-
PDF
- Split View
-
Views
-
Cite
Cite
Atsushi Hoshino, Yasumasa Morita, Jeong-Doo Choi, Norio Saito, Kenjiro Toki, Yoshikazu Tanaka, Shigeru Iida, Spontaneous Mutations of the Flavonoid 3′-hydroxylase Gene Conferring Reddish Flowers in the Three Morning Glory Species, Plant and Cell Physiology, Volume 44, Issue 10, 15 October 2003, Pages 990–1001, https://doi.org/10.1093/pcp/pcg143
Close - Share Icon Share
Abstract
Among the Ipomoea plants, both Ipomoea nil and Ipomoea tricolor display bright blue flowers, and Ipomoea purpurea exhibits dark purple flowers. While all of these flowers contain cyanidin-based anthocyanin pigments, the mutants of I. nil, I. purpurea, and I. tricolor carrying the magenta, pink, and fuchsia alleles, respectively, produce reddish flowers containing pelargonidin derivatives, and all of them are deficient in the gene for flavonoid 3′-hydroxylase (F3′H). The magenta allele in I. nil is a nonsense mutation caused by a single C to T base transition generating the stop codon TGA, and the cultivar Violet carries the same mutation. Several tested pink mutants in I. purpurea carry inserts of the 0.55-kb DNA transposable element Tip201 belonging to the Ac/Ds superfamily at the identical site. No excision of Tip201 from the F3′H gene could be detected, and both splicing and polyadenylation patterns of the F3′H transcripts were affected by the Tip201 integration. The fuchsia allele in I. tricolor is a single T insertion generating the stop codon TAG, and the accumulation of the F3′H transcripts was drastically reduced by the nonsense-mediated RNA decay. Spontaneous mutations in Ipomoea, including a possible founder mutation in the pink allele, are also discussed.
(Received July 15, 2003; accepted August 22, 2003)
Introduction
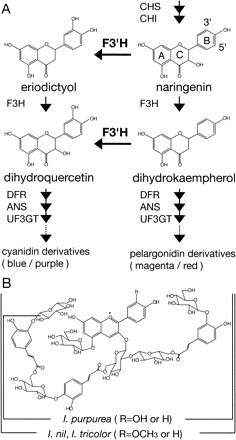
Considerable attention has been paid recently to the morning glory genus Ipomoea in experimental versatility of floral biology including genetics of floral variation, flavonoid biosynthesis and transposon-induced mutations (Clegg and Durbin 2003). Some of the Ipomoea plants exhibiting blue or purple flowers, which contain cyanidin-derived anthocyanins, are rather exceptional; most blue or purple flowers in plants generally contain anthocyanin pigments derived from the purple delphinidin, whereas many reddish flowers contain anthocyanins from the scarlet pelargonidin or the crimson cyanidin (Harborne 1994). As Fig. 1 and 2 show, both the Japanese morning glory (Ipomoea nil) and Ipomoea tricolor display bright blue flowers that contain the peonidin (3′-methoxyl cyanidin) derivative named Heavenly Blue Anthocyanin (HBA) (Kondo et al. 1987, Lu et al. 1992b), and the common morning glory (Ipomoea purpurea) produces dark purple flowers containing a cyanidin derivative that lacks one glucose molecule and a methyl residue from HBA (Saito et al. 1995). Among the morning glories, I. nil and I. purpurea belong to the same subgenera Ipomoea, whereas I. tricolor was classified into another subgenera, Quamoclit (Austin and Huáman 1996). Genetic studies on the color of I. nil have shown that blue flower coloration was mainly controlled by two genetic loci, Magenta and Purple (Hagiwara 1931, Imai 1931). Recessive magenta and purple mutants have magenta and purple flowers, respectively (Fig. 1). Various I. nil mutants displaying reddish flowers and/or carrying the magenta allele accumulate pelargonidin-based anthocyanin pigments instead of peonidin derivatives (Lu et al. 1992a, Toki et al. 2001b, N. Saito unpublished), and the cultivar Violet extensively used for studies of the photoperiodic induction of flowering (Imamura 1967, Vince-Prue and Gressel 1985) also has darker red flowers containing pelargonidin derivatives (Toki et al. 2001a). These results indicate that the Magenta locus in I. nil is highly likely to control flavonoid 3′-hydroxylase (F3′H) activity. The F3′H enzyme in plants hydroxylates the 3′ position of the B-ring of naringenin and dihydrokaempferol to produce eriodictyol and dihydroquercetin, respectively, and the latter two products are precursors of cyanidin (Fig. 2A) (Harborne 1994, Holton and Cornish 1995, Mol et al. 1998, Shirley 2001). The F3′H enzyme belongs to the cytochrome P450 superfamily, and its cDNA was first isolated from petunia (Brugliera et al. 1999). The other Purple gene was shown to encode a vacuolar Na+/H+ exchanger (InNHX1) that increases the vacuolar pH during flower opening to create the usual blue petals (Fukada-Tanaka et al. 2000, Yamaguchi et al. 2001). Based on the studies of red flower mutants in I. purpurea, the locus controlling the dark purple pigmentation was designated as Blue by Barker (1917) and later as Pink by M. Clegg and his collaborators (Ennos and Clegg 1983, Schoen et al. 1984, Epperson and Clegg 1988). Although Imai and his collaborator (Imai 1927a, Imai and Tabuchi 1935) also used mutants exhibiting red flowers, they did not give any designation to their mutation. Since chemical analysis showed that the red flower pigments are pelargonidin derivatives (Saito et al. 1996), it was also postulated that the Pink (Blue) locus in I. purpurea controls the F3′H activity. No such analysis has been conducted on the I. tricolor mutant Wedding Bells, which produces fuchsia red flowers (Fig. 1).
In this study, we first analyzed the red flower pigments of I. tricolor cv. Wedding Bells and found that it is also deficient in the F3′H activity. Subsequently, we cloned the F3′H cDNAs from all three Ipomoea species, I. nil, I. purpurea, and I. tricolor, and characterized the genomic structures of their F3′H genes, all of which comprise three exons. By comparing these, we identified a new small mobile element-like sequence (MELS) within intron 1 of the I. nil F3′H gene. The structural features of the MELS elements resemble MITEs (miniature inverted-repeat transposable elements) (Feschotte et al. 2002), and their possible roles in the divergence of genes before and after speciation were discussed by comparing the genomic CHS and DFR sequences in I. nil and I. purpurea (Hoshino et al. 2001). We then identified the F3′H mutations in these Ipomoea species. The magenta mutation is a nonsense mutation in exon 3, the pink allele is caused by an insertion of a novel DNA transposon Tip201 (Transposon Ipomoea purpurea 201) into exon 3, and the mutation of Wedding Bells named fuchsia is a single bp insertion at exon 2 of the F3′H genes. We also examined the effects of these mutations on the F3′H mRNA production.
Results
Anthocyanins in the petals of Wedding Bells
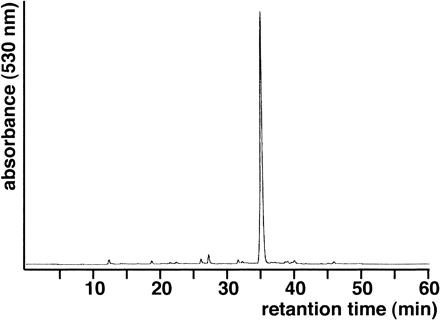
HPLC analysis of the anthocyanin components extracted from the petals of Wedding Bells gave virtually a single major peak (Fig. 3), which contains a pelargonidin derivative with a polyacylated glucoside moiety that is identical to that of HBA, named Pharbitis red anthocyanin 5 or PRA-5 (Lu et al. 1992a, Toki et al. 2001a) (Fig. 2B). The HPLC profile of Wedding Bells was in striking contrast with those of Violet and Hama-no-kagayaki, in which more than 10 anthocyanin peaks could easily be detected and PRA-5 amounted to only 14–24% (Toki et al. 2001a, data not shown). It is clear that the I. tricolor mutant Wedding Bells is deficient in F3′H activity.
