-
PDF
- Split View
-
Views
-
Cite
Cite
Jessica L. Metcalf, Matthew R. Siegle, Andrew P. Martin, Hybridization Dynamics between Colorado's Native Cutthroat Trout and Introduced Rainbow Trout, Journal of Heredity, Volume 99, Issue 2, March/April 2008, Pages 149–156, https://doi.org/10.1093/jhered/esm118
Close - Share Icon Share
Abstract
Newly formed hybrid populations provide an opportunity to examine the initial consequences of secondary contact between species and identify genetic patterns that may be important early in the evolution of hybrid inviability. Widespread introductions of rainbow trout (Oncorhynchus mykiss) into watersheds with native cutthroat trout (Oncorhynchus clarkii) have resulted in hybridization. These introductions have contributed to the decline of native cutthroat trout populations. Here, we examine the pattern of hybridization between introduced rainbow trout and 2 populations of cutthroat trout native to Colorado. For this study, we utilized 7 diagnostic, codominant nuclear markers and a diagnostic mitochondrial marker to investigate hybridization in a population of greenback cutthroat trout (Oncorhynchus clarkii stomias) and a population of Colorado River cutthroat trout (Oncorhynchus clarkii pleuriticus). We infer that cutthroat–rainbow trout hybrid swarms have formed in both populations. Although a mixture of hybrid genotypes was present, not all genotype combinations were detected at expected frequencies. We found evidence that mitochondrial DNA introgression in hybrids is asymmetric and more likely from rainbow trout than from cutthroat trout. A difference in spawning time of the 2 species or differences in the fitness between the reciprocal crosses may explain the asymmetry. Additionally, the presence of intraspecific cytonuclear associations found in both populations is concordant with current hypotheses regarding coevolution of mitochondrial and nuclear genomes.
Throughout western North America, rainbow trout (Oncorhynchus mykiss) have been introduced into environments supporting a closely related native species, the cutthroat trout (Oncorhynchus clarkii). When the 2 species interact, they often hybridize. Surveys of populations where the 2 species are sympatric have revealed differences in the dynamics of hybridization; in some cases, hybrid swarms exist (Leary et al. 1983, 1984; Rubidge and Taylor 2004; Bettles et al. 2005), whereas in others, there appears to be some degree of reproductive isolation (Young et al. 2001; Ostberg et al. 2004). One important parameter that may determine the outcome of the interaction is whether sympatry is natural or the result of introductions of hatchery-reared rainbow trout. Where the 2 species are naturally sympatric, hybridization is often limited (Young et al. 2001; Ostberg et al. 2004). By contrast, in places where rainbow trout have been introduced, hybridization occurs perhaps because hatchery-reared rainbow trout have lost prezygotic isolating mechanisms (Docker et al. 2003; Bettles et al. 2005). A potential consequence of hybridization is the localized loss of the native species by genetic introgression. Interestingly, in places where extensive introgression occurs, hybrid genotypes occur at various unexpected frequencies (Young et al. 2001; Rubidge and Taylor 2004; Bettles et al. 2005; Ostberg and Rodriguez 2006), suggesting that natural selection may play an active role in determining the characteristics of hybridization.
Hybridization of rainbow and cutthroat trout has become increasingly problematic because many populations of inland cutthroat trout species are declining. The last remaining Alvord cutthroat trout (Oncorhynchus clarkii alvordensis) populations went extinct in the mid-1980s from hybridization with rainbow trout (Behnke 2002), and many other subspecies are actively managed to reduce the threat of hybridization. Moreover, because of widespread introductions of rainbow trout from both privately and publicly funded hatcheries, the native ranges of cutthroat trout have steadily declined (Young and Harig 2001) and many populations persist in marginal habitats (Young and Guenther-Gloss 2004; Young et al. 2005).
Rainbow trout have been introduced within the range of the federally threatened greenback cutthroat trout (Oncorhynchus clarkii stomias). Our objective in this study was to assess the extent and direction of hybridization between rainbow and cutthroat trout in 2 similar stream systems and comment on implications to conservation management of the native cutthroat trout of Colorado.
Methods
Study Site and Collection Methods
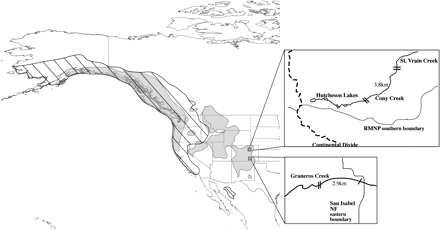
We sampled 2 cutthroat trout populations in the native range of greenback cutthroat trout, Cony Creek in the South Platte River basin and Graneros Creek in the Arkansas River basin. Sections of both creeks were known or suspected to contain introduced rainbow trout (Figure 1). Both creeks are divided by a waterfall that acts as a natural barrier by preventing upstream migration. In both streams, the section thought to contain rainbow trout alleles was below the waterfall barrier with pure cutthroat trout persisting upstream. Stocking records of Cony Creek show that the system was chemically treated to remove invasive rainbow trout and subsequently restocked with hatchery-reared greenback cutthroat trout in upper Cony Creek (above the barrier) and Lower Hutcheson Lake in the 1980s (Rosenlund et al. 2001). Thus, restored cutthroat trout were able to migrate downstream into the putative hybrid sections. The stocking history of Graneros Creek is unknown, but recent work by Metcalf et al. (2007) revealed that the population contains Colorado River cutthroat trout (Oncorhynchus clarkii pleuriticus), which is a subspecies of cutthroat trout native to the west slope of Colorado's Continental Divide. Colorado River cutthroat trout likely were introduced into Graneros Creek in the early 1900s during a time of intensive fish stocking (Metcalf et al. 2007).
