-
PDF
- Split View
-
Views
-
Cite
Cite
Zhuowei Li, Shijie Ma, Huan Song, Zheng Yang, Cuizhu Zhao, David Taylor, Meng Zhang, A 3-ketoacyl-CoA synthase 11 (KCS11) homolog from Malania oleifera synthesizes nervonic acid in plants rich in 11Z-eicosenoic acid, Tree Physiology, Volume 41, Issue 2, February 2021, Pages 331–342, https://doi.org/10.1093/treephys/tpaa125
Close - Share Icon Share
Abstract
Nervonic acid (24:1) is a major component in nerve and brain tissues and it has important applications in food and pharmaceutical industries. Malania oleifera seeds contain about 40% nervonic acid. However, the mechanism of nervonic acid biosynthesis and accumulation in seeds of this endangered tree species remains unknown. In this study, developmental changes in fatty acid composition within embryos and their pericarps were investigated. Nervonic acid proportions steadily increased in developing embryos but 24:1 was not detected in pericarps at any stage. Two 3-ketoacyl-CoA synthase (KCS) homologs have been isolated from M. oleifera developing seeds by homologous cloning methods. Both KCSs are expressed in developing embryos but not detected in pericarps. Based on a phylogenetic analysis, these two KCSs were named as MoKCS4 and MoKCS11. Seed-specific expression of the MoKCS11 in Arabidopsis thaliana led to about 5% nervonic acid accumulation, while expression of the MoKCS4 did not show an obvious change in fatty acid composition. It is noteworthy that the transformation of the same MoKCS11 construct into two Brassica napus cultivars with high erucic acid did not produce the expected accumulation of nervonic acid, although expression of MoKCS11 was detected in the developing embryos of transgenic lines. In contrast, overexpression of MoKCS11 results in similar level of nervonic acid accumulation in camelina, a species which contains a similar level of 11Z-eicosenoic acid as does Arabidopsis thaliana. Taken together, the MoKCS11 may have a substrate preference for 11Z-eicosenoic acid, but not for erucic acid, in planta.
Introduction
Nervonic acid (15Z-tetracosenoic acid, 24:1, n-9), also known as selacholeic acid, is a very-long-chain monounsaturated fatty acid. Malania oleifera Chun et Lee (named Garlic-fruit), a perennial broad-leaved evergreen arbor, contains about 40% nervonic acid in its seeds (Tang et al. 2013). Malania oleifera is a monotypic species within the Olacaceae family, naturally distributed within the Karst regions in the southeast of Yunnan and the west of Guangxi, China. Wild populations of M. oleifera are greatly reduced due to continuous logging and natural habitat clearance. At present, M. oleifera has been identified as an endangered plant species requiring urgent conservation (Ma et al. 2013). In addition, because of niche specialization (Xie et al. 2009b), a low germination rate (Xie et al. 2009a), lack of pollinator and pathogen susceptibility during seed storage or sowing (Xiong et al. 2003), M. oleifera has been assigned to the IUCN Red List (Sun 1998). Clearly, it is unsustainable and impractical to grow M. oleifera as a supply source of nervonic acid. Alternatively, investigating the molecular mechanism of nervonic acid biosynthesis in M. oleifera embryos may provide information for bioengineering to produce nervonic acid in oilseed crops.
Nervonic acid was initially isolated from the cerebrosides of sharks and is ubiquitously distributed in the sphingolipids of vertebrate animals. Nervonic acid is particularly enriched in the white matter of animal brains and in the tissues of the central and peripheral nervous system, where nervonyl sphingolipids are the principal components of white matter and the myelin sheath of nerve fibers (Martínez and Mougan 1998); the nervonoyl moiety is connected by an amide bond to a sphingosine base (Poulos 1995, Merrill et al. 1997, Ma et al. 2013). Nervonic acid has many applications in human health, including the symptomatic treatment of many neuro-degenerative diseases (e.g., X-linked adrenoleukodystrophy, multiple sclerosis and Parkinson’s disease), in treating psoriasis, as well as in the manufacture of supplement formulations for enhancing infant nutrition and maternal health (Taylor et al. 2009). On the industrial front, like high erucic oils, high nervonic oils have potential as feedstocks in the manufacture of bioplastics, enhanced oil recovery surfactants and biojet fuels (Marillia et al. 2014).
