-
PDF
- Split View
-
Views
-
Cite
Cite
Cecilia Rodriguez-Furlan, Elena A. Minina, Glenn R. Hicks, Remove, Recycle, Degrade: Regulating Plasma Membrane Protein Accumulation, The Plant Cell, Volume 31, Issue 12, December 2019, Pages 2833–2854, https://doi.org/10.1105/tpc.19.00433
Close - Share Icon Share
Abstract
Interactions between plant cells and the environment rely on modulation of protein receptors, transporters, channels, and lipids at the plasma membrane (PM) to facilitate intercellular communication, nutrient uptake, environmental sensing, and directional growth. These functions are fine-tuned by cellular pathways maintaining or reducing particular proteins at the PM. Proteins are endocytosed, and their fate is decided between recycling and degradation to modulate localization, abundance, and activity. Selective autophagy is another pathway regulating PM protein accumulation in response to specific conditions or developmental signals. The mechanisms regulating recycling, degradation, and autophagy have been studied extensively, yet we are just now addressing their regulation and coordination. Here, we (1) provide context concerning regulation of protein accumulation, recycling, or degradation by overviewing endomembrane trafficking; (2) discuss pathways regulating recycling and degradation in terms of cellular roles and cargoes; (3) review plant selective autophagy and its physiological significance; (4) focus on two decision-making mechanisms: regulation of recycling versus degradation of PM proteins and coordination between autophagy and vacuolar degradation; and (5) identify future challenges.
INTRODUCTION
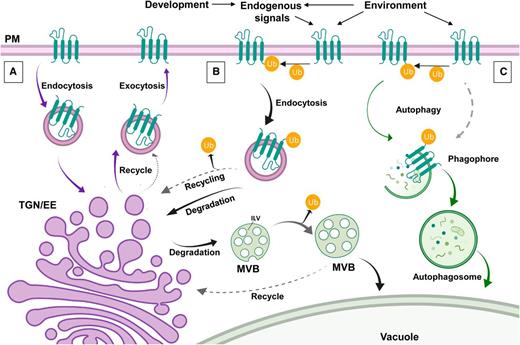
Approximately 25% of nuclear genes in Arabidopsis (Arabidopsis thaliana) are predicted to encode membrane proteins (Schwacke et al., 2003). Many are located at the plasma membrane (PM) and facilitate an array of cellular functions including sensing of environmental and hormonal signals, nutrient transport, cell–cell communication, establishment and maintenance of cell polarity, regulation of division, and cell wall anchoring. The distribution of PM proteins is actively regulated by removal, recycling, and/or degradation (Figure 1; Table1). PM protein removal occurs via clathrin-dependent or clathrin-independent endocytosis (for review, see Fan et al., 2015). Internalized proteins are sorted toward the vacuole (plant lytic organelle) for degradation or are delivered back to the PM through recycling (Figure 1A; Table1). Internalized vesicles carrying PM proteins first reach the trans-Golgi network (TGN)/early endosome (EE). In plants, the TGN and EE converge in a single network directing both synthetic and endocytic trafficking (for review, see Rosquete et al., 2018). Proteins within the network are recycled back to the PM or are sorted into endosomal organelles that mature along the endocytic pathway, eventually fusing with the lytic vacuole (Figure 1B).
PM Proteins: To Degrade or Not to Degrade?
(A) Many PM proteins are constitutively endocytosed to maintain an active pool of proteins. Once they are endocytosed, they travel to the TGN/EE were they are recycled back to the PM by exocytosis.
(B) Environmental or endogenous signals can trigger ubiquitination (Ub) of proteins that are then endocytosed by clathrin-dependent or -independent pathways. Proteins can be directed for degradation or deubiquitinated and returned to the recycling pathways (broken arrows). Proteins that continue through the degradation pathway are sorted at the TGN where LEs emerge. LEs mature by forming ILVs and become MVBs. At MVBs, proteins can be sent to the TGN/EE for recycling and saved from degradation. Alternatively, MBVs carrying proteins for degradation will fuse with the plant lytic compartment, the vacuole.
(C) Under stress conditions, proteins at the PM can be degraded by autophagy. Autophagy can be nonselective or selective. During selective autophagy, the cargo to be internalized is either ubiquitin dependent (solid arrow) or ubiquitin independently tagged (broken arrow). Material selectively internalized is sequestered in a phagophore that matures into an autophagosome that will also fuse with the vacuole for degradation.
Examples of PM Protein Dynamics
| PM Protein . | Recycling . | Degradation . | Notes . | References . |
|---|---|---|---|---|
| Auxin facilitator | ||||
| PIN2 (PIN-FORMED 2) | PM accumulation maintained by clathrin-mediated endocytosis and recycling | Degradation triggered by the covalent addition of ubiquitin | Rapid re-localization in response to different stimuli allows to re-direct auxin flux, e.g., gravity | (Kleine‐Vehn et al., 2011; Leitner et al., 2012; Baster et al., 2013) |
| ABCB19 (ATP-binding cassette subfamily B19) | Stable at the PM, not observed to be recycled by endocytosis | ND | ABCB19 interacts with PIN1 stabilizing it at PM microdomains, enhancing auxin transport activity | (Blakeslee et al., 2007; Rojas-Pierce et al., 2007; Titapiwatanakun et al., 2009) |
| Leu-rich repeat receptor-like kinase | ||||
| BRI1 (BRASSINOSTEROID INSENSITIVE1) | Ligand-independent recycling to maintain the PM protein pool | Upon brassinosteroid binding is eventually ubiquitinated and degraded at the vacuole | BRI1 degradation modulates the brassinosteroid response | (Geldner et al., 2003; Russinova et al., 2004; Geldner et al., 2007; Irani et al., 2012; Di Rubbo et al., 2013; Zhou et al., 2018) |
| FLS2 (FLAGELLIN SENSING2) | Continuously endocytosed and recycled in a ligand-independent manner | Sent to the vacuole for degradation upon flagellin binding | FLS2 recognizes the bacterial protein flagellin (flg22), setting on plant defense responses | (Li et al., 2002; Chinchilla et al., 2006; Schulze et al., 2010; Lu et al., 2011) |
| CVL1 (CLAVATA1) | ND | Upon CLV3 peptide sensing is degraded | Active CLV1 degradation attenuates the response of the stem cell population | (Rojo et al., 2002; Nimchuk et al., 2011) |
| Nutrient transporter | ||||
| PHT1 (PHOSPHATE TRANSPORTER1) | Recycles at the PM during phosphate sufficient conditions or starvation | Ubiquitinated and degraded when phosphate is supplied to a starving plant | PHT1 is ubiquitinated by the E3 ubiquitin ligase NITROGEN LIMITATION ADAPTATION (NLA) | (Bayle et al., 2011; Lin et al., 2013) |
| BOR1 (BORON TRANSPORTER1) | Polarization maintained by clathrin-mediated endocytosis and recycling | At high concentrations of boron is ubiquitinated and sent to the vacuole for degradation | Boric acid over-accumulation is cytotoxic. | (Takano et al., 2002; Takano et al., 2005; Takano et al., 2010; Wang et al., 2017b; Yoshinari et al., 2019) |
| IRT1 (IRON-REGULATED TRANSPORTER1) | Pool of polarized proteins maintained by endocytosis and protein recycling | Ubiquitinated and degraded by low iron levels and excess of Mn2+, Zn2+, or Co2+ | Mn2+, Zn2+, or Co2+ can be cytotoxic when in excess | (Vert et al., 2002; Barberon et al., 2014; Dubeaux et al., 2018) |
| Water channel | ||||
| PIP2;1/PIP2;7 (PM INTRINSIC PROTEIN 2.1 and 2.7) | Recycled by a Flotillin1 pathway, independent of clathrin | Selectively degradated by autophagy upon ABA treatment | During salt stress, endocytosed, but not degraded, impacting root hydraulic conductivity | (Li et al., 2011; Hachez et al., 2014b; Pou et al., 2016; Junková et al., 2018) |
| PM Protein . | Recycling . | Degradation . | Notes . | References . |
|---|---|---|---|---|
| Auxin facilitator | ||||
| PIN2 (PIN-FORMED 2) | PM accumulation maintained by clathrin-mediated endocytosis and recycling | Degradation triggered by the covalent addition of ubiquitin | Rapid re-localization in response to different stimuli allows to re-direct auxin flux, e.g., gravity | (Kleine‐Vehn et al., 2011; Leitner et al., 2012; Baster et al., 2013) |
| ABCB19 (ATP-binding cassette subfamily B19) | Stable at the PM, not observed to be recycled by endocytosis | ND | ABCB19 interacts with PIN1 stabilizing it at PM microdomains, enhancing auxin transport activity | (Blakeslee et al., 2007; Rojas-Pierce et al., 2007; Titapiwatanakun et al., 2009) |
| Leu-rich repeat receptor-like kinase | ||||
| BRI1 (BRASSINOSTEROID INSENSITIVE1) | Ligand-independent recycling to maintain the PM protein pool | Upon brassinosteroid binding is eventually ubiquitinated and degraded at the vacuole | BRI1 degradation modulates the brassinosteroid response | (Geldner et al., 2003; Russinova et al., 2004; Geldner et al., 2007; Irani et al., 2012; Di Rubbo et al., 2013; Zhou et al., 2018) |
| FLS2 (FLAGELLIN SENSING2) | Continuously endocytosed and recycled in a ligand-independent manner | Sent to the vacuole for degradation upon flagellin binding | FLS2 recognizes the bacterial protein flagellin (flg22), setting on plant defense responses | (Li et al., 2002; Chinchilla et al., 2006; Schulze et al., 2010; Lu et al., 2011) |
| CVL1 (CLAVATA1) | ND | Upon CLV3 peptide sensing is degraded | Active CLV1 degradation attenuates the response of the stem cell population | (Rojo et al., 2002; Nimchuk et al., 2011) |
| Nutrient transporter | ||||
| PHT1 (PHOSPHATE TRANSPORTER1) | Recycles at the PM during phosphate sufficient conditions or starvation | Ubiquitinated and degraded when phosphate is supplied to a starving plant | PHT1 is ubiquitinated by the E3 ubiquitin ligase NITROGEN LIMITATION ADAPTATION (NLA) | (Bayle et al., 2011; Lin et al., 2013) |
| BOR1 (BORON TRANSPORTER1) | Polarization maintained by clathrin-mediated endocytosis and recycling | At high concentrations of boron is ubiquitinated and sent to the vacuole for degradation | Boric acid over-accumulation is cytotoxic. | (Takano et al., 2002; Takano et al., 2005; Takano et al., 2010; Wang et al., 2017b; Yoshinari et al., 2019) |
| IRT1 (IRON-REGULATED TRANSPORTER1) | Pool of polarized proteins maintained by endocytosis and protein recycling | Ubiquitinated and degraded by low iron levels and excess of Mn2+, Zn2+, or Co2+ | Mn2+, Zn2+, or Co2+ can be cytotoxic when in excess | (Vert et al., 2002; Barberon et al., 2014; Dubeaux et al., 2018) |
| Water channel | ||||
| PIP2;1/PIP2;7 (PM INTRINSIC PROTEIN 2.1 and 2.7) | Recycled by a Flotillin1 pathway, independent of clathrin | Selectively degradated by autophagy upon ABA treatment | During salt stress, endocytosed, but not degraded, impacting root hydraulic conductivity | (Li et al., 2011; Hachez et al., 2014b; Pou et al., 2016; Junková et al., 2018) |
ND, no data.
