Abstract
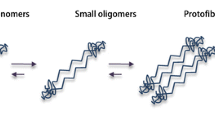
This review highlights the modern perception of anomalous folding of the prion protein and the role of chaperones therein. Special attention is paid to prion proteins from mammalian species, which are prone to amyloid-like prion diseases due to a unique aggregation pathway. Despite being a significantly popular current subject of investigations, the etiology, structure, and function of both normal and anomalous prion proteins still hold many mysteries. The most interesting of those are connected to the interaction with chaperone system, which is responsible for stabilizing protein structure and disrupting aggregates. In the case of prion proteins the following question is of the most importance — can chaperones influence different stages of the formation of pathological aggregates (these vary from intermediate oligomers to mature amyloid-like fibrils) and the whole transition from native prion protein to its amyloid-like fibril-enriched form? The existing inconsistencies and ambiguities in the observations made so far can be attributed to the fact that most of the investigations did not take into account the type and functional state of the chaperones. This review discusses in detail our previous works that have demonstrated fundamental differences between eukaryotic and prokaryotic chaperones in the action exerted on the amyloid-like transformation of the prion protein along with the dependence of the observed effects on the functional state of the chaperone.
Similar content being viewed by others
References
Prusiner, S. B. (1982) Novel proteinaceous infectious particles cause scrapie, Science, 216, 136–144.
Prusiner, S. B., Groth, D. F., Bolton, D. C., Kent, S. B., and Hood, L. E. (1984) Purification and structural studies of a major scrapie prion protein, Cell, 38, 127–134.
Chesebro, B., Race, R., Wehrly, K., Nishio, J., Bloom, M., Lechner, D., Bergstrom, S., Robbins, K., Mayer, L., Keith, J. M., Garon, C., and Haase, A. (1985) Identification of scrapie prion protein-specific mRNA in scrapie-infected and uninfected brain, Nature, 315, 331–333.
Oesch, B., Westaway, D., Walchli, M., McKinley, M. P., Kent, S. B., Aebersold, R., Barry, R. A., Tempst, P., Teplow, D. B., Hood, L. E., Prusiner, S. B., and Weissmann, C. (1985) A cellular gene encodes scrapie PrP 27–30 protein, Cell, 40, 735–746.
Basler, K., Oesch, B., Scott, M., Westaway, D., Walchli, M., Groth, D. F., McKinley, M. P., Prusiner, S. B., and Weissmann, C. (1986) Scrapie and cellular PrP isoforms are encoded by the same chromosomal gene, Cell, 46, 417–428.
Bueler, H., Aguzzi, A., Sailer, A., Greiner, R. A., Autenried, P., Aguet, M., and Weissmann, C. (1993) Mice devoid of PrP are resistant to scrapie, Cell, 73, 1339–1347.
Kushnirov, V. V., and Ter-Avanesyan, M. D. (1998) Structure and replication of yeast prions, Cell, 94, 13–16.
Paushkin, S. V., Kushnirov, V. V., Smirnov, V. N., and Ter-Avanesyan, M. D. (1997) In vitro propagation of the prionlike state of yeast Sup35 protein, Science, 277, 381–383.
Kushnirov, V. V., Ter-Avanesyan, M. D., Telckov, M. V., Surguchov, A. P., Smirnov, V. N., and Inge-Vechtomov, S. G. (1988) Nucleotide sequence of the SUP2 (SUP35) gene of Saccharomyces cerevisiae, Gene, 66, 45–54.
Harris, D. A. (2001) in Advances in Protein Chemistry (Byron, C., ed.) Vol. 57, Academic Press, pp. 203–228.
Huang, Z., Prusiner, S. B., and Cohen, F. E. (1995) Scrapie prions: a three-dimensional model of an infectious fragment, Fold. Des., 1, 13–19.
Lysek, D. A., Schorn, C., Nivon, L. G., Esteve-Moya, V., Christen, B., Calzolai, L., von Schroetter, C., Fiorito, F., Herrmann, T., Guntert, P., and Wuthrich, K. (2005) Prion protein NMR structures of cats, dogs, pigs, and sheep, Proc. Natl. Acad. Sci. USA, 102, 640–645.