Characterization of cDNAs encoding F3′H and their genomic sequences from the Ipomoea species
Since the Ipomoea species of I. nil, I. purpurea, and I. tricolor displaying reddish flowers were assumed to lack F3′H activity, we isolated F3′H cDNA clones from wild-type plants exhibiting blue or dark purple flowers. The I. nil F3′H cDNA was first isolated from young flower buds using petunia F3′H cDNA (Brugliera et al. 1999) as a probe, and the obtained I. nil F3′H cDNA was then employed as a probe for screening the F3′H cDNAs from I. purpurea and I. tricolor. To determine their 5′ ends, 5′-RACE (rapid amplification of cDNA ends) was performed. The cDNA sequences from I. nil and I. purpurea indicated that their F3′H genes encode polypeptides of 519 amino acids and that they are different in 18 amino acid residues (data not shown). The F3′H gene of I. tricolor encodes a polypeptide of 522 amino acids, which contains 32 and 34 residues different from those of I. nil and I. purpurea, respectively. The amino acid sequences of the deduced F3′H enzymes in these Ipomoea species show about 72–74% identity and 82–84% similarity with the petunia F3′H sequence, supporting the notion that these cDNAs encode the F3′H enzymes of the morning glories.
Southern blot analysis using the F3′H cDNAs as probes indicated that all of the wild-type Ipomoea species tested contain only one copy of the F3′H gene (data not shown). We cloned and characterized the genomic F3′H gene regions from the wild-type plants of these Ipomoea species. Comparison of the sequences of these genomic F3′H gene regions with those of their cDNAs revealed that all of them consist of three exons and two introns and that the 9-bp segment encoding three additional amino acids in I. tricolor resides at the 3′ end of exon 2 (Fig. 4A). The sequences at the transcriptional initiation sites of the F3′H gene in I. nil and I. tricolor are identical, and their transcription starts at G or A (12 bp or 11 bp upstream of the ATG initiation codon, respectively), whereas dinucleotide TT instead of G was found at the corresponding site in I. purpurea, and its transcription initiates predominantly at A (11 bp upstream of the ATG initiation codon). The transcriptional termination sites appeared to vary in these morning glories.
Various deletions and insertions were detected in the intron regions by comparing the genomic F3′H sequences, and one of them, which resides in the first I. nil intron, is a newly identified MELS element of 1,086 bp named MELS9-1[J] (Fig. 4A). MELS9-1[J] is an AT-rich (70.4%) element with 7-bp terminal inverted repeats (TIRs) that is flanked by 9-bp direct repeats including a diverged terminal 3-bp sequence (Fig. 4C). Like other MELS elements found in the genomic CHS and DFR regions in I. nil and I. purpurea (Hoshino et al. 2001), MELS9-1[J] does not appear to affect the activity of the I. nil F3′H gene. Out of the remaining 21 non-homologous regions, 10 segments [N/P3, 5, 6; N/T1, 2, 4; P/T1, 3 (=P/N1); T/P1 (=T/N1), 2 in Fig. 4A] were flanked by short direct repeats of 2–30 bp, and one copy of the repeat sequences was found at the corresponding deletion sites, indicating that recombination events between these repeats are likely to have produced these deletions.
Expression of the F3′H genes in the mutants displaying reddish flowers
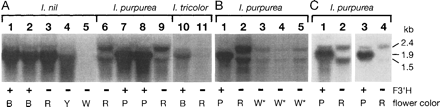
Since the mutants displaying reddish flowers are postulated to be deficient in F3′H activity, we used Northern blot analysis to examine whether the mutations affected the expression of the F3′H genes in the young flower buds. No apparent difference was observed between the wild-type Magenta lines (KK/ZSK-2 and Hama-no-sora) and the magenta mutants (Hama-no-kagayaki and 54Y) in I. nil, whereas a significant reduction was detected in I. tricolor Wedding Bells compared with the wild-type Heavenly Blue (Fig. 5A). In I. purpurea, pink mutants (KK/FR-35 and KK/VR-40) mainly showed mRNAs of about 2.4-kb and 1.5-kb, with slightly reduced amounts, in contrast to the 1.9-kb F3′H mRNAs produced in the wild-type lines (KK/FP-36 and YO/FP-39). It is clear that the expression of the F3′H gene was affected, at least in the I. purpurea and I. tricolor mutants. As is described below, the pink mutation in I. purpurea affects both the splicing and polyadenylation patterns of the F3′H transcripts and the fuchsia mutation in I. tricolor causes the F3′H mRNA degradation.
We previously showed that mRNA accumulation of the structural genes encoding the CHS-D, CHI, F3H, DFR-B, ANS, and UF3GT enzymes for anthocyanin biosynthesis in Fig. 2 was significantly reduced in the mutant 78WWc-1 displaying white flowers (Abe et al. 1997, Fukada-Tanaka et al. 1997, Hoshino et al. 1997, Y. Morita et al. unpublished). Recently, we identified that the C-1 gene encodes a MYB transcriptional activator for anthocyanin biosynthesis, a homologue of the petunia An2 gene (Quattrocchio et al. 1999), and that several c-1 mutants, including 78WWc-1, carry a deletion within the coding region (Y. Morita et al. unpublished results). The reduced F3′H mRNA accumulation in 78WWc-1 (Fig. 5A) suggests that the F3′H gene is also regulated by the C-1 protein. The gene in I. purpurea corresponding to the C-1 gene of I. nil also appears to control the F3′H gene expression because the F3′H mRNA accumulation was found to be drastically reduced (Fig. 5B) in an I. purpurea mutant carrying two discrete deletions in the coding region of the gene for the MYB transcriptional activator (M. Rausher personal communication, Y. Morita et al. unpublished results). Interestingly, the mutation confers white flowers with pigmented spots in the rays, and the major residual F3′H transcripts appear to be ascribed to the expression in the pigmented spots of the rays. The same mutation in the MYB gene was reported to affect several structural genes for anthocyanin biosynthesis (Tiffin et al. 1998). Since the petunia An2 gene product was not shown to control the Ht1 gene encoding the F3′H enzyme (Brugliera et al. 1999), the results described here indicate that the action of the MYB transcriptional activator on the F3′H gene in petunia and the morning glories are different. It is noted that the An2 protein controls the Hf1 gene for flavonoid 3′,5′-hydroxylase (F3′5′H) for flower pigmentation in petunia (Mol et al. 1998).
The magenta mutation of I. nil is a single-base change generating a stop codon in the F3′H gene
To examine whether the magenta mutation of I. nil resides within the F3′H gene, we characterized F3′H cDNA from the mutant Hama-no-kagayaki, which exhibits reddish flowers, by reverse transcription-PCR (RT-PCR) amplification and 5′-RACE and found a single base transition from C to T generating the stop codon TGA at position 1,282 of the F3′H cDNA (Fig. 4A, B). Moreover, the authentic magenta mutants, 54Y and 78WWc-1, also carried the same transition mutation. The results were further confirmed by the fact that an identical nonsense mutation was found in all of the genomic sequences of the additional 13 different I. nil mutants, including Violet (data not shown), which have reddish flowers and/or bear the magenta allele (see Materials and Methods). These mutants must produce truncated F3′H peptides lacking C-terminal 92 amino acids containing the prerequisite heme-binding domain conserved among the P450 superfamily (Chapple 1998). Based on these results, we can conclude that the F3′H gene corresponds to the Magenta locus in the Japanese morning glory.
