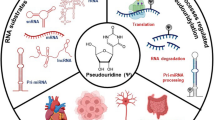
Abstract
This review considers approaches for detection of modified monomers in the RNA structure of living organisms. Recently, some data on dynamic alterations in the pool of modifications of the key RNA species that depend on external factors affecting the cells and physiological conditions of the whole organism have been accumulated. The recent studies have presented experimental data on relationship between the mechanisms of formation of modified/minor nucleotides of RNA in mammalian cells and the development of various pathologies. The development of novel methods for detection of chemical modifications of RNA nucleotides in the cells of living organisms and accumulation of knowledge on the contribution of modified monomers to metabolism and functioning of individual RNA species establish the basis for creation of novel diagnostic and therapeutic approaches. This review includes a short description of routine methods for determination of modified nucleotides in RNA and considers in detail modern approaches that enable not only detection but also quantitative assessment of the modification level of various nucleotides in individual RNA species.
Similar content being viewed by others
Abbreviations
- CMC:
-
N-cyclohexyl-N′-(2-(4-methyl)morpholinoethyl)carbodiimide tosylate
- DMS:
-
dimethyl sulfate
- ds:
-
double-stranded
- HTS:
-
high-throughput sequencing
- I:
-
inosine
- 1M7:
-
1-methyl-7-nitroisatoic anhydride
- m5C:
-
5-methylcytidine
- m6A:
-
N6-methyladenosine
- mN:
-
nucleotide modified at the base
- MS:
-
mass spectrometry
- Nm:
-
nucleotide methylated at the 2′-O-position
- NMIA:
-
N-methylisatoic anhydride
- 2′-O-Me:
-
2′-O-methyl group
- pre-mRNA:
-
precursor of mRNA
- RNP:
-
ribonucleoprotein
- RT:
-
reverse transcription
- RT-PCR:
-
reverse transcription followed by polymerase chain reaction
- snRNA:
-
small nuclear RNA
- snoRNA:
-
small nucleolar RNA
- ss:
-
single-stranded
- Ψ:
-
pseudouridine
References
Rozenski, J., Crain, P. F., and McCloskey, J. A. (1999) The RNA Modification Database: 1999 update, Nucleic Acids Res., 27, 196–197.
Piekna-Przybylska, D., Decatur, W. A., and Fournier, M. J. (2008) The 3D rRNA modification maps database: with interactive tools for ribosome analysis, Nucleic Acids Res., 36, D178–D183.
Machnicka, M. A., Milanowska, K., Osman Oglou, O., Purta, E., Kurkowska, M., Olchowik, A., Januszewski, W., Kalinowski, S., Dunin-Horkawicz, S., Rother, K. M., Helm, M., Bujnicki, J. M., and Grosjean, H. (2013) MODOMICS: a database of RNA modification pathways–2013 update, Nucleic Acids Res., 41, D262–267.
Helm, M. (2006) Post-transcriptional nucleotide modification and alternative folding of RNA, Nucleic Acids Res., 34, 721–733.
Kawai, G., Yamamoto, Y., Kamimura, T., Masegi, T., Sekine, M., Hata, T., Iimori, T., Watanabe, T., Miyazawa, T., and Yokoyama, S. (1992) Conformational rigidity of specific pyrimidine residues in tRNA arises from posttran-scriptional modifications that enhance steric interaction between the base and the 2′-hydroxyl group, Biochemistry, 31, 1040–1046.
Yarian, C. S., Basti, M. M., Cain, R. J., Ansari, G., Guenther, R. H., Sochacka, E., Czerwinska, G., Malkiewicz, A., and Agris, P. F. (1999) Structural and func-tional roles of the N1-and N3-protons of psi at tRNA’s position 39, Nucleic Acids Res., 27, 3543–3549.
Desaulniers, J. P., Chang, Y. C., Aduri, R., Abeysirigunawardena, S. C., SantaLucia, J., Jr., and Chow, C. S. (2008) Pseudouridines in rRNA helix 69 play a role in loop stacking interactions, Org. Biomol. Chem., 6, 3892–3895.
Charette, M., and Gray, M. W. (2000) Pseudouridine in RNA: what, where, how, and why, IUBMB Life, 49, 341–351.
Hudson, G. A., Bloomingdale, R. J., and Znosko, B. M. (2013) Thermodynamic contribution and nearest-neighbor parameters of pseudouridine-adenosine base pairs in oligoribonucleotides, RNA, 19, 1474–1482.
Kierzek, E., Malgowska, M., Lisowiec, J., Turner, D. H., Gdaniec, Z., and Kierzek, R. (2014) The contribution of pseudouridine to stabilities and structure of RNAs, Nucleic Acids Res., 42, 3492–3501.
Kimura, S., and Suzuki, T. (2010) Fine-tuning of the ribosomal decoding center by conserved methyl-modifications in the Escherichia coli 16S rRNA, Nucleic Acids Res., 38, 1341–1352.
Helm, M., Giege, R., and Florentz, C. (1999) A Watson–Crick base-pair-disrupting methyl group (m1A9) is sufficient for cloverleaf folding of human mitochondrial tRNALys, Biochemistry, 38, 13338–13346.