The native range of cutthroat trout is shown as a filled gray color, and the native range of rainbow trout is shown overlaid in slanted lines. Magnified schematics of the sample sites, Cony Creek and Graneros Creek, are shown with hybrid populations between hatch marks. Double hatch marks signify a natural waterfall barrier is present.
Fish were sampled from Cony Creek in summer 2003 with 2-pass electrofishing of the first 25 m of every 100 m for more than 5 km; 343 tissue samples were collected below the natural barrier and 99 above it. The hybrid section of Cony Creek spans 3.8 km. Cony Creek has a minimum elevation of 2904 m, a mean gradient of 10.5%, and a flow rate of 0.08 cm. Above the hybrid population, a 1.2-km length of creek between the upper natural barrier and Lower Hutcheson Lake hosts a putative pure cutthroat population.
Graneros Creek samples were collected by electrofishing a 2.9-km section below the natural barrier in summer 2002 (N = 25) and in 2004 (N = 80). Thirty samples were also collected above the natural barrier in 2002 by electrofishing. Graneros Creek has a minimum elevation of 2365 m, a mean gradient of 5.8%, and a flow rate of 0.07 cm.
Molecular Methods
DNA was extracted with Qiagen DNeasy extraction kits for each adipose fin, 442 tissue samples from Cony Creek and 135 tissue samples from Graneros Creek (Qiagen, Valencia, CA). We used a diagnostic mitochondrial marker and 7 diagnostic, codominant nuclear markers to assess presence and extent of introgression from each species in these specimens. We tested the mitochondrial marker for diagnostic properties using more than 250 putatively native, historical greenback cutthroat individuals from 10 populations spanning the South Platte and Arkansas River drainages. Furthermore, we tested the mitochondrial marker on 50 rainbow trout individuals from 3 populations, including 2 strains that are commonly stocked in Colorado. The nuclear markers, OCC16, OCC34, OCC35, OCC36, OCC37, OCC38, and OCC42, were originally identified using microsatellite primers to amplify arbitrary gene regions with species-specific insertion/deletions (Ostberg and Rodriguez 2002). These biparentally inherited markers were demonstrated to be diagnostic in a large number of cutthroat trout and rainbow trout individuals, including controlled hybrid crosses (Ostberg and Rodriguez 2002; Ostberg and Rodriguez 2004). The mitochondrial marker, cytochrome oxidase 1, was amplified with the following primers: COI F6 (5′-atc tct cag taccaa acc cc-3′) and COI- aH redo (5′-cac agt gtr tag gcg tct gg-3′). The cytochrome oxidase 1 gene was amplified using 1× buffer, 4 mM deoxynucleoside triphosphates, 0.25 units Taq polymerase (New England Biolabs, Ipswich, MA) at 94 °C for 2 min, at 94 °C for 30 s, at 54 °C for 45 s, and at 72 °C for 75 s for 35 cycles and a 72 °C extension for 5 min. Subsequently, the polymerase chain reaction was digested with the enzyme Bsu361 according to New England Biolaboratories specifications. The mitochondrial gene digest fragments and the OCC16 nuclear fragment were visualized on 2% agarose gel. The remaining nuclear markers were separated using a 3.5% acrylamide gel on a LiCor 4200.
Analysis Methods
The percentage of rainbow trout alleles in each population was calculated for both nuclear and mitochondrial DNA (mtDNA). Fish were assigned to 1 of 4 classes: pure cutthroat, pure rainbow, first generation cross (F1), and post-F1 backcrosses. Individuals assigned to a pure parent class were homozygous at every nuclear locus for the pure parent and had the COI mitochondrial haplotype of the parent. Individuals were designated F1 if they were heterozygous at every nuclear locus. Finally, individuals that were not F1 crosses but had at least one allele from each parent were grouped into the category of post-F1 hybrid. We did not distinguish among post-F1 genotypes because the frequency distributions for the various post-F1 classes overlap greatly with 7 nuclear markers (Boecklen and Howard 1997).
We measured linkage disequilibrium between nuclear markers for both hybrid populations according to Weir (1979) using the software program Genetix (Belkhir et al. 2001). Estimates of whether genotype frequencies conformed to Hardy–Weinberg expectations (HWE) were calculated using Arlequin 2.0 (Schneider et al. 2000). A Bonferroni-corrected critical value of 0.007 was used to detect significant departures from expectations. Deviations from HWE were expressed as the inbreeding coefficient FIS (Nei 1987).
Asymmetric hybridization, in which the progeny from the female of 1 species and male of the second species are more abundant than in the reciprocal cross, can be driven by assortative mating or selection. To determine the degree of asymmetry between nuclear type and mitochondrial type in each population, we calculated ratios of cutthroat nuclear DNA and cutthroat mtDNA for each population. We tested for independence between the proportion of nuclear alleles and mitochondrial haplotypes of each species using a 2 × 2 contingency table. The 2 × 2 matrix included the observed count number of cutthroat and rainbow trout mtDNA haplotypes and nuclear alleles. A Fisher's 2-sided P value was used to evaluate independence.