Knaus, K. J., Morillas, M., Swietnicki, W., Malone, M., Surewicz, W. K., and Yee, V. C. (2001) Crystal structure of the human prion protein reveals a mechanism for oligomerization, Nat. Struct. Biol., 8, 770–774.
Sawaya, M. R., Sambashivan, S., Nelson, R., Ivanova, M. I., Sievers, S. A., Apostol, M. I., Thompson, M. J., Balbirnie, M., Wiltzius, J. J., McFarlane, H. T., Madsen, A. O., Riekel, C., and Eisenberg, D. (2007) Atomic structures of amyloid cross-β spines reveal varied steric zippers, Nature, 47, 453–457.
Haire, L. F., Whyte, S. M., Vasisht, N., Gill, A. C., Verma, C., Dodson, E. J., Dodson, G. G., and Bayley, P. M. (2004) The crystal structure of the globular domain of sheep prion protein, J. Mol. Biol., 336, 1175–1183.
Riek, R., Hornemann, S., Wider, G., Glockshuber, R., and Wuthrich, K. (1997) NMR characterization of the full-length recombinant murine prion protein, mPrP(23–231), FEBS Lett., 413, 282–288.
Brockes, J. P. (1999) Topics in prion cell biology, Curr. Opin. Neurobiol., 9, 571–577.
Aguzzi, A., Sigurdson, C., and Heikenwaelder, M. (2008) Molecular mechanisms of prion pathogenesis, Annu. Rev. Pathol., 3, 11–40.
Bujdoso, R., Burke, D. F., and Thackray, A. M. (2005) Structural differences between allelic variants of the ovine prion protein revealed by molecular dynamics simulations, Proteins, 61, 840–849.
Gossert, A. D., Bonjour, S., Lysek, D. A., Fiorito, F., and Wuthrich, K. (2005) Prion protein NMR structures of elk and of mouse/elk hybrids, Proc. Natl. Acad. Sci. USA, 102, 646–650.
Huang, Z., Gabriel, J. M., Baldwin, M. A., Fletterick, R. J., Prusiner, S. B., and Cohen, F. E. (1994) Proposed three-dimensional structure for the cellular prion protein, Proc. Natl. Acad. Sci. USA, 91, 7139–7143.
Prusiner, S. B. (1994) Neurodegeneration in humans caused by prions, West. J. Med., 161, 264–272.
Prusiner, S. B., Scott, M. R., DeArmond, S. J., and Cohen, F. E. (1998) Prion protein biology, Cell, 93, 337–348.
Kaneko, K., Ball, H. L., Wille, H., Zhang, H., Groth, D., Torchia, M., Tremblay, P., Safar, J., Prusiner, S. B., DeArmond, S. J., Baldwin, M. A., and Cohen, F. E. (2000) A synthetic peptide initiates Gerstmann-Straussler-Scheinker (GSS) disease in transgenic mice, J. Mol. Biol., 295, 997–1007.
Kitamoto, T., Iizuka, R., and Tateishi, J. (1993) An amber mutation of prion protein in Gerstmann-Straussler syndrome with mutant PrP plaques, Biochem. Biophys. Res. Commun., 192, 525–531.
Supattapone, S., Bosque, P., Muramoto, T., Wille, H., Aagaard, C., Peretz, D., Nguyen, H. O., Heinrich, C., Torchia, M., Safar, J., Cohen, F. E., DeArmond, S. J., Prusiner, S. B., and Scott, M. (1999) Prion protein of 106 residues creates an artificial transmission barrier for prion replication in transgenic mice, Cell, 96, 869–878.
Ghetti, B., Piccardo, P., Spillantini, M. G., Ichimiya, Y., Porro, M., Perini, F., Kitamoto, T., Tateishi, J., Seiler, C., Frangione, B., Bugiani, O., Giaccone, G., Prelli, F., Goedert, M., Dlouhy, S. R., and Tagliavini, F. (1996) Vascular variant of prion protein cerebral amyloidosis with τ-positive neurofibrillary tangles: the phenotype of the stop codon 145 mutation in PRNP, Proc. Natl. Acad. Sci. USA, 93, 744–748.
Govaerts, C., Wille, H., Prusiner, S. B., and Cohen, F. E. (2004) Evidence for assembly of prions with left-handed β-helices into trimers, Proc. Natl. Acad. Sci. USA, 101, 8342–8347.