Ishitani, R., Yokoyama, S., and Nureki, O. (2008) Structure, dynamics, and function of RNA modification enzymes, Curr. Opin. Struct. Biol., 18, 330–339.
Byrne, R. T., Waterman, D. G., and Antson, A. A. (2009) in DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution: Enzyme–RNA Substrate Recognition in RNA-Modifying Enzymes (Grosjean, H., ed.) Landes Bioscience, Austin, pp. 303–327.
Kiss-Laszlo, Z., Henry, Y., Bachellerie, J. P., Caizergues-Ferrer, M., and Kiss, T. (1996) Site-specific ribose methy-lation of preribosomal RNA: a novel function for small nucleolar RNAs, Cell, 85, 1077–1088.
Cavaille, J., Nicoloso, M., and Bachellerie, J. P. (1996) Targeted ribose methylation of RNA in vivo directed by tai-lored antisense RNA guides, Nature, 383, 732–735.
Ganot, P., Bortolin, M. L., and Kiss, T. (1997) Site-specif-ic pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs, Cell, 89, 799–809.
Makarova, J. A., Ivanova, S. M., Tonevitsky, A. G., and Grigoriev, A. I. (2013) New functions of small nucleolar RNAs, Biochemistry (Moscow), 78, 638–650.
Dupuis-Sandoval, F., Poirier, M., and Scott, M. S. (2015) The emerging landscape of small nucleolar RNAs in cell biology, Wiley Interdiscip. Rev. RNA, 6, 381–397.
Shubina, M. Y., Musinova, Y. R., and Sheval, E. V. (2016) Nucleolar methyltransferase fibrillarin: evolution of struc-ture and functions, Biochemistry (Moscow), 81, 941–950.
Omer, A. D., Ziesche, S., Ebhardt, H., and Dennis, P. P. (2002) In vitro reconstitution and activity of a C/D box methylation guide ribonucleoprotein complex, Proc. Natl. Acad. Sci. USA, 99, 5289–5294.
Johansen, S. K., Maus, C. E., Plikaytis, B. B., and Douthwaite, S. (2006) Capreomycin binds across the ribo-somal subunit interface using tlyA-encoded 2′-O-methyla-tions in 16S and 23S rRNAs, Mol. Cell, 23, 173–182.
Okamoto, S., Tamaru, A., Nakajima, C., Nishimura, K., Tanaka, Y., Tokuyama, S., Suzuki, Y., and Ochi, K. (2007) Loss of a conserved 7-methylguanosine modification in 16S rRNA confers low-level streptomycin resistance in bacte-ria, Mol. Microbiol., 63, 1096–1106.
Esguerra, J., Warringer, J., and Blomberg, A. (2008) Functional importance of individual rRNA 2′-O-ribose methylations revealed by high-resolution phenotyping, RNA, 14, 649–656.
Karijolich, J., and Yu, Y. T. (2010) Spliceosomal snRNA modifications and their function, RNA Biol., 7, 192–204.
Jackman, J. E., and Alfonzo, J. D. (2013) Transfer RNA modifications: nature’s combinatorial chemistry play-ground, Wiley Interdiscip. Rev. RNA, 4, 35–48.
Giege, R., Juhling, F., Putz, J., Stadler, P., Sauter, C., and Florentz, C. (2012) Structure of transfer RNAs: similarity and variability, Wiley Interdiscip. Rev. RNA, 3, 37–61.
Jenner, L. B., Demeshkina, N., Yusupova, G., and Yusupov, M. (2010) Structural aspects of messenger RNA reading frame maintenance by the ribosome, Nat. Struct. Mol. Biol., 17, 555–560.
Das, G., Thotala, D. K., Kapoor, S., Karunanithi, S., Thakur, S. S., Singh, N. S., and Varshney, U. (2008) Role of 16S ribosomal RNA methylations in translation initia-tion in Escherichia coli, EMBO J., 27, 840–851.
Liu, B., Liang, X. H., Piekna-Przybylska, D., Liu, Q., and Fournier, M. J. (2008) Mis-targeted methylation in rRNA can severely impair ribosome synthesis and activity, RNA Biol., 5, 249–254.
Liang, X. H., Liu, Q., and Fournier, M. J. (2009) Loss of rRNA modifications in the decoding center of the ribosome impairs translation and strongly delays pre-rRNA process-ing, RNA, 15, 1716–1728.
Sonenberg, N., and Hinnebusch, A. G. (2009) Regulation of translation initiation in eukaryotes: mechanisms and biological targets, Cell, 136, 731–745.
Dominissini, D., Moshitch-Moshkovitz, S., Schwartz, S., Salmon-Divon, M., Ungar, L., Osenberg, S., Cesarkas, K., Jacob-Hirsch, J., Amariglio, N., Kupiec, M., Sorek, R., and Rechavi, G. (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq, Nature, 485, 201–206.
Zinshteyn, B., and Nishikura, K. (2009) Adenosine-to-inosine RNA editing, Wiley Interdiscip. Rev. Syst. Biol. Med., 1, 202–209.