Cytonuclear disequilibrium provides a measure of associations between cytoplasmic and nuclear alleles. Significant cytonuclear disequilibrium may indicate nonrandom mating or selection (Rawson and Burton 2002). Nonrandom association between genotypes and mtDNA haplotypes was tested using cytonuclear disequilibrium statistics (Clark 1984; Asmussen et al. 1987; Asmussen and Basten 1994; Asmussen and Christopher 1996). The statistic (D) measures the departure from expectation between mitochondrial and nuclear genotypic frequencies (DAM, DAAM, DAaM, and DaaM), where “A” represents the cutthroat trout nuclear allele, “a” represents the rainbow trout nuclear allele, and “M” represents the cutthroat mtDNA haplotype (Asmussen et al. 1987). The statistics were calculated on post-F1 hybrids only because we were interested in nonrandom associations between nuclear and mitochondrial types in the progeny of hybrids. For Cony Creek, asymptotic P values were used to assess significance of disequilibrium values. In Graneros Creek, however, the sample size was less than 100 and the marginal frequencies were extreme (<0.20). Thus, Fisher's exact test P values were employed as suggested by Asmussen and Basten (1994). Cytonuclear disequilibrium values were generated using a program provided by C. Basten at http://statgen.ncsu.edu/brcwebsite/software_BRC.php. The P value was Bonferroni corrected for multiple simultaneous tests (each locus) and critical value set at 0.007.
Results
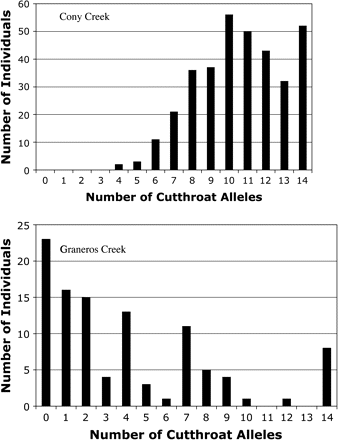
Cony Creek and Graneros Creek both contained hybrids in the putative hybrid section and pure cutthroat trout above the upstream barrier (Table 1). The nuclear allele frequency distribution profiles of hybrid sections in both creeks revealed hybrid fish that were the progeny from various crosses between hybrids and pure cutthroat trout or rainbow trout, indicating that these populations are hybrid swarms (Figure 2). In the Cony Creek hybrid population, cutthroat trout nuclear alleles were more common than rainbow trout nuclear alleles. This bias toward pure cutthroat trout alleles was likely caused by the partially successful chemical removal of rainbow trout from the stream in 1984 and subsequent restocking of several thousand greenback cutthroat trout (Rosenlund et al. 2001). In the Graneros Creek hybrid population, rainbow trout nuclear alleles were more common than cutthroat trout nuclear alleles. Additionally, there was a greater proportion of pure parents and F1 hybrids in Graneros Creek than in Cony Creek (Table 1). Both hybrid populations exhibited significant linkage disequilibrium among loci, although the magnitude of disequilibrium was substantially less in Cony Creek than in Graneros Creek (Table 2). The allele frequency distributions and disequilibrium values suggest that hybridization has been underway for longer in Cony Creek than in Graneros Creek.
The number of pure fish, F1 hybrids, and post-F1 hybrids in each Cony Creek and Graneros Creek. Both hybrid populations had a pure cutthroat trout population upstream above a large waterfall barrier
| Pure cutthroat | Pure rainbow | F1 hybrid | Post-F1 hybrid | |
| Cony Creek hybrid population | 50 | 0 | 4 | 289 |
| Graneros Creek hybrid population | 8 | 23 | 11 | 63 |
| Cony—above second barrier | 99 | 0 | 0 | 0 |
| Graneros—above barrier | 30 | 0 | 0 | 0 |
| Pure cutthroat | Pure rainbow | F1 hybrid | Post-F1 hybrid | |
| Cony Creek hybrid population | 50 | 0 | 4 | 289 |
| Graneros Creek hybrid population | 8 | 23 | 11 | 63 |
| Cony—above second barrier | 99 | 0 | 0 | 0 |
| Graneros—above barrier | 30 | 0 | 0 | 0 |
The number of pure fish, F1 hybrids, and post-F1 hybrids in each Cony Creek and Graneros Creek. Both hybrid populations had a pure cutthroat trout population upstream above a large waterfall barrier
| Pure cutthroat | Pure rainbow | F1 hybrid | Post-F1 hybrid | |
| Cony Creek hybrid population | 50 | 0 | 4 | 289 |
| Graneros Creek hybrid population | 8 | 23 | 11 | 63 |
| Cony—above second barrier | 99 | 0 | 0 | 0 |
| Graneros—above barrier | 30 | 0 | 0 | 0 |
| Pure cutthroat | Pure rainbow | F1 hybrid | Post-F1 hybrid | |
| Cony Creek hybrid population | 50 | 0 | 4 | 289 |
| Graneros Creek hybrid population | 8 | 23 | 11 | 63 |
| Cony—above second barrier | 99 | 0 | 0 | 0 |
| Graneros—above barrier | 30 | 0 | 0 | 0 |
Pairwise linkage disequilibrium (LD) values for Cony Creek (above the diagonal) and Graneros Creek (below the diagonal) measured using Genetix (Belkhir 2001). Pairwise loci that exhibited significant LD are designated with an asterisk and P values below the LD value (critical value = 0.007)