The pink mutation of I. purpurea is caused by integration of a novel transposon Tip201 within the F3′H gene
Since DNA rearrangements were detected in the F3′H gene of the pink mutants by Southern blot analysis (data not shown), the genomic F3′H gene regions in the wild-type Pink plant YO/FP-39 and the pink mutant KK/FR-35 were amplified by long and accurate-PCR (LA-PCR) using primer sets IPF3′H-E1F1 and IPF3′H-E3R1 (Fig. 4A). The PCR-amplified fragment obtained from KK/FR-35 was about 0.5 kb longer than that from YO/FP-39, indicating that the pink mutation is caused by an insertion in the F3′H gene (data not shown). To examine the pink mutation further, appropriate HindIII fragments from KK/FR-35 were cloned, and the genomic F3′H gene sequence of KK/FR-35 was compared from that of YO/FP-39. The 512-bp insertion named Tip201-1 was found to reside within exon 3. Furthermore, we carried out co-segregation analysis of the pink allele and the red flower phenotype using the LA-PCR protocol to distinguish the wild-type Pink allele and the mutant pink allele in which Tip201-1 had been inserted (see Materials and Methods). The results support the notion that the recessive pink allele conferring red flowers indeed corresponds to the F3′H gene carrying Tip201-1.
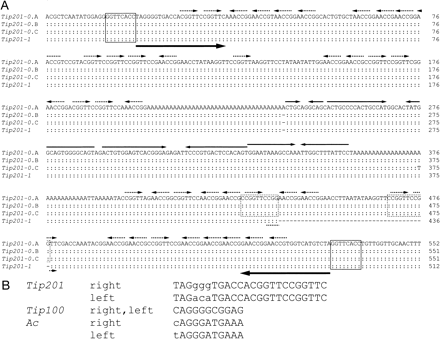
We also analyzed six additional pink mutants of I. purpurea by sequencing the LA-PCR-amplified fragments. Four of them, KK/VR-37, KK/VR-40, KK/WR-321, and CR/FR-L1, contain an identical 551-bp insertion, which we designated as Tip201-0. The remaining two mutants, YO/FR-34 and CR/FR-I1, carry Tip201-0-related elements with 1-bp insertion and 1-bp substitution, respectively (Fig. 6A). All of these insertion sites in the F3′H gene are identical to that of Tip201-1 in KK/FR-35. Tip201-1 is a 39-bp deletion apparently derived from Tip201-0, and the deletion occurred between the 10-bp direct repeats within the element. The Tip201-0 elements carry imperfect TIRs of 24 bp, generate identical 8-bp target site duplications (TSDs) in the F3′H gene, and contain multiple copies of the sequence motif CGGTT in their subterminal regions: 23 copies within 200 bp at the 5′ end, 18 copies within 150 bp at the 3′ terminus (15 copies in Tip201-1), and two copies within TIRs (Fig. 6A). The Tip201 elements also carry two long A stretches and four inverted repeats between the A stretches, but no apparent open reading frames for putative transposase. Neither any data supporting the excision of the Tip201 elements from the F3′H gene nor sectors due to somatic reversions of the pink alleles could be detected, indicating that the integrated Tip201 elements are quite stable.
Both the characteristic TIR sequence and generation of 8-bp TSD indicate that Tip201 is a member of the Ac/Ds superfamily (Fig. 6B). In I. purpurea, the active autonomous DNA transposon Tip100 belonging to the Ac/Ds superfamily was identified as an insert in the CHS-D gene, and the somatic excision of Tip100 from the CHS-D gene was shown to be responsible for the flower variegation observed in KK/VR-37 and KK/VR-40 (Habu et al. 1998, Ishikawa et al. 2002). Both the TIR and internal sequences of Tip100 are quite different from those of Tip201. Based upon the structural and functional features, we conclude that Tip201 is a novel nonautonomous DNA transposon belonging to the Ac/Ds superfamily (Hoshino et al. 2001, Kunze and Weil 2002). Southern blot analysis revealed that at least 20 copies of Tip201-related elements are present in the I. nil and I. purpurea genomes (data not shown).
Integration of Tip201 can affect both the splicing and polyadenylation patterns of the F3′H transcripts in the pink mutants
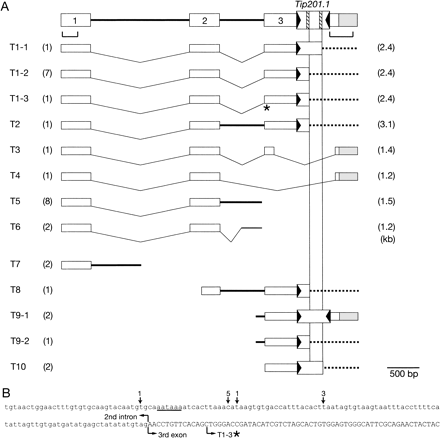
To examine the structures of the aberrant F3′H transcripts (mainly 1.5 kb and 2.4 kb) accumulated in the flower buds of the pink mutants (Fig. 5), we isolated and sequenced 30 F3′H cDNA clones from KK/FR-35, and the results are summarized in Fig. 7A. None of the cDNAs obtained could encode an intact F3′H enzyme, and the occurrence of shorter aberrant F3′H transcripts of about 1.5 kb was due to altered splicing and polyadenylation processes. The 3′ ends of the 10 clones were polyadenylated within intron 2, around 60 bp upstream of exon 3 (T5 and T6 in Fig. 7A). No A stretches that could anneal with the oligo(dT) primer for reverse transcription were present at the polyadenylated sites in intron 2, which excludes the possibility that their polyadenylations are artifacts generated by annealing of the oligo(dT) primer with A stretches in the precursor mRNA (pre-mRNA) molecules containing introns (Lorkovic et al. 2000, Reddy 2001). Moreover, a hexanucleotide, AATAAA, was found at 93 bp upstream of exon 3, and 9 out of the 10 cDNAs were polyadenylated within 11–33 bp downstream of the hexanucleotide sequence, AATAAA (Fig. 7B), which has been reported as a weak conserved near-upstream element acting as a polyadenylation signal in plants (Rothnie 1996, Li and Hunt 1997, Graber et al. 1999). The results suggest that at least one third of the transcripts was polyadenylated and terminated within the second intron.
Out of 20 remaining clones analyzed, 18 were polyadenylated within exon 3 (T1-T4 and T8-T10 in Fig. 7A). Four of them were terminated at the original polyadenylation sites of the F3′H gene, whereas the polyadenylation sites of 14 coincided with one of the two A stretches within Tip201. Since the longer aberrant F3′H transcripts of about 2.4 kb are about 0.5 kb longer than the wild-type F3′H transcripts, we assumed that they were normally spliced F3′H mRNAs carrying the Tip201-1 sequence and that the reverse transcription for the first strand cDNA synthesis had initiated by annealing the oligo(dT) primer with the A stretches in Tip201-1 of the mRNA templates. This assumption was confirmed by the fact that the 3′ half of the exon 3 sequence to the right of the Tip201-1 insertion site (indicated under the F3′H gene structure in Fig. 7A) was shown to be hybridizable with the 2.4-kb aberrant F3′H transcripts in the pink mutants by Northern blot analysis (Fig. 5C). The remaining two clones terminated in intron 1 (T7 in Fig. 7A) are likely to be artifacts by annealing the oligo(dT) primer with an A stretch in intron 1 of the mRNA templates.
A single bp insertion at exon 2 of the F3′H gene of Wedding Bells affects mRNA stability
To identify the fuchsia allele in the F3′H gene of Wedding Bells, we isolated its 8.3-kb EcoRI fragment and compared the F3′H gene sequence from Wedding Bells with that of the wild-type plant Heavenly Blue. Among the four differences found between the two lines, three were length polymorphisms of T or AT stretches within introns 1 and 2 or the 3′ untranslated region (data not shown). The remaining one alteration was caused by a single nucleotide insertion of T at position 82 bp upstream of the F3′H intron 2 in Wedding Bells, which resulted in changing the arginine codon AGA into the termination codon TAG and in producing a truncated F3′H peptide that lacks the C-terminal 241 amino acid region (Fig. 4A, B). The same mutation also abolishes the CTTAG DdeI site in the wild-type F3′H gene by changing into the sequence CTTTAG, which can be used to detect the mutation (Fig. 4B).
