-
PDF
- Split View
-
Views
-
Cite
Cite
Robert C. Gensure, Bhaskar Ponugoti, Yasemin Gunes, Madhusudhan R. Papasani, Beate Lanske, Murat Bastepe, David A. Rubin, Harald Jüppner, Identification and Characterization of Two Parathyroid Hormone-Like Molecules in Zebrafish, Endocrinology, Volume 145, Issue 4, 1 April 2004, Pages 1634–1639, https://doi.org/10.1210/en.2003-0964
Close - Share Icon Share
Abstract
Zebrafish (Danio rerio) have receptors homologous to the human PTH (hPTH)/PTHrP receptor (PTH1R) and PTH-2 receptor (PTH2R) and an additional receptor (PTH3R) with high homology to the PTH1R. To find natural ligands for zPTH1R and zPTH3R, we searched the zebrafish genomic database and discovered two distinct regions that, when translated (zPTH1 and zPTH2), showed high homology to hPTH. Isolation of cDNAs and determination of the intron/exon boundaries revealed genomic structures which were similar to known PTHs. Peptides consisting of the first 34 amino acids after the pre- and prosequences of the zebrafish PTHs (zPTHs) were synthesized and were shown to be fully active at the hPTH1R. zPTH2(1–34) was, however, approximately 30-fold less potent at the zPTH1R than hPTH(1–34), hPTHrP(1–36), and zPTH1(1–34). When tested with zPTH3R, zPTH1(1–34) and hPTHrP(1–36) showed similar potencies, whereas the potency of zPTH2(1–34) was moderately (3-fold) reduced. To determine whether other fishes have multiple PTHs, we searched the genomic database of the Japanese pufferfish (Takifugu rubripes) and identified zPTH1 and zPTH2 homologs. Phylogenetic analysis showed that PTHs from zebrafish and pufferfish are more closely related to each other than to known mammalian PTH homologs or to PTHrP and tuberoinfundibular peptide of 39 residues. This is consistent with evolution of two teleost PTH-like peptides occurring after the evolutionary divergence between fishes and mammals. Overall, the PTH system appears more complex in fishes than in mammals, providing evidence of continued evolution in nontetrapod species. The availability of multiple forms of fish PTH and their receptors provide additional tools for PTH ligand/receptor structure-function studies.
PTH AND PTHrP MEDIATE their biological effects by acting as ligands for the same receptor (1, 2), the PTH/PTHrP receptor (PTH1R). Peptides containing the first 34 amino acids of PTH and PTHrP are capable of fully activating the PTH1R (1, 2), and interest in generating potent PTH analogs has increased since the report of PTH(1–34) as an effective treatment for osteoporosis (3–5). Structure-function studies have suggested that PTH(1–34) and PTHrP(1–34) both have two principal functional domains: the amino- terminal domain, which is important for receptor activation; and the carboxyl-terminal domain, which is important for high-affinity binding (2, 6). A second receptor, the PTH2R, is activated in humans by PTH but not PTHrP (7–9). Based on these findings, it was initially postulated that the PTH2R may be responsible for mediating some PTH-specific responses; however, it has since been discovered that the PTH2R has a distinct ligand, tuberoinfundibular peptide, which appears to function in hypothalamic neuroendocrine regulation (10), but may also have roles in renal blood flow regulation (11) and spermatogenesis (12). Furthermore, PTH is only a partial agonist for the PTH2R in other species (13), including zebrafish (Danio rerio) (14), and it therefore seems unlikely that this receptor has a major role in PTH signaling.
Homologs to the human PTH (hPTH)/PTHrP receptor have been found in a variety of other species, both mammalian and nonmammalian. In zebrafish, homologs to the hPTH1R (15) and PTH2R (16) have been described. A third PTH/PTHrP receptor (zPTH3R) has also been described in zebrafish; this receptor bears greater homology to zPTH1R than to zPTH2R (15). Despite this homology, the pharmacological properties of this receptor differ from those of the zPTH1R, in that hPTHrP is more potent at the zPTH3R than hPTH (15). Despite several attempts in this laboratory, a mammalian homolog to the zPTH3R has not been cloned, nor did a search of the human genome reveal any DNA regions that are homologous to the zPTH3R (17). It therefore appears likely that the PTH1R and the PTH3R evolved through a gene duplication event that occurred in fishes and potentially other species, but either never occurred in mammals or has been lost in these species. Biological roles mediated by zPTH1R and zPTH3R in zebrafish are currently unknown.