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| OCC16 | — | 0.049 | 0.033 | 0.045 | 0.026 | 0.021 | 0.028 |
| 0.000* | 0.000* | 0.000* | 0.003* | 0.035 | 0.001* | ||
| OCC34 | 0.138 | — | 0.052 | 0.055 | 0.044 | 0.039 | 0.035 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC35 | 0.165 | 0.166 | — | 0.054 | 0.049 | 0.040 | 0.038 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC36 | 0.181 | 0.171 | 0.164 | — | 0.025 | 0.041 | 0.020 |
| 0.000* | 0.000* | 0.000* | 0.007* | 0.000* | 0.035 | ||
| OCC37 | 0.163 | 0.144 | 0.176 | 0.162 | — | 0.035 | 0.023 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC38 | 0.184 | 0.174 | 0.187 | 0.178 | 0.180 | — | 0.049 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC42 | 0.146 | 0.141 | 0.146 | 0.139 | 0.140 | 0.154 | — |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* |
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| OCC16 | — | 0.049 | 0.033 | 0.045 | 0.026 | 0.021 | 0.028 |
| 0.000* | 0.000* | 0.000* | 0.003* | 0.035 | 0.001* | ||
| OCC34 | 0.138 | — | 0.052 | 0.055 | 0.044 | 0.039 | 0.035 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC35 | 0.165 | 0.166 | — | 0.054 | 0.049 | 0.040 | 0.038 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC36 | 0.181 | 0.171 | 0.164 | — | 0.025 | 0.041 | 0.020 |
| 0.000* | 0.000* | 0.000* | 0.007* | 0.000* | 0.035 | ||
| OCC37 | 0.163 | 0.144 | 0.176 | 0.162 | — | 0.035 | 0.023 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC38 | 0.184 | 0.174 | 0.187 | 0.178 | 0.180 | — | 0.049 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC42 | 0.146 | 0.141 | 0.146 | 0.139 | 0.140 | 0.154 | — |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* |
Pairwise linkage disequilibrium (LD) values for Cony Creek (above the diagonal) and Graneros Creek (below the diagonal) measured using Genetix (Belkhir 2001). Pairwise loci that exhibited significant LD are designated with an asterisk and P values below the LD value (critical value = 0.007)
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| OCC16 | — | 0.049 | 0.033 | 0.045 | 0.026 | 0.021 | 0.028 |
| 0.000* | 0.000* | 0.000* | 0.003* | 0.035 | 0.001* | ||
| OCC34 | 0.138 | — | 0.052 | 0.055 | 0.044 | 0.039 | 0.035 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC35 | 0.165 | 0.166 | — | 0.054 | 0.049 | 0.040 | 0.038 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC36 | 0.181 | 0.171 | 0.164 | — | 0.025 | 0.041 | 0.020 |
| 0.000* | 0.000* | 0.000* | 0.007* | 0.000* | 0.035 | ||
| OCC37 | 0.163 | 0.144 | 0.176 | 0.162 | — | 0.035 | 0.023 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC38 | 0.184 | 0.174 | 0.187 | 0.178 | 0.180 | — | 0.049 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC42 | 0.146 | 0.141 | 0.146 | 0.139 | 0.140 | 0.154 | — |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* |
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| OCC16 | — | 0.049 | 0.033 | 0.045 | 0.026 | 0.021 | 0.028 |
| 0.000* | 0.000* | 0.000* | 0.003* | 0.035 | 0.001* | ||
| OCC34 | 0.138 | — | 0.052 | 0.055 | 0.044 | 0.039 | 0.035 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC35 | 0.165 | 0.166 | — | 0.054 | 0.049 | 0.040 | 0.038 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC36 | 0.181 | 0.171 | 0.164 | — | 0.025 | 0.041 | 0.020 |
| 0.000* | 0.000* | 0.000* | 0.007* | 0.000* | 0.035 | ||
| OCC37 | 0.163 | 0.144 | 0.176 | 0.162 | — | 0.035 | 0.023 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC38 | 0.184 | 0.174 | 0.187 | 0.178 | 0.180 | — | 0.049 |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | ||
| OCC42 | 0.146 | 0.141 | 0.146 | 0.139 | 0.140 | 0.154 | — |
| 0.000* | 0.000* | 0.000* | 0.000* | 0.000* | 0.000* |
The allele frequency distribution of number of cutthroat trout nuclear alleles per individual fish for Cony Creek (top) and Graneros Creek (bottom). Cony Creek has 2 peaks at 10 and 14 cutthroat trout alleles. Graneros Creek has multiple peaks at each pure species (0 and 14 alleles) and at F1 hybrids (7 alleles) and backcrosses between F1 hybrids and rainbow trout (4 alleles).
Calculations of the inbreeding coefficient, FIS, revealed opposite trends for Cony Creek and Graneros Creek hybrid populations. For Cony Creek, FIS values were negative, indicating an excess of heterozygotes at 6 of 7 loci, and 1 locus (OCC37) showed a significant excess of heterozygotes (Table 3). In contrast, FIS values were positive across loci in Graneros Creek fish, with the locus OCC35 showing a significant deficiency of heterozygotes.