Tomaselli, S., Bonamassa, B., Alisi, A., Nobili, V., Locatelli, F., and Gallo, A. (2013) ADAR enzyme and miRNA story: a nucleotide that can make the difference, Int. J. Mol. Sci., 14, 22796–22816.
Zust, R., Cervantes-Barragan, L., Habjan, M., Maier, R., Neuman, B. W., Ziebuhr, J., Szretter, K. J., Baker, S. C., Barchet, W., Diamond, M. S., Siddell, S. G., Ludewig, B., and Thiel, V. (2011) Ribose 2′-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5, Nat. Immunol., 12, 137–143.
Hull, C. M., and Bevilacqua, P. C. (2016) Discriminating self and non-self by RNA: roles for RNA structure, mis-folding, and modification in regulating the innate immune sensor PKR, Acc. Chem. Res., 49, 1242–1249.
Belin, S., Kindbeiter, K., Hacot, S., Albaret, M. A., Roca-Martinez, J. X., Therizols, G., Grosso, O., and Diaz, J. J. (2010) Uncoupling ribosome biogenesis regulation from RNA polymerase I activity during herpes simplex virus type 1 infection, RNA, 16, 131–140.
Krug, R. M., Morgan, M. A., and Shatkin, A. J. (1976) Influenza viral mRNA contains internal N6-methyladeno-sine and 5′-terminal 7-methylguanosine in cap structures, J. Virol., 20, 45–53.
Daffis, S., Szretter, K. J., Schriewer, J., Li, J., Youn, S., Errett, J., Lin, T. Y., Schneller, S., Zust, R., Dong, H., Thiel, V., Sen, G. C., Fensterl, V., Klimstra, W. B., Pierson, T. C., Buller, R. M., Gale, M., Jr., Shi, P. Y., and Diamond, M. S. (2010) 2′-O-methylation of the viral mRNA cap evades host restriction by IFIT family members, Nature, 468, 452–456.
Reich, S., Guilligay, D., Pflug, A., Malet, H., Berger, I., Crepin, T., Hart, D., Lunardi, T., Nanao, M., Ruigrok, R. W., and Cusack, S. (2014) Structural insight into cap-snatching and RNA synthesis by influenza polymerase, Nature, 516, 361–366.
Guilligay, D., Kadlec, J., Crepin, T., Lunardi, T., Bouvier, D., Kochs, G., Ruigrok, R. W., and Cusack, S. (2014) Comparative structural and functional analysis of orthomyxovirus polymerase cap-snatching domains, PLoS One, 9, e84973.
Belin, S., Beghin, A., Solano-Gonzalez, E., Bezin, L., Brunet-Manquat, S., Textoris, J., Prats, A. C., Mertani, H. C., Dumontet, C., and Diaz, J. J. (2009) Dysregulation of ribosome biogenesis and translational capacity is associated with tumor progression of human breast cancer cells, PLoS One, 4, e7147.
Kishore, S., and Stamm, S. (2006) The snoRNA HBII-52 regulates alternative splicing of the serotonin receptor 2C, Science, 311, 230–232.
Singh, M. (2013) Dysregulated A to I RNA editing and non-coding RNAs in neurodegeneration, Front. Genet., 3, 326.
Prestwich, E. G., Mangerich, A., Pang, B., McFaline, J. L., Lonkar, P., Sullivan, M. R., Trudel, L. J., Taghizedeh, K., and Dedon, P. C. (2013) Increased levels of inosine in a mouse model of inflammation, Chem. Res. Toxicol., 26, 538–546.
Stepanov, G. A., Filippova, J. A., Komissarov, A. B., Kuligina, E. V., Richter, V. A., and Semenov, D. V. (2015) Regulatory role of small nucleolar RNAs in human dis-eases, Biomed. Res. Int., 2015, 206849.
Therizols, G., Laforets, F., Marcel, V., Catez, F., Bouvet, P., and Diaz, J. J. (2015) in Epigenetic Cancer Therapy: Ribosomal RNA Methylation and Cancer (Gray, S. G., ed.) American Elsevier, N. Y., pp. 129–139.
Grosjean, H., Keith, G., and Droogmans, L. (2004) Detection and quantification of modified nucleotides in RNA using thin-layer chromatography, Methods Mol. Biol., 265, 357–391.
Silberklang, M., Gillum, A. M., and RajBhandary, U. L. (1977) The use of nuclease P1 in sequence analysis of end group labeled RNA, Nucleic Acids Res., 4, 4091–4108.
Chan, C. T., Dyavaiah, M., DeMott, M. S., Taghizadeh, K., Dedon, P. C., and Begley, T. J. (2010) A quantitative systems approach reveals dynamic control of tRNA modi-fications during cellular stress, PLoS Genet., 6, e1001247.
Basanta-Sanchez, M., Temple, S., Ansari, S. A., D’Amico, A., and Agris, P. F. (2016) Attomole quantification and global profile of RNA modifications: epitranscriptome of human neural stem cells, Nucleic Acids Res., 44, e26.