| PM Protein . | Recycling . | Degradation . | Notes . | References . |
|---|---|---|---|---|
| Auxin facilitator | ||||
| PIN2 (PIN-FORMED 2) | PM accumulation maintained by clathrin-mediated endocytosis and recycling | Degradation triggered by the covalent addition of ubiquitin | Rapid re-localization in response to different stimuli allows to re-direct auxin flux, e.g., gravity | (Kleine‐Vehn et al., 2011; Leitner et al., 2012; Baster et al., 2013) |
| ABCB19 (ATP-binding cassette subfamily B19) | Stable at the PM, not observed to be recycled by endocytosis | ND | ABCB19 interacts with PIN1 stabilizing it at PM microdomains, enhancing auxin transport activity | (Blakeslee et al., 2007; Rojas-Pierce et al., 2007; Titapiwatanakun et al., 2009) |
| Leu-rich repeat receptor-like kinase | ||||
| BRI1 (BRASSINOSTEROID INSENSITIVE1) | Ligand-independent recycling to maintain the PM protein pool | Upon brassinosteroid binding is eventually ubiquitinated and degraded at the vacuole | BRI1 degradation modulates the brassinosteroid response | (Geldner et al., 2003; Russinova et al., 2004; Geldner et al., 2007; Irani et al., 2012; Di Rubbo et al., 2013; Zhou et al., 2018) |
| FLS2 (FLAGELLIN SENSING2) | Continuously endocytosed and recycled in a ligand-independent manner | Sent to the vacuole for degradation upon flagellin binding | FLS2 recognizes the bacterial protein flagellin (flg22), setting on plant defense responses | (Li et al., 2002; Chinchilla et al., 2006; Schulze et al., 2010; Lu et al., 2011) |
| CVL1 (CLAVATA1) | ND | Upon CLV3 peptide sensing is degraded | Active CLV1 degradation attenuates the response of the stem cell population | (Rojo et al., 2002; Nimchuk et al., 2011) |
| Nutrient transporter | ||||
| PHT1 (PHOSPHATE TRANSPORTER1) | Recycles at the PM during phosphate sufficient conditions or starvation | Ubiquitinated and degraded when phosphate is supplied to a starving plant | PHT1 is ubiquitinated by the E3 ubiquitin ligase NITROGEN LIMITATION ADAPTATION (NLA) | (Bayle et al., 2011; Lin et al., 2013) |
| BOR1 (BORON TRANSPORTER1) | Polarization maintained by clathrin-mediated endocytosis and recycling | At high concentrations of boron is ubiquitinated and sent to the vacuole for degradation | Boric acid over-accumulation is cytotoxic. | (Takano et al., 2002; Takano et al., 2005; Takano et al., 2010; Wang et al., 2017b; Yoshinari et al., 2019) |
| IRT1 (IRON-REGULATED TRANSPORTER1) | Pool of polarized proteins maintained by endocytosis and protein recycling | Ubiquitinated and degraded by low iron levels and excess of Mn2+, Zn2+, or Co2+ | Mn2+, Zn2+, or Co2+ can be cytotoxic when in excess | (Vert et al., 2002; Barberon et al., 2014; Dubeaux et al., 2018) |
| Water channel | ||||
| PIP2;1/PIP2;7 (PM INTRINSIC PROTEIN 2.1 and 2.7) | Recycled by a Flotillin1 pathway, independent of clathrin | Selectively degradated by autophagy upon ABA treatment | During salt stress, endocytosed, but not degraded, impacting root hydraulic conductivity | (Li et al., 2011; Hachez et al., 2014b; Pou et al., 2016; Junková et al., 2018) |
| PM Protein . | Recycling . | Degradation . | Notes . | References . |
|---|---|---|---|---|
| Auxin facilitator | ||||
| PIN2 (PIN-FORMED 2) | PM accumulation maintained by clathrin-mediated endocytosis and recycling | Degradation triggered by the covalent addition of ubiquitin | Rapid re-localization in response to different stimuli allows to re-direct auxin flux, e.g., gravity | (Kleine‐Vehn et al., 2011; Leitner et al., 2012; Baster et al., 2013) |
| ABCB19 (ATP-binding cassette subfamily B19) | Stable at the PM, not observed to be recycled by endocytosis | ND | ABCB19 interacts with PIN1 stabilizing it at PM microdomains, enhancing auxin transport activity | (Blakeslee et al., 2007; Rojas-Pierce et al., 2007; Titapiwatanakun et al., 2009) |
| Leu-rich repeat receptor-like kinase | ||||
| BRI1 (BRASSINOSTEROID INSENSITIVE1) | Ligand-independent recycling to maintain the PM protein pool | Upon brassinosteroid binding is eventually ubiquitinated and degraded at the vacuole | BRI1 degradation modulates the brassinosteroid response | (Geldner et al., 2003; Russinova et al., 2004; Geldner et al., 2007; Irani et al., 2012; Di Rubbo et al., 2013; Zhou et al., 2018) |
| FLS2 (FLAGELLIN SENSING2) | Continuously endocytosed and recycled in a ligand-independent manner | Sent to the vacuole for degradation upon flagellin binding | FLS2 recognizes the bacterial protein flagellin (flg22), setting on plant defense responses | (Li et al., 2002; Chinchilla et al., 2006; Schulze et al., 2010; Lu et al., 2011) |
| CVL1 (CLAVATA1) | ND | Upon CLV3 peptide sensing is degraded | Active CLV1 degradation attenuates the response of the stem cell population | (Rojo et al., 2002; Nimchuk et al., 2011) |
| Nutrient transporter | ||||
| PHT1 (PHOSPHATE TRANSPORTER1) | Recycles at the PM during phosphate sufficient conditions or starvation | Ubiquitinated and degraded when phosphate is supplied to a starving plant | PHT1 is ubiquitinated by the E3 ubiquitin ligase NITROGEN LIMITATION ADAPTATION (NLA) | (Bayle et al., 2011; Lin et al., 2013) |
| BOR1 (BORON TRANSPORTER1) | Polarization maintained by clathrin-mediated endocytosis and recycling | At high concentrations of boron is ubiquitinated and sent to the vacuole for degradation | Boric acid over-accumulation is cytotoxic. | (Takano et al., 2002; Takano et al., 2005; Takano et al., 2010; Wang et al., 2017b; Yoshinari et al., 2019) |
| IRT1 (IRON-REGULATED TRANSPORTER1) | Pool of polarized proteins maintained by endocytosis and protein recycling | Ubiquitinated and degraded by low iron levels and excess of Mn2+, Zn2+, or Co2+ | Mn2+, Zn2+, or Co2+ can be cytotoxic when in excess | (Vert et al., 2002; Barberon et al., 2014; Dubeaux et al., 2018) |
| Water channel | ||||
| PIP2;1/PIP2;7 (PM INTRINSIC PROTEIN 2.1 and 2.7) | Recycled by a Flotillin1 pathway, independent of clathrin | Selectively degradated by autophagy upon ABA treatment | During salt stress, endocytosed, but not degraded, impacting root hydraulic conductivity | (Li et al., 2011; Hachez et al., 2014b; Pou et al., 2016; Junková et al., 2018) |
ND, no data.
Autophagy also participates in selective degradation of PM proteins, for example, under abiotic stress conditions (Figure 1C). Through the coordination of endocytosis, recycling, and/or degradation, the steady state of PM proteins is established, maintained, and modified in response to physiological and environmental changes (Luschnig and Vert, 2014). Here, we explore the molecular events that balance decisions between recycling versus endocytic and autophagic degradation and how and these processes are regulated.
Because of space limitations, we focus on nondividing cells. The regulation of membrane trafficking in dividing cells is a complex topic with significant literature worthy of separate reviews (for recent reviews, see Robatzek, 2014; Boruc and Van Damme, 2015; Müller and Jürgens, 2016; Frémont and Echard, 2018; Livanos and Müller, 2019).
ENDOSOME MATURATION AND TRAFFICKING TO THE VACUOLE
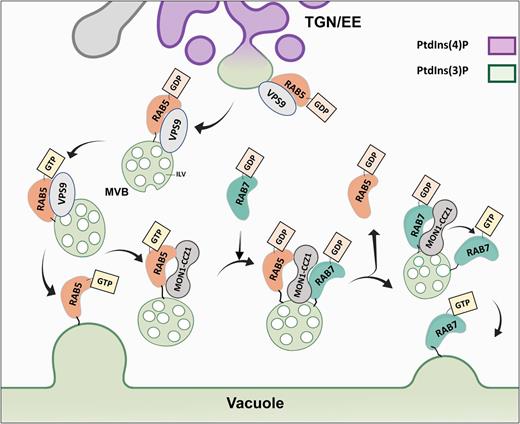
PM proteins are endocytosed and sorted into TGN/EE compartments that mature on their way to the vacuole and are recognized by three characteristics: associated phosphoinositides, RAB GTPase (hereafter referred to as RAB) conversion, and formation of intraluminal vesicles (ILVs; Figure 2).
Endosome Maturation: RAB Conversion and Phosphoinositols.
Endocytosed vesicles enriched in PtdIns(4)P arrive at the TGN/EE. Subdomains of the TGN/EE become decorated by PtdIns(3)P and RAB5-like proteins. RAB5-like endosomes detach from the TGN/EE (these endosomes are MVBs as they are described to have ILVs). RAB5-like endosomes are also enriched in PtdIns(3)P. VPS9 acts as a GEF exchanging GDP for GTP, activating the RAB5-like GTPases. RAB5-GTP then triggers the fusion of MVBs with the vacuole. Alternatively, RAB5-GTP can bind the complex MON1-CCZ1 that catalyzes GTP hydrolysis and RAB5-GDP exchange for RAB7-GDP. The MON1-CCZ1 complex also acts as a GEF for RAB7-like exchanging GDP for GTP. RAB7-like activated (GTP bound) can now trigger the fusion of the MVB to the vacuole. RAB5 and RAB7-like proteins are anchored to the membrane via a prenylated tail (black line).