Biosynthesis of very-long-chain fatty acids (VLCFAs) has been intensively studied. Fatty acids are de novo synthesized in the plastid using the photosynthesis product acetyl-CoA as a substrate. After the hydrolysis of acyl-ACP by plastid membrane-localized FATA/B thioesterase/s, 16 or 18 carbon fatty acyl-CoAs are synthesized (the precise subcellular location for Long-Chain Acyl-CoA Synthetases is currently under debate) and exported to the endoplasmic reticulum (Ohlrogge 1995). These fatty acyl-CoAs can be elongated by a microsomal fatty acid elongation (FAE) complex/metabolon, comprised of four enzymes catalyzing four successive reactions: the condensation of malonyl-CoA with an acyl-CoA catalyzed by a 3-ketoacyl-CoA synthase (KCS), the reduction of 3-ketoacyl-CoA by a 3-ketoacyl-CoA reductase (KCR), the dehydration of 3-hydroxyacyl-ACP by a 3-hydroxyacyl-CoA dehydratase (HCD) and the reduction of enoyl-CoA by an enoyl-CoA reductase (ECR). The resulting acyl-CoA with two more carbons in the chain can be further elongated up to 32 carbons (Lassner et al. 1996, Lai et al. 2007). KCS is the rate-limiting enzyme in the FAE complex (Millar and Kunst 1997, Haslam and Kunst 2013). About 20% of 11Z-eicosenoic acid (20:1) accumulates in Arabidopsis seeds, and only a trace amount of 20:1 is detected in fae1, an Arabidopsis mutant of KCS18 (James et al. 1995, Millar and Kunst 1997). Twenty-one KCSs in total have been identified in the Arabidopsis genome (Beisson et al. 2003), and KCSs show distinct substrate preferences and produce fatty acyl-CoAs of specific lengths (Blacklock and Jaworski 2006). KCSs play a key role in the biosynthesis and accumulation of VLCFAs, and many KCSs have been characterized in other plant species (Joubès et al. 2008). BnFAE1 (KCS) has been identified in Brassica napus (Rossak et al. 2001) and a TmFAE has been isolated from nasturtium (Tropaeolum majus) (Rossak et al. 2001, Mietkiewska et al. 2004), and they are the key enzymes for the accumulation of erucic acid in seeds of both species. Nervonic acid is also rich in the seed oils of Lunaria annua (honesty), Borago officinalis (borage), Cannabis sativa (hemp), Acer truncatum (Purple-blow maple), Tropaeolum speciosum (flame flower) and Cardamine graeca (southern bittercress) (Bettger et al. 2001, Wang and Wang 2005, Guo et al. 2009, Taylor et al. 2009); the KCS genes involved in the biosynthesis of nervonic acid have been isolated from L. annua (Guo et al. 2009) and C. graeca (Guo et al. 2009, Taylor et al. 2009). Interestingly, the other three enzymes in FAE complex, namely the KCR, HCD and ECR, do not show substrate preference (Kunst and Samuels 2009), and their overexpression show no effects on the accumulation of VLCFAs (Huai et al. 2015). Therefore, KCS is of primary interest for understanding the molecular mechanism of VLCFA biosynthesis in M. oleifera.
To date, none of the lipid-related genes has been functionally characterized from M. oleifera, although this species could be an important source of nervonic acid. In this study, fatty acid compositions of developing embryos and pericarps were investigated first. Two KCS genes, MoKCS4 and MoKCS11, were cloned from developing embryos of M. oleifera, and their expression levels were tested in embryos and pericarps. Their evolutionary relationships were investigated by phylogenetic analysis and their conserved amino residues in the functional domain were compared with KCSs from other sources. Their specificity in the elongation process was investigated by overexpressing these KCSs in seeds of Arabidopsis, B. napus, and camelina (C. sativa). Recently, transcriptome analysis has also provided information that may be utilized to isolate nervonic acid synthesis-related genes and genome sequencing and gene annotation of M. oleifera has been reported (Xu et al. 2019), but lipid genes have not been further tested. Therefore, identification of the key KCS gene for the biosynthesis of nervonic acid in seeds of M. oleifera will facilitate its bioengineering in oil crops.
Materials and methods
Plant materials and growth conditions
Early (121 days-after-flowering (DAF), Stage I) and middle (151 DAF, Stage II) developing and mature fruits (186 DAF, Stage III) of M. oleifera were collected in Yachang, Leye County, Guangxi Province, China (24°48′N, 106°34′E). Once fruits were collected, embryos and pericarps were immediately frozen in liquid nitrogen and stored at −80 °C for fatty acid analysis, DNA and RNA isolation. Arabidopsis ecotype Col-0, camelina cultivar Som and B. napus cultivars (GJ2 and GY416) were used as wild-type hosts for gene expression. The plant culture medium and growth conditions were as described in previous studies (Zhang et al. 2016).
Preparation of DNA, RNA and cDNA synthesis
The pericarp and embryos were isolated by dissection and ground with polyvinylpyrrolidone in liquid nitrogen. Genomic DNA from M. oleifera and B. napus was extracted using the cetyltrimethylammonium bromide method (Webb and Knapp 1990, Allen et al. 2006). RNA from M. oleifera and B. napus was extracted by following the DNA-free RNA isolation protocol prescribed for Arabidopsis seeds and siliques (Yang et al. 2020). First-strand cDNA synthesis for full-length gene amplification was prepared by using 0.2 μg total RNA and the PrimeScript 1st strand cDNA Synthesis Kit (Takara Cat. No. 6110A).
Cloning conserved regions of KCS
Degenerate primers were designed according to amino acid sequences conserved among the following KCSs: C. graeca (GenBank accession No. ACJ61778.1), L. annua (GenBank accession No. ACJ61777.1), Glycine max (soybean) (GenBank accession No. AFY23861.1), Arachis hypogaea (peanut) (GenBank accession No. ACZ51240.1), Gossypium hirsutum (upland cotton) (GenBank accession No. ABA01490.1), Medicago truncatula (barrel clover) (GenBank accession No. AES89124.1), Arabidopsis thaliana (GenBank accession No. NP_195178.1) and Teesdalia nudicaulis (GenBank accession No. ABM65880.1), using the CODEHOP (COnsensus-DEgenerate Hybrid Oligonucleotide Primer) strategy. Two pairs of degenerate primers, FAE1 737-F/FAE1 737-R and KCS18 524-F/KCS18 524-R (Table S1 available as Supplementary Data at Tree Physiology Online), were designed for cloning two conserved regions, respectively. To have the binding site for one of the adapter primers in RACE (Rapid Amplification of cDNA Ends), an internal part of the elongase sequence transcript was amplified.
3′-RACE and 5′-RACE, thermal asymmetric interlaced PCR (TAIL-PCR) and cloning the CDS of MoKCSs
Total RNA was reverse-transcribed with an anchor-tagged switching template (SMART II oligo-nucleotide, Clontech, Takara Bio USA) and oligo-dT primer (for 5′-RACE, 30 dT + NV, Clontech) or with an anchor-tagged oligo-dT primer (for 3′-RACE, anchor sequence + 30 dT + NV, Clontech) using PrimeScript Reverse Transcriptase (Takara Cat. No. 6110A) to prepare the 3′-RACE-ready cDNA and 5′-RACE-ready cDNA. All primers are listed in Table S1 available as Supplementary Data at Tree Physiology Online.