Der-Sarkissian, A., Jao, C. C., Chen, J., and Langen, R. (2003) Structural organization of α-synuclein fibrils studied by site-directed spin labeling, J. Biol. Chem., 278, 37530–37535.
Petkova, A. T., Ishii, Y., Balbach, J. J., Antzutkin, O. N., Leapman, R. D., Delaglio, F., and Tycko, R. (2002) A structural model for Alzheimer’s β-amyloid fibrils based on experimental constraints from solid state NMR, Proc. Natl. Acad. Sci. USA, 99, 16742–16747.
Sunde, M., Serpell, L. C., Bartlam, M., Fraser, P. E., Pepys, M. B., and Blake, C. C. (1997) Common core structure of amyloid fibrils by synchrotron X-ray diffraction, J. Mol. Biol., 273, 729–739.
Luhrs, T., Ritter, C., Adrian, M., Riek-Loher, D., Bohrmann, B., Dobeli, H., Schubert, D., and Riek, R. (2005) 3D structure of Alzheimer’s amyloid-β (1–42) fibrils, Proc. Natl. Acad. Sci. USA, 102, 17342–17347.
Wasmer, C., Lange, A., Van Melckebeke, H., Siemer, A. B., Riek, R., and Meier, B. H. (2008) Amyloid fibrils of the HET-s (218–289) prion form a β-solenoid with a triangular hydrophobic core, Science, 319, 1523–1526.
Gajdusek, D. C. (1988) Transmissible and non-transmissible amyloidoses: autocatalytic posttranslational conversion of host precursor proteins to β-pleated sheet configurations, J. Neuroimmunol., 20, 95–110.
Come, J. H., Fraser, P. E., and Lansbury, P. T., Jr. (1993) A kinetic model for amyloid formation in the prion diseases: importance of seeding, Proc. Natl. Acad. Sci. USA, 90, 5959–5963.
Kelly, J. W. (2000) Mechanisms of amyloidogenesis, Nat. Struct. Biol., 7, 824–826.
Eghiaian, F., Daubenfeld, T., Quenet, Y., van Audenhaege, M., Bouin, A. P., van der Rest, G., Grosclaude, J., and Rezaei, H. (2007) Diversity in prion protein oligomerization pathways results from domain expansion as revealed by hydrogen/deuterium exchange and disulfide linkage, Proc. Natl. Acad. Sci. USA, 104, 7414–7419.
Rezaei, H., Eghiaian, F., Perez, J., Doublet, B., Choiset, Y., Haertle, T., and Grosclaude, J. (2005) Sequential generation of two structurally distinct ovine prion protein soluble oligomers displaying different biochemical reactivities, J. Mol. Biol., 347, 665–679.
Singh, J., Sabareesan, A. T., Mathew, M. K., and Udgaonkar, J. B. (2012) Development of the structural core and of conformational heterogeneity during the conversion of oligomers of the mouse prion protein to worm-like amyloid fibrils, J. Mol. Biol., 423, 217–231.
Aguzzi, A., and Polymenidou, M. (2004) Mammalian prion biology: one century of evolving concepts, Cell, 116, 313–327.
Vassallo, N., and Herms, J. (2003) Cellular prion protein function in copper homeostasis and redox signaling at the synapse, J. Neurochem., 86, 538–544.
Tsiroulnikov, K., Rezaei, H., Dalgalarrondo, M., Chobert, J. M., Grosclaude, J., and Haertle, T. (2006) Cu(II) induces small-size aggregates with amyloid characteristics in two alleles of recombinant ovine prion proteins, Biochim. Biophys. Acta, 1764, 1218–1226.
Wong, E., Thackray, A. M., and Bujdoso, R. (2004) Copper induces increased β-sheet content in the scrapie-susceptible ovine prion protein PrPVRQ compared with the resistant allelic variant PrPARR, Biochem. J., 380, 273–282.
Roucou, X., Gains, M., and LeBlanc, A. C. (2004) Neuroprotective functions of prion protein, J. Neurosci. Res., 75, 153–161.