The native zebrafish ligands for zPTH1R and zPTH3R have not been described, although a single fugu (Takifugu rubripes) analog has recently been described (18). Fishes do not have an anatomical structure that corresponds to the mammalian parathyroid glands. In fact, these glands only become apparent in amphibians, i.e. with the development of the terrestial form of life. In this study, we sought to discover PTH-like ligand(s) in fishes that interact with the zPTH1R and zPTH3R. We found that zebrafish and pufferfish each have two distinct PTH peptides. The genomic structure, cDNA, amino acid sequence, and some pharmacological properties of these peptides are described.
Materials and Methods
Identification of putative zPTH transcripts
hPTH cDNA (GenBank accession no. NM_000315) was used as a probe to search the zebrafish genome web site (http://134.174.23.160/HumanblastZebrafish/) for homologous sequences. Two putative zebrafish genomic DNA regions (zPTH1, zfish35939-334d11.p1c; zPTH2, zfishJ-a492f09.p1c) that showed sequence homology to the rat PTH transcript were identified. Gene-specific primers for zPTH1 and zPTH2 were designed for RT-PCR and rapid amplification of cDNA ends (RACE) reactions.
Total RNA isolation, RT-PCR, and DNA sequencing
Total zebrafish RNA was obtained using the microRNA isolation kit (Promega, Madison, WI) following the manufacturer’s protocols. Approximately 1 μg of deoxyribonuclease-treated RNA was reverse transcribed using Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA), and PCR amplification was performed using gene-specific primers for zPTH1 (forward: 5′-AATGAAGTGCAGCTGATGCAT-3′ and 5′-CAGGACTGGCTGCAGATGAA-3′; reverse: 5′-CGATGGGTTCATGA-GCTTTTC-3′), and zPTH2 (forward: 5′-GAAGTTCAGCTGATGC-ACAATGT-3′ and 5′-AGGACTGGCTTCAGCTGAAG-3′, reverse: 5′-GTTGCACCTTCCCCCTTTCT-3′). DNA amplicons were electro-phoresed, purified, ligated to pGEM-Teasy as previously described (15), and used to transform Escherichia coli TOP10 cells (Invitrogen). Bacterial colonies were screened by PCR using the same gene-specific primers. Plasmids containing cDNAs encoding zPTH were purified using Concert Midiprep (Invitrogen), and manually sequenced according to manufacturer’s protocols (ABI, PerkinElmer Corp., Shel-ton, CT). DNA sequence analysis and comparisons were performed using basic local alignment and search tool (BLAST), translation, and alignment algorithms (http://www.ncbi.nlm.nih.gov/BLAST/, http://searchlauncher.bcm.tmc.edu/seq-util/seq-util.html).
5′ and 3′ RACE reactions
To identify the 5′ end of the cDNAs encoding zPTH1 and zPTH2, total RNA isolated from zebrafish head and body was reverse transcribed by using Superscript II reverse transcriptase and specific reverse zPTH primers (zPTH1, 5′-TGAGTTCATCCCATGCGCTGT-3′; zPTH2, 5′-CTGACGTTAGTTCATTAGCTTTTC-3′). Poly (deoxy-GTP) 5′-tailing of the first-strand cDNA was performed according to manufacturer’s specifications (Promega). The initial PCR for 5′-tailed cDNA used the provided AAP primer (Invitrogen) and specific reverse PTH primers (zPTH1, 5′-CATCCTAATCTGTTTTCACGA-3′; zPTH2, 5′-TGACAATATCAGTAAAA-GCTCCA-3′). The PCR profile included an initial denaturation at 95 C for 3 min, followed by 35 cycles with denaturation at 94 C for 1 min, annealing at 54 C for 1 min, polymerization at 72 C for 2 min, and a final extension at 72 C for 10 min. Nested PCR was performed on a 1:100 dilution of the first amplified product using cycling conditions identical with those described above, except annealing at 60 C, using forward Abridged Universal Amplification Primer (AUAP) (Invitrogen) and reverse zPTH primers (zPTH1, 5′-CGATGGGTTCATGAGCTTTTC-3′; zPTH2, 5′-GTTGCACCTTCCCCCTTTCT-3′). 5′-RACE cDNAs encoding zPTH1 and zPTH2 were purified by agarose gel electrophoresis, isolated, cloned into pGEM-Teasy, screened, and sequenced as described above.