The cDNAs were amplified according to the methods presented in the SMART RACE cDNA Amplification Kit User Manual (Clontech, Takara Bio USA, Inc.). The 5′- and 3′-RACEs were performed using the primer sets of the forward/reverse primer and an adaptor primer (Clontech). All reaction mixtures were formulated following the manufacturer’s recommendations (Clontech). The 3′-RACE PCR was carried out with forward gene-specific primers (MoKCS11 3′-RACE GSP and MoKCS11 3′-RACE NGSP) and the Universal Primer A Mix (UPM; Clontech, Takara Bio USA) as the reverse primer. For the 5′-RACE, UPM was used as the forward primer in PCR in conjunction with the reverse gene-specific primers (MoKCS11 5′-RACE GSP and MoKCS11 5′-RACE NGSP). PCR amplification conditions for both the 3′- and 5′-RACE were as follows: 5 cycles at 94 °C for 30 s and 72 °C for 3 min; 5 cycles at 94 °C for 30 s, 70 °C for 30 s and 72 °C for 3 min; and 20 cycles at 94 °C for 30 s, 68 °C for 30 s and 72 °C for 3 min. The second-round PCR products were cloned into the pEASY- T1 Cloning Vector (TransGen Biotech, Beijing, China) for sequencing.
To clone the 3′ and 5′ untranslated region sequence of MoKCS4, two rounds of TAIL-PCR were performed with target-specific primers (MoKCS4 3′ TAIL GSP, MoKCS4 3′ TAIL NGSP, MoKCS4 5′ TAIL GSP and MoKCS4 5′ TAIL NGSP) and degenerate TAIL-PCR primers (AD1, AD2, AD3 and AD4). The whole TAIL-PCR process included two sequential PCR. Briefly, the first TAIL-PCR was performed in a total volume of 20 μl which contained: 1× PCR buffer, 2.5 mM dNTPs, 0.2 μmol l−1 target-specific primers, 2 μmol l−1 arbitrary degenerate primers (AD1, AD2, AD3 and AD4), 2.5 U Taq DNA Polymerase (Takara Cat. No. R001A) and 200 ng M. oleifera genomic DNA. The method of the primary PCR program is presented in Table S2 available as Supplementary Data at Tree Physiology Online. The second TAIL-PCR amplification was carried out in a total volume of 20 μl that contained: 1× PCR buffer, 2.5 mM dNTPs, 0.2 μM l−1 nest primers, 2 μM l−1 arbitrary degenerate primers (AD1, AD2, AD3 and AD4), 2.5 U Taq DNA Polymerase (Takara Cat. No. R001A) and 1 μl of primary TAIL-PCR product (diluted to 1/100). The method of the secondary PCR program is presented in Table S2 available as Supplementary Data at Tree Physiology Online. The secondary TAIL-PCR products were cloned into the pEASY-T1 Cloning Vector (TransGen Biotech, Beijing, China) for sequencing.
Based on 5′- hyphenate: 3′-sequences, MoKCS11-F and MoKCS11-R were used for cloning CDS of MoKCS11. MoKCS4-F and MoKCS4-R were used for cloning the CDS of MoKCS4. The MoKCS4 and MoKCS11 were amplified with 34 cycles and the two genes were subsequently cloned into the pENTER-TOPO vector (Invitrogen, Carlsbad, California, USA).
Bioinformatic analysis
The KCS protein amino acid sequences from M. oleifera, and several other species, including B. napus, Crambe abyssinica, T. nudicaulis, C. graeca, L. annua, Nicotiana attenuata, jojoba (Simmondsia chinensis), Gossypium raimondii, nasturtium (T. majus) and Arabidopsis, were used for building the neighbor-joining phylogenetic tree using MEGA6 (Version 6.0) with default indexes.
The amino acid sequences of the M. oleifera KCSs and related KCSs from other plant species were compared using DNAMAN software (Version 7.0.2). The alignment contained the KCSs of B. napus (BnFAE1, GenBank accession no. AHM24976), T. nudicaulis (TnFAE, GenBank accession No. ABM65880), Arabidopsis (AtFAE1, AtKCS18, AT4G34520, GenBank accession No. NP_195178), C. graeca (CgFAE, GenBank accession No. ACJ61778), L. annua (LaFAE, GenBank accession No. ACJ61777) and T. majus (TmFAE, GenBank accession No. AAL99199). The alignment of cDNA sequences with genomic DNA sequences was analyzed by the DNAMAN software (Version 7.0.2).
Gene expression analysis of MoKCSs
To study the gene expression of MoKCSs, developing embryo (Stage II) and pericarp (Stage II) cDNAs were used as templates for semi-quantitative RT-PCR. Gene-specific primers, MoKCS11-F2 and MoKCS11-R, were used to detect the expression level of MoKCS11, while gene-specific primers, MoKCS4-F2 and MoKCS4-R2, were designed to detect the expression level of MoKCS4. MoGAPDH was used as a reference gene for comparing the mRNA expression levels in embryos and pericarps. MoGAPDH gene-specific primers, MoGAPDH-F and MoGAPDH-R, were used for semi-quantitative RT-PCR.
To detect the gene expression of MoKCS11 in B. napus transgenic lines, cDNA of developing seeds was used as the template for RT-PCR. BnActin was used as a reference gene for comparing the mRNA expression levels in B. napus transgenic lines. BnActin, gene-specific primers, MoKCS11-F2 and MoKCS11-R2, were used to test the expression level of MoKCS11.