Cai, W. M., Chionh, Y. H., Hia, F., Gu, C., Kellner, S., McBee, M. E., Ng, C. S., Pang, Y. L. J., Prestwich, E. G., Lim, K. S., Babu, I. R., Begley, T. J., and Dedon, P. C. (2015) A platform for discovery and quantification of mod-ified ribonucleosides in RNA: application to stress-induced reprogramming of tRNA modifications, Methods Enzymol., 560, 29–71.
Thuring, K., Schmid, K., Keller, P., and Helm, M. (2016) Analysis of RNA modifications by liquid chromatography-tandem mass spectrometry, Methods, 107, 48–56.
Pang, B., Zhou, X., Yu, H., Dong, M., Taghizadeh, K., Wishnok, J. S., Tannenbaum, S. R., and Dedon, P. C. (2007) Lipid peroxidation dominates the chemistry of DNA adduct formation in a mouse model of inflammation, Carcinogenesis, 28, 1807–1813.
Behm-Ansmant, I., Helm, M., and Motorin, Y. (2011) Use of specific chemical reagents for detection of modified nucleotides in RNA, J. Nucleic Acids, 2011, 408053.
Loverix, S., Winqvist, A., Stromberg, R., and Steyaert, J. (2000) Mechanism of RNase T1: concerted triester-like phosphoryl transfer via a catalytic three-centered hydrogen bond, Chem. Biol., 7, 651–658.
Guymon, R., Pomerantz, S. C., Ison, J. N., Crain, P. F., and McCloskey, J. A. (2007) Post-transcriptional modifica-tions in the small subunit ribosomal RNA from Thermotoga maritima, including presence of a novel modified cytidine, RNA, 13, 396–403.
Stepinski, J., Waddell, C., Stolarski, R., Darzynkiewicz, E., and Rhoads, R. E. (2001) Synthesis and properties of mRNAs containing the novel “anti-reverse” cap analogs 7-methyl(3′-O-methyl)GpppG and 7-methyl (3′-deoxy)GpppG, RNA, 7, 1486–1495.
Deshpande, R. A., and Shankar, V. (2002) Ribonucleases from T2 family, Crit. Rev. Microbiol., 28, 79–122.
Gaur, R., Bjork, G. R., Tuck, S., and Varshney, U. (2007) Diet-dependent depletion of queuosine in tRNAs in Caenorhabditis elegans does not lead to a developmental block, J. Biosci., 32, 747–754.
Volkin, E., and Cohn, W. E. (1953) On the structure of ribonucleic acids. II. The products of ribonuclease action, J. Biol. Chem., 205, 767–782.
Glitz, D. G., and Dekker, C. A. (1964) Studies on a ribonu-clease from Ustilago sphaerogenna. II. Specificity of the enzyme, Biochemistry, 3, 1399–1406.
Addepalli, B., Lesner, N. P., and Limbach, P. A. (2015) Detection of RNA nucleoside modifications with the uri-dine-specific ribonuclease MC1 from Momordica charan-tia, RNA, 21, 1746–1756.
Morse, D. P., and Bass, B. L. (1997) Detection of inosine in messenger RNA by inosine-specific cleavage, Biochemistry, 36, 8429–8434.
Morse, D. P. (2004) Identification of substrates for adeno-sine deaminases that act on RNA, Methods Mol. Biol., 265, 199–218.
Nishikura, K. (2010) Functions and regulation of RNA edit-ing by ADAR deaminase, Annu. Rev. Biochem., 79, 321–349.
Mengel-Jorgensen, J., and Kirpekar, F. (2002) Detection of pseudouridine and other modifications in tRNA by cya-noethylation and MALDI mass spectrometry, Nucleic Acids Res., 30, e135.
Chan, C. T., Pang, Y. L., Deng, W., Babu, I. R., Dyavaiah, M., Begley, T. J., and Dedon, P. C. (2012) Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins, Nat. Commun., 3, 937.
Ross, R., Cao, X., Yu, N., and Limbach, P. A. (2016) Sequence mapping of transfer RNA chemical modifica-tions by liquid chromatography tandem mass spectrometry, Methods, 107, 73–78.
Kirpekar, F., Douthwaite, S., and Roepstorff, P. (2000) Mapping posttranscriptional modifications in 5S ribosomal RNA by MALDI mass spectrometry, RNA, 6, 296–306.
Yoshida, M., and Ukita, T. (1968) Modification of nucleo-sides and nucleotides. VII. Selective cyanoethylation of inosine and pseudouridine in yeast transfer ribonucleic acid, Biochim. Biophys. Acta, 157, 455–465.
Emmerechts, G., Herdewijn, P., and Rozenski, J. (2005) Pseudouridine detection improvement by derivatization with methyl vinyl sulfone and capillary HPLC-mass spec-trometry, J. Chromatogr. B. Analyt. Technol. Biomed. Life Sci., 825, 233–238.
Ofengand, J., Del Campo, M., and Kaya, Y. (2001) Mapping pseudouridines in RNA molecules, Methods, 25, 365–373.