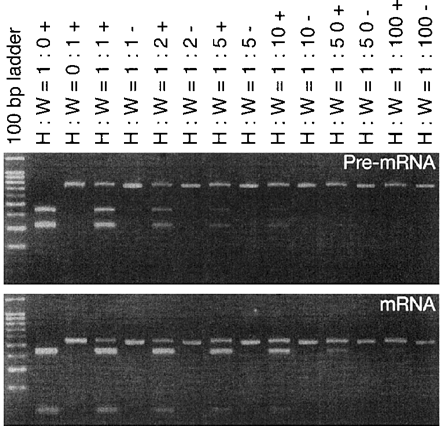
Since the newly created premature stop codon is located more than 50 bp upstream of the last exon-exon junction in the spliced F3′H mRNA in Wedding Bells, the occurrence of nonsense-mediated RNA decay (NMD) is likely to result in the reduction of F3′H mRNA accumulation, as shown in Fig. 5A (Schell et al. 2002). If this is the case, the accumulation of the pre-mRNA molecules containing introns in both Fuchsia and fuchsia plants will not be so different. Alternatively, the F3′H mRNA reduction in the mutant might be due to the reduction of its transcriptional initiation, and the pre-mRNA accumulation in the fuchsia mutants could be reduced consequently. To evaluate these possibilities, we examined the accumulation of pre-mRNAs containing introns (Lorkovic et al. 2000, Reddy 2001), which is considered to be reflected from the transcriptional initiation. To compare pre-mRNA accumulation between the wild-type and mutant flower buds, we performed competitive RT-PCR analysis using various mixtures of total RNAs from Heavenly Blue and Wedding Bells. The first-strand cDNAs synthesized from the mixtures were subjected to PCR amplification using the primers to be annealed with the first and second intron sequences, and the amplified fragments were cleaved with DdeI in order to distinguish the cDNAs derived from the pre-mRNAs in Wedding Bells from those in Heavenly Blue (Fig. 8A). We simultaneously examined the accumulation of spliced mRNA molecules by competitive RT-PCR analysis with the primers to be annealed with the first and third exon sequences (Fig. 8B). Although we noticed that the relative amounts of the pre-mRNAs in Wedding Bells were slightly lower than those in Heavenly Blue, the amounts of the spliced mRNAs in Wedding Bells were much more drastically reduced than those in Heavenly Blue. The results indicated that the major cause of the F3′H mRNA reduction is due to NMD triggered by the single base insertion into the second exon.
Discussion
Although various plants exhibiting reddish flowers are postulated to carry mutations controlling the F3′H activity (Forkmann 1994), none of them have been identified and only a few F3′H mutations identified are those affecting seed coloration. The tt7 mutation in Arabidopsis conferring pale brown seeds and reduced anthocyanin content to the whole plant is caused by a single C to T base transition generating the stop codon TAA (Schoenbohm et al. 2000), and the recessive t mutant in soybean affecting pigmentation in its seed coats and trichome hairs is a frameshift (a single C deletion) mutation (Toda et al. 2002, Zabala and Vodkin 2003). We describe here three spontaneous F3′H mutations that confer reddish flowers without affecting seed coloration in three Ipomoea species. The magenta and fuchsia alleles in I. nil and I. tricolor are a nonsense mutation (a single C to T base transition) and a single T insertion mutation, both of which generate the stop codons TGA and TAG in the F3′H gene, respectively (Fig. 4). We examined 16 different I. nil mutants with various genetic backgrounds, which display reddish flowers and/or produce pelargonidin-based pigments and/or carry genetically characterized magenta mutations, and found that all of them carry an identical nonsense mutation. Since most of the spontaneous mutations identified in I. nil are caused by the insertion of Tpn1 and its relatives (Iida et al. 1999, Hoshino et al. 2001, Y. Morita and A. Hoshino unpublished results), the magenta mutation described here is an exceptional point mutation. Probably, the magenta mutation is one of the early spontaneous mutations isolated before the massive generation of floriculturally important traits by spontaneous transposon mutagenesis with the Tpn1-related elements in the early 19th century (Imai 1927b, Iida et al. 1999), since an I. nil mutant with red flowers was documented in an early floricultural book in Japan (Kafu by Ekiken Kaibara 1695).
We also identified the pink mutations of I. purpurea as insertion mutations of the Tip201 elements in the F3′H gene (Fig. 4). The Tip201 elements carry two long A stretches where the sequence alterations occurred frequently, and Tip201-1 must derive from its parental Tip201-0 element by deletion of 39 bp (Fig. 6A). A mutant of I. purpurea displaying red flowers was first recorded by Curtis (1790), and recessive mutations conferring red flowers were first studied by Barker (1917), also used by Imai and his collaborator (Imai 1927a, Imai and Tabuchi 1935), and characterized later by Clegg and his collaborators who obtained their mutants from natural populations in Georgia (Ennos and Clegg 1983, Schoen et al. 1984). The lines KK/FR-35, KK/VR-37, and KK/VR-40 are direct descendants of the mutants used by Imai, and the lines CR/FR-I1 and CR/FR-L1 are the authentic pink mutants obtained from Clegg. Because all of them actually carry the same mutation, these pink mutations are likely to have originated from the founder allele. The divergence appeared to occur primarily within Tip201, and no nucleotide polymorphism other than Tip201 could be detected in the 4.4-kb F3′H gene regions between primers IPF3′H-E1F1 and IPF3′H-E3R1 used in KK/FR-34, KK/FR-35, KK/VR-37, KK/VR-40, and KK/WR-321 (Fig. 4A).
The two major F3′H transcripts of 2.4 kb and 1.5 kb are accumulated in the flower buds of the pink mutants (Fig. 5). The 2.4-kb transcripts are regarded to be the F3′H mRNAs containing the intact Tip201-1 sequence, whereas around 1.5-kb molecules contain aberrantly spliced mRNAs and transcripts with an unspliced intron 2 that is anomaly polyadenylated (T3 and T5 in Fig. 7A). Integration of the AT-rich Tip201-1 element into exon 3 is likely to modulate exonic splicing enhancers residing within exon 3 to lead to mRNAs containing intron 2 (Lorkovic et al. 2000, Reddy 2001). Consequently, the polyadenylation machinery recognizes the hexanucleotide AATAAA 93 bp upstream of exon 3 as a near-upstream element for polyadenylation (Rothnie 1996, Li and Hunt 1997, Graber et al. 1999), and polyadenylation at 11–33 bp downstream of the hexanucleotide sequence AATAAA produces the 1.5-kb transcripts T5 in Fig. 7A. Although Tip201-1 carries the 39-bp internal deletion, the sequence deleted probably does not affect the splicing and polyadenylation patterns of the F3′H transcripts because both 2.4-kb and 1.5-kb mRNAs were detected as two major transcripts from the F3′H gene containing Tip201-0 in KK/VR-40 (Fig. 5A). Integration of a retrotransposon leading to create an additional polyadenylation site at 5′ upstream of its insertion site was also reported in the wxG allele of maize (Marillonnet and Wessler 1997). The accumulation of the F3′H transcripts is significantly reduced in the fuchsia mutant of I. tricolor, whereas no reduction of the F3′H mRNAs was observed in the magenta mutant of I. nil (Fig. 5A). The difference in these mRNA stabilities is due to the positions of the newly generated termination codons in these mutations: exon 2 in the fuchsia mutant and exon 3 in the magenta mutant. Thus, NMD occurs only in the fuchsia mutant of I. tricolor because of the premature translational termination at the termination codon more than 50 bp upstream of the junction of the second and third exons in a spliced mRNA (Schell et al. 2002).