Phosphatidylinositols (PtdIns) are negatively charged lipids formed by phosphorylation of the inositol ring and are a minority of acidic phospholipids. At the cell membranes, specific kinases and phosphatases dynamically interconvert PtdIns. PtdIns are recognized by proteins involved in trafficking through protein lipid binding domains such as PHOX (PX), PLECKSTRIN-HOMOLOGY (PH), or Fab1-YOTB-Vac1-EEA1 (FYVE; Simon et al., 2014). PtdIns have restricted localizations. For example, phosphatidylinositol 4-phosphate [PtdIns(4)P] and, in lower quantities, PtdIns(4,5)P2 accumulate at the PM and TGN/EE (for review, see Noack and Jaillais, 2017). After endocytosis, endosomes gradually lose PtdIns(4)P, becoming enriched in PtdIns(3)P found at late endosomes (LEs), autophagosomes, and vacuoles (Figure 2).
RABs are another indicator of endosome maturity. In Arabidopsis, two RAB5-like groups represent distinct endosome populations involved in vacuolar trafficking (Ueda et al., 2001; Sohn et al., 2003; Kotzer et al., 2004; Ebine et al., 2011; Bottanelli et al., 2012): RAB5-like canonical (RABF2B/ARA7, RABF2a/RHA1) and plant-specific noncanonical (ARA6/RABF1) endosomes. Both groups are prenylated at C-terminal Cys residues by a RAB geranylgeranyltransferase (Shi et al., 2016). The prenylated proteins are delivered to endosomes and then activated by a guanine nucleotide exchange factor (GEF), releasing GDP and favoring loading of more abundant GTP (Traut, 1994). GTP stabilizes RAB conformation, allowing the recruitment of specific effector proteins to the membranes (Vetter and Wittinghofer, 2001). RABs are inactivated by hydrolysis of GTP via GTPase-activating proteins. A GDP dissociation inhibitor then releases RAB-GDP from membranes into the cytosol until reactivated by GEFs (Zárský et al., 1997; Ueda et al., 1998). Both RAB5 groups share a common GEF protein, VACUOLAR PROTEIN SORTING 9a (VPS9a; Goh et al., 2007). VPS9a is recruited to the endosome by a PLANT-UNIQUE RAB5 EFFECTOR2 (PUF2; Ito et al., 2018). PUF2 also recruits inactive canonical RAB5-GDP at the endosome, enhancing its activation. Canonical RAB5, RABF2b/ARA7, becomes anchored to the membrane at a subdomain of the TGN/EE (Singh et al., 2014). The subdomain becomes decorated by PtdIns(3)P and upon scission is considered LE. The complex MON1(SAND1)-CZZ1 can interact with RAB5 endosomes and recruit a GDP-bound RAB7-like protein, RABG3f. MON1(SAND1)-CZZ1 acts as a GEF, activating RABG3f and as an effector (GTPase-activating protein) of RAB5 (Cui et al., 2014; Singh et al., 2014). The exchange of RAB5 to RAB7 occurs late in endosomal trafficking but before tonoplast fusion (Figure 2). Thus, RAB7 is an indicator of LEs.
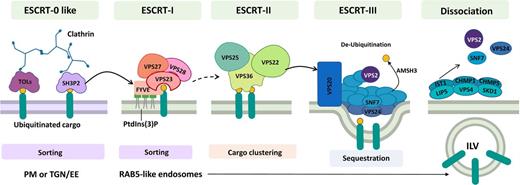
As endosomes mature, they form inwardly budding ILVs that cluster ubiquitinated proteins and form multivesicular bodies (MVBs) also termed prevacuolar compartments (Figures 1 and 2; Barberon et al., 2011; Kasai et al., 2011; Herberth et al., 2012; Leitner et al., 2012; Scheuring et al., 2012). A series of multimeric complexes, Endosomal Sorting Complex Required for Transport (ESCRT), controls sorting of ubiquitinated proteins into ILVs. In metazoans and fungi, ubiquitinated protein recognition at the PM and cargo concentration are performed by ESCRT-0 subunits that then recruit the downstream machinery (Mayers et al., 2011). Three other multi-subunit ESCRT complexes, ESCRT‐I, ESCRT-II, and ESCRT-III, sequentially induce membrane budding processes and form endosomal ILVs (Gao et al., 2017). Plants contain orthologs of ESCRT-I, ESCRT-II, and ESCRT-III subunits (Richardson et al., 2011). Plant ESCRT-0 orthologs have not been identified. Instead, TOM1-LIKE (TOL) protein complexes serve this function (Figure 3; Korbei et al., 2013). The Arabidopsis genome encodes nine TOLs (TOL1 to TOL9) ubiquitously expressed with the exception of TOL8, which is mostly expressed in siliques and flowers (Moulinier-Anzola et al., 2014). However, distinct subcellular distributions suggest functional compartmentalization. For example, TOL6 is associated with PM and EE where it binds ubiquitin moieties of cargo proteins, whereas TOL5 is enriched at MVBs (Korbei et al., 2013; Yoshinari et al., 2018).
Endosome Maturation: An ESCRT Point of View.
ESCRTs are multi-subunit complexes that act sequentially through the endocytic pathway, directing ubiquitinated proteins into ILVs at LEs. Plants do not have a conventional ESCRT-0 complex; instead, TOL and SH3P2 proteins appear to fulfill this function. Sorting of ubiquitinated cargo proteins is proposed to start at the PM or right after endocytosis at the TGN/EE. Both TOL and SH3P2 directly interact with ubiquitin and clathrin. SH3P2 then can interacts with the ESCRT-I component FYVE/FREE1 that is anchored at the TGN/EE by PtdIns(3)P. FVYEE interacts with another ESCRT-I component, VPS23, and together they bind to ubiquitinated cargo continuing the sorting process. VPS23 binds to VPS27 and VPS28 assembling the entire ESCRT-I complex. The transition between ESCRT-I and ESCRT-II is still unclear in plants (broken arrow). However, the ESCRT-II subunit VPS36 binds ubiquitinated cargo, and the rest of the complex, namely, VPS22 and VPS25. ESCRT-II acts by clustering ubiquitinated cargo. The ESCRT-II VPS22 subunit then recruits the ESCRT-III VPS20 subunit at sites of ubiquitin cargo enrichment. VPS20 coordinates polymerization of SNF7/VPS24 into highly curved rings that sequester proteins forming ILV. AMSH3 protein acts by de-ubiquitinating cargo. Finally, the accessory proteins CHMP1, CHMP5 and IST1 regulate VPS4/SKD1-LIP5 ATPase complex activity to trigger the disassembly of the ESCRT complex. RAB5-like endosomes already present in ILV become an indicator of LE maturity.
Proteins sorted for degradation by ESCRT-0–like complexes are recognized by ESCRT-I that in Arabidopsis is formed by subunits VPS23A, VPS23B, VPS37, VPS28, and FYVE domain protein required for endosomal sorting1 (FREEI/FYVE; Figure 3). SRC-HOMOLOGY-3 DOMAIN-CONTAINING PROTEIN2 (SH3P2), another ESCRT-0–like protein, binds ubiquitinated proteins and clathrin heavy chains, colocalizes with clathrin-coated vesicles, and interacts with ESCRT-I components VPS23A and FREEI/FVYE (Figure 3; Belda-Palazon et al., 2016; Nagel et al., 2017). Whether TOL proteins directly or indirectly recruit ESCRT-I complexes or whether SH3P2 acts together with TOL proteins, or as an alternative ESCRT-0 subunit, remains to be determined. ESCRT-I VPS23A binds ubiquitinated proteins at RAB5-LEs and recruits VPS37 and VPS28 (Spitzer et al., 2006). Additionally, the plant-specific subunit FREEI/FVYE is recruited to RAB5 endosomes through PtdIns(3)P binding. Once membrane associated, RAB5 specifically interacts with VPS23A, VPS23B, and ubiquitinated proteins (Gao et al., 2014; Kolb et al., 2015). Both FREEI/FYVE and VPS23A are involved in ILV formation, resulting in MVBs (Spitzer et al., 2006; Gao et al., 2014). The ESCRT-I complex colocalizes with ubiquitinated FLS2, PIN2, and IRT1 cargoes (Barberon et al., 2014; Belda-Palazon et al., 2016; Yu et al., 2016).
ESCRT-I probably delivers cargo to the ESCRT-II complex that in plants is formed by VPS22, VPS25, and VPS36 (Figure 3). VPS36 binds ubiquitin and interacts with VPS25 and VPS22 (Wang et al., 2017a). VPS25 and VPS36 are detected in proximity to the PM, EE/TGN, and in RAB5 endosomes (Richardson et al., 2011; Wang et al., 2017a). The VPS22 subunit is located at the EE/TGN, indicating a probable transient binding compared with other subunits (Scheuring et al., 2012). A VPS36 mutant results in defects in PIN1, PIN2, AUX1, and PIP1 trafficking to the vacuole (Wang et al., 2017a).
ESCRTI-III core subunits lack ubiquitin binding domains. In yeast, ESCRT-II initiates assembly of ESCRT-III by binding the Vps20 subunit at sites enriched with ubiquitinated cargo, coordinating Sucrose non-fermenting7 (Snf7) polymerization into highly curved ESCRT-II/ESCRT-III supercomplexes (Figure 3; Fyfe et al., 2011; Henne et al., 2012). These complexes favor formation of highly curved rings that physically sequester transmembrane cargoes at distinct regions of the endosome surface prior to packaging into ILVs (Buono et al., 2017). Accordingly, Arabidopsis dominant negative overexpression ESCRT-III mutants shows reduced ILVs in the LEs, indicating a similar process is occurring in plant cells (Cai et al., 2014). ESCRT-III complex (VPS20.1, SNF7.1, VPS24.1, and VPS2.1) disassembly requires accessory proteins (CHMP1, CHMP5, and IST1) and coordinated interactions with the VPS4/SKD1-LIP5 complex (Figure 3; Spitzer et al., 2009, 2015). The LIP5 cofactor and the flowering plant–specific positive regulator of SKD1 (PROS), which is involved in flowering, positively regulate the ATPase mechano-enzyme VPS4/SKD1. VPS4/SKD1 hydrolyzes ATP, dissociating the ESCRT-III complex from the endosomal membrane (Figure 3; Haas et al., 2007; Reyes et al., 2014; Buono et al., 2016, 2017). Endosomal maturation thus sorts proteins and prepares LEs for tonoplast fusion (Figures 2 and 3; Takemoto et al., 2018).