Plant binary vector construction
The MoKCS11 and MoKCS4 on pENTER-TOPO vectors were cloned downstream of the seed-specific phaseolin promoter in the binary vector pK7WG2D-Pha-dsRed with the DsRed marker (Du et al. 2018) through LR reaction. The final binary vectors were transformed into Agrobacterium tumefaciens strain GV3101 containing helper plasmid pMP90 (Koncz and Schell 1986) through the freeze–thaw method (Weigel and Glazebrook 2006).
Plant transformation
Arabidopsis ecotype Col-0 and camelina plants were transformed with Agrobacterium-containing MoKCS binary vectors by the floral dip method (Clough and Bent 1999). Transgenic T1 seeds were detected under green light (543 nm) through a red color filter and seeds showing red fluorescence were collected. Transgenic homozygous Arabidopsis plants were propagated for fatty acid analysis. Transgenic T1 camelina seeds were used for fatty acid analysis.
Brassica napus transformation followed previous methods (Bhalla and Singh 2008, Boszoradova et al. 2011) with some modifications: hypocotyls of 7-day seedlings of two cultivars, GJ2 and GY416, were cut into 7–10 mm segments and these explants were pre-cultured on MS medium (Sigma-Aldrich, St Louis, MO, USA) for 2 days and then co-cultured with Agrobacterium containing MoKCS binary vectors (OD 0.6–0.8) for 10–15 min. Agrobacterium-infected explants were transferred to MS medium and incubated in the dark for 2 days. Calli were induced on MS medium with 1 mg l−1 2,4-d (Sigma-Aldrich). Shoots were induced on B5 medium (Sigma-Aldrich) with 4.5 mg l−1 6-BA, 0.3 mg l−1 TDZ, 200 mg l−1 cephalosporin and 10 mg l−1 kanamycin. Induced shoots were cut from callus and transferred to B5 medium with 150 mg l−1 cephalosporin and 10 mg l−1 hygromycin for 2 weeks. Shoots were induced for rooting on B5 medium with 2 mg l−1 IBA for 2–4 weeks. Regenerated, transformed T0 seedlings were transferred to soil. DNA was extracted from T1 seedlings, and MoKCS11-F and MoKCS11-R were used to confirm the MoKCS11 transgenic lines by PCR.
Lipid and fatty acid analysis
The M. oleifera embryos, pericarps and B. napus seeds were homogenized in liquid nitrogen. Chloroform:methanol (2:1, v/v) was used as an extracting solvent for total lipids. One milliliter of H2O was added to form phases. The organic phase (chloroform with lipids) was transferred to vials and dried under nitrogen for fatty acid analysis.
The fatty acids were analyzed by the method described previously (Zhang et al. 2016) with minor modifications. Arabidopsis seeds were transmethylated at 85 °C for 2 h in 2 ml 5% methanolic H2SO4 solution containing 30% toluene. Heptadecanoic acid (Sigma-Aldrich) (17:0) was added as the internal standard. Fatty acid methyl-esters were extracted by adding 2 ml hexane and 2 ml 0.9% (w/v) NaCl, and the organic phase was analyzed by GC (model Clarus® 680, PerkinElmer, Norwalk, Connecticut, USA). The fatty acids of M. oleifera fruit materials and camelina seeds were measured with the same method as that used for Arabidopsis seeds (Zhang et al. 2016, Du et al. 2018).
Results
Fatty acid analysis of M. oleifera developing and mature fruits
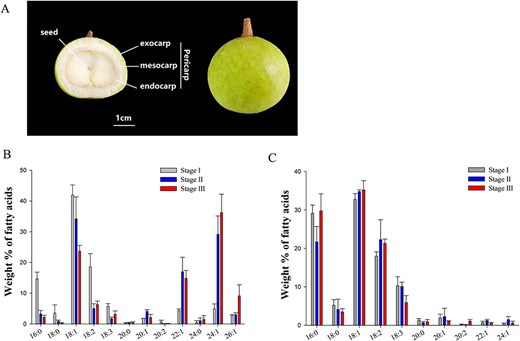
Fruits of M. oleifera develop slowly and flowering to mature fruit set takes over 6 months. To characterize the dynamic change in nervonic acid accumulation, fruits from three different developmental stages, namely early, middle and mature, were collected and whole embryos were extracted for fatty acid analysis (Figure 1A). Results showed that oleic acid (18:1), linoleic acid (18:2) and palmitic acid (16:0) were dominant at the early stage (Stage I), while VLCFAs, including 11Z-eicosenoic acid (20:1), erucic acid (22:1) and nervonic acid (24:1), were minor fatty acids. The three dominant fatty acids (18:1, 18:2 and 16:0) dramatically decreased with further development of the embryo, while VLCFAs, especially nervonic acid, greatly increased from <10% to about 65% when the embryo became mature. Of note, there was about 15% of 22:1 and 10% of ximenic acid (26:1) present in mature seeds, while there was a very low level of 20:1 (Figure 1B). These results indicated that biosynthesis of VLCFAs actively progressed with the development of the embryo, which could be driven by high activities of the fatty acid elongase complex to produce VLCFAs >20 carbons.
Fruits of M. oleifera and their fatty acid composition. (A) Cross section of developing fruit (Stage I: 121DAF; left); mature fruit (Stage III: 186DAF; right); (B) fatty acid composition of M. oleifera embryos; and (C) fatty acid composition of M. oleifera pericarps. Total lipid was extracted from embryos and pericarps at different development stages (Stage I: 121DAF; Stage II: 151DAF; Stage III: 186DAF). Error bars indicate ±SE (n = 3).
