-
PDF
- Split View
-
Views
-
Cite
Cite
Xiaobing Kou, Peng Cao, Qianke He, Peng Wang, Shaoling Zhang, Juyou Wu, PbrROP1/2-elicited imbalance of cellulose deposition is mediated by a CrRLK1L-ROPGEF module in the pollen tube of Pyrus, Horticulture Research, Volume 9, 2022, uhab034, https://doi.org/10.1093/hr/uhab034
Close - Share Icon Share
Abstract
Pollen tube growth is critical for the sexual reproduction of flowering plants. Catharanthus roseus receptor-like kinases (CrRLK1L) play an important role in plant sexual reproduction, pollen tube growth, and male and female gametophyte recognition. Here, we identified a CrRLK1L protein in pear (Pyrus bretschneideri), PbrCrRLK1L13, which is necessary for normal tip growth of the pollen tube. When PbrCrRLK1L13 was knocked down, the pollen tube grew faster. Interaction analysis showed that the kinase domain of PbrCrRLK1L13 interacted with the C-terminal region of PbrGEF8, and PbrCrRLK1L13 activated the phosphorylation of PbrGEF8 in vitro. Furthermore, PbrROP1 and PbrROP2 were the downstream targets of PbrCrRLK1L13-PbrGEF8. When we knocked down the expression of PbrCrRLK1L13, PbrGEF8, or PbrROP1/2, the balance of cellulose deposition in the pollen tube wall was disrupted. Considering these factors, we proposed a model for a signaling event regulating pear pollen tube growth. During pear pollen tube elongation, PbrCrRLK1L13 acted as a surface regulator of the PbrROP1 and PbrROP2 signaling pathway via PbrGEF8 to affect the balance of cellulose deposition and regulate pear pollen tube growth.
Introduction
Fruit trees reproduce sexually with the pollen tube transporting the sperm to the ovule for fertilization. The germination of a pollen grain and the subsequent rapid elongation of the pollen tube are regulated by multiple factors, such as the extracellular peptide, hormone, intracellular signals of calcium ion, and reactive oxygen species, etc. [1–3]. The plasma membrane protein receptor-like kinases (RLKs) have been demonstrated to play critical roles in pollen germination and pollen tube growth by transducing the extracellular signal into an intracellular signal in pollen [4–6]. A type of RLKs, the Catharanthus roseus receptor-like kinase (CrRLK1L) family, has been identified as the regulator for sexual reproduction in both male and female gametophytes. For example, FERONIA mediates pollen tube rupture in the female gametophyte to discharge sperm for fertilization [4, 7]. ANXUR1 (ANX1) and ANXUR1 (ANX2) are the orthologs of FERONIA and are necessary for the regulation of pollen tube integrity [8, 9]. BUPS1 and BUPS2 are another two RLKs that belong to the CrRLK1L family [5, 10]. The bups1 and bups1/bups2 loss-of-function mutants show a precocious pollen tube rupture phenotype similar to that of the anx1/anx2 double mutant [5]. However, the signaling pathways for the structural changes in pollen tube cell walls or rupture of pollen tubes in these mutations are poorly understood.
As a transmembrane protein, RLKs will phosphorylate intracellular proteins after receiving extracellular signals, causing signal cascades. Intracellular protein RopGEFs has been reported to receive RLKs signal to regulate the polar growth of plant cells. For example, the Arabidopsis pollen-specific receptor-like kinases (AtPRK2) transduce ROP signaling via RopGEF12 to regulate the growth of the pollen tube [11]. CrRLK1L receptor kinase FERONIA mediates RopGEF4 and RopGEF10 to regulate root hair growth in Arabidopsis [12]. RopGEFs possess 14 members in Arabidopsis and a highly conserved PRONE domain, which is flanked by variable N- and C-termini, and are known to activate Rho GTPases of plants (ROP) by promoting the conversion of Rho GTPases from their GDP-bound inactive form to the GTP-bound active form [13–16]. ROP GTPases play important roles in several cellular activities such as the formation of F-actin, production of reactive oxygen species, and regulation of the intracellular Ca2+concentration gradient [17–19]. ROPs participate in multiple physiological reactions and signal transduction processes. For example, ROP11 negatively regulates the abscisic acid (ABA) response in Arabidopsis [20, 21]; ROP9 and ROP10 respectively regulate auxin and ABA responses [22]; ROP2 plays important roles in the formation of F-actin [23]. In addition, several ROPs are highly expressed in pollen tubes and control their growth. ROP1, pollen-specific expression in Arabidopsis, plays a central role in regulating pollen tube tip growth. ROP1 promotes pollen tube polar growth via the RIC4 or RIC3 pathways, which are opponent [17]. ROP1 promotes apical F-actin assembly via the RIC4 pathway. In contrast, it increases Ca2+ signaling at the tip and subsequently promotes the disassembling of apical F-actin via the RIC3 pathway [17]. Maize ZmROP2 and ZmROP9 are orthologs of ROP1 and are highly expressed in pollen [18, 24]. The loss-of-function mutants of Zmrop2 display transmission defects in the male gametophyte [24]. Previous reports have revealed that the ROPs are the downstream targets of RLK [25–27]. Considering the relationships among RLK, RopGEF, and ROP, we assume that RLK acts as an upstream regulator of ROP signaling via RopGEF to regulate tip growth.