Patteson, K. G., Rodicio, L. P., and Limbach, P. A. (2001) Identification of the mass-silent post-transcriptionally modified nucleoside pseudouridine in RNA by matrix-assisted laser desorption/ionization mass spectrometry, Nucleic Acids Res., 29, E49-9.
Durairaj, A., and Limbach, P. A. (2008) Improving CMC-derivatization of pseudouridine in RNA for mass spectro-metric detection, Anal. Chim. Acta, 612, 173–181.
Popova, A. M., and Williamson, J. R. (2014) Quantitative analysis of rRNA modifications using stable isotope label-ing and mass spectrometry, J. Am. Chem. Soc., 136, 2058–2069.
Kellner, S., Neumann, J., Rosenkranz, D., Lebedeva, S., Ketting, R. F., Zischler, H., Schneider, D., and Helm, M. (2014) Profiling of RNA modifications by multiplexed sta-ble isotope labelling, Chem. Commun. (Camb.), 50, 3516–3518.
Taoka, M., Nobe, Y., Hori, M., Takeuchi, A., Masaki, S., Yamauchi, Y., Nakayama, H., Takahashi, N., and Isobe, T. (2015) A mass spectrometry-based method for comprehen-sive quantitative determination of post-transcriptional RNA modifications: the complete chemical structure of Schizosaccharomyces pombe ribosomal RNAs, Nucleic Acids Res., 43, e115.
Nakayama, H., Takahashi, N., and Isobe, T. (2011) Informatics for mass spectrometry-based RNA analysis, Mass Spectrom. Rev., 30, 1000–1012.
Sample, P. J., Gaston, K. W., Alfonzo, J. D., and Limbach, P. A. (2015) RoboOligo: software for mass spectrometry data to support manual and de novo sequencing of post-transcriptionally modified ribonucleic acids, Nucleic Acids Res., 43, e64.
Ho, N. W., and Gilham, P. T. (1971) Reaction of pseudouridine and inosine with N-cyclohexyl-N′-β-(4-methylmorpholinium)ethylcarbodiimide, Biochemistry, 10, 3651–3657.
Bakin, A., and Ofengand, J. (1993) Four newly located pseudouridylate residues in Escherichia coli 23S ribosomal RNA are all at the peptidyltransferase center: analysis by the application of a new sequencing technique, Biochemistry, 32, 9754–9762.
Bakin, A., and Ofengand, J. (1998) Mapping of pseudouri-dine residues in RNA to nucleotide resolution, Methods Mol. Biol., 77, 297–309.
Carlile, T. M., Rojas-Duran, M. F., Zinshteyn, B., Shin, H., Bartoli, K. M., and Gilbert, W. V. (2014) Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells, Nature, 515, 143–146.
Lovejoy, A. F., Riordan, D. P., and Brown, P. O. (2014) Transcriptome-wide mapping of pseudouridines: pseudouridine synthases modify specific mRNAs in S. cere-visiae, PLoS One, 9, e110799.
Schwartz, S., Bernstein, D. A., Mumbach, M. R., Jovanovic, M., Herbst, R. H., Leon-Ricardo, B. X., Engreitz, J. M., Guttman, M., Satija, R., Lander, E. S., Fink, G., and Regev, A. (2014) Transcriptome-wide map-ping reveals widespread dynamic-regulated pseudouridyla-tion of ncRNA and mRNA, Cell, 159, 148–162.
Li, X., Zhu, P., Ma, S., Song, J., Bai, J., Sun, F., and Yi, C. (2015) Chemical pulldown reveals dynamic pseudouridylation of the mammalian transcriptome, Nat. Chem. Biol., 11, 592–597.
Clark, S. J., Harrison, J., Paul, C. L., and Frommer, M. (1994) High sensitivity mapping of methylated cytosines, Nucleic Acids Res., 22, 2990–2997.
Gu, W., Hurto, R. L., Hopper, A. K., Grayhack, E. J., and Phizicky, E. M. (2005) Depletion of Saccharomyces cere-visiae tRNA(His) guanylyltransferase Thg1p leads to uncharged tRNAHis with additional m5C, Mol. Cell. Biol., 25, 8191–8201.
Schaefer, M., Pollex, T., Hanna, K., and Lyko, F. (2009) RNA cytosine methylation analysis by bisulfite sequenc-ing, Nucleic Acids Res., 37, e12.
Edelheit, S., Schwartz, S., Mumbach, M. R., Wurtzel, O., and Sorek, R. (2013) Transcriptome-wide mapping of 5-methylcytidine RNA modifications in bacteria, archaea, and yeast reveals m5C within archaeal mRNAs, PLoS Genet., 9, e1003602.
Khoddami, V., and Cairns, B. R. (2013) Identification of direct targets and modified bases of RNA cytosine methyl-transferases, Nat. Biotechnol., 31, 458–464.
Hussain, S., Sajini, A. A., Blanco, S., Dietmann, S., Lombard, P., Sugimoto, Y., Paramor, M., Gleeson, J. G., Odom, D. T., Ule, J., and Frye, M. (2013) NSun2-medi-ated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs, Cell Rep., 4, 255–261.