Both I. nil and I. purpurea belong to the same subgenera Ipomoea, and I. tricolor is in a different subgenera, Quamoclit (Austin and Huáman 1996). Indeed, amino acid sequences of the F3′H enzymes characterized here confirm the notion that I. tricolor is distantly related to I. nil and I. purpurea among the three morning glories. However, both I. tricolor and I. nil produce the same anthocyanin (HBA) as a major flower pigment, while I. purpurea gives a slightly different anthocyanin that lacks one glucose molecule and a methyl residue (Fig. 2B), suggesting that certain genes for glycosyltransferase and methyltransferase for HBA production are inactivated. Comparison of their genomic F3′H gene regions revealed that various deletions and insertions occurred in the intron regions (Fig. 4A), indicating that these genes underwent structural divergence without changing their function. Only minor portions of the intron regions including the dMELS6-8 element retain the homologous sequences, even though these introns contain the various cis elements essential for efficient splicing (Lorkovic et al. 2000, Reddy 2001). It would be interesting to see whether the I. purpurea genes for glycosyltransferase and methyltransferase responsible for production of the flower pigment HBA are inactivated by the processes associated with the generation of the intron divergence or point mutations found in the F3′H genes in I. nil and I. tricolor. Similarly, we are wondering whether the three Ipomoea species we examined carry inactivated F3′5′H genes for the purple delphinidin biosynthesis, since no delphinidin-derived anthocyanins can be detected in their blue or purple flowers. Such analyses would shed light on evolutionary processes and diversity of floral pigmentation variation in the morning glories (Clegg and Durbin 2003). In addition, the F3′H mutations identified here provide PCR-based molecular markers for diagnosing the genotypes of the F3′H genes in cotyledons, which will serve for breeding flower color varieties in I. nil, I. purpurea and I. tricolor.
Materials and Methods
Plant materials
The cultivars of I. nil, Scarlet O’Hara, Dainichi-rin, Sun Smile Red, Yaguruma, and Akatsuki-no-beni, exhibit reddish flowers, and their flower pigments have been described before (Lu et al. 1992a). The red flower pigments of the cultivar Violet (Imamura 1967, Vince-Prue and Gressel 1985) have also been analyzed (Toki et al. 2001a). The I. nil cv. Hama-no-kagayaki displays reddish flowers (Fig. 1B). The I. nil lines Danjuro, which displays maroon (dusky red) flowers, Q853 (duskish-1), Q854 (duskish-2), and Q857 (dingy) are thought to carry the magenta allele genetically, and their flower pigments have been described before (Saito et al. 1994, Toki et al. 2001b). All of these plants listed above contain pelargonidin-based pigments. The I. nil lines Akatsuki-no-mai, Q173, and Q265-2 (Beni-chidori) also display reddish flowers. The I. nil lines 54Y and 78WWc-1 have pale yellow and white flowers due to the mutations speckled and c-1, respectively (Abe et al. 1997, Hoshino et al. 1997). Both 54Y and 78WWc-1 carry the authentic magenta allele studied by Hagiwara (1931), and their F1 hybrids display red flowers (Abe et al. 1997) that contain pelargonidin-based pigments (N. Saito et al. unpublished results). The I. nil lines KK/ZSK-2 and Hama-no-sora, which display blue flowers (Abe et al. 1997, Hoshino et al. 1997), were used as control plants carrying the wild-type Magenta allele. The I. purpurea lines KK/FP-36 and YO/FP-39 carry the wild-type Pink allele, whereas YO/FR-34, KK/FR-35, KK/VR-37, KK/VR-40, and KK/WR-321 bear the pink allele (Habu et al. 1998). The authentic pink mutants CR/FR-I1 (AAppWWII) and CR/FR-L1 (AAppWWii), which display dark pink and light pink flowers, respectively (Epperson and Clegg 1988), were provided by M. Durbin and M. Clegg (UC Riverside, CA, U.S.A.). The I. purpurea line KK/WRSP-1 displays white flowers with pigmented spots in the rays; they were designated as “spotted flowers” by Imai (1927a). The mutation conferring the spotted flowers was named u by Imai (1927a) and was later independently called w by Ennos and Clegg (1983). The gene U or W in I. purpurea corresponds to the C-1 gene in I. nil and encodes a MYB transcriptional activator for anthocyanin biosynthesis (Y. Morita et al. unpublished results, M. Rausher personal communication). The I. tricolor cv. Heavenly Blue and Wedding Bells, which produce blue and reddish purple flowers, respectively, were obtained from Hem Zaden (Hemmerbuurt, Holland) and David Dubowski (Chapel Hill, NC, U.S.A.), respectively. All plants were grown in a pollinator-free greenhouse.
Co-segregation analysis of the pink allele and flower phenotypes in I. purpurea
An F1 plant producing dark purple flowers was obtained by crossing the wild-type line YO/FP-39 with the pink mutant KK/FR-35, and selfed progeny of the F1 plant were scored for their flower phenotypes and their F3′H gene structures using PCR analysis, as described below. Out of 70 F2 plants examined, 20, 33, and 17 plants were found to be homozygotes with the wild-type allele producing purple flowers, heterozygotes with purple flowers, and homozygotes with red flowers carrying the Tip201-inserted F3′H gene, respectively.
Preparation and characterization of nucleic acids
General nucleic acid procedures, including plant DNA and RNA preparation, Southern and Northern blot analyses, and DNA sequencing analysis, were performed as described before (Habu et al. 1998, Yamaguchi et al. 2001).
All of the cDNA libraries used were prepared from young flower buds. The cDNA library from the I. nil line KK/ZSK-2 was screened with the petunia F3′H cDNA clone (Brugliera et al. 1999), and the isolated I. nil cDNA was used as a probe for screening of the cDNA libraries from the I. purpurea lines KK/FR-35 and YO/FP-39 and from I. tricolor cv. Heavenly Blue. The F3′H cDNA of the magenta mutant Hama-no-kagayaki was obtained by RT-PCR amplification using the primers IF3′H-E1F1 (5′-AGTATGGCTACCTTAACC-3′) and INF3′H-E3R1 (5′-ACAACAAATACACTGGGTTTACC-3′). The reaction was carried out for the initial denaturation at 94°C for 5 min, 30 cycles consisting of denaturation (94°C for 30 s), annealing (58°C for 30 s), and extension (72°C for 2 min). To determine the transcriptional initiation sites for the F3′H genes in I. nil and I. purpurea, we employed 5′-RACE using the 5′-RACE System for Rapid Amplification of cDNA Ends, Version 2.0 (Invitrogen, Carlsbad, CA, U.S.A.). The primers used for 5′RACE were IN-GSP1 (5′-GCCGTTGGAGAAGTTCGAG-3′); IP-GSP1 (5′-GCCGTTGGAGAAGTTCGCG-3′); Common-GSP2 (5′-CAAGAACTGAGCGGCCACGG-3′); IN-GSP3 (5′-CGGCCACAACCACGTCCACG-3′); and IP-GSP3 (5′-CGGCCACAACCACATCCACG-3′). We used GeneRacer (Invitrogen) to determine the transcriptional initiation sites for the F3′H genes in I. tricolor. The primer and subsequent nested primer used were IT-GSP1 (5′-TCAGTCCCAGCTGTGAACAA-3′) and IT-GSP2 (5′-GTTCGTGGTGCACACGTTCA-3′), respectively. For Northern blot analysis, either I. nil or I. purpureaF3′H cDNA was generally used as a probe. The probe for exon 1 in I. purpurea was prepared by PCR amplification with primers IF3′H-E1F2 (5′-ATGGCTACCTTAACCTTAATTTTCTGC-3′) and IF3′H-E1R1 (5′-GCCGGTTGGAGAAGTTCGCG-3′), and the probe for 3′ part of exon 3 was prepared by PCR amplification with primers IF3′H-E2F2 (5′-GGTTCACCTGTTGGTTGCAACTTTGG-3′) and IPF3′H-E3R1 (5′-CCTTGGTAGTCTTCATGAGGTGTTTCGGAC-3′) (Fig. 5, 7).