FIS values and P values for each locus are shown for Cony Creek and Graneros Creek. Significant values are identified with an asterisk (critical value = 0.007)
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| Cony Creek | −0.0778 P = 0.1793 | −0.0861 0.1386 | −0.0537 0.3827 | −0.0039 1.0000 | −0.1814 0.0009* | 0.0620 0.7932 | −0.0509 0.2615 |
| Graneros Creek | 0.2568 0.0076 | 0.0710 0.6382 | 0.3050 0.0013* | 0.1421 0.2040 | 0.1275 0.3163 | 0.2259 0.0249 | 0.0469 0.6625 |
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| Cony Creek | −0.0778 P = 0.1793 | −0.0861 0.1386 | −0.0537 0.3827 | −0.0039 1.0000 | −0.1814 0.0009* | 0.0620 0.7932 | −0.0509 0.2615 |
| Graneros Creek | 0.2568 0.0076 | 0.0710 0.6382 | 0.3050 0.0013* | 0.1421 0.2040 | 0.1275 0.3163 | 0.2259 0.0249 | 0.0469 0.6625 |
FIS values and P values for each locus are shown for Cony Creek and Graneros Creek. Significant values are identified with an asterisk (critical value = 0.007)
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| Cony Creek | −0.0778 P = 0.1793 | −0.0861 0.1386 | −0.0537 0.3827 | −0.0039 1.0000 | −0.1814 0.0009* | 0.0620 0.7932 | −0.0509 0.2615 |
| Graneros Creek | 0.2568 0.0076 | 0.0710 0.6382 | 0.3050 0.0013* | 0.1421 0.2040 | 0.1275 0.3163 | 0.2259 0.0249 | 0.0469 0.6625 |
| OCC16 | OCC34 | OCC35 | OCC36 | OCC37 | OCC38 | OCC42 | |
| Cony Creek | −0.0778 P = 0.1793 | −0.0861 0.1386 | −0.0537 0.3827 | −0.0039 1.0000 | −0.1814 0.0009* | 0.0620 0.7932 | −0.0509 0.2615 |
| Graneros Creek | 0.2568 0.0076 | 0.0710 0.6382 | 0.3050 0.0013* | 0.1421 0.2040 | 0.1275 0.3163 | 0.2259 0.0249 | 0.0469 0.6625 |
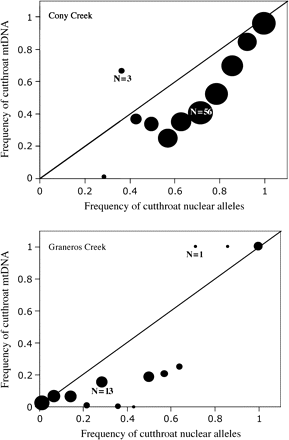
The frequency of cutthroat mtDNA haplotypes in hybrid fish was significantly lower than the frequency of cutthroat nuclear alleles for samples from Cony Creek (P < 0.001) and was also lower for samples in Graneros Creek (Figure 3, Table 4). The lower than expected frequency of cutthroat mtDNA was most evident in F1 hybrids and, to a lesser extent, in post-F1 hybrids.
Ratios of cutthroat trout mtDNA to nuclear DNA (nuDNA) in pure parents, F1 hybrids, and post-F1 hybrids are shown. Actual proportions of cutthroat mtDNA and nuDNA are given in parentheses. P values are from Fisher's exact test of 2 × 2 contingency tables of mitochondrial and nuclear allele count data (critical value = 0.025)
| All individuals cutthroat mtDNA/cutthroat nuDNA | Post-F1 cutthroat mtDNA/cutthroat nuDNA | F1 cutthroat mtDNA/cutthroat nuDNA | |
| Cony Creek | 0.73 (0.55/0.76) P = 0.000* | 0.68 (0.48/0.72) P = 0.000* | 0 (0/0.50) P = 0.079 |
| Graneros Creek | 0.60 (0.17/0.28) P = 0.049 | 0.49 (0.13/0.26) P = 0.013 | 0.36 (0.18/0.50) P = 0.043 |
| All individuals cutthroat mtDNA/cutthroat nuDNA | Post-F1 cutthroat mtDNA/cutthroat nuDNA | F1 cutthroat mtDNA/cutthroat nuDNA | |
| Cony Creek | 0.73 (0.55/0.76) P = 0.000* | 0.68 (0.48/0.72) P = 0.000* | 0 (0/0.50) P = 0.079 |
| Graneros Creek | 0.60 (0.17/0.28) P = 0.049 | 0.49 (0.13/0.26) P = 0.013 | 0.36 (0.18/0.50) P = 0.043 |
Ratios of cutthroat trout mtDNA to nuclear DNA (nuDNA) in pure parents, F1 hybrids, and post-F1 hybrids are shown. Actual proportions of cutthroat mtDNA and nuDNA are given in parentheses. P values are from Fisher's exact test of 2 × 2 contingency tables of mitochondrial and nuclear allele count data (critical value = 0.025)
| All individuals cutthroat mtDNA/cutthroat nuDNA | Post-F1 cutthroat mtDNA/cutthroat nuDNA | F1 cutthroat mtDNA/cutthroat nuDNA | |
| Cony Creek | 0.73 (0.55/0.76) P = 0.000* | 0.68 (0.48/0.72) P = 0.000* | 0 (0/0.50) P = 0.079 |
| Graneros Creek | 0.60 (0.17/0.28) P = 0.049 | 0.49 (0.13/0.26) P = 0.013 | 0.36 (0.18/0.50) P = 0.043 |
| All individuals cutthroat mtDNA/cutthroat nuDNA | Post-F1 cutthroat mtDNA/cutthroat nuDNA | F1 cutthroat mtDNA/cutthroat nuDNA | |
| Cony Creek | 0.73 (0.55/0.76) P = 0.000* | 0.68 (0.48/0.72) P = 0.000* | 0 (0/0.50) P = 0.079 |
| Graneros Creek | 0.60 (0.17/0.28) P = 0.049 | 0.49 (0.13/0.26) P = 0.013 | 0.36 (0.18/0.50) P = 0.043 |
The frequency of cutthroat trout mtDNA versus nuclear DNA in Cony Creek (top) and Graneros Creek (bottom). The area of the circle is proportional to the sample size at that point. The black line in each graph represents the expected trend under random mating (slope = 1). Points below the line signify a lower than expected proportion of cutthroat mtDNA.
Significant cytonuclear associations in post-F1 hybrids were detected for 2 of 7 loci in Cony Creek, OCC16 and OCC35, and for 1 of the same loci in Graneros Creek, OCC35 (Table 5). Cony Creek and Graneros Creek both showed a general trend of homozygote genotypes associated with the matching parental mitochondrial types. The consistent trend with OCC35 deviating from expectations in both populations provides evidence that cytonuclear interactions influence allele frequencies of hybrid populations of greenback cutthroat trout and rainbow trout.