To identify the 3′ end of the cDNAs encoding zPTH1 and zPTH2, total zebrafish RNA was reverse transcribed using Superscript II reverse transcriptase and an oligo-dT adapter primer (5′-GGCCACGCGTCGACTAGTACTTTTTTTTTTTTTTTTT-3′) containing nucleotide sequences that contain several recognition sites for infrequently cutting endonucleases (Invitrogen). The first strand was then directly amplified by PCR using forward zPTH primers (zPTH1, 5′-AATGAAGTGCAGCTGATGCAT-3′; zPTH2, 5′-GAAGTTCAGCTGATGCACAAT-GT-3′) and an adapter primer (AUAP) by following the same PCR profile described above except for an annealing temperature of 59 C for 40 cycles. This reaction was followed by a nested amplification using forward zPTH primers (zPTH1, 5′-CAGGACTGGCTGCAGATGAA-3′; zPTH2, 5′-GTTGCACCTTCCCCCTTTCT-3′) and an adapter AUAP primer at an annealing temperature of 60 C for 40 cycles. 3′-RACE cDNAs encoding zPTH1 and zPTH2 were isolated, subcloned, screened, and sequenced as described above.
Determination of putative intron/exon boundaries
The intron/exon structure of the zPTH1 and zPTH2 genes was determined using Grail Experimental Gene Discovery Suite (Baylor College of Medicine Human Genome Sequencing Center, http://searchlauncher.bcm.tmc.edu/seq-search/gene-search.html) to search for putative introns and the Splice Site Prediction by Neural Networks (Berkeley Drosophila Genome Project, http://www.fruitfly.org/seq_tools/splice.html) to predict the locations of RNA splice sites.
Peptide synthesis
The peptides zPTH1(1–34)amide [zPTH1(1–34)], zPTH2(1–34)amide [zPTH2(1–34)], human [Tyr34]PTH(1–34)amide [hPTH(1–34)], and human [Tyr36]PTHrP(1–36)amide [hPTHrP(1–36)] were synthesized by the Protein and Peptide Core Facility at Massachusetts General Hospital (Boston, MA) by the solid-phase method on a PerkinElmer Model 430A and 431A synthesizers. Peptides were purified by reverse-phase chromatography and the composition was confirmed by amino acid analysis and mass spectroscopy.
cAMP accumulation assays
COS-7 cells were transiently transfected with cDNA encoding hPTH1R, zPTH1R, or zPTH3R using the Effectene reagent (QIAGEN, Valencia, CA). After 72 h, cells were treated with 0–10−6m of peptide and incubated in 3-isobutyl-1-methylxanthine-containing buffer for 60 min at room temperature. After rinsing, intracellular cAMP accumulation was measured by RIA as previously described (19).
Results
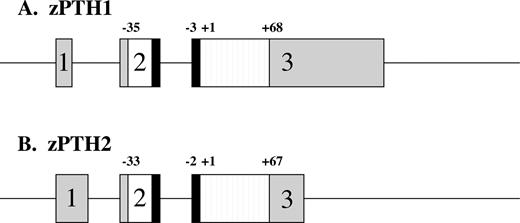
The cDNA encoding hPTH was used as a probe to search the zebrafish genome web site (http://134.174.23.160/HumanblastZebrafish/) for homologous sequences. Two putative zebrafish genomic DNA regions (zPTH1, zfish35939–334d11.p1c; and zPTH2, zfishJ-a492f09.p1c) were identified which showed sequence homology to the hPTH transcript. To determine whether these putative genes are expressed, gene-specific primers for zPTH1 and zPTH2 were designed for amplification by RT-PCR, and subsequently 5′ and 3′ RACE reactions were performed to obtain the full-length cDNAs (see Materials and Methods). These PCR-amplified cDNAs were then cloned into pGEM-Teasy for nucleotide sequence analysis. Each sequence contained a polyadenylation signal and a large open reading frame with considerable amino acid homology with hPTH. Genomic regions with sequence identical with the cDNA sequences were determined by BLAST, and together with analysis to predict intron-exon boundaries, the putative intron-exon structure for each zPTH gene was determined (see Fig. 1).
