to inactivate endogenous peroxidases. Rabbit anti-PITX2 (1:400; Dr. Tord Hjalt, Lund University, Sweden), mouse anti-ISL1 (1:600; Developmental Studies Hybridoma Band, University of Iowa), and rabbit anti-TLE3 (1:150; Millipore, Billerica, MA) were incubated overnight at 4 C. The following day, the sections were washed and then incubated with either a biotinylated antimouse IgG (1:200, Vectastain Mouse on Mouse kit; Vector Laboratories, Burlingame, CA) or a biotinylated antirabbit IgG (1:200; Jackson ImmunoResearch Laboratories, Inc., West Grove, PA). The antibodies were detected using the reagents and protocols provided in the tyramide signal amplification fluorescein kit (PerkinElmer, Boston, MA). Mouse anti-NR5A1 (1:1500; Dr. Ken-ichirou Morohashi, National Institute for Basic Biology, Okazaki, Japan), monkey anti-GH [1:1000; National Hormone Pituitary Program (NHPP), Torrance, CA], guinea pig anti-LH (1:500; National Hormone Pituitary Program, NHPP), rabbit anti-TSH (1:1000; NHPP), rabbit anti-proopiomeanocortin (1:1000; NHPP), rabbit anti-FSH (1:400; NHPP), and rabbit anti-αGSU (1:200; NHPP) were also incubated for 2 h at room temperature. Biotinylated secondary antibodies were used in conjunction with avidin and biotinylated peroxidase (Vectastain Mouse on Mouse kit, Vectastain Rabbit Elite kit, Vectastain Human Elite kit, and Vectastain Guinea Pig kit; Vector). With this experiment, diaminobenzidine was used as the chromogen (Sigma, St. Louis, MO). All immunostaining experiments were done in duplicate in any one experiment, and experiments were repeated to sample regions throughout the pituitary gland.
Plasmid constructs for transfection
Human PROP1 cDNA cloned into pCDNA3.1 was a gift from Dr. Simon Rhodes (Indiana University School of Medicine, Indianapolis, IN) (41). Human HESX1 cDNA was generated from clones encoding GAL4-HESX1 fusion proteins (42), released with EcoRI and BamHI and subcloned into pCDNA3.1. GAL4-HESX1 WT, GAL4-HESX1(Δ1-50), POU1F1 promoter coupled to luciferase and (P3)6E4 luciferase plasmids were gifts from Dr. Mehul Dattani (Biochemistry, Endocrinology, and Metabolism Unit, Institute of Child Health, London, UK) (19). Human pCMV2-FLAG-GroTLE1 and TLE3 cloned into the pCIneo mammalian expression vector were gifts from Dr. Stefano Stifani (Center for Neuronal Survival, Montreal Neurological Institute, McGill University, Montreal, Canada) (43). pCDNA3.1 was used as a negative control vector (Invitrogen). pRL-TK plasmid DNA (thymidine kinase promoter-driven renilla luciferase; Promega) was used as an internal control for the transfection.
Cell culture, transient transfection, and dual luciferase reporter assay
The αT3-1 (mouse pituitary pre-gonadotroph) (45) was a gift from Dr. Pamela Mellon (University of San Diego, La Jolla, CA). These and 293T cells (human embryonic kidney) were maintained at 37 C, 5% CO2 in DMEM (Invitrogen) supplemented with 10% heat-inactivated fetal bovine serum (Hyclone, Logan, UT). Cells were plated onto 24-well plastic plates (Fisher Scientific, Fair Lawn, NJ) at a density of 3 × 104 (293T) and 6 × 104 (αT3-1) such that the cells were 50–80% confluent on the day of transfection. Three reporter constructs coupled to luciferase were used for transfections: a 15-kb mouse Pou1f1 promoter and two artificial promoters constructed with three (PRDQ3) or six (P36E4) paired homeodomain transcription factor binding sites. A DNA cocktail consisting of a combination of the following was transfected: 100 ng of the reporter construct, 10 ng pRL-TK renilla as an internal control, 50 ng PROP1, 50 ng HESX1, 50 ng TLE1, 50 ng TLE3, and pcDNA 3.1 empty expression vector ranging from 50–200 ng to normalize up to 310 ng/well. Optimal DNA concentrations were determined by titrating increasing amounts of HESX1 (25, 50, 75, and 100 ng) and constant Prop1 at 50 ng (data not shown). A dose-response assay was carried out by transfecting variable quantities of TLE1 or TLE3 individually with 50 ng PROP1 expression vector. The range of corepressor expression vector amounts were 50, 100, 150, 200, and 250 ng of TLE1 or TLE3 cotransfected with PROP1, and pcDNA 3.1 empty expression vector ranging from 0–200 ng to generate a total of 310 ng/well. DNA was transfected in 500 μl serum-free DMEM (Invitrogen) into culture cells using Fugene 6 (Roche) at a ratio of 3:1 according to the manufacturer’s protocol. For each transfection experiment, the pcDNA 3.1 empty expression vector was used as a negative control, and measurement of the basal level of luciferase activity. A total of 300 ng DNA was transfected to keep the total amount of DNA transfected consistent. Forty-eight hours after transfection, dual-luciferase assays (Promega) were performed according to the manufacturer’s protocol and quantified using the Lmax Microplate Illuminometer (Molecular Devices, Sunnyvale, CA) with the SOFT max Pro software (Molecular Devices). All assays were done in triplicate, and the results were repeated at least three times. Results were averaged and expressed as fold change of the percent activity of the experimental plasmids over basal.