The cell wall provides both shape and protection for the plant cell. The pollen tube cell wall is mainly composed of pectin, hemicellulose, and cellulose. Cellulose is involved in the establishment of the growth direction of pollen tubes and plays an important role in the cell wall construction during pollen tube development [28, 29]. Cellulose synthesis and the distribution of the cell wall at the apical tip enable the pollen tube to steadily elongate and help the pollen tube maintain cell wall integrity [30]. In the present study, we identified a CrRLK1L family protein in pear, PbrCrRLK1L13, essential for pear pollen tube growth and cell wall integrity maintenance. We here present a mechanism of CrRLK1L regulating ROP-signaled cellulose distribution and mediating pollen tube growth.
Results
PbrCrRLK1L13 acts as the regulator of cellulose deposition to control pear pollen tube growth
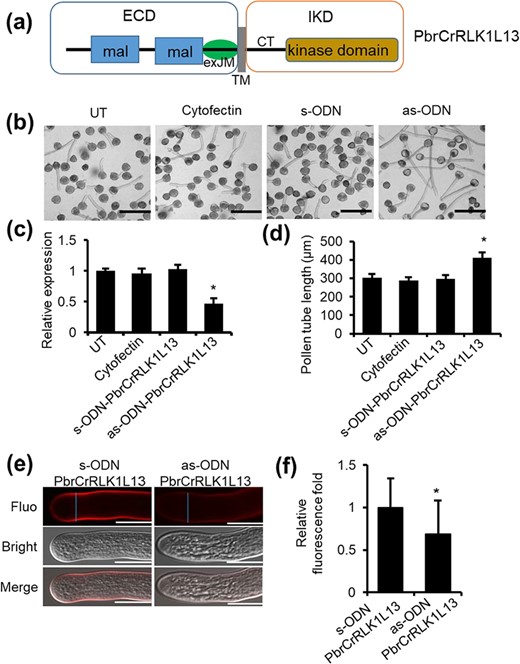
PbrCrRLK1L13 encodes one member of the C. roseus RLK1-like (CrRLK1L) family and is closely related to BUPS in Arabidopsis [5] and RUPO in rice [31, 32]. PbrCrRLK1L13 is a membrane protein comprising an extracellular domain (ECD) known as the extracellular juxtamembrane region (exJM), a transmembrane domain (TM), and an intracellular kinase domain (IKD) (Fig. 1a).
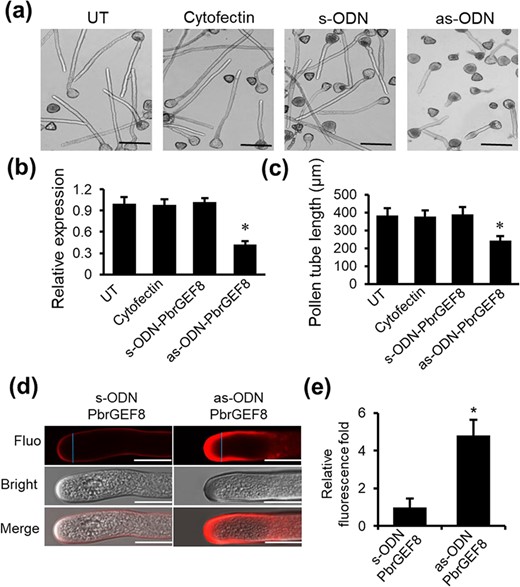
PbrCrRLK1L13 inhibits pear pollen tube growth by controlling the cellulose content. (a) The predicted structure of PbrCrRLK1L13, including extracellular domain (ECD), exJM, transmembrane domain (TM), and intracellular kinase domain (IKD). (b) PbrCrRLK1L13 inhibits pear pollen tube growth. UT represents no-treatment controls, and cytofectin and s-ODN are used as negative controls. Images shown were acquired 2 h after ODN treatment. The bar = 200 μm. (c) qRT-PCR analysis of PbrCrRLK1L13 expression level under ODN treatment. Differences were identified using Student’s t-test and were considered significant when P < 0.05, represented by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (d) Statistical analysis of the pollen tube length. Differences were identified using Student’s t-test and were considered significant when P < 0.05, represented by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (e) As-ODN-PbrCrRLK1L13 treatments decrease the cellulose contents in the pear pollen tube wall. The bar = 50 μm. (f) The cellulose content was quantified by the cellulose fluorescence staining. The pollen tube marked by a blue line was used to quantify the cellulose content in the cell wall.