Schematic representation of zPTH1 (A) and zPTH2 (B) genes. Boxes depict exons, with gray regions indicating untranslated regions. White and black regions represent pre- and prosequences, respectively. Striped regions depict mature peptides. The boundaries of the pre-, pro-, and mature sequences are indicated; +1 indicates the start of the tested peptide.
Similar to the mammalian PTHs, each zPTH amino acid sequence contained a hydrophobic presequence and a putative prosequence (Fig. 2), suggesting that the zPTHs are secreted peptides. Each sequence also contained the dibasic putative cleavage site that is conserved in mammalian and chicken PTH genes. After cleavage, the mature zPTH1 was predicted to contain 68 amino acids, and zPTH2 was predicted to be 67 amino acids in length. The first 34 amino acids of the mature peptides showed the greatest similarity to hPTH, with several key resi-dues known to be critical for hPTH(1–34) binding (Arg20, Trp23, Leu24, Leu28) and activity (residues 1–9) (2) showing a high degree of conservation.
Alignment of the amino acid sequences of zPTH1, zPTH2, and hPTH. The presequences are outlined with dotted lines; prosequences are outlined with solid lines. The first 34 amino acids of the mature peptide are in bold type. *, Identical residues; :, conservative substitutions;., partially conservative substitutions.
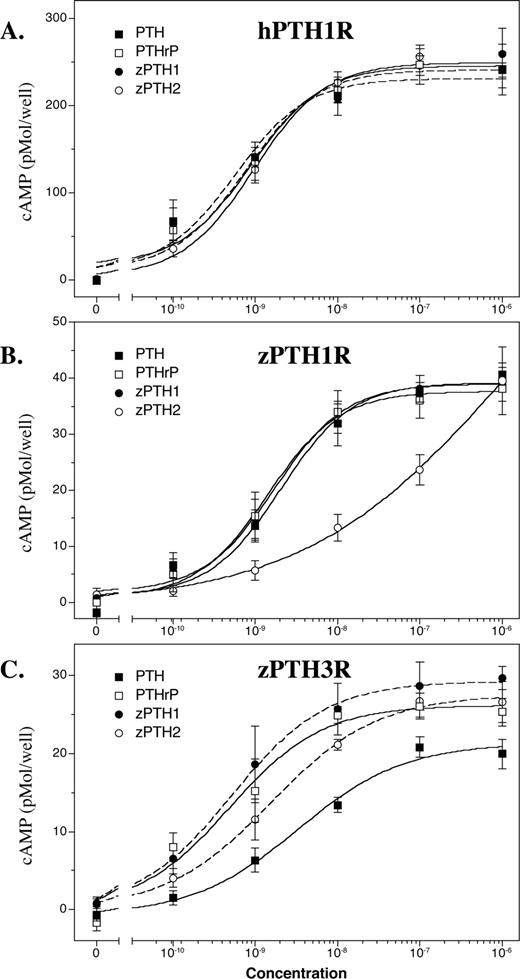
We next tested whether the first 34 amino acids of each zPTH could fully activate hPTH1R and the two zPTH receptors, zPTH1R, and zPTH3R. Both zPTH1(1–34) and zPTH2(1–34) activated hPTH1R with EC50 values comparable to those of hPTH(1–34) and hPTHrP(1–36) (see Table 1). When tested with zPTH1R, zPTH1(1–34) again exhibited an EC50 comparable to that of hPTH(1–34) and of hPTHrP(1–36); however, the EC50 of zPTH2(1–34) was approximately 30-fold lower (Fig. 3 and Table 1). When tested with zPTH3R, both zPTH1(1–34) and hPTHrP(1–36) showed similar EC50s, whereas the EC50 of zPTH2 was only slightly (∼3-fold) reduced. The EC50 of hPTH(1–34) was approximately 6-fold lower than that of PTHrP(1–36), as previously described (15). The combined results for the zPTH receptors indicate that zPTH1 activates zPTH1R and zPTH3R with similar potencies, whereas zPTH2 is more selective for zPTH3R.
cAMP accumulation induced by zPTH1(1–34) and zPTH2(1–34) treatment. COS-7 cells transiently expressing hPTH1R (A), zPTH1R (B), or zPTH3R (C) were evaluated for agonist-stimulated cAMP production (▪, hPTH(1–34); □, hPTHrP(1–36); •, zPTH1(1–34); ○, zPTH2(1–34). Data are expressed as cAMP accumulation in picomoles/well. The results shown are the average of at least three independent transfections.