EMSA and coimmunoprecipitation studies
Gel mobility shift assays were performed to search for TLE DNA interactions on a radiolabeled annealed synthetic PROP1 recognition element (PRDQ3 oligonucleotide). PRDQ3 oligonucleotide was labeled with [32P]deoxy-CTP. PROP1, TLE1, and TLE3 were transcribed and translated using the TNT-coupled reticulocyte lysate system with T7 polymerase (Promega). The protein-DNA complexes were analyzed by electrophoresis through a 5% polyacrylamide gel containing 2.5% glycerol in a 0.5× Tris-borate buffer at 4 C.
For the coimmunoprecipitation study, human HEK 293 cells were transfected with PROP1 and/or TLE1 (10 μg total DNA, with 10 μg PROP1 alone, 10 μg TLE1 alone, or 5 μg PROP1 and 5 μg TLE1). Whole-cell lysates were prepared and subjected to immunoprecipitation as described previously (43), followed by Western blotting with antibodies against the myc epitope (Invitrogen) or the Gro/TLE proteins (pan-TLE antibodies, a generous gift from Dr. Stefano Stifani) (44). The blot was visualized with the Odyssey infrared imaging system (LI-COR Biosciences, Lincoln, NE).
Statistical analysis
SigmaStat statistical software version 2.0 (SPSS Inc., Chicago, IL) was used for data analysis. Statistically significant differences between the input groups were considered as P < 0.001 in the t test and one-way ANOVA.
Acknowledgments
We acknowledge Wanda Filipiak, D.V.M.; Dr. Maggie Van Keuren; and Galina Gavrilina, M.S., for the preparation of transgenic mice and Dr. Thomas L. Saunders of the Transgenic Animal Model Core in the University of Michigan’s Biomedical Research Core Facilities. We thank Mary Anne Potok for assistance with analysis of transgenic animals. We thank the following individuals and programs for providing reagents: Drs. Mehul Dattani, Tord Hjalt, Pamela Mellon, Ken-ichirou Morohashi, Takashi Okamoto, Simon Rhodes, Paul Thomas, Stefano Stifani, and the National Hormone Pituitary Program. We thank Drs. Bob Lyons and the University of Michigan DNA Sequencing Core, Stefano Stifani of McGill University, Alexander Augusto Lima Jorge (Clinicas’ Hospital-FMUSP, Brazil), Christopher Krebs (University of Michigan, Department of Human Genetics), and Travis Maure (University of Michigan, Department of Molecular and Integrative Physiology) for scientific contributions and valuable discussions.
This work was supported by National Institutes of Health (NIH) R37HD30428 and R01HD34283 (S.A.C.); The Endocrine Society International Scholars Program (L.R.C.); Novo-Nordisk, Societe Française d’endocrinologie (Novartis-Ipsen), and ADEREM (F.C.); F32 HD046300 (B.S.E.); NIH Center Grants CA46592, AR20557, DK34933, DK34933, and P30AG013283 (University of Michigan Transgenic Animal Model Core); and Grant 085P1000815 (University of Michigan Center for Organogenesis, the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor).
Transgenic animal analysis was done by M.L.B. Cell culture studies with the (P3)6E4 promoter were done by L.R.C. and repeated by B.S.E., whereas the PRDQ3 and Pou1f1 promoter transfections were done by F.C. The EMSA and coimmunoprecipitation were by F.C. The paper was written by M.L.B., F.C., L.R.C., and S.A.C.
Current address for L.R.C.: Unidade de Endocrinologia do Desenvolvimento, Laboratório de Hormônios e Genética Molecular LIM/42, Disciplina de Endocrinologia, Hospital das Clínicas da Faculdade de Medicina da Universidade de São Paulo, Brasil.
Disclosure Summary: None of the authors have conflicts of interests to disclose.
L.R.C., M.L.B., and F.C. are equally contributing authors.
Abbreviations:
- CGA,
Chromogranin A;
- CPHD,
combined pituitary hormone deficiency;
- e9.0,
embryonic d 9.0;
- αGSU,
glycoprotein hormone α-subunit promoter;
- WT,
wild type.