Cytonuclear disequilibrium statistics were calculated for Cony Creek and Graneros Creek. Measures of D, the departure from expected between nuclear genotype and mitochondrial type, and asymptotic P values are shown in parentheses (critical value = 0.007)
| DAM (P value) | DAAM | DAaM | DaaM | |
| Cony Creek | ||||
| OCC16 | 0.035* (0.000) | 0.063* (0.000) | −0.057* (0.000) | −0.006 (0.621) |
| OCC34 | 0.020 (0.074) | 0.035 (0.056) | −0.031 (0.109) | −0.004 (0.798) |
| OCC35 | 0.037* (0.000) | 0.051* (0.005) | −0.021 (0.373) | −0.027* (0.001) |
| OCC36 | 0.027 (0.013) | 0.033 (0.076) | −0.012 (0.711) | −0.021 (0.028) |
| OCC37 | 0.005 (0.811) | 0.011 (0.759) | −0.011 (0.743) | 0.000 (0.996) |
| OCC38 | 0.011 (0.483) | 0.016 (0.562) | −0.009 (0.844) | −0.007 (0.702) |
| OCC42 | 0.011 (0.426) | 0.014 (0.637) | −0.006 (0.922) | −0.008 (0.358) |
| Graneros Creek | ||||
| OCC16 | 0.022 (0.125) | 0.020 (0.164) | 0.005 (1.000) | −0.025 (0.272) |
| OCC34 | 0.022 (0.125) | 0.028 (0.014) | −0.011 (0.716) | −0.017 (0.474) |
| OCC35 | 0.057* (0.000) | 0.051* (0.002) | 0.011 (0.678) | −0.063* (0.004) |
| OCC36 | 0.011 (0.348) | 0.012 (0.240) | −0.001 (1.000) | −0.011 (0.707) |
| OCC37 | 0.026 (0.053) | 0.012 (0.240) | 0.029 (0.247) | −0.041 (0.065) |
| OCC38 | 0.015 (0.234) | 0.006 (0.505) | 0.019 (0.434) | −0.025 (0.272) |
| OCC42 | 0.011 (0.418) | 0.006 (0.505) | 0.011 (0.716) | −0.017 (0.473) |
| DAM (P value) | DAAM | DAaM | DaaM | |
| Cony Creek | ||||
| OCC16 | 0.035* (0.000) | 0.063* (0.000) | −0.057* (0.000) | −0.006 (0.621) |
| OCC34 | 0.020 (0.074) | 0.035 (0.056) | −0.031 (0.109) | −0.004 (0.798) |
| OCC35 | 0.037* (0.000) | 0.051* (0.005) | −0.021 (0.373) | −0.027* (0.001) |
| OCC36 | 0.027 (0.013) | 0.033 (0.076) | −0.012 (0.711) | −0.021 (0.028) |
| OCC37 | 0.005 (0.811) | 0.011 (0.759) | −0.011 (0.743) | 0.000 (0.996) |
| OCC38 | 0.011 (0.483) | 0.016 (0.562) | −0.009 (0.844) | −0.007 (0.702) |
| OCC42 | 0.011 (0.426) | 0.014 (0.637) | −0.006 (0.922) | −0.008 (0.358) |
| Graneros Creek | ||||
| OCC16 | 0.022 (0.125) | 0.020 (0.164) | 0.005 (1.000) | −0.025 (0.272) |
| OCC34 | 0.022 (0.125) | 0.028 (0.014) | −0.011 (0.716) | −0.017 (0.474) |
| OCC35 | 0.057* (0.000) | 0.051* (0.002) | 0.011 (0.678) | −0.063* (0.004) |
| OCC36 | 0.011 (0.348) | 0.012 (0.240) | −0.001 (1.000) | −0.011 (0.707) |
| OCC37 | 0.026 (0.053) | 0.012 (0.240) | 0.029 (0.247) | −0.041 (0.065) |
| OCC38 | 0.015 (0.234) | 0.006 (0.505) | 0.019 (0.434) | −0.025 (0.272) |
| OCC42 | 0.011 (0.418) | 0.006 (0.505) | 0.011 (0.716) | −0.017 (0.473) |
Cytonuclear disequilibrium statistics were calculated for Cony Creek and Graneros Creek. Measures of D, the departure from expected between nuclear genotype and mitochondrial type, and asymptotic P values are shown in parentheses (critical value = 0.007)
| DAM (P value) | DAAM | DAaM | DaaM | |
| Cony Creek | ||||
| OCC16 | 0.035* (0.000) | 0.063* (0.000) | −0.057* (0.000) | −0.006 (0.621) |
| OCC34 | 0.020 (0.074) | 0.035 (0.056) | −0.031 (0.109) | −0.004 (0.798) |
| OCC35 | 0.037* (0.000) | 0.051* (0.005) | −0.021 (0.373) | −0.027* (0.001) |
| OCC36 | 0.027 (0.013) | 0.033 (0.076) | −0.012 (0.711) | −0.021 (0.028) |
| OCC37 | 0.005 (0.811) | 0.011 (0.759) | −0.011 (0.743) | 0.000 (0.996) |
| OCC38 | 0.011 (0.483) | 0.016 (0.562) | −0.009 (0.844) | −0.007 (0.702) |
| OCC42 | 0.011 (0.426) | 0.014 (0.637) | −0.006 (0.922) | −0.008 (0.358) |
| Graneros Creek | ||||
| OCC16 | 0.022 (0.125) | 0.020 (0.164) | 0.005 (1.000) | −0.025 (0.272) |
| OCC34 | 0.022 (0.125) | 0.028 (0.014) | −0.011 (0.716) | −0.017 (0.474) |
| OCC35 | 0.057* (0.000) | 0.051* (0.002) | 0.011 (0.678) | −0.063* (0.004) |
| OCC36 | 0.011 (0.348) | 0.012 (0.240) | −0.001 (1.000) | −0.011 (0.707) |
| OCC37 | 0.026 (0.053) | 0.012 (0.240) | 0.029 (0.247) | −0.041 (0.065) |
| OCC38 | 0.015 (0.234) | 0.006 (0.505) | 0.019 (0.434) | −0.025 (0.272) |
| OCC42 | 0.011 (0.418) | 0.006 (0.505) | 0.011 (0.716) | −0.017 (0.473) |
| DAM (P value) | DAAM | DAaM | DaaM | |
| Cony Creek | ||||
| OCC16 | 0.035* (0.000) | 0.063* (0.000) | −0.057* (0.000) | −0.006 (0.621) |
| OCC34 | 0.020 (0.074) | 0.035 (0.056) | −0.031 (0.109) | −0.004 (0.798) |