To test whether VLCFA accumulated in other tissues of M. oleifera, pericarp, a tissue proximal to the developing embryo, was examined. Fatty acid compositional analysis showed that relative contents of all VLCFAs were very low at all three stages and the dominant fatty acids were 16:0, 18:1 and 18:2, indicating that the activity of the fatty acid elongase complex was drastically lower in pericarps than in embryos (Figure 1C). The former showed a profile more similar to that of vegetative tissue. Therefore, pericarps were used as a negative control for VLCFA accumulation in the following experiments.
Gene cloning of M. oleifera KCS homologs and their expression analysis
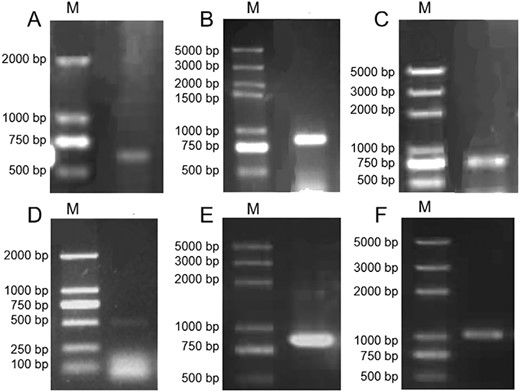
In order to identify the conserved region of M. oleifera candidate KCS(s), the KCS protein amino acid sequences with known FAE activities were first collected from several plants and aligned (see Figure S2 available as Supplementary Data at Tree Physiology Online). Two pairs of degenerate primers were designed to clone the central conserved region, and two fragments of 737 and 521 bp were generated by PCR from the cDNA of M. oleifera embryos, respectively (Figure 2A and D). About 79% similarity of these two fragments suggested that they were from two KCSs and they were preliminarily named as MoKCSx (renamed as MoKCS11 after phylogenetic analysis) and MoKCSy (renamed as MoKCS4 after phylogenetic analysis), respectively.
Gene cloning of M. oleifera KCS homologs. (A) Conserved region of MoKCS11; (B) RACE-PCR cloning of 3′- end of MoKCS11; (C) RACE-PCR cloning of 5′- end of MoKCS11; (D) conserved region of MoKCS4; (E) TAIL-PCR cloning of 3′- end of MoKCS4 and (F) TAIL-PCR cloning of 5′- end of MoKCS4. M: DNA marker.
Based on the sequences of the two conserved regions, 3′- and 5′-RACE or TAIL-PCR were carried out to clone the 5′ and 3′ ends. Two fragments of 835 and 768 bp were amplified by 3′- and 5′-RACE of MoKCSx, respectively (Figure 2B and C). Primers were designed to amplify the coding region and a 1539 bp CDS of MoKCSx was cloned. Then, two fragments of 782 and 1123 bp were cloned from 5′ and 3′ ends of MoKCSy by TAIL-PCR, respectively (Figure 2E and F). Primers were designed to amplify the coding region and a 1557 bp full-length MoKCSy was cloned. Additionally, genomic MoKCSx and MoKCSy were cloned and found to be identical to the corresponding cDNA sequences, which indicated that there are no introns in the coding regions of MoKCSx and MoKCSy.
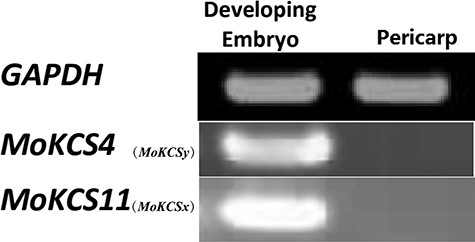
Expressions of MoKCSx (MoKCS11) and MoKCSy (MoKCS4) were compared in developing embryos and pericarps. RT-PCR suggested that both MoKCSx (MoKCS11) and MoKCSy (MoKCS4) were strongly expressed in embryos but were not detected in pericarps (Figure 3), suggesting that they may be involved in the biosynthesis of embryo VLCFAs.
Gene expression analysis of MoKCSs in developing embryos and pericarps of M. oleifera. Total RNA was isolated from embryos and pericarps of developing fruits (Stage II). GAPDH was used as a reference gene for gauging the mRNA expression levels in embryos and pericarps.
Phylogenetic and conserved amino acid analysis of MoKCSx and MoKCSy
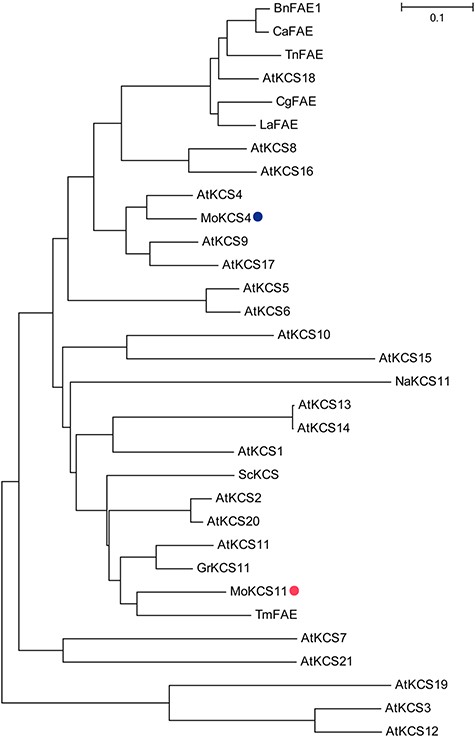
The MoKCSx encodes a polypeptide of 513 amino acids with a predicted molecular mass of 57.7 kDa, while the MoKCSy encodes a polypeptide of 519 amino acids, with a predicted molecular mass of 57.48 kDa. To characterize their relationship with other KCSs, protein sequences of all 21 Arabidopsis KCSs and some functionally identified KCSs from other species were collected for phylogenetic analysis. Results suggested that MoKCSx, as well as TmFAE and GrKCS11, was closely related to AtKCS11. Therefore, the MoKCSx was renamed as MoKCS11. MoKCSy was phylogenetically grouped with AtKCS4, and so, the former was renamed as MoKCS4. It was interesting to note that these KCSs, identified as functioning in VLCFA biosynthesis in seeds such as BnFAE1 (Han et al. 2001), TmFAE (Mietkiewska et al. 2004), TnFAE (Mietkiewska et al. 2007b) and LaFAE (Guo et al. 2009), fell into different KCS subfamilies, which indicated the complicated relationship between KCS structure and function (Figure 4).
