Abstract
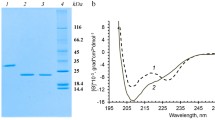
Potato virus A (PVA) protein coat contains on its surface partially unstructured N-terminal domain of the viral coat protein (CP), whose structural and functional characteristics are important for understanding the mechanism of plant infection with this virus. In this work, we investigated the properties and the structure of intact PVA and partially trypsinized PVAΔ32 virions using small-angle X-ray scattering (SAXS) and complimentary methods. It was shown that after the removal of 32 N-terminal amino acids of the CP, the virion did not disintegrate and remained compact, but the helical pitch of the CP packing changed. To determine the nature of these changes, we performed ab initio modeling, including the multiphase procedure, with the geometric bodies (helices) and restoration of the PVA structure in solution using available high-resolution structures of the homologous CP from the PVY potyvirus, based on the SAXS data. As a result, for the first time, a low-resolution structure of the filamentous PVA virus, both intact and partially degraded, was elucidated under conditions close to natural. The far-UV circular dichroism spectra of the PVA and PVAΔ32 samples differed significantly in the amplitude and position of the main negative maximum. The extent of thermal denaturation of these samples in the temperature range of 20-55°C was also different. The data of transmission electron microscopy showed that the PVAΔ32 virions were mostly rod-shaped, in contrast to the flexible filamentous particles typical of the intact virus, which correlated well with the SAXS results. In general, structural analysis indicates an importance of the CP N-terminal domain for the vital functions of PVA, which can be used to develop a strategy for combating this plant pathogen.








Similar content being viewed by others
Abbreviations
- CP:
-
coat protein
- PVA:
-
potato virus A
- SAXS:
-
small angle X-ray scattering
- TEM:
-
transmission electron microscopy
References
Ksenofontov, A. L., Paalme, V., Arutyunyan, A. M., Semenyuk, P. I., Fedorova, N. V., et al. (2013) Partially disordered structure in intravirus coat protein of potyvirus potato virus A, PLoS One, 8, e67830, https://doi.org/10.1371/journal.pone.0067830.
Zamora, M., Mendez-Lopez, E., Agirrezabala, X., Cuesta, R., Lavin, J. L., et al. (2017) Potyvirus virion structure shows conserved protein fold and RNA binding site in ssRNA viruses, Sci. Adv., 3, eaao2182, https://doi.org/10.1126/sciadv.aao2182.
Kezar, A., Kavcic, L., Polak, M., Novacek, J., Gutierrez-Aguirre, I., et al. (2019) Structural basis for the multitasking nature of the potato virus Y coat protein, Sci. Adv., 5, eaaw3808, https://doi.org/10.1126/sciadv.aaw3808.
Cuesta, R., Yuste-Calvo, C., Gil-Carton, D., Sanchez, F., Ponz, F., and Valle, M. (2019) Structure of Turnip mosaic virus and its viral-like particles, Sci. Rep., 9, 15396, https://doi.org/10.1038/s41598-019-51823-4.
Uversky, V. N. (2013) Unusual biophysics of intrinsically disordered proteins, Biochim. Biophys. Acta, 1834, 932-951, https://doi.org/10.1016/j.bbapap.2012.12.008.
Ksenofontov, A. L., Parshina, E. Y., Fedorova, N. V., Arutyunyan, A. M., Rumvolt, R., et al. (2016) Heating-induced transition of Potyvirus Potato Virus A coat protein into beta-structure, J. Biomol. Struct. Dyn., 34, 250-258, https://doi.org/10.1080/07391102.2015.1022604.
Ksenofontov, A. L., Dobrov, E. N., Fedorova, N. V., Arutyunyan, A. M., et al. (2018) Structure of Potato Virus A coat protein particles and their dissociation, Mol. Biol. (Mosk.), 52, 1055-1065, https://doi.org/10.1134/S0026898418060101.
Ksenofontov, A. L., Dobrov, E. N., Fedorova, N. V., Serebryakova, M. V., Prusov, A. N., et al. (2018) Isolated Potato Virus A coat protein possesses unusual properties and forms different short virus-like particles, J. Biomol. Struct. Dyn., 36, 1-11, https://doi.org/10.1080/07391102.2017.1333457.
Anindya, R., and Savithri, H. S. (2003) Surface-exposed amino- and carboxy-terminal residues are crucial for the initiation of assembly in Pepper vein banding virus: a flexuous rod-shaped virus, Virology, 316, 325-336, https://doi.org/10.1016/s0042-6822(03)00593-2.
Tatineni, S., McMechan, A. J., and Hein, G. L. (2018) Wheat streak mosaic virus coat protein is a determinant for vector transmission by the wheat curl mite, Virology, 514, 42-49, https://doi.org/10.1016/j.virol.2017.10.018.
Voloudakis, A. E., Malpica, C. A., Aleman-Verdaguer, M. E., Stark, D. M., Fauquet, C. M., and Beachy, R. N. (2004) Structural characterization of Tobacco etch virus coat protein mutants, Arch. Virol., 149, 699-712, https://doi.org/10.1007/s00705-003-0247-x.