Potencies and efficacies of zPTH1 and zPTH2 for stimulation of cAMP accumulation through human and zebrafish PTH receptors
| Peptide . | hPTH1R . | . | zPTH1R . | . | zPTH3R . | . | |||
|---|---|---|---|---|---|---|---|---|---|
| . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | |||
| hPTH(1–34) | 0.57 ± 0.25 | 231 ± 18 | 2.0 ± 0.7 | 39.2 ± 2.2 | 3.3 ± 1.2 | 21.3 ± 1.7 | |||
| hPTHrP(1–36) | 0.77 ± 0.22 | 241 ± 10 | 1.4 ± 0.2 | 37.7 ± 1.0 | 0.49 ± 0.18 | 26.2 ± 1.8 | |||
| zPTH1(1–34) | 0.88 ± 0.33 | 245 ± 14 | 1.7 ± 0.3 | 39.0 ± 0.9 | 0.52 ± 0.06 | 29.2 ± 0.5 | |||
| zPTH2(1–34) | 0.96 ± 0.16 | 249 ± 6 | 67 ± 28 | 40.9 ± 4.5 | 1.6 ± 0.20 | 27.5 ± 0.6 | |||
| Peptide . | hPTH1R . | . | zPTH1R . | . | zPTH3R . | . | |||
|---|---|---|---|---|---|---|---|---|---|
| . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | |||
| hPTH(1–34) | 0.57 ± 0.25 | 231 ± 18 | 2.0 ± 0.7 | 39.2 ± 2.2 | 3.3 ± 1.2 | 21.3 ± 1.7 | |||
| hPTHrP(1–36) | 0.77 ± 0.22 | 241 ± 10 | 1.4 ± 0.2 | 37.7 ± 1.0 | 0.49 ± 0.18 | 26.2 ± 1.8 | |||
| zPTH1(1–34) | 0.88 ± 0.33 | 245 ± 14 | 1.7 ± 0.3 | 39.0 ± 0.9 | 0.52 ± 0.06 | 29.2 ± 0.5 | |||
| zPTH2(1–34) | 0.96 ± 0.16 | 249 ± 6 | 67 ± 28 | 40.9 ± 4.5 | 1.6 ± 0.20 | 27.5 ± 0.6 | |||
Emax, Maximal cAMP accumulation observed.
Potencies and efficacies of zPTH1 and zPTH2 for stimulation of cAMP accumulation through human and zebrafish PTH receptors
| Peptide . | hPTH1R . | . | zPTH1R . | . | zPTH3R . | . | |||
|---|---|---|---|---|---|---|---|---|---|
| . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | |||
| hPTH(1–34) | 0.57 ± 0.25 | 231 ± 18 | 2.0 ± 0.7 | 39.2 ± 2.2 | 3.3 ± 1.2 | 21.3 ± 1.7 | |||
| hPTHrP(1–36) | 0.77 ± 0.22 | 241 ± 10 | 1.4 ± 0.2 | 37.7 ± 1.0 | 0.49 ± 0.18 | 26.2 ± 1.8 | |||
| zPTH1(1–34) | 0.88 ± 0.33 | 245 ± 14 | 1.7 ± 0.3 | 39.0 ± 0.9 | 0.52 ± 0.06 | 29.2 ± 0.5 | |||
| zPTH2(1–34) | 0.96 ± 0.16 | 249 ± 6 | 67 ± 28 | 40.9 ± 4.5 | 1.6 ± 0.20 | 27.5 ± 0.6 | |||
| Peptide . | hPTH1R . | . | zPTH1R . | . | zPTH3R . | . | |||
|---|---|---|---|---|---|---|---|---|---|
| . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | EC50 (nm) . | Emax (pmol/well) . | |||
| hPTH(1–34) | 0.57 ± 0.25 | 231 ± 18 | 2.0 ± 0.7 | 39.2 ± 2.2 | 3.3 ± 1.2 | 21.3 ± 1.7 | |||
| hPTHrP(1–36) | 0.77 ± 0.22 | 241 ± 10 | 1.4 ± 0.2 | 37.7 ± 1.0 | 0.49 ± 0.18 | 26.2 ± 1.8 | |||
| zPTH1(1–34) | 0.88 ± 0.33 | 245 ± 14 | 1.7 ± 0.3 | 39.0 ± 0.9 | 0.52 ± 0.06 | 29.2 ± 0.5 | |||
| zPTH2(1–34) | 0.96 ± 0.16 | 249 ± 6 | 67 ± 28 | 40.9 ± 4.5 | 1.6 ± 0.20 | 27.5 ± 0.6 | |||
Emax, Maximal cAMP accumulation observed.