Phylogenetic tree of M. oleifera KCSs and KCSs from other species. The MoKCS4 and MoKCS11 from M. oleifera are indicated with a blue and a red dot, respectively. BnFAE1 is from B. napus; CaFAE is from C. abyssinica; TnFAE is from T. nudicaulis; CgFAE is from C. graeca; LaFAE is from L. annua; NaKCS11 is from N. attenuata; ScKCS is from jojoba (S. chinensis); GrKCS11 is from G. raimondii; TmFAE is from nasturtium (T. majus) and all others are from A. thaliana. The Neighbor-joining phylogenetic tree was drawn using default indexes of MEGA6. The bar represents an evolutionary distance of 0.1.
To identify conserved and dissimilar amino acids among KCSs with known function, proteins sequences of MoKCS4 and MoKCS11 were compared with other KCSs, including AtFAE1 (Ghanevati and Jaworski 2002), BnFAE1 (Han et al. 2001), TnFAE (Mietkiewska et al. 2007b), TmFAE (Mietkiewska et al. 2004), CgFAE (Taylor et al. 2009) and LaFAE (Guo et al. 2009). Like all other putative ER-localized plant KCSs characterized to date, the MoKCS4 and MoKCS11 possess all six conserved cysteines (Cys109, Cys239, Cys285, Cys328, Cys404 and Cys475; Figure 5). A previous study showed that four histidine residues are essential for A. thaliana FAE1 activity, and MoKCS4 has all four conserved histidine residues (His329, His413, His417 and His456). However, in MoKCS11, the fourth histidine is replaced with tyrosine (Tyr445) (Figure 5). This distinction led us to speculate that MoKCS4 might have a higher probability to play a role in VLCFA biosynthesis in M. oleifera embryos.
Amino acid sequence alignment of M. oleifera KCSs and fatty acid elongases (FAEs, KCSs) from other species. These include KCSs from B. napus (BnFAE1), T. nudicaulis (TnFAE), A. thaliana (AtFAE1, AtKCS18, AT4G34520), C. graeca (CgFAE), L. annua (LaFAE) and nasturtium T. majus (TmFAE). Conserved Cys (C) and His (H) residues are labeled with diamonds and arrows, respectively. The Tyr (H) at residue position 429 of MoKCS11 is indicated by a star.
Heterologous expression of the M. oleifera KCS11 and KCS4 in Arabidopsis
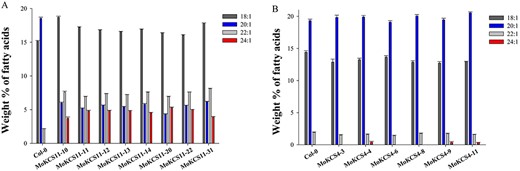
To identify the functions of these two MoKCSs in planta, each, driven by phaseolin promoter, was inserted into a plant binary vector and transferred into Arabidopsis. Homozygous transgenic lines were screened and seeds were analyzed for fatty acid composition. Seed-specific expression of the MoKCS11 resulted in the accumulation of erucic acid (22:1) and nervonic acid (24:1). Eight out of 31 lines showed the highest accumulation of VLCFAs and nervonic acid accumulated to as high as 5%. It is worth noting that the increased proportions of erucic acid and nervonic acid were correlated with a concomitant decrease in eicosenoic acid (20:1). However, the primary elongation substrate, oleic acid (18:1), did not show a decrease; on the contrary, it was somewhat increased (Figure 6A). Additionally, an increased proportion of tetracosanoic acid (24:0) at the expense of its precursors, 18:0 and 20:0, was observed (data not shown). In contrast, the MoKCS4 transgenic T3 lines showed only minor changes in the proportions of eicosenoic, erucic and nervonic acids. The level of nervonic acid was increased from 0 in the Col-0 control line to only 0.2–0.4% in three MoKCS4 transgenic lines (lines 4, 9 and 11) (Figure 6B). These results suggested that it is MoKCS11, rather than the MoKCS4, that plays a key role in the biosynthesis of nervonic acid and other VLCFAs in planta.
Fatty acid composition of MoKCS transgenic Arabidopsis seeds. (A) Relative proportions of 18:1, 20:1, 22:1 and 24:1 in seeds of wild-type Col-0 and eight A. thaliana T3 homozygous transgenic lines expressing the MoKCS11 gene under control of the phaseolin promoter; (B) relative proportions of 18:1, 20:1, 22:1 and 24:1 in seeds of wild-type Col-0 and six A. thaliana T3 homozygous transgenic lines expressing the MoKCS4 gene under control of the phaseolin promoter. Error bars indicate ±SE (n = 3).