Characterization of the genomic F3′H gene regions
Using the I. nil F3′H cDNA as a probe, the HindIII fragments of the genomic F3′H gene regions from I. nil and I. purpurea plants were cloned into λZAP Express, and the clones were subsequently converted into pBK-CMV phagemid derivatives by in vivo excision (Stratagene, La Jolla, CA, U.S.A.). From the I. nil line KK/ZSK-2, the 5.5-kb, 3.7-kb, and 3.3-kb fragments containing exon 1, exon 2, and a 3′ part of exon 3, respectively, were isolated (Fig. 4A). The 1.3-kb segment of intron1 containing two HindIII sites was obtained by PCR amplification with primers INF3′H-I1F1 (5′-TGTAAATAGCCTAAGCCACG-3′) and INF3′H-I1R1 (5′-AAGGTACATGCATGCACGTC-3′), and the obtained fragment was sequenced without further cloning. The 0.4-kb segment of exon 3 containing a HindIII site was amplified by PCR with primers IF3′H-E3F1 (5′-GGAGACCTTCAGGCTACACC-3′) and IF3′H-E3R1 (5′-GATGTGGTTGTAATCTGGGC-3′), and the amplified products were cloned into pGEM-T Easy vector (Promega, Madison, WI, U.S.A.). Since the 0.4-kb segment of exon 3 also contains the magenta mutation (Fig. 4A), we also sequenced the segments from the other 13 I. nil lines that include 54Y and 78WWc-1 carrying the authentic magenta mutation as well as Hama-no-kagayaki.
The 8.5-kb fragment containing the major part of the F3′H gene and the 1.0-kb fragment from the 3′ part of exon 3 were isolated from the wild-type I. purpurea line YO/FP-39 (Fig. 4A) using the I. nil F3′H cDNA as a probe, and the 9.0-kb fragment containing Tip201-1 and the 1.0-kb fragment were obtained from the pink mutant KK/FR-35. The genomic segments of the F3′H gene region of 4.4 kb from the I. purpurea lines, YO/FR-34, KK/VR-37, KK/VR-40, KK/WR-321, CR/FR-I1, and CR/FR-L1, were obtained by LA-PCR amplification using LA Taq polymerase (Takara Biomedicals, Otsu, Japan) with primers IPF3′H-E1F1 (5′-AACAACGTTACCCTTTACCTCTCCCACCCG-3′) and IPF3′H-E3R1. The LA-PCR reaction was performed as follows: initial denaturation at 94°C for 1 min, 14 cycles of denaturation (94°C for 10 s), annealing and extension (68°C for 15 min), and then 16 cycles of denaturation (98°C for 10 s) and annealing and extension (68°C for 15 min with the autoextension feature to add 15 s per cycle). The LA-PCR-amplified fragments were cloned into pCRII vector (Invitrogen). A slightly modified LA-PCR reaction conditions (annealing and extension at 60°C for 5 min) were used with the primers IF3′H-E3F1 and IF3′H-E3R1 in order to distinguish the wild-type Pink allele and the mutant pink allele having Tip201-1 inserted in the co-segregation analysis described above. Nearly equal amounts of the 0.45-kb fragment from the Pink allele and the 1.0-kb bands from the pink allele could be detected under the reaction condition used.
The presence of two long A stretches within the Tip201 elements created difficulty in sequencing the elements. To sequence the Tip201 elements precisely, we employed the restriction sites AccI, NcoI, and PstI, which reside in the segment between the two long A stretches. For sequencing Tip201-1 from KK/FR-35, two pBK-CMV phagemid derivatives carrying the 9.0-kb fragment containing Tip201-1 in a different orientation were cleaved with PstI and self-ligated to generate the clones with either end of Tip201-1. For sequencing the Tip201-0 elements, two sets of pCRII derivatives carrying the 4.4-kb PCR-amplified fragments containing Tip201-0 in a different orientation were cleaved with AccI and self-ligated to generate the clones with either end of Tip201-1. Using the resulting clones, we sequenced either the left or right half part of Tip201 flanking a single copy of the A stretch. To determine the A stretch sequences of Tip201, segments of about 1.0 kb containing the Tip201 elements were amplified by PCR with primers IF3′H-E3F1 and IF3′H-E3R1, and the amplified fragments were cleaved with NcoI, isolated an appropriate fragment and sequenced without further cloning.
We also cloned the 8.3-kb EcoRI fragments containing the entire F3′H gene of I. tricolor cv. Heavenly Blue and Wedding Bells into λZAP Express.
Competitive RT-PCR analysis of the F3′H transcripts in I. tricolor
Total RNAs (2.5 µg) from Wedding Bells were mixed with different amounts of total RNAs (2.5 µg, 1.25 µg, 0.5 µg, 0.25 µg, 0.05 µg, and 0.025 µg) from Heavenly Blue separately, and first-strand cDNA synthesis was performed in the individual mixtures by SuperScriptII Reverse Transcriptase Kit (Invitrogen). The PCR reaction was carried out for initial denaturation at 94°C for 3 min, 30 cycles of denaturation (94°C for 30 s), annealing (60°C for 30 s), and extension (72°C for 2 min). The primers for the premature F3′H mRNAs were ITF3′H-I1F1 (5′-TAATTGCTCAATTAACTCAAGCTT-3′) in intron 1 and ITF3′H-I2R1 (5′-TGCGCGTGTGCATATGTATC-3′) in intron 2, and the primers for the spliced F3′H mRNAs were ITF3′H-E1F1 (5′-GGCCAAGGCGTTGGATGACT-3′) in exon 1 and IT-GSP1 (see 5′-RACE mentioned above) in exon 3. Total RNAs (5.0 µg) from either Heavenly Blue alone or Wedding Bells alone were also treated in the same way as controls. Ten µl of the amplified fragments was cleaved with DdeI and separated by 2.0% agarose gel.
Acknowledgments
We thank Miwako Matsumoto and Kyoko Ikegaya for their technical assistance, Filippa Brugliera for providing the petunia F3′H cDNA, Eiji Nitasaka, David Dubowski, Mary Durbin, Michael Clegg, and Kichiji Kasahara for the Ipomoea seeds, and Kyeung-Il Park for discussions. We also thank Yasuhiro Kikuchi and Yoshiki Habu for their suggestion at the initial stage of this study, and Mark Rausher for sharing unpublished results. This work was supported in part by grants from the Ministry of Education, Culture, Sports, Science and Technology in Japan.
These authors contributed equally to this work.
Corresponding author: E-mail, shigiida@nibb.ac.jp; Fax, +81-564-55-7685.
Fig. 1 Flower variation in Ipomoea. I. nil: (A) Hama-no-sora, (B) Hama-no-kagayaki. I. purpurea: (C) YO/FP-39, (D) KK/FR-35. I. tricolor: (E) Heavenly Blue, (F) Wedding Bells.
Fig. 2 Simplified pathway for anthocyanin biosynthesis (A) and structures of anthocyanins in the petals of morning glories (B). The wild-type blue flowers of I. nil and I. tricolor contain peonidin derivatives (R = OCH3) named Heavenly Blue Anthocyanin (HBA) after I. tricolor cv. Heavenly Blue and the wild-type dark purple flowers of I. purpurea contain cyanidin derivatives (R = OH). The reddish flowers of their F3′H mutants contain pelargonidin derivatives (R = H). It would be more appropriate to refer to the major flower pigment as Wedding Bells Anthocyanin (WBA) than as Pharbitis red anthocyanin 5 (PRA-5) because I. tricolor cv. Wedding Bells contains virtually only WBA in the flower petals (see Fig. 3).
Fig. 3 HPLC analysis of the flower pigments in Wedding Bells. The major peak contains pelargonidin 3-O-(2-O-(6-O-(3-O-glucosyl-caffeoyl)-glucosyl)-6-O-(4-O-(6-O-(3-O-glucosyl-caffeoyl)-glucosyl)-caffeoyl)-glucoside)-5-O-(glucoside). The identification of the pigments was performed as described previously (Lu et al. 1992a, Toki et al. 2001a).