THE FINAL STEP: REGULATION OF VACUOLE FUSION
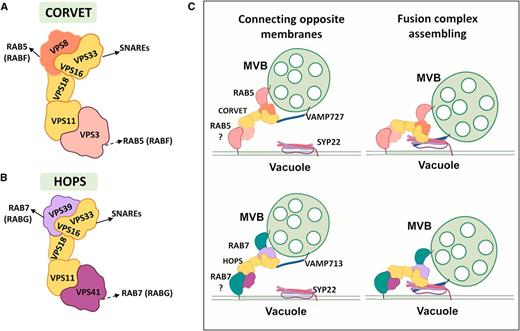
The final step in membrane protein degradation is MVB-vacuole fusion. For fusion two membranes must be in close proximity and displace the hydration layer (for review, see Kozlov and Chernomordik, 2015). Fusion is facilitated by SOLUBLE N-ETHYLMALEIMIDE–SENSITIVE FACTOR ATTACHMENT RECEPTOR (SNARE) proteins found on both membranes (Figure 4 for review, see Bonifacino and Glick, 2004; El Kasmi et al., 2013). Based on a Gln (Q) or Arg (R) residue in the SNARE interaction domain, SNAREs can be classified as Q-SNAREs or R-SNAREs (for review, see Lipka et al., 2007). Q-SNAREs are divided into three subgroups: syntaxins of plants (SYP)/Qa-SNAREs, Qb-SNAREs, and Qc-SNAREs (Sanderfoot et al., 2000). Functional SNARE complexes consist of a vesicle-localized R-SNARE or vesicle-associated membrane protein (VAMP) and target membrane localized Q-SNAREs. Regulation of endomembrane fusions also requires a conserved machinery that consists of RABs and their interacting tethering complexes, which mediate initial contact. Two tethering complexes are related to MVB-tonoplast fusion in Arabidopsis: homotypic fusion and protein sorting (HOPS), and class C core vacuole/endosome tethering (CORVET; for review, see Balderhaar and Ungermann, 2013). These complexes share a common core consisting of VPS11, VPS16, VPS18, and VPS33 (Figures 4A and 4B). Likewise, both complexes are distinguished by two specific subunits: VPS3 and VPS8 (CORVET) and VPS39 and VPS41 (HOPS). In yeast, GTP-activated RAB7-like (Ypt7) proteins interact with both Vps41 and Vps39 (Lürick et al., 2017). HOPS forms a bridge between two RAB7s: one at the endosome and the other at the tonoplast, bringing the membranes in close proximity. Vps33 and Vps16 then interact with the SNAREs, triggering fusion. Similarly, it is proposed that CORVET subunits Vps3 and Vps8 bind two RAB5-like proteins (Vps21; Lürick et al., 2017). Molecular architectures of the plant complexes are undetermined; however, it is described that the CORVET complex interacts with RAB5-like proteins (RABF family) in LEs, vesicle R-SNARE VAMP727, and tonoplast Qa-SNARE SYP22 (Figure 4C; Takemoto et al., 2018). HOPS interacts with RAB7-like proteins (RABG family), the R-SNARE VAMP713, and Qa-SNARE SYP22 (Figure 4C; Brillada et al., 2018; Takemoto et al., 2018). Activation of RAB5 by its GEF (VPS9) is required to assemble CORVET (Takemoto et al., 2018). Similarly, activation of RAB7 by its GEF, MON1(SAND1)-CZZ1, is necessary to assemble HOPS. Dexamethasone-inducible artificial micro-RNA knockdown mutants for HOPS- or CORVET-specific subunits reveal CORVET deficiency does not influence vacuolar phenotype, whereas lack of HOPS leads to round fragmented vacuoles, suggesting that only HOPS acts in biogenesis (Brillada et al., 2018; Takemoto et al., 2018). Additionally, HOPS and CORVET knockdown mutants are defective in vacuolar localization of different cargo proteins (Takemoto et al., 2018; Zhang et al., 2019), indicating that CORVET and HOPS regulate different fusion pathways.
Tether to Fuse: LEs to Vacuole.
(A) to (C) Fusion of MVBs with the vacuole is facilitated by two tethering complexes CORVET (A) and HOPS (B). Core subunits VPS33, VPS16, VPS18, and VPS11 are part of both complexes. CORVET has two specific subunits, VPS8 and VPS3. In yeast, both VPS8 and VPS3 bind RAB5-like GTPases (arrows). In plants, VPS8 has proven to bind the plant RABF family members (solid arrow), while VPS3 remains to be tested (broken arrow). The HOPS complex also has two specific subunits, VPS41 and VPS39. In yeast, VP39 binds to a RAB7-like protein located at endosomes and VPS41 to another RAB7-like protein located at the vacuole (arrows). In plants, VPS39 has been characterized to bind RABG family (solid arrow), while VPS41 association to RAB7 remains to be tested (broken arrow). VPS33 from both subunits interacts with SNARE proteins. CORVET and HOPS act at different endosome populations connecting opposite membranes. CORVET interacts with the R-SNARE VAMP727 and with the Qa-SNARE SYP22, favoring the formation of a four-subunit oligomeric SNARE complex allowing fusion (see [C], top). Similarly, HOPS interacts with VAMP713 and VPS22, allowing formation of the fusion complex (see [C], bottom).
Overall, our knowledge of vacuole fusion indicates that two sets of RABs, together with multiple tethering complexes and distinct SNARE combinations, facilitate vacuole fusion of subpopulations of MVBs (Figure 4C). Accumulating evidence supports the idea that endosome–vacuole fusion shares common mechanisms across kingdoms with specialized roles in plants.
ESCAPING DEGRADATION: PROTEIN RECYCLING PATHWAYS
Not all endocytosed membrane proteins are packaged into MVBs and degraded at the vacuole. Proteins can escape degradation utilizing cargo recycling pathways from endosomes back to the PM (Figures 1 and 5).
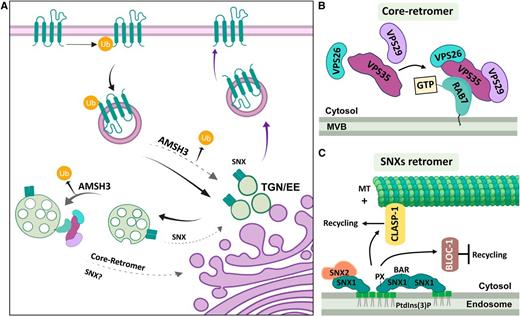
Recycling Retromers.
(A) SNXs are believed to act by retrieving proteins for recycling at the TGN/EE and at RAB5-like endosomes. It is still debated whether SNX1 also acts at RAB7-like endosomes. The core retromer is anchored to RAB7-like endosomes and acts by recycling proteins before fusion with the vacuole. Proteins selected for retrieval by endosomes can be de-ubiquitinated by AMSH3. This de-ubiquitinating enzyme acts at the TGN/EE and at MBVs before the assembly of the ESCRT-III complex. Proteins are identified by colors as in (B) and (C).
(B) The core retromer is anchored to MBVs by direct interaction of subunit VPS35 with RAB7-like proteins (RABG3f). The rest of the complex is then assembled by recruiting VPS26 and VPS29.
(C) SNX1 can bind PtdIns(3)P by its PX domain, which anchors the protein to endosomal membranes (TGN/EE; MVBs). SNX1 can form homodimers or heterodimers when interacting with SNX2 through their BAR domains. SNX1 can also interact with CLASP-1, a plus-end microtubule (MT +) binding protein directing endosomes to the cell edges. SNX1 alternatively can interact with BLOC-1, which inhibits protein recycling. AMSH3, ASSOCIATED MOLECULE with the SH3 DOMAIN of STAM 3; BAR, Bin/Amphiphysin/Rvs domain; CLASP, MICROTUBULE-ASSOCIATED PROTEIN CYTOPLASMIC LINKER-ASSOCIATED PROTEINS; MT+, plus end microtubules; MVB, Multivesicular Bodies; PtdIns(3)P, phosphatidylinositol 3-phosphate; PX, PHOX phospholipid binding domain; SNX, SORTIN NEXIN; TGN/EE, Trans Golgi Network/Early Endosomes; Ub, Ubiquitin; VPS, VACUOLAR PROTEIN SORTING.
Protein Sorting for Recycling
Protein retrieval is directed by the assembly of coat complexes that (1) recognize and concentrate specific cargo, (2) drive membrane remodeling, (3) elicit scission, and (4) direct retrograde transport. In yeast, these activities are regulated by the retromer complex core subunits Vps35, Vps29, and Vps26 and sorting nexins (SNXs; for review, see Simonetti and Cullen, 2018). SNX proteins associated with membrane phosphoinositols then recruit the rest of the core complex. In plants, the core retromer complex (VPS35A, VPS35B, VPS35C; VPS26A, VPS26B; and VPS29) is conserved (Jha et al., 2018) and assembles independently of SNX subunits (SNX1, SNX2a, and SNX2b; Zelazny et al., 2013b). The core retromer binds MVBs through interaction with RAB7-like RABG3f (Zelazny et al., 2013a, 2013b). SNXs are detected at the TGN/EE (Jaillais et al., 2006; Niemes et al., 2010a, 2010b; Stierhof et al., 2013; Ivanov et al., 2014; Wijdeven et al., 2016), RAB5-like endosomes (Pourcher et al., 2010; Heard et al., 2015), and RAB7-like endosomes (Kleine-Vehn et al., 2008; Yamazaki et al., 2008). The differential distribution suggests SNX proteins act earlier in the endocytic pathway than the core retromer (Figure 5A).
The triple mutant vps35a1b2c1, double mutant vps26a1b1, and single mutant vps29 show strong developmental phenotypes (Jaillais et al., 2007; Yamazaki et al., 2008; Zelazny et al., 2013b). A VPS35A mutant (pat3) exhibits intracellular accumulations of PIN1-GFP similar to vps29-3 (Jaillais et al., 2007; Nodzynski et al., 2013). Interestingly, all VPS35 isoforms interact with VPS26 and VPS29, suggesting multiple subcomplexes (Zelazny et al., 2013b). Unlike mutants of core retromer subunits, SNX mutants exhibit mild developmental phenotypes. They accumulate PIN1, PIN2, and IRT1 in the vacuole and have reduced accumulation at the PM, indicating that nonrecycled, internalized proteins are sorted for degradation (Pourcher et al., 2010; Ivanov et al., 2014). Thus, SNXs regulates recycling of several PM proteins distinct from the VPS26-VPS29-VPS35 core retromer.
SNX1 and SNX2b become membrane associated by interaction with PtdIns(3)P and PtdIns(3,5)P2 through phospholipid binding domains (PHOX and PX; Figure 5C; Hirano et al., 2015). These PtdIns reside at the intersection between TGN/EE and RAB5 endosomes and remain at RAB7 endosomes, which may explain the broad localization of SNXs (Poteryaev et al., 2010). SNX1 forms homodimers and heterodimers with SNX2a and SNX2b through their BIN/AMPHIPHYSIN/RVS (BAR) domain (Pourcher et al., 2010). In other organisms, SNXs acts in a multimeric complex directing protein retrieval (for review, see Kvainickas et al., 2017). In plants, SNX1 interacts with the microtubule-associated protein cytoplasmic linker-associated proteins (CLASPs) through PX and BAR domains (Ambrose et al., 2013). CLASP binds microtubules, stabilizing the plus end proximal to the PM (Ambrose et al., 2013; Lindeboom et al., 2019) and directing SNX1 endosomes/cargo (Figure 5C; Salanenka et al., 2018). clasp-1 mutants exhibit decreased endosomal SNX1 association and increased cytoplasmic signal, indicating a role in vesicle anchoring. Accordingly, clasp-1 displays enhanced PIN2 and BRI1 degradation at the vacuole (Ambrose et al., 2013; Ruan et al., 2018). Treatments with the microtubule-depolarizing drug oryzalin increase PIN2 degradation at the vacuole, indicating that cortical microtubules are necessary for recycling (Hirano et al., 2015). Therefore, SNX-CLASP directs proteins toward recycling by rescuing them from degradation. Interestingly, interaction of SNXs with the BIOGENESIS OF LYSOSOME-RELATED ORGANELLES COMPLEX-1 (BLOC-1) inhibits PIN1 and PIN2 recycling and favors degradation by an unknown mechanism (Figure 5C; Cui et al., 2010). Similarly, brassinosteroid treatment stimulates SNX-1/BLOC-1 interaction, decreasing BRI-1 recycling and favoring degradation (Ruan et al., 2018).