Heterologous expression of the MoKCS11 in B. napus
Considering that erucic acid is a direct precursor of nervonic acid and there is high erucic acid content in classical B. napus, we speculated that the overexpression of MoKCS11 in a high erucic acid B. napus cultivar could result in a much higher accumulation of nervonic acid than what was observed in Arabidopsis. The same MoKCS11 binary vector was transferred into two B. napus cultivars, each with about 30% erucic acid naturally present in their seed oils. Nine transgenic lines were obtained. Surprisingly, seed fatty acid analysis of these T1 and T2 lines indicated that there was no obvious accumulation of nervonic acid (Figure 7A and Figure S2 available as Supplementary Data at Tree Physiology Online). To test possible gene silencing of MoKCS11 in these lines, its expression was tested. RT-PCR showed that MoKCS11 was indeed expressed at transcription level in the developing transgenic B. napus embryos (see Figure S3 available as Supplementary Data at Tree Physiology Online). Although we could not rule out the possibilities of translational or posttranslational gene silencing of MoKCS11 or some other kind of inhibition, this result suggested that MoKCS11 may not synthesize nervonic acid in B. napus, even in the presence of rich proportions of erucic acid, the direct nervonic precursor.
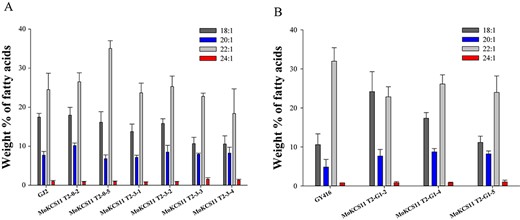
Fatty acid composition of MoKCS11 transgenic in seeds of B. napus. Relative proportions of 18:1, 20:1, 22:1 and 24:1 in seed oils from the host cultivar GJ2 (A) or GY614 (B) and their T2 independent transgenic lines expressing the MoKCS11 gene under control of the phaseolin promoter. Error bars indicate ±SE (n = 3).
Heterologous expression of the MoKCS11 in camelina
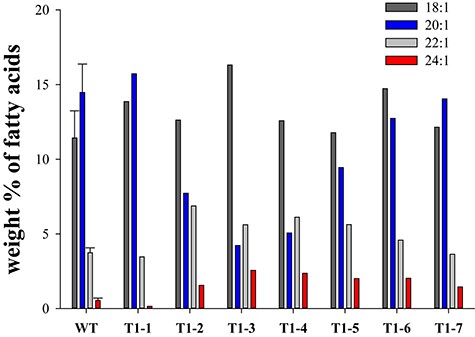
Based on the above unexpected phenotype of MoKCS11 B. napus lines, we postulated that MoKCS11 might prefer 20:1, rather than 22:1, as an elongation substrate. To test this hypothesis, the same MoKCS11 binary vector was transferred into camelina, a species whose seeds are rich in 20:1, having a proportion similar to that found in Arabidopsis seeds. T1 seeds were screened for DsRed fluorescence and positive seeds were collected. Fatty acid analysis of these T1 seeds showed that nervonic acid accumulated in six out of seven lines. On average, the proportion of nervonic acid in T1 seeds increased from 0.5% in the non-transformed control line (CK) to as high as 2.5% in the seeds expressing the MoKCS11. The erucic acid content also increased from 3.7% in the control line to 5.1%, on average, in MoKCS11 transgenic T1 lines; in lines 2, 22:1 was as high as 6.9%. The increases of erucic and nervonic proportions were correlated with a decrease in 20:1. Moreover, none of lines showed a decrease in proportions of 18:1; on the contrary, all showed increases in 18:1 (Figure 8). Taken together, these results suggest that MoKCS11 functions best for synthesis of VLCFAs when 20:1 is presented as a precursor.
Fatty acid composition of MoKCS11 transgenic seeds of camelina. Relative proportions of 18:1, 20:1, 22:1 and 24:1 in seeds of wild type and T1 transgenic seeds expressing the MoKCS11 gene under control of the phaseolin promoter. Error bars indicate ±SE (n = 3).
Discussion
Malania oleifera embryos can contain more than 40% nervonic acid (Figure 1) and its seeds are used for producing edible oil by local populations. Because this endangered tree is only adapted to a very small region (Xu et al. 2019), it is impractical to commercialize it widely for sustainable pharmaceutical or nutritional feedstocks. Understanding the molecular mechanism of nervonic acid biosynthesis in this species may provide information to allow bioengineering for new high nervonic crops. Heterologous seed-specific expression of MoKCS in Brassicaceae oil crops, such as camelina, is a potential way to engineer high nervonic acid oils for medical and food applications. Since the identification of FAE1 (KCS) in Arabidopsis (James et al. 1995, Rossak et al. 2001), KCSs from several plant families have been characterized for producing VLCFAs of different chain lengths. For example, jojoba FAE, T. majus FAE and C. abyssinica FAE mainly produce 22:1 (Lassner et al. 1996, Mietkiewska et al. 2004, 2007b). In previous studies, two KCS genes involved in biosynthesis of VLCFA, specifically with respect to wax biosynthesis, have been characterized in the Brassicaceae (Millar et al. 1999, Todd et al. 1999). These results provided important background information for identifying KCSs for the biosynthesis of nervonic acid in M. oleifera.
In this study, investigation of fatty acid composition indicated that nervonic acid increased steadily from 4% to nearly 40% in developing embryos of M. oleifera (Figure 1). Two MoKCSs cloned from M. oleifera exhibit high homology and structural similarity to the KCS sequences of various other plant species possessing high VLCFAs in seeds (Figure 4). Expression of both MoKCSs was detected in developing embryos but not in pericarps (Figure 3); the latter has near negligible proportions of nervonic acid (Figure 1C). While conducting this study, both a transcriptome and an assembled genome of M. oleifera developing embryos have been released (Yang et al. 2018, Xu et al. 2019). Two MoKCSs are present in the transcriptome and show identical sequences as the two MoKCSs in this study (Yang et al. 2018). By comparing our CDSs with genomic sequences, we confirmed the absence of introns in these two KCSs, as is the case in those from T. majus, C. abyssinica, C. graeca and L. annua (Mietkiewska et al. 2004, 2007b, Guo et al. 2009, Taylor et al. 2009). Overexpression of MoKCS11 produced more than 5% of de novo synthesized nervonic acid in transgenic Arabidopsis seeds, while the overexpression of MoKCS4 produced only a trace amount of nervonic acid in a parallel experiment (Figure 6). These results indicate that it is MoKCS11, and not MoKCS4, which plays a key role in the biosynthesis of nervonic acid in M. oleifera.