Fig. 4 Genomic structures of the F3′H gene regions. (A) Comparison of the genome structures of the F3′H gene regions in Ipomoea. The horizontal boxes indicate the sequenced genomic F3′H segments. The open boxes with the numerals represent exons, and the shadowed parts within the exon 3 boxes indicate the 3′ untranslated regions. The transcription starts at 11 or 12 bp upstream of the ATG initiation codon in all three F3′H genes. The rectangles with filled arrowhead(s) indicate the MELS elements. The hatched boxes indicate the repetitive sequences Rep.E and Rep.F. Rep.E found within the MELS9-1[J] box showed significant homology to the segment found in the second intron of both the InNHX1 gene and the InNHX1 pseudogene, and Rep.F also resides at the 5′ upstream region of the mgsdcg1 (S-adenosylmethionine decarboxylase gene of I. nil) gene (Accession No. U64927). The other MELS and Rep elements were described before (Hoshino et al. 2001). The inverted and direct repeats are represented by open arrowheads. The brackets and horizontal bars under the genomic structures with numerals represent non-homologous regions with and without short direct repeats of 2–30 bp at both ends, respectively. The symbols N/P and N/T indicate the I. nil segments heterologous to I. purpurea and I. tricolor regions, respectively, and other symbols (P/N, P/T, T/N, and T/P) conform to the same designation system. The vertical arrows with C to T, Tip201, and T indicate the single base transition site in the magenta mutants, the Tip201 insertion site in the pink mutants, and the single bp insertion site in Wedding Bells, respectively. The filled vertical arrowhead with 9 bp at the 3′ end of exon 2 in I. tricolor indicates the site for three additional amino acids. Only the HindIII restriction sites used for cloning, indicated by the small open arrowheads under the I. nil and I. purpurea genomic structures, are shown. The filled horizontal arrowheads with numerals under the genomic structures represent the site and orientation of the primers used. The primers are 1, IF3′H-E1F1; 2, INF3′H-I1F1; 3, INF3′H-I1R1; 4, IF3′H-E3F1; 5, IF3′H-E3R1; 6, INF3′H-E3R1; 7, IPF3′H-E1F1; 8, IPF3′H-E3R1; 9, ITF3′H-E1F1; 10, ITF3′H-I1F1; 11, ITF3′H-I2R1; 12, IT-GSP1. (B) Sequence alterations in the magenta and fuchsia mutations. In the magenta allele, a single C to T base transition results in generation of the TGA stop codon in exon 3. In the fuchsia allele, a single T insertion generates the TAG stop codon in exon 2. The altered nucleotides are printed in bold, and the DdeI site is underlined. (C) Junction sequences of the MELS9-1[J] insertion site in I. nil and its corresponding sequence in I. purpurea. The TSDs are underlined, and the divergent 3-bp sequences are shown in italics. The uppercase letters marked with horizontal arrows represent 7-bp TIRs.
Fig. 5 The F3′H transcripts accumulated in the Ipomoea flower buds. (A) Various Ipomoea lines. RNAs from floral tissues were obtained at 2 d before flower opening. The total RNAs (10 µg) obtained are (lane 1) KK/ZSK-2; (lane 2) Hama-no-sora; (lane 3) Hama-no-kagayaki; (lane 4) 54Y; (lane 5) 78WWc-1; (lane 6) KK/FR-35; (lane 7) KK/FP-36; (lane 8) YO/FP-39; (lane 9) KK/VR-40; (lane 10) Heavenly Blue; (lane 11) Wedding Bells. The probes used were I. nil F3′H cDNA (lanes 1–5); I. purpurea F3′H cDNA (lanes 6–11). The flower phenotypes blue, purple, reddish, pale yellow, white, and white with pigmented rays are indicated by B, P, R, Y, W, and W*, respectively. The plants carrying the wild-type and mutant F3′H genes are indicated by + and –, respectively. (B) Characterization of the I. purpureaF3′H transcripts in line KK/WRSP-1. Total RNAs (10 µg) obtained are (lane 1) YO/FP-39; (lane 2) KK/FR-35; (lane 3) KK/WRSP-1, whole floral limb; (lane 4) KK/WRSP-1, limb removing pigmented rays; (lane 5) KK/WRSP-1, pigmented rays. Both I. nil line 78WWc-1 and I. purpurea mutant KK/WRSP-1 bear the mutations in the genes encoding MYB transcriptional activators for anthocyanin biosynthesis. Since KK/WRSP-1 displays white flowers with pigmented spots in the rays, RNAs were prepared from whole limbs including pigmented rays, limbs from which the pigmented spots were removed, and pigmented rays alone. The probe used was I. purpurea F3′H cDNA. (C) Characterization of the I. purpureaF3′H transcripts in the pink mutant line KK/FR-35. Total RNAs (10 µg) obtained are (lanes 1 and 3) YO/FP-39; (lanes 2 and 4) KK/FR-35. The probes used were I. purpurea F3′H exon 1 (lanes 1 and 2); 3′ part of I. purpurea F3′H exon 3 (lanes 3 and 4). The segments of these probes are shown in Fig. 7A.
Fig. 6 Structure of Tip201. (A) Sequence of the Tip201 elements and their flanking regions. The 24-bp imperfect TIRs are indicated by large horizontal arrows under the Tip201 sequences. The four inverted repeats between the A stretch sequences are indicated by the horizontal arrows above the sequence, and the CGGTT motifs are indicated by the broken horizontal arrows. The 8-bp TSDs flanking the Tip201 elements are boxed, and the 10-bp direct repeats within Tip201-0 are surrounded by the broken boxes. Nucleotides identical to those of Tip201-0.A are marked with a colon (:), and deleted nucleotides are marked with a hyphen (–). Tip201-0.A is the longest Tip201-0 element of 552 bp found in YO/FR-34, and Tip201-0.B of 551 bp was found in KK/VR-37, KK/VR-40, KK/WR-321, and CR/FR-L1. Tip201-0.C of 551 bp was obtained from CR/FR-I1, and Tip201-1 of 512 bp was obtained from KK/FR-35. (B) Alignment of TIRs of Tip201, Tip100, and Ac. Nucleotide sequences that are not identical between the right and left TIRs are shown by lowercase letters. The sequences of Tip100 and Ac were taken from the literature (Kunze and Weil 2002).
Fig. 7 Characterization of the F3′H transcripts in a pink mutant. (A) Structure of the F3′H cDNAs from KK/FR-35. The genomic structure of the F3′H gene having Tip201-1 integrated in the pink mutant KK/FR-35 is drawn at the top, and the exons are indicated by open boxes with numerals. The brackets under the exons indicate the segments used as probes for Northern blot analysis in Fig. 5C. The two hatched boxes within the Tip201-1 box represent the two A stretches, and the two vertical lines represent the positions corresponding to these A stretches. Among the 30 cDNAs characterized, the molecules T7-T10 probably do not represent the mRNAs in the young flower buds: first-strand cDNA synthesis started at an A stretch in intron 1 (T7) or prematurely stopped at the indicated sites (T8-T10). As indicated by the broken horizontal lines, the T1 and T2 transcripts are assumed to terminate at the same site as the wild-type F3′H transcripts, and the first-strand cDNA synthesis started at one of the two A stretches within Tip201-1. The numerals in the parentheses at the left and right columns represent the numbers of the clones examined and the expected sizes of the transcripts. (B) Sequence at the junction region of the intron 2 (lowercase letters) and exon 3 (uppercase letters). The underline indicates a putative polyadenylation signal, and numerals indicate the numbers of the obtained clones polyadenylated at the sites pointed by the vertical arrows. The asterisk indicates the 5′ end of exon 3 in the T1-3 cDNA.
Fig. 8 Analysis of precursor and spliced transcripts from the F3′H gene in Heavenly Blue to Wedding Bells. The leftmost lanes are size markers (100-bp DNA ladder) purchased from Promega (Madison, WI, U.S.A.). The relative amounts of total RNAs from Heavenly Blue to Wedding Bells in the reaction mixtures used are indicated above the lanes. The other symbols are (H) Heavenly Blue; (W) Wedding Bells; (+) cleaved with DdeI; (–) uncleaved.
Abbreviations
- ANS
anthocyanidin synthase
- CHI
chalcone isomerase
- CHS
chalcone synthase
- DFR
dihydroflavonol 4-reductase
- F3H
flavanone 3-hydroxylase
- F3′5′H
flavonoid 3′,5′-hydroxylase
- F3′H
flavonoid 3′-hydroxylase
- HBA
Heavenly Blue Anthocyanin
- MELS
mobile element-like sequence
- NMD
nonsense-mediated RNA decay
- pre-mRNA
precursor mRNA
- TIR
terminal inverted repeat
- TSD
target site duplication
- UF3GT
UDP-glucose:flavonoid 3-O-glycosyltransferase.