To test whether PbrCrRLK1L13 is important for pollen tube polar growth, we knocked down gene expression using an antisense oligonucleotide (as-ODN) approach. When the expression of PbrCrRLK1L13 was knocked down (Fig. 1c), we observed a significant promotion of pollen tube length in as-ODN-PbrCrRLK1L13, but not in the corresponding sense oligonucleotides (Fig. 1b and d), suggesting that PbrCrRLK1L13 could inhibit pear pollen tube growth. Moreover, when we knocked down the expression of PbrCrRLK1L13 in pear pollen, the cellulose contents at the pollen tube tips were significantly increased (Fig. 1e and f), indicating that PbrCrRLK1L13 could cause the imbalance of cellulose distribution in pollen tube tips and thus regulate pear pollen tube growth.
PbrGEF8 is one of the downstream targets of PbrCrRLK1L13
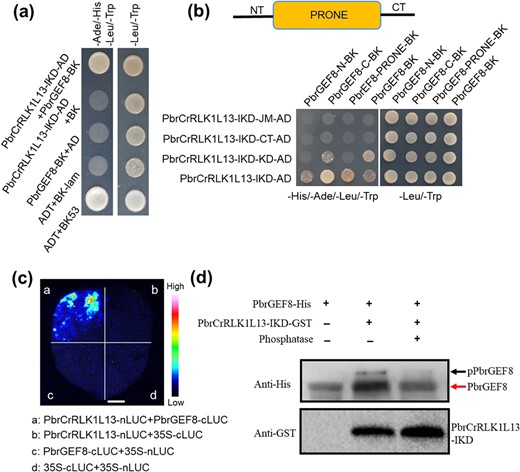
To identify intracellular downstream signaling molecules of PbrCrRLK1L13 in pollen tubes, we used the PbrCrRLK1L13 kinase domain as the bait to screen a cDNA library of pear pollen tube by a yeast two-hybrid (Y2H) screen. Here, an interactor of PbrCrRLK1L13, PbrGEF8, which belongs to the guanine nucleotide exchange factor (RopGEF) family, was identified. We confirmed the physical interaction between PbrGEF8 and PbrCrRLK1L13 with two approaches. A Y2H assay revealed that PbrGEF8 directly interacted with PbrCrRLK1L13 in vitro (Fig. 2a). No obvious interaction between PbrCrRLK1L13 and the other five PbrGEF proteins (PbrGEF1, PbrGEF9, PbrGEF10, PbrGEF11, and PbrGEF12) was found (Fig. S1). In addition, the luciferase complementation imaging (LCI) assay revealed the binding of PbrCrRLK1L13 and PbrGEF8 in vivo (Fig. 2c and S4a). Deletion analysis in Y2H was used to identify the interaction sites between PbrCrRLK1L13 and PbrGEF8, and the results showed that the kinase domain (KD) of PbrCrRLK1L13 interacted with the C-terminal region (CT) domain of PbrGEF8 (Fig. 2b), indicating that the KD domain of PbrCrRLK1L13 and the CT domain of PbrGEF8 contributed to the interaction between PbrCrRLK1L13 and PbrGEF8.