Kawahara, M., Kuroda, Y., Arispe, N., and Rojas, E. (2000) Alzheimer’s β-amyloid, human islet amylin, and prion protein fragment evoke intracellular free calcium elevations by a common mechanism in a hypothalamic GnRH neuronal cell line, J. Biol. Chem., 275, 14077–14083.
Moore, R. C., Lee, I. Y., Silverman, G. L., Harrison, P. M., Strome, R., Heinrich, C., Karunaratne, A., Pasternak, S. H., Chishti, M. A., Liang, Y., Mastrangelo, P., Wang, K., Smit, A. F., Katamine, S., Carlson, G. A., Cohen, F. E., Prusiner, S. B., Melton, D. W., Tremblay, P., Hood, L. E., and Westaway, D. (1999) Ataxia in prion protein (PrP)-deficient mice is associated with upregulation of the novel PrP-like protein doppel, J. Mol. Biol., 292, 797–817.
Collinge, J., Whittington, M. A., Sidle, K. C., Smith, C. J., Palmer, M. S., Clarke, A. R., and Jefferys, J. G. (1994) Prion protein is necessary for normal synaptic function, Nature, 370, 295–297.
Jeffrey, M., Goodsir, C., McGovern, G., Barmada, S. J., Medrano, A. Z., and Harris, D. A. (2009) Prion protein with an insertional mutation accumulates on axonal and dendritic plasmalemma and is associated with distinctive ultrastructural changes, Am. J. Pathol., 175, 1208–1217.
Stys, P. K., You, H., and Zamponi, G. W. (2012) Copper-dependent regulation of NMDA receptors by cellular prion protein: implications for neurodegenerative disorders, J. Physiol., 590, 1357–1368.
Maglio, L. E., Martins, V. R., Izquierdo, I., and Ramirez, O. A. (2006) Role of cellular prion protein on LTP expression in aged mice, Brain Res., 1097, 11–18.
Khosravani, H., Zhang, Y., Tsutsui, S., Hameed, S., Altier, C., Hamid, J., Chen, L., Villemaire, M., Ali, Z., Jirik, F. R., and Zamponi, G. W. (2008) Prion protein attenuates excitotoxicity by inhibiting NMDA receptors, J. Gen. Physiol., 131, i5.
Gimbel, D. A., Nygaard, H. B., Coffey, E. E., Gunther, E. C., Lauren, J., Gimbel, Z. A., and Strittmatter, S. M. (2010) Memory impairment in transgenic Alzheimer mice requires cellular prion protein, J. Neurosci., 30, 6367–6374.
Gunther, E. C., and Strittmatter, S. M. (2010) β-Amyloid oligomers and cellular prion protein in Alzheimer’s disease, J. Mol. Med. (Berl.), 88, 331–338.
Lauren, J., Gimbel, D. A., Nygaard, H. B., Gilbert, J. W., and Strittmatter, S. M. (2009) Cellular prion protein mediates impairment of synaptic plasticity by amyloid-β oligomers, Nature, 457, 1128–1132.
Um, J. W., and Strittmatter, S. M. (2013) Amyloid-β induced signaling by cellular prion protein and Fyn kinase in Alzheimer disease, Prion, 7, 37–41.
Bukau, B., and Horwich, A. L. (1998) The Hsp70 and Hsp60 chaperone machines, Cell, 92, 351–366.
Carrell, R. W., and Lomas, D. A. (1997) Conformational disease, Lancet, 350, 134–138.
Polyakova, O. V., Roitel, O., Asryants, R. A., Poliakov, A. A., Branlant, G., and Muronetz, V. I. (2005) Misfolded forms of glyceraldehyde-3-phosphate dehydrogenase interact with GroEL and inhibit chaperonin-assisted folding of the wild-type enzyme, Protein Sci., 14, 921–928.
Welch, W. J., and Gambetti, P. (1998) Chaperoning brain diseases, Nature, 392, 23–24.
Dulle, J. E., Bouttenot, R. E., Underwood, L. A., and True, H. L. (2013) Soluble oligomers are sufficient for transmission of a yeast prion but do not confer phenotype, J. Cell. Biol, 203, 197–204.
Silveira, J. R., Raymond, G. J., Hughson, A. G., Race, R. E., Sim, V. L., Hayes, S. F., and Caughey, B. (2005) The most infectious prion protein particles, Nature, 437, 257–261.