Increasing evidence indicates that SNX subcomplexes act, at least in part, in pathways other than core retromer. However, SNX1 colocalizes with retromer subunits VPS35A at MVBs and VPS29 at TGN/EE (Kleine-Vehn et al., 2008; Niemes et al., 2010b; Stierhof et al., 2013), and SNX1 and VPS29 interact genetically (Jaillais et al., 2007). Nevertheless, no direct interaction of core retromer and SNXs has been detected (Pourcher et al., 2010; Nodzynski et al., 2013; Zelazny et al., 2013a). Perhaps at RAB7 endosomes, SNXs and core retromer interact indirectly to retrieve proteins. The complexes may also perform different tasks at MVBs by interacting with and retrieving specific cargoes (Figure 5A). The latter suggests a regulatory role of RAB7 in mating SNX subcomplexes and core retromer, and RAB7 may selectively activate one or the other complex. Another exciting possibility that requires further research is that SNX is involved in fast or slow recycling depending on the associations and/or compartments where it is acting.
Exocytosis of Recycled Proteins
Proteins sorted for recycling at endosomes are secreted by exocytosis back to the PM. Exocytosis is coordinated by small GTPases, the exocyst vesicle tethering complex, and the pairing of cognate SNAREs in opposing membranes (Figure 6). In yeast and mammals, the exocyst is formed by eight subunits: Sec3, Sec5, Sec6, Sec8, Sec10, Sec15, Exo70, and Exo84 (Ahmed et al., 2018; Lepore et al., 2018). In plants, the core subunits SEC3, SEC5, SEC6, SEC8, and SEC10 are present as one or two gene copies. By contrast, SEC15 and EXO84 have two to eight genes, while the EXO70 subunits are encoded by a gene family (23 genes in Arabidopsis; Cvrčková et al., 2012). This expansion could indicate regulated secretion of specific cargoes (Žárský et al., 2013). For example, the EXO84B subunit regulates lateral secretion of the ATP BINDING CASSETTE TRANSPORTER PENETRATION 3 (PEN3) and the boric acid transporter NODULIN 26-LIKE INTRINSIC PROTEIN (NIP1;5; Figure 6B; Mao et al., 2016). EXO70A1 functions in the recycling of PIN2, PIN1, and BRI1, but not aquaporin PIP2a (Drdová et al., 2013; Zhang et al., 2016). Interestingly, the small molecule Endosidin2 (ES2) binds the N terminus of EXO70A1, interfering with PIN2-GFP secretion, which accumulates in RAB5-like (ARA7) compartments and is re-routed toward the vacuole (Zhang et al., 2016). Similarly, the exo70a-1 mutant accumulates PIN3-GFP at the vacuole (Janková Drdová et al., 2019). This indicates again that failure to recycle results in sorting for degradation. ES2 affects EXO70A1 localization at the PM, but not EXO84B localization, supporting their independent assembly and function in the complex. EXO70A1 interacts with PtdIns(4)P and PtdIns(4,5)P2 at the PM, showing protein enrichment at apical and basal domains of epidermal root cells (Figures 6A and C; Wu et al., 2017). Mutants defective in phosphoinositol synthesis show alterations in PIN trafficking and localization (Ischebeck et al., 2013; Tejos et al., 2014; Ugalde et al., 2016). EXO70A1 specifically recruits the protein PATELLIN3 (PATL3) to the PM (Figure 6C; Wu et al., 2017), where its role in exocytosis is unknown. However, quadruple mutants of PATL genes show defects in PIN1 trafficking and polarity (Tejos et al., 2017).
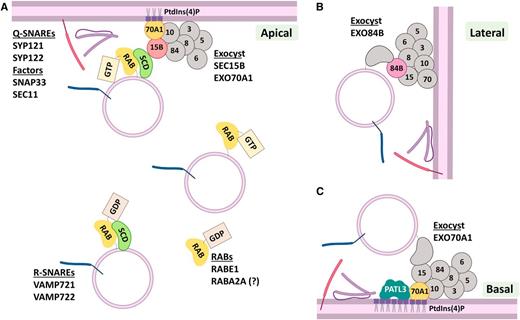
Exocytosis of Recycled Proteins, the Final Step.
(A) Apical PM domains can receive exocytic material coming from the recycling pathway. RABE1 and RABA2A specifically regulate this pathway. RABE1 is attached to the membrane at the TGN/EE where the SCD complex (SCD1-SCD2) acts as a GEF exchanging GDP for GTP, which activates the GTPase. The SCD complex can also interact with SEC15B, a subunit of the EXOCYST complex. EXO15B specifically binds EXO70A1 at the apical PM. EXO70A1 is anchored to the membrane by interaction with PtdIns(4)P. The EXOCYST acts as a multi-subunit tethering complex bringing together R-SNAREs (VAMP721, VAMP722) and Qa-SNAREs (SYP121, SYP122) to form the fusion complex. Once the quadruple SNARE helix is formed, the factors SNAP33 and SEC11 regulate the fusion and complex disassembly.
(B) Lateral recycling and exocytosis require the polar localization and action of the EXOCYST subunit EXO84B.
(C) Basal secretion is specifically regulated by EXO70A1 that is anchored to PtdIns(4)P-enriched regions at the basal membrane. EXO70A1 interacts with Patelin3 (PATL3) that is related to regulation of exocytosis of recycling material and is also anchored to PtdIns(4)P. PALT3 function remains unknown.
RABE1 functions in regulation of assembly of the exocyst during PIN2 recycling (Mayers et al., 2017). At the TGN, RABE1 is activated by interaction with the ARF-GEF complex STOMATAL CYTOKINESIS DEFECTIVE1 and 2 (SCD1-SCD2), that in turn interacts with the exocyst subunit SEC15B (Figure 6A). Supporting a role for SEC15B in secretion of recycling cargo, loss-of-function mutants display PIN3-GFP mis-localization to the vacuole (Janková Drdová et al., 2019). Other members of the family, RABA1B and RABA2A, are implicated in exocytosis of recycling cargoes (Chow et al., 2008; Feraru et al., 2012; Choi et al., 2013; Li et al., 2017). Interestingly, RABA2A alters PIN2 exocytosis, but not that of PIN1, indicating specific activity in apical/shootward secretion (Figure 6A; Li et al., 2017). Nevertheless, the target mechanisms regulated by RABA GTPases have not been elucidated.
Exocytosis can be triggered by the interaction of vesicle-associated SNAREs (VAMPs) with SYP1 family members at the PM (Figure 6A). For example, SYP121 and SYP122 have different roles in the secretion of biosynthetic and recycled cargoes (Assaad et al., 2004; Zhang et al., 2007; Waghmare et al., 2018), but both interact directly with VAMP721 or VAMP722 (Kwon et al., 2008; Uemura et al., 2019). SYP121 participates in biosynthetic secretion and recycling of both aquaporin PIP5;7 and K+ channels by interaction with VAMP721, SNAP33, and SEC11, which regulate assembly of the SNARE complex (Kwon et al., 2008; Besserer et al., 2012; Hachez et al., 2014a; Karnik et al., 2015; Zhang et al., 2015, 2017, 2019). In pollen tubes, SYP131 is involved only in secretion of de novo–synthetized material, whereas SYP125 and SYP124 support exocytosis of recycled material, indicating specialization of these Q-SNAREs (Richter et al., 2011; Ul-Rehman et al., 2011; Slane et al., 2017). Similarly, in root hairs SYP123—but not SYP132—is polarly distributed, continuously cycled by endocytosis, and exhibits specific functions (Kalde et al., 2007; Enami et al., 2009; Reichardt et al., 2011; Rodriguez-Furlán et al., 2016).
Natural selection probably led specific SNARE-adaptor and cargo interactions, as this would facilitate transport along equally specific, restricted routes. Further investigation will dissect the specificity of closely related SYP1 proteins acting in SNARE-tethering complexes in combination with different cargoes and/or as part of a response to endogenous or exogenous signals.
BIG DECISIONS, BIG QUESTIONS
Key Questions
Our knowledge regarding the decision to recycle or degrade endocytic cargo molecules is limited. Two key questions are (1) how are proteins tagged for endocytosis and (2) how are proteins sorted dynamically between recycling and degradation?
The PIN auxin efflux carriers are internalized by clathrin-mediated endocytosis (Cheung and de Vries, 2008; Kitakura et al., 2011; Kleine-Vehn et al., 2011). In mammalian cells, Tyr- and Phe-based motifs present in cargo proteins are recognized by clathrin adaptor proteins (APs), facilitating endocytosis. In plants, AP2 is involved in the endocytosis of BRI1, NIP1;5, and BOR1, but not PIN1 or PIN2 (Takano et al., 2010; Di Rubbo et al., 2013; Sancho-Andrés et al., 2016). Interestingly, cytosolic Tyr motifs present in BOR1 and BRI1 are not essential for interaction with AP2 or internalization (Yoshinari et al., 2012, 2019), and their signals for clathrin-mediated endocytosis remain elusive. Once endocytosed these proteins are constitutively recycled back to the PM (Geldner et al., 2001) or degraded.
What Has BFA Taught Us?
The small molecule brefeldin A (BFA; a macrocyclic lactone produced by fungi in the genus Penicillium) has been a major tool used to study protein trafficking, generating detailed literature and leading to the identification of actively recycled proteins and regulatory mechanisms. Here, we overview BFA action in plants for those less familiar. BFA interferes with the activity of GEFs, specifically those acting on ARF-GTPases (ARF-GEFs) essential for recruitment of coat proteins in trafficking processes (Singh and Jürgens, 2018). BFA binds to the catalytic domain of ARF-GEFs, interfering with GDP-GTP exchange (Mossessova et al., 2003). BFA and its targets remain locked in a complex incapable of recruiting coat proteins, thus inhibiting the corresponding transport pathway.