Flusberg, B. A., Webster, D. R., Lee, J. H., Travers, K. J., Olivares, E. C., Clark, T. A., Korlach, J., and Turner, S. W. (2010) Direct detection of DNA methylation during single-molecule, real-time sequencing, Nat. Methods, 7, 461–465.
Vilfan, I. D., Tsai, Y. C., Clark, T. A., Wegener, J., Dai, Q., Yi, C., Pan, T., Turner, S. W., and Korlach, J. (2013) Analysis of RNA base modification and structural rearrangement by single-molecule real-time detection of reverse transcription, J. Nanobiotechnol., 11, 8.
Golovina, A. Y., Dzama, M. M., Petriukov, K. S., Zatsepin, T. S., Sergiev, P. V., Bogdanov, A. A., and Dontsova, O. A. (2014) Method for site-specific detection of m6A nucleoside presence in RNA based on high-resolu-tion melting (HRM) analysis, Nucleic Acids Res., 42, e27.
Jia, G., Fu, Y., and He, C. (2013) Reversible RNA adeno-sine methylation in biological regulation, Trends Genet., 29, 108–115.
Tycowski, K. T., Smith, C. M., Shu, M.-D., and Steitz, J. A. (1996) A small nucleolar RNA requirement for site-spe-cific ribose methylation of rRNA in Xenopus, Proc. Natl. Acad. Sci. USA, 93, 14480–14485.
Rebane, A., Roomere, H., and Metspalu, A. (2002) Locations of several novel 2′-O-methylated nucleotides in human 28S rRNA, BMC Mol. Biol., 3, 1.
Smith, J. D., and Dunn, D. B. (1959) An additional sugar component of ribonucleic acids, Biochim. Biophys. Acta, 31, 573–575.
Buchhaupt, M., Peifer, C., and Entian, K. D. (2007) Analysis of 2′-O-methylated nucleosides and pseudouridines in ribosomal RNAs using DNAzymes, Anal. Biochem., 361, 102–108.
Hengesbach, M., Meusburger, M., Lyko, F., and Helm, M. (2008) Use of DNAzymes for site-specific analysis of ribonucleotide modifications, RNA, 14, 180–187.
Gehrig, S., Eberle, M. E., Botschen, F., Rimbach, K., Eberle, F., Eigenbrod, T., Kaiser, S., Holmes, W. M., Erdmann, V. A., Sprinzl, M., Bec, G., Keith, G., Dalpke, A. H., and Helm, M. (2012) Identification of modifica-tions in microbial, native tRNA that suppress immunos-timulatory activity, J. Exp. Med., 209, 225–233.
Buchhaupt, M., Sharma, S., Kellner, S., Oswald, S., Paetzold, M., Peifer, C., Watzinger, P., Schrader, J., Helm, M., and Entian, K. D. (2014) Partial methylation at Am100 in 18S rRNA of baker’s yeast reveals ribosome het-erogeneity on the level of eukaryotic rRNA modification, PLoS One, 9, e89640.
Maden, B. E., Corbett, M. E., Heeney, P. A., Pugh, K., and Ajuh, P. M. (1995) Classical and novel approaches to the detection and localization of the numerous modified nucleotides in eukaryotic ribosomal RNA, Biochimie, 77, 22–29.
Maden, B. E. (2001) Mapping 2′-O-methyl groups in ribo-somal RNA, Methods, 25, 374–382.
Filippova, J. A., Stepanov, G. A., Semenov, D. V., Koval, O. A., Kuligina, E. V., Rabinov, I. V., and Richter, V. A. (2015) Modified method of rRNA structure analysis reveals novel characteristics of box C/D RNA analogues, Acta Naturae, 7, 64–73.
Qu, G., Van Nues, R. W., Watkins, N. J., and Maxwell, E. S. (2011) Mol. Cell. Biol., 31, 365–374.
Blatter, N., Bergen, K., Nolte, O., Welte, W., Diederichs, K., Mayer, J., Wieland, M., and Marx, A. (2013) Structure and function of an RNA-reading thermostable DNA poly-merase, Angew. Chem. Int. Ed. Engl., 52, 11935–11939.
Aschenbrenner, J., and Marx, A. (2016) Direct and site-specific quantification of RNA 2′-O-methylation by PCR with an engineered DNA polymerase, Nucleic Acids Res., 44, 3495–3502.
Karijolich, J., Kantartzis, A., and Yu, Y. T. (2010) Quantitative analysis of RNA modifications, Methods Mol. Biol., 629, 21–32.
Wu, G., Xiao, M., Yang, C., and Yu, Y. T. (2011) U2 snRNA is inducibly pseudouridylated at novel sites by Pus7p and snR81 RNP, EMBO J., 30, 79–89.
Yu, Y. T., Shu, M. D., and Steitz, J. A. (1997) A new method for detecting sites of 2′-O-methylation in RNA molecules, RNA, 3, 324–331.
Gonzales, B., Henning, D., So, R. B., Dixon, J., Dixon, M. J., and Valdez, B. C. (2005) The Treacher Collins syn-drome (TCOF1) gene product is involved in pre-rRNA methylation, Hum. Mol. Genet., 14, 2035–2043.