Simoneau, S., Rezaei, H., Sales, N., Kaiser-Schulz, G., Lefebvre-Roque, M., Vidal, C., Fournier, J. G., Comte, J., Wopfner, F., Grosclaude, J., Schatzl, H., and Lasmezas, C. I. (2007) In vitro and in vivo neurotoxicity of prion protein oligomers, PLoS Pathog., 3, e125.
Huang, P., Lian, F., Wen, Y., Guo, C., and Lin, D. (2013) Prion protein oligomer and its neurotoxicity, Acta Biochim. Biophys. Sin. (Shanghai), 45, 442–451.
Novitskaya, V., Bocharova, O. V., Bronstein, I., and Baskakov, I. V. (2006) Amyloid fibrils of mammalian prion protein are highly toxic to cultured cells and primary neurons, J. Biol. Chem., 281, 13828–13836.
Bailey, C. K., Andriola, I. F., Kampinga, H. H., and Merry, D. E. (2002) Molecular chaperones enhance the degradation of expanded polyglutamine repeat androgen receptor in a cellular model of spinal and bulbar muscular atrophy, Hum. Mol. Genet., 11, 515–523.
Hageman, J., Rujano, M. A., van Waarde, M. A., Kakkar, V., Dirks, R. P., Govorukhina, N., Oosterveld-Hut, H. M., Lubsen, N. H., and Kampinga, H. H. (2010) A DNAJB chaperone subfamily with HDAC-dependent activities suppresses toxic protein aggregation, Mol. Cell, 37, 355–369.
Waudby, C. A., Knowles, T. P., Devlin, G. L., Skepper, J. N., Ecroyd, H., Carver, J. A., Welland, M. E., Christodoulou, J., Dobson, C. M., and Meehan, S. (2010) The interaction of αB-crystallin with mature α-synuclein amyloid fibrils inhibits their elongation, Biophys. J., 98, 843–851.
DebBurman, S. K., Raymond, G. J., Caughey, B., and Lindquist, S. (1997) Chaperone-supervised conversion of prion protein to its protease-resistant form, Proc. Natl. Acad. Sci. USA, 94, 13938–13943.
Shorter, J., and Lindquist, S. (2004) Hsp104 catalyzes formation and elimination of self-replicating Sup35 prion conformers, Science, 304, 1793–1797.
Paushkin, S. V., Kushnirov, V. V., Smirnov, V. N., and Ter-Avanesyan, M. D. (1996) Propagation of the yeast prionlike [psi+] determinant is mediated by oligomerization of the SUP35-encoded polypeptide chain release factor, EMBO J., 15, 3127–3134.
Kryndushkin, D. S., Alexandrov, I. M., Ter-Avanesyan, M. D., and Kushnirov, V. V. (2003) Yeast [PSI+] prion aggregates are formed by small Sup35 polymers fragmented by Hsp104, J. Biol. Chem., 278, 49636–49643.
Naletova, I. N., Muronetz, V. I., and Schmalhausen, E. V. (2006) Unfolded, oxidized, and thermoinactivated forms of glyceraldehyde-3-phosphate dehydrogenase interact with the chaperonin GroEL in different ways, Biochim. Biophys. Acta, 1764, 831–838.
Stockel, J., and Hartl, F. U. (2001) Chaperonin-mediated de novo generation of prion protein aggregates, J. Mol. Biol., 313, 861–872.
Kiselev, G. G., Naletova, I. N., Sheval, E. V., Stroylova, Y. Y., Schmalhausen, E. V., Haertle, T., and Muronetz, V. I. (2011) Chaperonins induce an amyloid-like transformation of ovine prion protein: the fundamental difference in action between eukaryotic TRiC and bacterial GroEL, Biochim. Biophys. Acta, 1814, 1730–1738.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Biokhimiya, 2014, Vol. 79, No. 8, pp. 957–973.
Rights and permissions
About this article
Cite this article
Stroylova, Y.Y., Kiselev, G.G., Schmalhausen, E.V. et al. Prions and chaperones: Friends or foes?. Biochemistry Moscow 79, 761–775 (2014). https://doi.org/10.1134/S0006297914080045
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297914080045