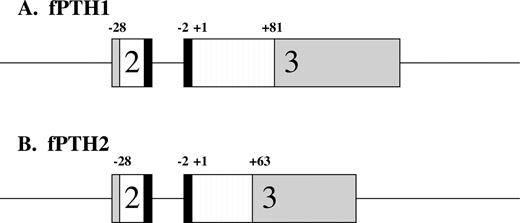
Japanese pufferfish, Takifugu rubripes, were recently reported to have a PTH-like peptide, discovered by PCR-amplification of genomic DNA (18). We sought to determine whether this species also has a second PTH-like peptide, similar to the zebrafish. Using zPTH1(1–34) or zPTH2(1–34) as a probe, we identified two distinct genomic DNA regions of high homology (scaffolds 847 and 81, respectively). The DNA sequence identified with zPTH1(1–34) was identical with that in the previous report (18). We have defined putative intron-exon boundaries for two coding exons based on homology to zebrafish and prediction of splice sites (Fig. 4). The corresponding amino acid sequence revealed a putative signal sequence and dibasic cleavage site similar to zPTH1 and zPTH2 (Fig. 5). The second genomic DNA region, identified with zPTH2(1–34) as a probe, revealed a potential pufferfish homolog for zPTH2. Putative intron-exon boundaries were identified for this DNA region (Fig. 4), and the corresponding amino acid sequence again revealed a putative signal sequence and dibasic cleavage site (Fig. 5). Pufferfish and zebrafish thus appear to have two similar PTH-like ligands.
Schematic representation of fPTH1 (A) and fPTH2 (B) genes. Boxes depict exons, with gray regions indicating untranslated regions. White and black regions represent pre- and prosequences, respectively. Striped regions depict mature peptides. The boundaries of the pre-, pro-, and mature sequences are indicated; +1 indicates the start of the mature peptide.
Alignment of the amino acid sequences of zPTHs, fugu PTHs, and several mammalian PTHs. The presequences are outlined with dotted lines; prosequences are outlined with solid lines. The first 34 amino acids of the mature peptide are in bold type. *, Identical residues; :, conservative substitutions;., partially conservative substitutions. Gray boxes denote residues (2, 11, and 12) with PTH2-specific substitutions.