The main target of BFA in plants was described to be the ARF-GEF GNOM, which is implicated in transport from TGN/EE back to the PM (Geldner et al., 2001, 2003; Friml et al., 2002). Interruption of GNOM activity generates so-called BFA bodies, which were thought to contain membrane agglomerations that trap proteins targeted for recycling. By using photoconvertible fluorescent-tagged PIN proteins (red for existing, green for newly synthetized), it was demonstrated that BFA bodies also accumulate proteins from the biosynthetic pathway (Jásik and Schmelzer, 2014; Mao et al., 2016). Using super-resolution and transmission electron microscopies, GNOM was proven to reside at the Golgi, and upon BFA treatment to associate with the TGN/EE, causing aggregation of these compartments (Naramoto et al., 2014). Additionally, GNOM activity maintains normal TGN/EE function, and interference with GNOM indirectly affects both PM protein secretion and recycling (Naramoto et al., 2014; Doyle et al., 2015).
GNOME-like1 (GNL1), the closest homolog to GNOM, shares some of these characteristics, but is resistant to BFA. Specifically, GNL1 does not accumulate in BFA bodies like GNOM (Teh and Moore, 2007; Doyle et al., 2015). GNL1 has been shown to regulate endoplasmic reticulum (ER)-to-Golgi trafficking, but not endocytosis, of ABCB19 (Titapiwatanakun et al., 2009). Additionally, other ARF-GEFs including BIG1 to BIG4 are in the same pathways; among them, BIG3 is resistant to BFA (Richter et al., 2014; Jonsson et al., 2017). Therefore, treatments with BFA in big3 and gnl1 mutant backgrounds should interfere with all the ARF-GEFs in the pathway.
Treatments with the protein synthesis inhibitor cycloheximide prior to BFA application can help to discriminate between newly synthesized and pre-existing proteins going through endocytic trafficking. By blocking protein synthesis including cargo proteins, cycloheximide inhibits the endomembrane transport of newly synthesized proteins (synthetic transport route), but not the transport of pre-existing proteins from the PM via endocytosis (Wang et al., 2013; Jásik and Schmelzer, 2014). After treatment, BFA can be washed out, releasing pre-existing proteins from BFA bodies. After their release and transport, these proteins can then be observed at the PM, vacuoles, or other compartments. Thus, this approach discriminates endocytosed proteins to be recycled to the PM from those trafficked to the vacuole for degradation (Kleine-Vehn et al., 2008). Using these and complementary approaches, it was later corroborated that PIN auxin carriers are endocytosed and constitutively recycled at the PM; however, under specific endogenous or environmental signals, they are degraded in the vacuole (Kleine-Vehn et al., 2008, 2011; Baster et al., 2013; Du et al., 2013). Many other proteins are characterized as constitutively recycled based solely on their presence in BFA bodies and should be re-assessed. Refining hypotheses based on additional evidence is thus an open task for the community.
Recycling versus Degradation
PIN protein phosphorylation has been proposed as a signal regulating recycling, re-localization, and polarization (Lee and Cho, 2006; Kleine-Vehn et al., 2009; Ding et al., 2011; Zourelidou et al., 2014; Karampelias et al., 2016; Weller et al., 2017). The role of PIN phosphorylation in recycling is indicated mainly by observations that the phosphorylation state of PIN fluorescent proteins can alter their retention in BFA bodies. Reduced retention can be observed as smaller BFA bodies, indicating that PIN fluorescent proteins destined for recycling or secretion to the PM are affected by phosphorylation state. It is unclear whether recycling and re-localization are triggered by PIN phosphorylation via the PINOID or D6PK kinases, secondary effects of BFA reorganization of TGN/EE, or subsequent defects in protein secretion. It is clear however that PINOID phosphorylation of PINs and ABC auxin transporters regulates auxin efflux, suggesting that their phosphorylation by PINIOD acts as a regulatory mechanism for polar accumulation (Friml et al., 2004; Michniewicz et al., 2007; Sukumar et al., 2009; Zourelidou et al., 2009; Huang et al., 2010; Zhang et al., 2010; Dai et al., 2012; Henrichs et al., 2012; Wang et al., 2012, 2019).
However, mutations in the three PINOID phosphorylation sites of PIN2 show only a delay in polarity establishment without affecting endocytosis or secretion (Glanc et al., 2018), suggesting that other signals are in place. In fact, mutants in the Ca2+/calmodulin-dependent kinase CRK5 decrease endocytosis and recycling of PIN2, resulting in alteration of its polar accumulation at the PM (Rigó et al., 2013), indicating another layer of complexity in regulation of PIN protein trafficking.
Phosphorylation is known to affect trafficking of other proteins as well. For example, phosphorylation of the boric acid channel NIP5;1 at Thr in a conserved Thr-Pro-Gly repeat increases clathrin-mediated endocytosis, whereas reduced endocytosis is observed using mutated Thr-to-Ala versions of NIP5;1 and cycloheximide plus BFA treatments (Wang et al. 2017b). Phosphorylation of the IRT1 transporter, which has high affinity for iron and the heavy metals magnesium, zinc, cobalt, and cadmium (Vert et al., 2002), by the kinase CIPK23 regulates ubiquitination and affects IRT1 membrane accumulation. Under non-iron and non-heavy metal conditions, IRT1 localizes to outer polar domains of epidermal cells (Barberon et al., 2011). IDF1 RING3 ubiquitin ligase then mono-ubiquitinates IRT1, triggering transporter endocytosis and recycling (Barberon et al., 2011, 2014; Zhu et al., 2018) . In media depleted of iron, but with an excess heavy metals, cells avoid accumulation of toxic metals by degrading IRT1 (Dubeaux et al., 2018). Metals in excess bind to the cytosolic loop of IRT and recruit the CIPK23 kinase that, in turn, phosphorylates IRT1. This phosphorylation increases the binding of IDF1 at endosomes, triggering polyubiquitination through ubiquitin Lys-63 residues (known as K63 ubiquitination) and re-direction to the vacuole for degradation. Similarly, polyubiquitinated PIN2 is directed to the vacuole. When ubiquitination is decreased by K17R and K12R mutations, endocytosed PIN2 is not directed to the vacuole for degradation, and PIN2 accumulation at the PM suggests that recycling is not affected (Leitner et al., 2012). It is probable but unknown whether PIN phosphorylation is also associated with ubiquitination and impacts on recycling/degradation cycles.
For BRI1, after brassinosteroid perception, the receptor autophosphorylates and then phosphorylates ubiquitin ligases PUB12 and PUB13. These ligases polyubiquitinate BRI1, directing it toward the vacuole for degradation (Zhou et al., 2018). PUB12 and PUB13 are associated with BRI1 before ligand interaction, indicating that basal ubiquitination may occur and regulate receptor recycling. Similarly, PUB12 and PUB13 ubiquitinate the receptor FLS2 in a ligand-independent manner (Table 1; Lu et al., 2011). However, when FLS2 binds its ligand flg22, it associates with the BAK1 kinase that then phosphorylates PUB12 and PUB13. This enhances their binding to FLS2, thus directing polyubiquitination of the receptor and its degradation (Li et al., 2002; Chinchilla et al., 2006; Schulze et al., 2010; Lu et al., 2011). Therefore, overall the literature suggests that the degree of ubiquitination determines the sorting of PM proteins for recycling or degradation (Figure 5A).
Conversely, in yeast, cargo deubiquitinating enzymes remove ubiquitin, prompting recycling either by a default pathway or a retromer-directed pathway (Strochlic et al., 2007; MacDonald et al., 2015). Similarly, in mammalian cells, the deubiquitinating enzyme ASSOCIATED MOLECULE with the SH3 DOMAIN of STAM (AMSH) attenuates degradation of the PM epidermal growth factor receptor that is recycled to the PM (Pareja et al., 2012). Similar to yeast and mammals, AMSH3, an AMSH ortholog in Arabidopsis, hydrolyzes Lys-63–linked ubiquitination (K63). As such, an amsh3 point mutant accumulates ubiquitinated membrane proteins (Isono et al., 2010), and dominant negative AMSH3 causes accumulation of CHITIN ELICITOR RECEPTOR KINASE1 (CERK1) at the PM (Katsiarimpa et al., 2014). AMSH3 interacts with the putative ESCRT-0 protein SH3P2 that binds K63-ubiquitinated cargo and colocalizes with clathrin at the PM. AMSH3-SH3P2 interaction occurs at the TGN/EE and includes interactions with ESCRT-I subunits VPS23.1 and FYVEI (Nagel et al., 2017). This indicates that SH3P2 directs ubiquitinated proteins to the TGN/EE, where they interact with the deubiquitinating enzyme for recycling. Whether SH3P2 cooperates with TOLs in cargo selection or parallel internalization routes exist remains to be determined.
Overall, the TGN/EE may function as a sorting endosome directing proteins for recycling or degradation. As detailed previously, our knowledge of endocytic sorting indicates that retromer and ESCRT subunits are located at the same compartments, at least for part of their lifetimes. Therefore, to efficiently sort proteins their activities must be controlled and segregated. In Caenorhabditis elegans, ESCRT-0 subunits interact with the cochaperone RME-8 that promotes assembly/disassembly of protein complexes (Walsh et al., 2004). RME-8 can then interact with the SNX1 subunit of retromer (Norris et al., 2017). Consequently, RME-8 can segregate ESCRT-0 subunits from the degradative ESCRT machinery, allowing interaction with retromer for recycling. Interestingly, the Arabidopsis RME-8 ortholog GRAVITROPISM DEFECTIVE2 (GRV2) is endosome localized, and grv2 mutants display enlarged aggregated endosomes, defective vacuole biogenesis and embryogenesis, and reduced gravitropic responses (Silady et al., 2004, 2008; Tamura et al., 2007). These phenotypes are consistent with defects in ESCRT subunits, core retromer, and the SNX machinery. While intriguing, further investigation is needed to establish whether RME-8 activity segregating recycling from degradation is conserved in plants. Segregation of recycling functions into microdomains is also described in mammalian cells. SNX subcomplexes interact with microtubule motors generating and maintaining subdomains within endosomes (Hunt et al., 2013). A similar role could be attributed to SNX1-CLASP interaction at TGN/EE in Arabidopsis. Accordingly, in plants, the ESCRT and retromer pathways appear to coexist at TGN/EE but drive opposing functions that must equilibrate continuously.
With so many unanswered basic questions, it is highly relevant to focus on mechanisms regulating PM protein endocytosis, recycling, and degradation in plants.