Saikia, M., Dai, Q., Decatur, W. A., Fournier, M. J., Piccirilli, J. A., and Pan, T. (2006) A systematic, ligation-based approach to study RNA modifications, RNA, 12, 2025–2033.
Liu, N., Parisien, M., Dai, Q., Zheng, G., He, C., and Pan, T. (2013) Probing N6-methyladenosine RNA modifi-cation status at single nucleotide resolution in mRNA and long noncoding RNA, RNA, 19, 1848–1856.
Liu, N., and Pan, T. (2016) Probing N6-methyladenosine (m6A) RNA modification in total RNA with SCARLET, Methods Mol. Biol., 1358, 285–292.
Mishima, E., Jinno, D., Akiyama, Y., Itoh, K., Nankumo, S., Shima, H., Kikuchi, K., Takeuchi, Y., Elkordy, A., Suzuki, T., Niizuma, K., Ito, S., Tomioka, Y., and Abe, T. (2015) Immuno-Northern blotting: detection of RNA modifications by using antibodies against modified nucle-osides, PLoS One, 10, e0143756.
Waghmare, S. P., and Dickman, M. J. (2011) Characterization and quantification of RNA post-tran-scriptional modifications using stable isotope labeling of RNA in conjunction with mass spectrometry analysis, Anal. Chem., 83, 4894–4901.
Russell, S. P., and Limbach, P. A. (2013) Evaluating the reproducibility of quantifying modified nucleosides from ribonucleic acids by LC-UV-MS, J. Chromatogr. B Analyt. Technol. Biomed. Life Sci., 923-924, 74–82.
Su, D., Chan, C. T., Gu, C., Lim, K. S., Chionh, Y. H., McBee, M. E., Russell, B. S., Babu, I. R., Begley, T. J., and Dedon, P. C. (2014) Quantitative analysis of ribonu-cleoside modifications in tRNA by HPLC-coupled mass spectrometry, Nat. Protoc., 9, 828–841.
Kellner, S., Ochel, A., Thuring, K., Spenkuch, F., Neumann, J., Sharma, S., Entian, K. D., Schneider, D., and Helm, M. (2014) Absolute and relative quantification of RNA modifications via biosynthetic isotopomers, Nucleic Acids Res., 42, e142.
Meyer, K. D., Saletore, Y., Zumbo, P., Elemento, O., Mason, C. E., and Jaffrey, S. R. (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3′-UTRs and near stop codons, Cell, 149, 1635–1646.
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N., and Rechavi, G. (2013) Transcriptome-wide mapping of N6-methyladenosine by m6A-seq based on immunocapturing and massively paral-lel sequencing, Nat. Protoc., 8, 176–189.
Schwartz, S., Agarwala, S. D., Mumbach, M. R., Jovanovic, M., Mertins, P., Shishkin, A., Tabach, Y., Mikkelsen, T. S., Satija, R., Ruvkun, G., Carr, S. A., Lander, E. S., Fink, G. R., and Regev, A. (2013) High-res-olution mapping reveals a conserved, widespread, dynam-ic mRNA methylation program in yeast meiosis, Cell, 155, 1409–1421.
Squires, J. E., Patel, H. R., Nousch, M., Sibbritt, T., Humphreys, D. T., Parker, B. J., Suter, C. M., and Preiss, T. (2012) Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA, Nucleic Acids Res., 40, 5023–5033.
Chen, K., Lu, Z., Wang, X., Fu, Y., Luo, G. Z., Liu, N., Han, D., Dominissini, D., Dai, Q., Pan, T., and He, C. (2015) High-resolution N6-methyladenosine (m6A) map using photo-crosslinking-assisted m6A sequencing, Angew. Chem. Int. Ed. Engl., 54, 1587–1590.
Birkedal, U., Christensen-Dalsgaard, M., Krogh, N., Sabarinathan, R., Gorodkin, J., and Nielsen, H. (2015) Profiling of ribose methylations in RNA by high-through-put sequencing, Angew. Chem. Int. Ed. Engl., 54, 451–455.
Krogh, N., Jansson, M. D., Hafner, S. J., Tehler, D., Birkedal, U., Christensen-Dalsgaard, M., Lund, A. H., and Nielsen, H. (2016) Profiling of 2′-O-Me in human rRNA reveals a subset of fractionally modified positions and provides evidence for ribosome heterogeneity, Nucleic Acids Res., 44, 7884–7895.
Cattenoz, P. B., Taft, R. J., Westhof, E., and Mattick, J. S. (2013) Transcriptome-wide identification of A >I RNA editing sites by inosine specific cleavage, RNA, 19, 257–270.
Merino, E. J., Wilkinson, K. A., Coughlan, J. L., and Weeks, K. M. (2005) RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE), J. Am. Chem. Soc., 127, 4223–4231.
Wilkinson, K. A., Merino, E. J., and Weeks, K. M. (2006) Selective 2′-hydroxyl acylation analyzed by primer exten-sion (SHAPE): quantitative RNA structure analysis at sin-gle nucleotide resolution, Nat. Protoc., 1, 1610–1616.