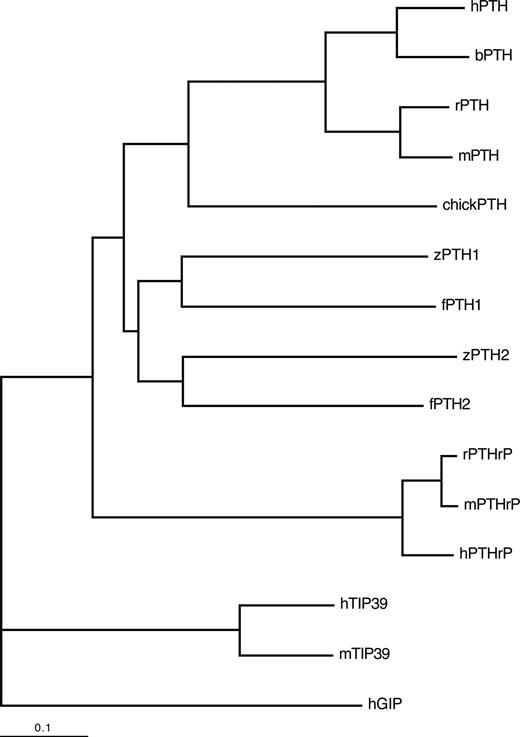
We next performed a phylogenetic analysis comparing the zebrafish and putative pufferfish PTHs to known PTH, PTHrP, and tuberoinfundibular peptide of 39 residues (TIP39) from other species (Fig. 6). The fish PTHs were more closely related to other PTH peptides than they were to PTHrP or TIP39 peptides. The fish PTH molecules were more related to each other than they were to any of the other PTH peptides analyzed. Comparisons between the two species of fishes showed that the two PTH1 molecules were more closely related to each other than they were to the PTH2 molecule from the same species. The phylogenetic analysis is thus consistent with the hypothesis that teleost PTH2 evolved after the evolutionary divergence between fishes and mammals, but before the evolutionary divergence between the Cypriniformes (zebrafish) and Perciformes (pufferfish); two distant orders of teleosts.
Phylogenetic analysis of PTH, PTHrP, and TIP39 from various species. Phylogentic analysis was performed on the full-length sequences, including pre- and prosequences, of the indicated PTH, PTHrP, and TIP39 molecules. The phylogenetic tree is rooted with human gastrin inhibitory peptide (hGIP). The length of the horizontal lines indicates the degree of evolutionary divergence, according to the indicated scale.
Discussion
In this report, we have described two distinct PTH ligands in the zebrafish. The genomic sequence with putative intron/exon boundaries, the cDNA sequence, and the amino acid sequence were described. Both peptides have considerable homology to hPTH and were highly potent at the hPTH1R. However, when tested with the zPTH1R, the potency of zPTH1(1–34) was significantly greater than that of zPTH2(1–34) (Fig. 3B). Zebrafish have a third receptor, the zPTH3R, and the potencies of the zPTH molecules at this receptor were more similar (Fig. 3C). The results thus indicate that zPTH2(1–34) is more selective for zPTH3R, whereas zPTH1(1–34) nonselectively activates zPTH1R and zPTH3R. It is therefore likely that zPTH1 and zPTH2 have distinct physiologic roles in the zebrafish. The existence of a similar putative PTH2-like molecule in the fugu genomic database suggests these findings may generalize to other fishes.
The phylogenetic analysis of PTH suggests that the presence of two PTH ligands is the result of a gene duplication which occurred after the evolutionary divergence between fishes and mammals, but before the evolutionary divergence between zebrafish and pufferfish. Analysis of the adjacent intergenetic DNA sequence and comparison of nearby genes for the zPTH1 and zPTH2 genes suggests that the duplicated region was limited to the PTH gene itself. Analysis of additional fishes and amphibians may yield further data regarding the timing of this event, and provide additional information to order the evolutionary tree. Of note, the PTH-like sequence comprising 10 amino acids which was PCR-amplified from rainbow trout DNA has nine residues in common with zPTH1, suggesting that this species also has at least one PTH-like peptide (20). It is interesting that the PTH endocrine system appears to have evolved to greater complexity in fishes than it has in mammals. This observation provides direct evidence that basal vertebrates have continued to evolve after the divergence between the Actinopterygii (bony ray-finned fishes including the teleosts) and the Sarcopterygii (lobe-finned fishes including the Dipnoi and the tetrapods). Thus, the assumption that derived vertebrates (i.e. mammals) have all of the gene constituents of basal vertebrates (fishes) is not universal. In support of this concept, a comparison of the human, fly, and worm genomes showed an equal likelihood for the presence in one genome of increased complexity from gene duplications that were not present in the other two genomes (17).
The increased complexity of the PTH endocrine system in fishes may relate to a more diverse physiological role involved with osmoregulation. Because some fishes are capable of enduring both salt water and fresh water, mechanisms must exist that regulate both serum Ca2+ and osmolality in varied environmental conditions (21). Previous teleost studies indicate that prolactin, GH, and 1,25-dihydroxyvitamin D3, in addition to their roles in calcium regulation, are also necessary for osmoregulation in fresh water and in salt water (22–26). Thus, it is possible that the PTH-like hormones might also have multiple physiological roles that are not restricted to Ca2+ regulation. Determination of these roles in fishes will require further study.