AUTOPHAGY IS INTEGRAL TO PLANT ENDOMEMBRANE TRAFFICKING
The autophagy pathway adds complexity to the interplay between the secretory and endocytic traffic routes. Autophagy (from Greek self-eating) is a catabolic process allowing eukaryotic cells to digest damaged or superfluous content and reutilize it (Mizushima, 2018). Again, for those not familiar with the complexity of autophagy, we provide a brief overview. Discovered as a process activated under starvation, autophagy plays important roles in biotic and abiotic stress responses, immunity, development, aging, fecundity, and programmed cell death (Minina et al., 2014; Avin-Wittenberg et al., 2018; Mizushima, 2018). Upon activation of a process known as macroautophagy (hereafter autophagy), a specialized cytoplasmic double-membrane vesicle, the autophagosome, engulfs cargo for delivery to lytic compartments (the vacuole in plants and yeast and the lysosome in animals) for degradation and recycling (Marshall and Vierstra, 2018; Mizushima, 2018). This process is regulated by highly conserved autophagy-related proteins (ATGs; for review, see Marshall and Vierstra, 2018). ATGs are structurally and functionally diverse, but core proteins are grouped into five functional clusters: the ATG1/ATG13 kinase complex, the ATG6/Vps34 complex, the ATG9 complex, and two ubiquitin-like systems acting sequentially and roughly corresponding to autophagosome initiation, nucleation, and elongation (Marshall and Vierstra, 2018; Mizushima, 2018). That is, assembly of ATG1/ATG13 is followed by ATG6/VPS34 and recruitment of ATG9-containing membranes to sites of autophagosome assembly. Next, lipidation of ATG8 by the two ubiquitin-like systems leads to elongation and maturation (Marshall and Vierstra, 2018; Mizushima, 2018). Interestingly, despite high conservation, some ATGs have kingdom-specific, autophagy-independent functions (Bestebroer et al., 2013; Cadwell and Debnath, 2018). ER or ER-mitochondria contact sites are the main source of membranes for autophagosomes, which are delivered to lytic vacuoles via a Golgi-independent route (Tooze and Yoshimori, 2010; Zhuang et al., 2016).
Although frequently overlooked, autophagy is closely intertwined with PM protein recycling or degradation. Key interaction points include LEs (Gao et al., 2014; Isono and Kalinowska, 2017), recycling endosomes (Puri et al., 2018), HOPS/CORVET complex (Rojo et al., 2001), retromer complex (Munch et al., 2015), exocyst complex (Pecenková et al., 2017), tethering factor TRAPPIII (Lynch-Day et al., 2010), clathrin-dependent endocytosis (Tooze et al., 2014), and PM-ER contact sites (Wang and Hussey, 2019). For instance, in mammals and yeast, ATG9-enriched vesicles crucial for autophagosome formation interact with Golgi and TGN, facilitating crosstalk between autophagy and endocytosis. Intriguingly, in plants, they are known thus far to be associated only with the ER (Zhuang et al., 2017; Suzuki and Emr, 2018; Judith et al., 2019). In animal cells, Rab11-positive recycling endosomes serve as a platform for autophagosome formation. These endosomes are implicated directly in the trafficking of transferrin and its receptor for degradation (Puri et al., 2018).
The multi-subunit retromer complex is an important component of the recycling pathway and is essential for lytic compartment function and thus directly impacts completion of autophagic activity. For example, loss-of-function mutations in retromer VPS35 subunits lead to pleiotropic phenotypes including autophagy-deficient phenotypes such as early senescence and decreased biomass (Yamazaki et al., 2008; Avin-Wittenberg et al., 2018). Loss of VPS35B also perturbs autophagy-dependent immune responses in Arabidopsis (Munch et al., 2015). Notably, retromer is crucial for yeast autophagy, where it recycles ATG27 from tonoplast back to Golgi (Suzuki and Emr, 2018). Other components of recycling or degradation, such as LEs, may interact directly with plant autophagosomes. For instance, loss-of-function mutants of plant ESCRT subunits AMSH1, VPS2.1, and FREE1 accumulate autophagosomes unable to fuse with the vacuole (for review, see Zhuang et al., 2015; Isono and Kalinowska, 2017). It remains to be investigated whether fusion of autophagosomes with LE is detrimental to plant autophagy, resulting in autophagosome accumulation. Disruption of LE maturation in ESCRT mutants may also alter the availability of SNARE and Rab proteins important for autophagosome fusion with the tonoplast. Notably, similar phenotypes are observed in ESCRT loss-of-function mutants in animal cells (Rusten et al., 2007; Murrow et al., 2015).
Understanding which SNAREs, Rabs, and tethering complexes are needed for plant autophagy is critical (Kwon et al., 2013; Zhuang et al., 2015). Data indicate autophagosome fusion with the tonoplast may require the same tethering complexes as the endocytic/degradation pathway. For example, loss of VPS16 function, which leads to the loss of HOPS and CORVET, results in accumulation of autophagosome-like structures in the cytoplasm (Rojo et al., 2001).
The secretory pathway is another important participant regulating turnover of PM proteins by delivering newly synthesized proteins and recycled proteins back to the PM (see “Exocytosis of Recycled Proteins”). Intriguingly, beyond catabolic activity, animal autophagy plays important nondegradative roles in conventional and unconventional secretion (Deretic et al., 2012; Ponpuak et al., 2015). Surprisingly, the plant exocyst complex subunits SEC5, EXO70B1, and EXO84B are directly implicated in autophagy (Kulich et al., 2013; Pecenková et al., 2017), thus providing another node for crosstalk between autophagy and recycling. Unconventional secretion can involve autophagy to exocytose cytosolic leaderless (lacking signal peptide) proteins or deliver transmembrane proteins to the cell surface via a Golgi-independent route (for review, see Pecenková et al., 2017). It is suggested that unconventional secretion during plant biotic stress responses is regulated by the exocyst with subunits EXO70B1 or EXO70E2, linking autophagy and secretion (Pecenková et al., 2017). In summary, a growing body of evidence indicates the contribution of autophagy to other endomembrane trafficking routes, adding a new layer of regulation and potential redundancy. Even from our fragmented knowledge, it is evident that crosstalk is evolutionarily conserved in eukaryotes.
SELECTIVE AUTOPHAGY AND ITS ROLE IN PROTEIN DEGRADATION
Originally, autophagy was seen as a bulk catabolic process, but further studies revealed the existence of selective autophagy. Subtypes of selective autophagy are generally named after the cargo plus the suffix phagy, for example, mitophagy, pexophagy, aggrephagy, xenophagy, proteaphagy, and chlorophagy (for review, see Marshall and Vierstra, 2018). Selective autophagy allows the rapid degradation of specific components and serves as a modulator of cellular responses to external stimuli, which is important for developmental programs and may ameliorate stresses that damage particular organelles such as mitochondria and ER (Gatica et al., 2018). However, it is increasingly evident that bulk autophagy exhibits selectivity to certain cargoes and may be a complex case of selective autophagy (Mathew et al., 2014).
Selective autophagy relies on recruitment of specific cargoes into autophagosomes with the help of receptor/adapter proteins (for review, see Gatica et al., 2018; Marshall and Vierstra, 2018), but only a few selective autophagy receptors have known cargoes (Table 2). A receptor recognizes cargo for autophagy-dependent degradation and sequesters it into forming autophagosomes by interaction with ATG8. Cargoes marked by receptors for degradation in proximity to autophagosomal membranes may trigger selective autophagy (for review, see Vlahakis and Debnath, 2016; Zaffagnini and Martens, 2016). ATG8 becomes stably anchored on the inner membrane of a growing autophagosome, ensuring cargo trapping.
Receptors of Plant Selective Autophagy with Confirmed Cargo
| Receptor Protein . | Type of Interaction . | Type of Cargo Recognition . | Cargo . | Reference . | |
|---|---|---|---|---|---|
| NBR1/Joka2 | AIM/LDS | Ubiquitin dependent | Ubiquitinated protein aggregates | (Svenning et al., 2011) | |
| TSPO | AIM/LDS | Ubiquitin independent | PIP2;7 | (Hachez et al., 2014b) | |
| ATI1 | AIM/LDS | Ubiquitin independent | Plastid proteins | (Michaeli et al., 2014) | |
| RPN10 | UIM/UDS | Ubiquitin dependent | Proteasome | (Marshall et al., 2015) | |
| PexRD54 | AIM/LDS | Unknown | Unknown | (Dagdas et al., 2016) | |
| DSK2a | AIM/LDS | Ubiquitin independent | Brassinosteroid-responsive transcription factor BES1 | (Nolan et al., 2017) | |
| ORM1 and 2 | AIM/LDS | Ubiquitin independent | FLS2 | (Yang et al., 2019) | |
| PUX proteins | UIM/UDS | Unknown | CDC48 | (Marshall et al., 2019) | |
| Receptor Protein . | Type of Interaction . | Type of Cargo Recognition . | Cargo . | Reference . | |
|---|---|---|---|---|---|
| NBR1/Joka2 | AIM/LDS | Ubiquitin dependent | Ubiquitinated protein aggregates | (Svenning et al., 2011) | |
| TSPO | AIM/LDS | Ubiquitin independent | PIP2;7 | (Hachez et al., 2014b) | |
| ATI1 | AIM/LDS | Ubiquitin independent | Plastid proteins | (Michaeli et al., 2014) | |
| RPN10 | UIM/UDS | Ubiquitin dependent | Proteasome | (Marshall et al., 2015) | |
| PexRD54 | AIM/LDS | Unknown | Unknown | (Dagdas et al., 2016) | |
| DSK2a | AIM/LDS | Ubiquitin independent | Brassinosteroid-responsive transcription factor BES1 | (Nolan et al., 2017) | |
| ORM1 and 2 | AIM/LDS | Ubiquitin independent | FLS2 | (Yang et al., 2019) | |
| PUX proteins | UIM/UDS | Unknown | CDC48 | (Marshall et al., 2019) | |
| Receptor Protein . | Type of Interaction . | Type of Cargo Recognition . | Cargo . | Reference . | |
|---|---|---|---|---|---|
| NBR1/Joka2 | AIM/LDS | Ubiquitin dependent | Ubiquitinated protein aggregates | (Svenning et al., 2011) | |
| TSPO | AIM/LDS | Ubiquitin independent | PIP2;7 | (Hachez et al., 2014b) | |
| ATI1 | AIM/LDS | Ubiquitin independent | Plastid proteins | (Michaeli et al., 2014) | |
| RPN10 | UIM/UDS | Ubiquitin dependent | Proteasome | (Marshall et al., 2015) | |
| PexRD54 | AIM/LDS | Unknown | Unknown | (Dagdas et al., 2016) | |
| DSK2a | AIM/LDS | Ubiquitin independent | Brassinosteroid-responsive transcription factor BES1 | (Nolan et al., 2017) | |
| ORM1 and 2 | AIM/LDS | Ubiquitin independent | FLS2 | (Yang et al., 2019) | |
| PUX proteins | UIM/UDS | Unknown | CDC48 | (Marshall et al., 2019) | |
| Receptor Protein . | Type of Interaction . | Type of Cargo Recognition . | Cargo . | Reference . | |
|---|---|---|---|---|---|
| NBR1/Joka2 | AIM/LDS | Ubiquitin dependent | Ubiquitinated protein aggregates | (Svenning et al., 2011) | |
| TSPO | AIM/LDS | Ubiquitin independent | PIP2;7 | (Hachez et al., 2014b) | |
| ATI1 | AIM/LDS | Ubiquitin independent | Plastid proteins | (Michaeli et al., 2014) | |
| RPN10 | UIM/UDS | Ubiquitin dependent | Proteasome | (Marshall et al., 2015) | |
| PexRD54 | AIM/LDS | Unknown | Unknown | (Dagdas et al., 2016) | |
| DSK2a | AIM/LDS | Ubiquitin independent | Brassinosteroid-responsive transcription factor BES1 | (Nolan et al., 2017) | |
| ORM1 and 2 | AIM/LDS | Ubiquitin independent | FLS2 | (Yang et al., 2019) | |
| PUX proteins | UIM/UDS | Unknown | CDC48 | (Marshall et al., 2019) | |
Receptor binding to ATG8 depends upon the four–amino acid motif W/F/Y-X-X-V/I/L (Noda et al., 2010), which forms hydrophobic bonds with two conserved hydrophobic pockets of the LIR-docking site (LDS) of ATG8 (Dagdas et al., 2016; Kim et al., 2016). Interestingly, affinity between LIR/AIM and LDS is weak, which may prevent receptor binding to ATG8 under conditions not requiring selective autophagy. It was proposed that receptor oligomerization is crucial for interactions strong enough to bind ATG8-enriched membranes to cargoes (Zaffagnini and Martens, 2016). Interaction between cargoes and receptors relies on ubiquitination or polyubiquitination of the cargo (Table 2; for review, see Veljanovski and Batoko, 2014). Notably, depending on the ubiquitin tag type, cargo is degraded via autophagy or the proteasome system (Zaffagnini and Martens, 2016), potentially impacting the scale and speed of degradation. For instance, K63-linked ubiquitin chains are signals for internalization and degradation of PM proteins (Isono and Kalinowska, 2017). Interestingly, p62, the mammalian ortholog of the plant selective autophagy receptor NBR1, prefers K63-linked chains during selective autophagy (Tan et al., 2008), but it is unknown whether this is conserved in plants. A recent study expands on the observation that ATG8 interacts with a protein lacking an AIM motif (Marshall et al., 2015), revealing a new class of selective autophagy receptors interacting with ATG8 via a ubiquitin-interacting motif (UIM). UIM is recognized by a docking site conserved in ATG8 orthologs across kingdoms (Marshall et al., 2019), expanding the potential receptors and thus cargo for selective autophagy in plants, yeast, and animals.