| OCC35 | 0.037* (0.000) | 0.051* (0.005) | −0.021 (0.373) | −0.027* (0.001) |
| OCC36 | 0.027 (0.013) | 0.033 (0.076) | −0.012 (0.711) | −0.021 (0.028) |
| OCC37 | 0.005 (0.811) | 0.011 (0.759) | −0.011 (0.743) | 0.000 (0.996) |
| OCC38 | 0.011 (0.483) | 0.016 (0.562) | −0.009 (0.844) | −0.007 (0.702) |
| OCC42 | 0.011 (0.426) | 0.014 (0.637) | −0.006 (0.922) | −0.008 (0.358) |
| Graneros Creek | ||||
| OCC16 | 0.022 (0.125) | 0.020 (0.164) | 0.005 (1.000) | −0.025 (0.272) |
| OCC34 | 0.022 (0.125) | 0.028 (0.014) | −0.011 (0.716) | −0.017 (0.474) |
| OCC35 | 0.057* (0.000) | 0.051* (0.002) | 0.011 (0.678) | −0.063* (0.004) |
| OCC36 | 0.011 (0.348) | 0.012 (0.240) | −0.001 (1.000) | −0.011 (0.707) |
| OCC37 | 0.026 (0.053) | 0.012 (0.240) | 0.029 (0.247) | −0.041 (0.065) |
| OCC38 | 0.015 (0.234) | 0.006 (0.505) | 0.019 (0.434) | −0.025 (0.272) |
| OCC42 | 0.011 (0.418) | 0.006 (0.505) | 0.011 (0.716) | −0.017 (0.473) |
Discussion
Rainbow trout introgressively hybridized with cutthroat trout in both streams we studied. Our results mirror those for other cutthroat trout subspecies and suggest that introduced rainbow trout pose a serious threat to greenback and Colorado River cutthroat trout (Leary et al. 1984; Hitt et al. 2003; Rubidge and Taylor 2004; Ostberg and Rodriguez 2006). Nevertheless, we noticed departures from expected genotype frequencies, which suggest that hybrid trout populations provide a fertile system for studying adaptation and natural selection on genomes in natural settings. In Cony Creek, we discovered an excess of heterozygote individuals at 6 of the 7 loci. This pattern may reflect nonrandom mating or higher fitness of heterozygous individuals. Numerous studies support a positive relationship between heterozygosity and fitness, suggesting that the excess heterozygosity may signal the action of selection. Conversely, Graneros Creek did not reveal any evidence of excess heterozygosity, and 1 locus (OCC35) showed a significant lack of heterozygote individuals. This opposite finding for Graneros Creek and Cony Creek is surprising as both systems contain pure cutthroat trout individuals and have an influx of pure cutthroat trout from the populations above the waterfall barrier. Cony Creek is approximately 600 m higher in elevation and twice as steep as Graneros Creek, which may contribute to different hybridization patterns within populations. Alternatively, differences in the evolutionary history of greenback cutthroat trout introduced into Cony Creek and Colorado River cutthroat trout introduced into Graneros Creek may generate different hybridization dynamics in the 2 populations. To discern the degree to which environmental characteristics and differences in subspecies’ evolutionary history affect hybridization dynamics in a population, additional hybrid populations need to be studied.
A second unexpected result was the excess of rainbow trout mitochondrial haplotypes relative to rainbow trout nuclear alleles in hybrid fish—the excess was most pronounced in F1 hybrids (Table 4). The asymmetric pattern may be initiated at the F1 stage followed by random mating in backcrosses. Similar to these findings, Ostberg et al. (2004) studied 2 populations of naturally sympatric coastal cutthroat trout and steelhead and found that F1s had only rainbow trout mtDNA and that the relative frequency of rainbow trout mtDNA decreased in post-F1 hybrids. Furthermore, Ostberg and Rodriguez (2006) observed a similar pattern of greater proportions of hybrid progeny from rainbow trout females and westslope cutthroat trout males. Bettles et al. (2005), however, surveyed 13 sympatric populations containing native and introduced coastal cutthroat and rainbow trout and found asymmetries in F1 hybrid haplotypes in several populations, but most populations had excess cutthroat trout mtDNA. Asymmetry patterns appear variable across hybrid systems of rainbow and cutthroat trout. Therefore, conservation management strategies that use estimates of percentage of hybridization to identify populations under a certain threshold of hybridization (e.g., 20%) may be inaccurate when only based on mtDNA.