The nucleotide sequences reported in this paper have been deposited to DDBJ, EMBL, and GenBank under accession numbers AB113261 to AB113272 (F3′H mRNA of I. nil, F3′H mRNA of I. purpurea, F3′H mRNA of I. tricolor, F3′H gene of I. nil, F3′H gene of I. purpurea, F3′H gene from the pink mutant of I. purpurea, F3′H gene of I. tricolor, F3′H gene from the fuchsia mutant of I. tricolor, Tip201-0.A, Tip201-0.B, Tip201-0.C, and Tip201-1, respectively).
References
Abe, Y., Hoshino, A. and Iida, S. (
Austin, D.F. and Huáman, Z. (
Barker, E.E. (
Brugliera, F., Barri-Rewell, G., Holton, T.A. and Mason, J.G. (
Chapple, C. (
Clegg, M.T. and Durbin, M.L. (
Ennos, R.A. and Clegg, M.T. (
Epperson, B.K. and Clegg, M.T. (
Feschotte, C., Zhang, X. and Wessler, S.R. (
Forkmann, G. (
Fukada-Tanaka, S., Hoshino, A., Hisatomi, Y., Habu, Y., Hasebe, M. and Iida, S. (
Fukada-Tanaka, S., Inagaki, Y., Yamaguchi, T., Saito, N. and Iida, S. (
Graber, J.H., Cantor, C.R., Mohr, S.C. and Smith, T.F. (
Habu, Y., Hisatomi, Y. and Iida, S. (
Hagiwara, T. (
Holton, T.A. and Cornish, E.C. (
Hoshino, A., Abe, Y., Saito, N., Inagaki, Y. and Iida, S. (
Hoshino, A., Johzuka-Hisatomi, Y. and Iida, S. (
Iida, S., Hoshino, A., Johzuka-Hisatomi, Y., Habu, Y. and Inagaki, Y. (
Imai, Y. (
Imai, Y. and Tabuchi, K. (
Imamura, S. (
Ishikawa, N., Johzuka-Hisatomi, Y., Sugita, K., Ebinuma, H. and Iida, S. (
Kondo, T., Kawai, T., Tamura, H. and Goto, T. (
Kunze, R. and Weil, C.F. (
Lorkovic, Z.J., Wieczorek Kirk, D.A., Lambermon, M.H.L. and Filipowicz, W. (
Lu, T.S., Saito, N., Yokoi, M., Shigihara, A. and Honoda, T. (
Lu, T.S., Saito, N., Yokoi, M., Shigihara, A. and Honoda, T. (
Marillonnet, S. and Wessler, S.R. (
Mol, J., Grotewold, E. and Koes, R. (
Quattrocchio, F., Wing, J., van der Woude, K., Souer, E., de Vetten, N., Mol, J. and Koes, R. (
Saito, N., Lu, T.S., Akaizawa, M., Yokoi, M., Shigihara, A. and Honda, T. (
Saito, N., Tatsuzawa, F., Yoda, K., Yokoi, M., Kasahara, K., Iida, S., Shigihara, A. and Honda, T. (
Saito, N., Tatsuzawa, F., Yokoi, M., Kasahara, K., Iida, S., Shigihara, A. and Honda, T. (
Schell, T., Kulozik, A.E. and Hentze, M.W. (
Schoen, D.J., Giannasi, D.E., Ennos, R.A. and Clegg, M.T. (
Schoenbohm, C., Martens, S., Eder, C., Forkmann, G. and Weisshaar, B. (
Shirley, B.W. (
Tiffin, P., Miller, R.E. and Rausher, M.D. (
Toda, K., Yang, D., Yamanaka, N., Watanabe, S., Harada, K. and Takahashi, R. (
Toki, K., Saito, N., Iida, S., Hoshino, A., Shigihara, A. and Honda, T. (
Toki, K., Saito, N., Iida, S., Hoshino, A., Shigihara, A. and Honda, T. (
Vince-Prue, D. and Gressel, J. (
Yamaguchi, T., Fukada-Tanaka, S., Inagaki, Y., Saito, N., Yonekura-Sakakibara, K., Tanaka, Y., Kusumi, T. and Iida, S. (




![Fig. 4 Genomic structures of the F3′H gene regions. (A) Comparison of the genome structures of the F3′H gene regions in Ipomoea. The horizontal boxes indicate the sequenced genomic F3′H segments. The open boxes with the numerals represent exons, and the shadowed parts within the exon 3 boxes indicate the 3′ untranslated regions. The transcription starts at 11 or 12 bp upstream of the ATG initiation codon in all three F3′H genes. The rectangles with filled arrowhead(s) indicate the MELS elements. The hatched boxes indicate the repetitive sequences Rep.E and Rep.F. Rep.E found within the MELS9-1[J] box showed significant homology to the segment found in the second intron of both the InNHX1 gene and the InNHX1 pseudogene, and Rep.F also resides at the 5′ upstream region of the mgsdcg1 (S-adenosylmethionine decarboxylase gene of I. nil) gene (Accession No. U64927). The other MELS and Rep elements were described before (Hoshino et al. 2001). The inverted and direct repeats are represented by open arrowheads. The brackets and horizontal bars under the genomic structures with numerals represent non-homologous regions with and without short direct repeats of 2–30 bp at both ends, respectively. The symbols N/P and N/T indicate the I. nil segments heterologous to I. purpurea and I. tricolor regions, respectively, and other symbols (P/N, P/T, T/N, and T/P) conform to the same designation system. The vertical arrows with C to T, Tip201, and T indicate the single base transition site in the magenta mutants, the Tip201 insertion site in the pink mutants, and the single bp insertion site in Wedding Bells, respectively. The filled vertical arrowhead with 9 bp at the 3′ end of exon 2 in I. tricolor indicates the site for three additional amino acids. Only the HindIII restriction sites used for cloning, indicated by the small open arrowheads under the I. nil and I. purpurea genomic structures, are shown. The filled horizontal arrowheads with numerals under the genomic structures represent the site and orientation of the primers used. The primers are 1, IF3′H-E1F1; 2, INF3′H-I1F1; 3, INF3′H-I1R1; 4, IF3′H-E3F1; 5, IF3′H-E3R1; 6, INF3′H-E3R1; 7, IPF3′H-E1F1; 8, IPF3′H-E3R1; 9, ITF3′H-E1F1; 10, ITF3′H-I1F1; 11, ITF3′H-I2R1; 12, IT-GSP1. (B) Sequence alterations in the magenta and fuchsia mutations. In the magenta allele, a single C to T base transition results in generation of the TGA stop codon in exon 3. In the fuchsia allele, a single T insertion generates the TAG stop codon in exon 2. The altered nucleotides are printed in bold, and the DdeI site is underlined. (C) Junction sequences of the MELS9-1[J] insertion site in I. nil and its corresponding sequence in I. purpurea. The TSDs are underlined, and the divergent 3-bp sequences are shown in italics. The uppercase letters marked with horizontal arrows represent 7-bp TIRs.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/pcp/44/10/10.1093_pcp_pcg143/2/m_pcg14304.gif?Expires=1716394053&Signature=PpI~sMhkYkVb5lML7AA3An18aUq8LqzvfCaTwj3sY~ob1cTFLQRKJGdm9qyUVC3I4XZU3jj6c-qOT4fLlOWuMjz5ZBfcAUhpkf~mzh3Grk7s-kkscyRuSTsxiq~qIM~-yxznjW4rrnE~wN3g1GV-ijjskQj0IgM~~5HstTMYmpqQQVRqOHbEjGoLgYrY2sG0kfI9iHR0bLNxMje3bbaR~pQ2D8zKBXcQZ5xYyUwAfaMuyZjutQ8lpM1YJgnXIhDm8cnvZJItUEbHX5j3Lhba2xM9NDfHUE~adQjMsASpL-0k8tcrFygPeR3JwNoD4IAflPfiFBowO1l7hRpU3k0jow__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)