Identification of selective autophagy receptors is challenging; nevertheless, high-throughput investigations of ATG8 interactomes under stress conditions should reveal the complexity of selective autophagy, identify participants, and uncover the details of crosstalk with other endomembrane pathways such as recycling and degradation in the vacuole. A better understating of selective autophagy will answer several important questions in the context of this review: does bulk autophagy exist or i it a complex form of selective autophagy; what is the role of selective autophagy in the turnover of PM proteins; and what are the redundancies between autophagy and endocytosis-dependent PM protein degradation and how are they regulated?
Dynamic Regulation of Autophagy and Protein Recycling/Degradation
As discussed in the section "Recycling versus Degradation", K63 ubiquitination is an important signal for internalization and is a signal for degradation (for review, see Isono and Kalinowska, 2017). A significant proportion of known selective autophagy receptors have ubiquitin binding domains (for review, see Gatica et al., 2018; Marshall and Vierstra, 2018). However, there are examples of selective autophagy regulating PM proteins in a ubiquitin-independent manner. For instance, aquaporin PIP2;7 is internalized and degraded via selective autophagy mediated by TRYPTOPHAN-RICH SENSORY PROTEIN/PERIPHERAL-TYPE BENZODIAZEPINE (TSPO; Hachez et al., 2014b), leading to decreased water permeability of the PM under stress. It is unclear whether TSPO-dependent autophagy sequesters PIP2;7 at the PM or from endosomes. Autophagy for the regulation of PIP2;7 may be important for responses requiring highly specific, transient, rapid, and massive degradation to prevent dehydration, which may not be achieved alone via endocytosis and vacuole sorting. Another example of selective autophagy at the PM is degradation of FLS2 receptor, which is recruited into autophagosomes by OROSOMUCOID1 (ORM1) and ORM2 proteins that are negative regulators of sphingolipid biosynthesis possessing ATG8-interacting motifs (Yang et al., 2019). Ubiquitination triggers FLS2 internalization and degradation (Stegmann et al., 2012); however, Yang et al. (2019) showed that it is dispensable for FLS2 degradation via autophagy, suggesting that autophagy-dependent removal of the inactive FLS2 has a role in maintaining a functional pool of receptors at the PM.
In mammalian cells, clathrin-dependent and -independent endocytosis involves, at least peripherally, the autophagy protein ATG16L1 (Moreau et al., 2012). The plant ortholog of ATG16 was discovered recently (Young et al., 2019), but its role in endocytosis has not been reported. Another example of crosstalk between PM protein maintenance and autophagy is the recycling of transferrin receptor in mammalian cells. Under normal conditions transferrin and its receptor are recovered back to the PM from Rab11A-positive recycling endosomes, whereas under starvation these endosomes serve as a platform for autophagosome formation (Puri et al., 2018).
So far, the role of autophagy in PM protein turnover is restricted to degradation upon internalization. Further studies will reveal whether autophagy plays more complex roles in maintaining PM protein pools beyond maintaining the functionality of the endomembrane system. Furthermore, the potential involvement of ER-PM contacts in endocytosis and autophagy needs to be investigated (Wang and Hussey, 2019). It is still an open question whether ER-PM contacts in plant cells indeed serve as hubs for endocytosis and autophagosome formation and thus provide physical locations where the two pathways cross.
FINAL REMARKS
We have presented examples of endocytosed PM proteins and the mechanisms involved in sorting proteins for recycling or degradation. These processes are complex and dynamic, and perturbations in single components can generate pleiotropic or lethal phenotypes (Hicks and Raikhel, 2012). Because plants typically possess large protein families, a lack of function can be overcome by redundancy. Thus, traditional forward genetic methods are complemented by additional approaches. Among them, the use of chemicals that interfere with specific pathways in specific tissues and at specific developmental stages has facilitated the discovery of cellular phenotypes during short controlled incubations. A growing toolbox of chemicals is available for the community, allowing the study of several specific endocytic pathways (for review, see Li et al., 2012; Hicks and Raikhel, 2014; Rodriguez-Furlan et al., 2017). With such chemicals, it is imperative to consider possible off-target effects, such as those discussed for BFA. To avoid incorrect conclusions, chemical approaches should be supported by complementary approaches including genetics and analytical methods to examine target–ligand interactions. Despite the challenges, there are successful outcomes applying collections of synthetic chemicals to examine the dynamics of endomembrane-specific fluorescent marker proteins using high-resolution live cell imaging. The identification of PM proteins that are endocytosed, recycled, or degraded as well as our understanding of the machinery and regulation have greatly relied on chemical treatments (for review, see Norambuena and Tejos, 2017). Therefore, expansion of the toolbox of chemicals specifically targeting trafficking pathways continues to be of central importance in the field.
We have only begun to unravel the contribution of autophagy to PM protein recycling and degradation in plants. Interestingly, while autophagy is crucial for normal development in animals, plant knockouts of the core ATGs do not show severe developmental aberrations. Rather, typical autophagy-deficiency phenotypes include decreased biomass and early senescence (Avin-Wittenberg et al., 2015; Minina et al., 2018). However, most studies are performed in controlled environments under favorable conditions that may not reveal redundancies required for plant development and survival under changing conditions and stresses in field environments. Furthermore, most of the current plant autophagy studies still rely on constitutive loss-of-function mutants that may result in upregulation of compensatory pathways, masking the potential contribution of autophagy to turnover of essential PM proteins. Crosstalk between autophagy and the endocytic/recycling pathways raises important questions about changes in PM protein turnover upon autophagy upregulation and the physiological relevance of these changes. As autophagy interlaces with endocytic/recycling at several points, this may provide mechanisms to fine-tune PM protein turnover to optimize responses. In addition, utilizing redundancy may be a potent mechanism to maximize PM protein degradation under complex stresses. Selective autophagy may provide yet an additional point of regulation via the specific degradation of PM proteins that are harmful under particular stress conditions.
There is still much to explore and understand about the complexity of mechanisms involved in regulating trafficking between endosomes, PM, autophagosomes, and the vacuole. Many open questions can be and most likely will be addressed with new emerging tools including conditional mutants, genome editing, better imaging systems, fluorescent tools, proximity tagging, and tools for chemical target identification. A remaining challenge is the development of next-generation technologies to overcome our current limitations in studying the apoplast cell wall and cell wall–PM interfaces that are uncharted territory. It is also important as a community to re-visit acquired knowledge to maintain an updated interpretation of data. For example, the localization of SNX1 is still controversial. One of the reasons is the assumption that the fluorescent marker for RAB5-like (ARA7) is only visible at LEs. As noted in the section "Endosome Maturation and Trafficking to the Vacuole," this marker is also present at subdomains of the TGN/EE. Re-assessing existing hypotheses using new technologies uncovered that BFA bodies not only accumulate recycling proteins but also newly synthesized proteins. Therefore, a constant revision of our ever-growing knowledge is an important challenge that we must address.
In conclusion, the study of PM protein endocytosis, recycling, and degradation has come a long way. The first approaches focused on translating knowledge from yeast and mammalian systems to plant cells due to the remarkable conservation of the endocytic pathways across kingdoms. As discussed throughout this review, our understanding of plant endomembrane pathways indicates that one cannot assume that plant counterparts in yeast or animals have similar functions. Research has uncovered new and distinctive plant-specific mechanisms, specializations, and differential regulation. This knowledge in plants has become reciprocal. It was recently found that yeast lacks a recycling/sorting endosome and instead has a TGN/EE association similar to plant cells (Hohmann et al., 2017). These are exciting times to propose plant-specific hypotheses that can lead us to new and unexpected results. Controlling PM protein levels represents a hub of regulation physically linking the extracellular environment to intracellular signaling in order to manage diverse cellular responses. Several pathways can directly or indirectly converge to regulate PM protein abundance, suggesting that this hub can act as a complex network through which many downstream pathophysiological effects can be mediated. New hypotheses and knowledge can help us design biotechnological approaches to generate plants that efficiently interact with an increasingly challenging environment. The future looks bright and challenging for plant cell biologists.
DIVE Curated Terms
The following phenotypic, genotypic, and functional terms are of significance to the work described in this paper:
ACKNOWLEDGMENTS
We apologize to those colleagues whose important work we were unable to incorporate due to space restrictions. We thank Brandon Le and Stephanie Robert for critical reading of the article. This work was supported by U.S. Department of Energy (Grant DE-FG02-11ER15295 to Natasha V. Raikhel and G.R.H.), the Marie Skłodowska-Curie Actions (grant 799433 to E.A.M.), and Carl Tryggers Foundation (Grant 2018 to E.A.M.).
REFERENCES
Author notes
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantcell.org) is: Glenn R. Hicks (ghicks@ucr.edu).