Several hypotheses have been proposed to explain an asymmetry in mitochondrial type in rainbow–cutthroat trout hybrids. Ostberg et al. (2004) suggested that females of the rare species may become less discriminating and spawn with the abundant male species. This could explain the trend found in Cony Creek, but in Graneros Creek, the rare species’ mtDNA was underrepresented. Ostberg and Rodriguez (2006) suggest that rainbow trout males may complete spawning before cutthroat females become gravid, but female rainbow trout may overlap in spawning time with cutthroat males, which supports the trend found in this study. Alternatively, selection may explain the asymmetry. Hawkins and Foote (1998) showed that the cross between a rainbow trout female and cutthroat trout male resulted in a shorter time to hatching and that progeny had a faster growth rate and a greater abundance of yolk at hatch and emergence than the hybrids from the reciprocal cross. In the high-altitude streams of the Rocky Mountains, the combination of earlier emergence in the spring, a faster growth rate, and a larger yolk sac may increase survivorship relative to the reciprocal cross. Because the mismatch between egg size and developmental rate should decrease in progeny of hybrid fish as recombination shuffles the nuclear genomes of the 2 species, the asymmetric trend of mitochondrial haplotypes in hybrids should be less severe in post-F1 fish, which is consistent with the trend discovered in this study and in studies by Ostberg and colleagues (Ostberg and Rodriguez 2004, 2006).
Additionally, we discovered cytonuclear disequilibrium in the hybrid populations. In particular, both populations surveyed showed the same pattern, namely, there was positive disequilibrium between nuclear marker OCC35 and mitochondrial type, suggesting that the marker may be linked to loci involved in cytonuclear epistatic interactions influencing fitness (Asmussen et al. 1987). With so few loci, it was surprising that we detected significant cytonuclear disequilibrium; nonetheless, if the model of nuclear and cytoplasmic compensatory coadaptation of Rand et al. (2004) is correct, then we should observe cytonuclear disequilibrium in hybrids from crosses between these 2 species because they diverged long enough ago (at least 2 million years) for the accumulation of a significant number of deleterious changes in the mitochondrial-encoded genes and a corresponding number of compensatory changes in nuclear-encoded proteins that interact with the mitochondrial-encoded proteins. Because so few generations of recombination have elapsed since the inception of the hybrid population, each one of the nuclear markers may be linked to a large number of loci, increasing the chances of detecting disequilibrium.
Our study supports the notion that natural waterfall barriers provide refuges for pure native cutthroat trout genomes across their range. In the westslope cutthroat trout system, Hitt et al. (2003) found that migration of cutthroat–rainbow trout hybrids upstream in drainage systems was an important factor in the spread of rainbow trout alleles to previously pure cutthroat trout populations. Therefore, barriers that halt the spread of rainbow trout alleles are important. In a second study of westslope cutthroat and rainbow hybrids, pure cutthroat trout were only found above natural waterfall barriers as in our system (Ostberg and Rodriguez 2006). The management strategy of introducing restoration populations above natural waterfall barriers appears successful on the short term if the waters were previously barren of fish or if nonnative fish were completely removed by chemical treatment. Yet, chemical treatment is not always successful.
Although barriers may protect native trout from invaders, several factors should be considered with this management strategy. First, the habitat in these high alpine refuges may be of marginal quality for cutthroat trout and not support fish overtime (Harig and Fausch 2002; Young and Guenther-Gloss 2004; Young et al. 2005; Fausch et al. 2006). Second, the ecosystem of historically fishless lakes and streams in the Rocky Mountains changes with the introduction of fish. For example, in the Sierra Nevadas, introductions of trout into historically fishless lakes and streams have been associated with declines in amphibians, changes in invertebrate communities, and changes in nutrient cycling (Knapp and Matthews 2000; Schindler et al. 2001). Therefore, the overall conservation goals for native communities should also be considered with respect to cutthroat trout restoration practices.
Our results indicate that greenback cutthroat trout, Colorado River cutthroat trout, and rainbow trout readily hybridize, resulting in a hybrid swarm and the loss of the integrity of the cutthroat trout gene pool. Although management agencies view these hybrid populations as irreconcilable with the goals of native fish restoration projects and target hybrids for destruction, the observation of significant departures of genotype frequencies from the expectations of a hybrid swarm suggests that these populations have intrinsic value for understanding the evolution of trout. Information derived from research on hybrid populations may be relevant for implementing effective long-term conservation strategies.
Funding
National Science Foundation Graduate Fellowship Program; U.S. Fish and Wildlife Service; Colorado Division of Wildlife; U.S. Forest Service; National Park Service; Colorado Mountain Club; University of Colorado's Beverly Sears Graduate Grant; Department of Ecology and Evolutionary Biology Graduate Research Grants (including RC Averett memorial grant); University of Colorado Museum's Walker Van Riper Graduate Research Endowments; and the Universtiy of Colorado Undergraduate Research Opportunities Program (supporting K. Woolanin and M.R.S.).
K. Woolanin and K. Ballare contributed to data generation. C. Kennedy (US Fish and Wildlife), J. Melby (Colorado Division of Wildlife) and Trout Unlimited, and CU volunteers collected samples. J. B. Mitton and B. R. Kreiser provided modified COI sequence primers. O. C. Ostberg, E. A. Bean, and J. Rigor provided expertize. R. D. Noyes, S. B. Wise, and R. T. Jones provided critical comments.
References
Author notes
Corresponding Editor: William Modi