Lusvarghi, S., Sztuba-Solinska, J., Purzycka, K. J., Rausch, J. W., and Le Grice, S. F. (2013) RNA secondary structure prediction using high-throughput SHAPE, J. Vis. Exp., 75, e50243.
Weeks, K. M., and Mauger, D. M. (2011) Exploring RNA structural codes with SHAPE chemistry, Acc. Chem. Res., 44, 1280–1291.
Steen, K. A., Siegfried, N. A., and Weeks, K. M. (2011) Selective 2′-hydroxyl acylation analyzed by protection from exoribonuclease (RNase-detected SHAPE) for direct analysis of covalent adducts and of nucleotide flexibility in RNA, Nat. Protoc., 6, 1683–1694.
Mortimer, S. A., and Weeks, K. M. (2007) A fast-acting reagent for accurate analysis of RNA secondary and terti-ary structure by SHAPE chemistry, J. Am. Chem. Soc., 129, 4144–4145.
Wilkinson, K. A., Vasa, S. M., Deigan, K. E., Mortimer, S. A., Giddings, M. C., and Weeks, K. M. (2009) Influence of nucleotide identity on ribose 2′-hydroxyl reactivity in RNA, RNA, 15, 1314–1321.
Vasa, S. M., Guex, N., Wilkinson, K. A., Weeks, K. M., and Giddings, M. C. (2008) ShapeFinder: a software sys-tem for high-throughput quantitative analysis of nucleic acid reactivity information resolved by capillary elec-trophoresis, RNA, 14, 1979–1990.
Kladwang, W., VanLang, C. C., Cordero, P., and Das, R. (2011) Understanding the errors of SHAPE-directed RNA structure modeling, Biochemistry, 50, 8049–8056.
Watts, J. M., Dang, K. K., Gorelick, R. J., Leonard, C. W., Bess, J. W., Jr., Swanstrom, R., Burch, C. L., and Weeks, K. M. (2009) Architecture and secondary structure of an entire HIV-1 RNA genome, Nature, 460, 711–716.
Novikova, I. V., Hennelly, S. P., and Sanbonmatsu, K. Y. (2012) Structural architecture of the human long non-coding RNA, steroid receptor RNA activator, Nucleic Acids Res., 40, 5034–5051.
Loopez-Carrasco, A., and Flores, R. (2016) Dissecting the secondary structure of the circular RNA of a nuclear viroid in vivo: a “naked” rod-like conformation similar but not identical to that observed in vitro, RNA Biol., 1–9, doi: 10.1080/15476286.2016.1223005.
Watters, K. E., and Lucks, J. B. (2016) Mapping RNA structure in vitro with SHAPE chemistry and next-genera-tion sequencing (SHAPE-Seq), Methods Mol. Biol., 1490, 135–162.
Steen, K. A., Malhotra, A., and Weeks, K. M. (2010) Selective 2′-hydroxyl acylation analyzed by protection from exoribonuclease, J. Am. Chem. Soc., 132, 9940–9943.
Ding, Y., Tang, Y., Kwok, C. K., Zhang, Y., Bevilacqua, P. C., and Assmann, S. M. (2014) In vivo genome-wide pro-filing of RNA secondary structure reveals novel regulatory features, Nature, 505, 696–700.
Picardi, E., Gallo, A., Galeano, F., Tomaselli, S., and Pesole, G. (2012) A novel computational strategy to iden-tify A-to-I RNA editing sites by RNA-Seq data: de novo detection in human spinal cord tissue, PLoS One, 7, e44184.
Lokhov, P. G., Balashova, E. E., Voskresenskaya, A. A., Trifonova, O. P., Maslov, D. L., and Archakov, A. I. (2016) Mass spectrometric signatures of the blood plasma metabolome for disease diagnostics, Biomed. Rep., 4, 122–126.
Warren, L., Manos, P. D., Ahfeldt, T., Loh, Y.-H., Li, H., Lau, F., Ebina, W., Mandal, P. K., Smith, Z. D., Meissner, A., Daley, G. Q., Brack, A. S., Collins, J. J., Cowan, C., Schlaeger, T. M., and Rossi, D. J. (2010) Highly efficient reprogramming to pluripotency and directed differentia-tion of human cells with synthetic modified mRNA, Cell Stem Cell, 7, 618–630.
Snead, N. M., and Rossi, J. J. (2012) RNA interference trigger variants: getting the most out of RNA for RNA interference-based therapeutics, Nucleic Acid Ther., 22, 139–146.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © J. A. Filippova, D. V. Semenov, E. S. Juravlev, A. B. Komissarov, V. A. Richter, G. A. Stepanov, 2017, published in Biokhimiya, 2017, Vol. 82, No. 11, pp. 1557–1576.
Rights and permissions
About this article
Cite this article
Filippova, J.A., Semenov, D.V., Juravlev, E.S. et al. Modern approaches for identification of modified nucleotides in RNA. Biochemistry Moscow 82, 1217–1233 (2017). https://doi.org/10.1134/S0006297917110013
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297917110013