In addition to these pharmacological and phylogenetic insights, detailed analysis of PTH sequences of fishes may provide further insights into structure/function relationships of hPTH. Importantly, we determined that each of the zPTH molecules activate the hPTH1R with high potency and are fully efficacious (Fig 2). Comparison of the zPTH1 and zPTH2 with hPTH showed interesting differences and similarities (Fig 5). The amino-terminal portion of hPTH(1–34) is most critical for receptor activation (6), and most residues in this region of zPTH1 and zPTH2 are either identical or have conservative substitutions. The substitution of Val2 to Ile in zPTH2 and fPTH2 is unexpected; Val2 is a highly conserved residue in PTH, and the tested substitutions at this location disrupt agonist activity (27). This substitution is conservative, but at such a critical location the substitution may be contributing to the ligand selectivity of zPTH1. Residue 2 of PTH has been shown by cross-linking and mutational studies to interact with the TM5/TM6 region of the hPTH1R (28, 29), and differences exist in the amino acid sequence between zPTH1R and zPTH3R in this region (15). Two other substitutions unique to the fishes PTH2 molecules are Leu->Val at position 11 and Gly->Arg at position 12. Both residues in hPTH(1–34) are tolerant to substitution with alanine when tested with an hPTH1R mutant lacking the amino-terminal extracellular domain (30), but the zPTH1R may be more sensitive to changes at these positions. These residues have been found to be critical for antagonist binding (31) and inverse agonist activity (32) in hPTH. Gly12 has been postulated as the boundary residue for a flexible region in the PTH molecule (33), and the Arg substitution seen in the fishes peptides would be predicted to have a helix-stabilizing effect. Helix stabilization may in turn enhance ligand potency, similar to the effect seen with the Glu19->Arg substitution (34, 35). In the more carboxyl-terminal portion of PTH(1–34), substitutions at residues 20, 23, 24, 28, and 31 have been shown to significantly reduce ligand binding affinity (36). The PTHs in fishes have no substitutions at these residues, with the exception of residue 31 that shows a conservative substitution of Val->Ile. The remaining residues form an amphipathic helix, similar to hPTH and hPTHrP. Although we did not directly assess binding affinity of the zPTHs in these studies, no substitutions are present that would be expected to significantly affect binding affinity. Examination of the carboxyl-terminal region of PTH reveals very little homology between fish PTHs and mammalian PTHs (see Fig. 5). Although there is evidence that mammalian PTHs signal through a distinct carboxyl- terminal receptor that has not yet been isolated (37), the poor sequence homology in this region might suggest that fishes lack such receptors.
Overall, we have identified two distinct PTH molecules in the zebrafish. These molecules are each capable of activating the hPTH/PTHrP receptor, although they show different patterns of receptor selectivity among the zPTH receptors. Similar PTH molecules are present in the fugufish genomic database, at least one of which has been recently shown to activate the rat PTH1R (18). Phylogenetic analysis suggests that the additional PTH gene evolved through gene duplication. Comparisons between the amino acid sequences of zPTHs and hPTH revealed several interesting differences in the activation domain of the ligand, and further study may serve to enhance our understanding of PTH structure-function relationships at the hPTH1R.
Abbreviations:
- AUAP,
Abridged Universal Amplification Primer;
- BLAST,
basic local alignment and search tool;
- hPTH,
human PTH;
- PTH1R,
hPTH/PTHrP receptor;
- PTH2R or PTH3R,
PTH-2 or -3 receptor;
- zPTH,
zebrafish PTH;
- RACE,
rapid amplification of cDNA ends;
- TIP,
tuberoinfundibular peptide of 39 residues.
Acknowledgments
We would like to thank Dr. Ashok Khatri for synthesis of peptides used in these studies.
This work was supported by NIH Grants DK11794 and DK60513.
The sequences have been submitted to the GenBank database under accession nos. AY275670 (zebrafish PTH2), AY275669 (zebrafish PTH1), and AY302221 (Fugu PTH2).
References