Svergun, D. I., Koch, M. H. J., Timmins, P. A., and May, R. P. (2013) Small Angle X-Ray and Neutron Scattering from Solutions of Biological Macromolecules, First Edn., Oxford University Press, Oxford, https://doi.org/10.1093/acprof:oso/9780199639533.001.0001.
Franke, D., Petoukhov, M. V., Konarev, P. V., Panjkovich, A., Tuukkanen, A., et al. (2017) ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions, J. Appl. Crystallogr., 50, 1212-1225, https://doi.org/10.1107/S1600576717007786.
Ksenofontov, A. L., Petoukhov, M. V., Prusov, A. N., Fedorova, N. V., and Shtykova, E. V. (2020) Characterization of tobacco mosaic virus virions and repolymerized coat protein aggregates in solution by small-angle x-ray scattering, Biochemistry (Moscow), 85, 310-317, https://doi.org/10.1134/s0006297920030062.
Laemmli, U. K. (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4, Nature, 227, 680-685, https://doi.org/10.1038/227680a0.
Ksenofontov, A. L., Kozlovskii, V. S., Kordiukova, L. V., Radiukhin, V. A., Timofeeva, A. V., and Dobrov, E. N. (2006) Determination of concentration and aggregate size in influenza virus preparations using the true UV-absorption spectra, Mol. Biol., 40, 152-158, https://doi.org/10.1134/S0026893306010201.
Blanchet, C. E., Spilotros, A., Schwemmer, F., Graewert, M. A., Kikhney, A., et al. (2015) Versatile sample environments and automation for biological solution X-ray scattering experiments at the P12 beamline (PETRA III, DESY), J. Appl. Crystallogr., 48, 431-443, https://doi.org/10.1107/S160057671500254X.
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J., and Svergun, D. I. (2003) PRIMUS: a Windows PC-based system for small-angle scattering data analysis, J. Appl. Crystallogr., 36, 1277-1282, https://doi.org/10.1107/s0021889803012779.
Petoukhov, M. V., and Svergun, D. I. (2015) Ambiguity assessment of small-angle scattering curves from monodisperse systems, Acta Crystallogr. D Biol. Crystallogr., 71, 1051-1058, https://doi.org/10.1107/s1399004715002576.
Svergun, D. I. (1992) Determination of the regularization parameter in indirect-transform methods using perceptual criteria, J. Appl. Crystallogr., 25, 495-503, https://doi.org/10.1107/s0021889892001663.
Svergun, D. I. (1999) Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing, Biophys. J., 76, 2879-2886, https://doi.org/10.1016/S0006-3495(99)77443-6.
Konarev, P. V., Petoukhov, M. V., and Svergun, D. I. (2001) MASSHA – a graphics system for rigid-body modelling of macromolecular complexes against solution scattering data, J. Appl. Crystallogr., 34, 527-532, https://doi.org/10.1107/s0021889801006100.
Svergun, D., Barberato, C., and Koch, M. H. J. (1995) CRYSOL – a program to evaluate x-ray solution scattering of biological macromolecules from atomic coordinates, J. Appl. Crystallogr., 28, 768-773, https://doi.org/10.1107/s0021889895007047.
Artimo, P., Jonnalagedda, M., Arnold, K., Baratin, D., Csardi, G., et al. (2012) ExPASy: SIB bioinformatics resource portal, Nucleic Acids Res., 40, W597-603, https://doi.org/10.1093/nar/gks400.
Shtykova, E. V. (2015) Shape determination of polydisperse and polymorphic nanoobjects from small-angle X-ray scattering data (computer simulation), Nanotechnol. Russia, 10, 408-419, https://doi.org/10.1134/S1995078015030155.
Tardieu, A. (1994) Neutron and synchrotron radiation for condensed matter studies. Applications to Soft Condensed Matter and Biology, vol III (Les editions de Physique (France)) (Berlin: Springer), pp. 145-160.
Feigin, L. A., and Svergun, D. (1987) Structure analysis by small-angle x-ray and neutron scattering, Plenum Press, New York, p. 335.
Amari, K., Lerich, A., Schmitt-Keichinger, C., Dolja, V. V., and Ritzenthaler, C. (2011) Tubule-guided cell-to-cell movement of a plant virus requires class XI myosin motors, PLoS Pathog., 7, e1002327, https://doi.org/10.1371/journal.ppat.1002327.
Funding
This work was supported by the Russian Foundation for Basic Research (project no. 18-04-00525a), by the Ministry of Science and Higher Education of the Russian Federation (agreement 075-15-2019-1653), and by the Ministry of Science and Higher Education of the Russian Federation within the State assignment FSRC “Crystallography and Photonics” RAS (SAXS experiments).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflict of interest in financial or any other sphere. This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Shtykova, E.V., Petoukhov, M.V., Fedorova, N.V. et al. The Structure of the Potato Virus A Particles Elucidated by Small Angle X-Ray Scattering and Complementary Techniques. Biochemistry Moscow 86, 230–240 (2021). https://doi.org/10.1134/S0006297921020115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297921020115