A phylogenetic analysis shows that all KCSs from the Brassicaceae, which accumulate VLCFAs in seed oils, including BnFAE1 (Yan et al. 2015), CaFAE (Mietkiewska et al. 2007b), TnFAE (Mietkiewska et al. 2007a), CgFAE (Taylor et al. 2009) and LaFAE (Guo et al. 2009), are in the group designated as KCS18. All other KCSs identified for storage VLCFAs from families other than Brassicaceae, including TmFAE, ScKCS (Lassner et al. 1996) and the MoKCS11 in this study, are close to, or in the group of, KCS11 (Figure 4). More importantly, the latter plant species show a distant genetic lineage. The biological classifications of these plants are as follows: T. majus belongs to the order Geraniales, S. chinensis to the order Caryophyllales, G. raimondii to the order Malvales and M. oleifera to the order Santalales. Although more examples are needed to validate the following generalization, it may be that when it comes to storage VLCFAs, researchers will need to focus on KCS18s in Brassicaceae and to focus on other KCSs closer to KCS11 in other plant families.
In a previous study, it was shown that six residues of cysteine and four residues of histidine are conserved in the functional domain of KCSs (Ghanevati and Jaworski 2001). Cysteine 223 and all histidines are essential for elongation activity (Lassner et al. 1996, Ghanevati and Jaworski 2002). In this study, both MoKCSs show the six conserved residues of cysteine, while they show one residue difference regarding the four conserved histidine residues. MoKCS4 contains all four residues of histidine, while MoKCS11 contains three of the His residues with a Tyr substitution on the 429th histidine (Figure 5). It is believed that substitution of histidine by another amino acid is rare in nature and an occasionally safe substitution is by the structurally similar lysine (Bordo and Argos 1991). In the early stages of this study, the residue substitution on MoKCS11 led us to speculate that MoKCS4 might function in nervonic acid biosynthesis. Additionally, our phylogenetic analysis suggested that MoKCS4 is closer to two KCSs within the Brassicaceae, CgFAE (Taylor et al. 2009) and LaFAE (Guo et al. 2009), both of which have been shown to function in nervonate biosynthesis. However, our finding that nervonic acid was synthesized in seed oils of MoKCS11 transgenic Arabidopsis and camelina indicated that MoKCS11 is responsible for the accumulation of VLCFAs in M. oleifera (Figures 6A and8). This suggests that the Tyr substitution for histidine on residue 429 does not undermine elongation activity. TmFAE also contains a Tyr 429 substitution (Figure 5) (Mietkiewska et al. 2004). Taken together, the histidine 429 residue of KCS11 may be replaceable by a non-conserved amino acid, which needs to be further tested by substitution experiments.
It is interesting to note that the overexpression of MoKCS11 triggers de novo accumulation of nervonic acid in seeds of Arabidopsis as well as in camelina (Figures 6A and8). However, when the same vector was transferred into two high erucic acid B. napus cultivars, no further nervonic acid was accumulated beyond the low proportion already present in the base cultivars (Figure 7 and Figure S2 available as Supplementary Data at Tree Physiology Online). It may be argued that only limited transgenic lines have been tested and it would be possible to find a line with higher nervonic acid if more transgenic lines had been explored. However, this argument is weakened by the finding that MoKCS11 is indeed expressed in the developing seeds of all of these transgenic lines and there is no variation in 24:1 accumulation (see Figure S3 available as Supplementary Data at Tree Physiology Online). It is worth noting that a common feature of Arabidopsis and camelina is the similar proportion of 20:1 as the predominant VLCFA in their mature seed oils, which is distinct from high accumulation of 22:1 in the host B. napus cultivars we tested. We propose that MoKCS11 might have a substrate preference for 20:1, but not 22:1, in planta. This speculation is also supported by the very low proportion of 20:1 in all developmental stages of M. oleifera embryos. More study is required to confirm this hypothesis.
Acknowledgments
We thank Prof. Jianhua Tian (Hybrid rape research center of Shaanxi province) and Dr Chaofu Lu (Montana State University) who kindly provided two B. napus cultivars (GJ2 and GY416) and the camelina cultivar Som, respectively. This work was supported by the National Natural Science Foundation of China (31972964), the Key International Cooperation Project of Shaanxi province (2020KWZ-012) and the Programme of Introducing Talents of Innovative Discipline to Universities (Project 111) from the State Administration of Foreign Experts Affairs (#B18042) ‘Crop breeding for disease resistance and genetic improvement’.
Conflict of interest
None declared.
Authors’ contributions
Z.L. and M.Z. conceived the original screening and research plans. M.Z. supervised the experiments. Z.L., S.M., H.S., Z.Y. and C.Z. performed the experiments and analyzed the data. Z.L., S.M., D.T. and M.Z. conceived the project and wrote the manuscript with contributions of all the authors.
References
Ghanevati M, Jaworski JG (
Millar AA, Clemens S, Zachgo S, Giblin EM, Taylor DC, Kunst L (
Todd J, Post-Beittenmiller D, Jaworski JG (
Author notes
Zhuowei Li and Shijie Ma contributed equally to this work.