PbrCrRLK1L13 physically interacts with PbrGEF8 and phosphorylated PbrGEF8. (a) PbrGEF8 interacts with the intracellular kinase domain of PbrCrRLK1L13 in yeast. PbrGEF8 fused to the pGBKT7 vector was used as the bait to test its interaction with the intracellular kinase domain of PbrCrRLK1L13 fused to the pGADT7 vector. Yeast growth in the medium that lacks Trp, Leu, His, and Ade indicates protein–protein interaction. (b) The IKD of PbrCrRLK1L13 interacts with the CT domain of PbrGEF8. The predicted structure of PbrGEF8 includes the N-terminal region (NT), PRONE domain, and C-terminal region (CT). (c) PbrGEF8 interacts with PbrCrRLK1L13 in the leaves of N. benthamiana, as shown by LCI assays. Fluorescence on the leaf surface indicates protein–protein interaction. The bar = 1 cm. (d) PbrCrRLK1L13 phosphorylates PbrGEF8. His-tagged PbrGEF8 was incubated with PbrCrRLK1L13 (PbrCrRLK1L13-IKD) and analyzed by Phos-tag mobility shift assays. Black arrows indicate phosphorylated PbrGEF8 proteins. Red arrows indicate non-phosphorylated PbrGEF8 proteins.
We further examined whether the phosphorylation level of PbrGEF8 is affected by PbrCrRLK1L13 using a Phos-tag mobility shift assay. In the absence of PbrCrRLK1L13-IKD-GST, PbrGEF8-His was detected as a single band in non-phosphorylated form. When PbrCrRLK1L13-IKD-GST was added, the phosphorylated form of PbrGEF8-His was detected in Phos-tag gels (Fig. 2d), indicating PbrGEF8 is phosphorylated by PbrCrRLK1L13.
Moreover, when PbrGEF8 was knocked down using the antisense oligonucleotide approach, we observed a significant inhibition of pollen tube length (Fig. 3a–c), but not in the control groups (no-treatment groups and negative control groups). In addition, when PbrGEF8 was knocked down, the cellulose contents at the pollen tube tips were also significantly increased (Fig. 3d and e). It can be confirmed that PbrGEF8 acted as a downstream component of PbrCrRLK1L13 to mediate cellulose distribution and promote pollen tube growth.
Knockdown of the expression of PbrGEF8 inhibited pear pollen tube growth. (a) Knockdown of the expression of PbrGEF8 using the as-ODN approach inhibits pear pollen tube growth. UT suggests no-treatment controls, and cytofectin and s-ODN are used as negative controls. The bar = 200 μm. (b) The expression level after ODN treatment is shown in qRT-PCR. The value for each sample is the mean of its three replicates. Differences were identified using Student’s t-test and were considered significant when P < 0.05, indicated by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (c) Statistical analysis of the pollen tube length. Differences were identified using Student’s t-test and were considered significant when P < 0.05, represented by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (d) As-ODN-PbrGEF8 treatments increased the cellulose contents in the pear pollen tube wall. The bar = 50 μm. The value for each sample is the mean of three replicates. (e) The cellulose content was quantified by the cellulose fluorescence staining. The pollen tube marked by the blue line was used to quantify the cellulose content in the cell wall.
PbrCrRLK1L13-PbrGEF8 mediates the ROP signaling pathway to regulate pear pollen tube growth
Previous studies have shown that ROP GTPases play important roles in pollen tube growth. To further explore RLK regulation of ROP signaling, we first identified 112 members as candidate small GTPase genes in the pear genome (Table S1). According to the phylogenetic tree, the small GTPase proteins were divided into four large subgroups: Rab, Ran, Rop, and Arf/Sar (Fig. S2). The Rab proteins formed the largest subfamily of small GTP-binding proteins, including 70 members from pear, and the Rop subgroup was the smallest, containing six Rop GTPases (Fig. S2). Based on the transcript data (Fig. 4a) and quantitative RT-PCR (Fig. 4b), we found that the expression levels of PbrROP1 and PbrROP2 were significantly higher than those of other small GTPase genes during pear pollen tube growth. Furthermore, we investigated the expression of PbrROPs in pear root, stem, leaf, pollen, and pistil using reverse transcription PCR (RT-PCR) (Fig. S3) and found that the expression levels of PbrROP1 and PbrROP2 were the highest in pollen, indicating that PbrROP1 and PbrROP2 may primarily function in pollen tube growth.
The expression profile of small GTPase genes during pear pollen tube growth. (a) Heat map of RNA-Seq expression of small GTPase genes in the four stages of pollen tube growth [mature, dry pollen (MP), hydrated pollen grains (HP), pollen tubes (PT), and stopped-growth pollen tubes (SPT)]. (b) The qRT-PCR analysis of the expression of eight small GTPase genes in four stages of pear pollen tube growth. PbrTUB was used as the reference gene.
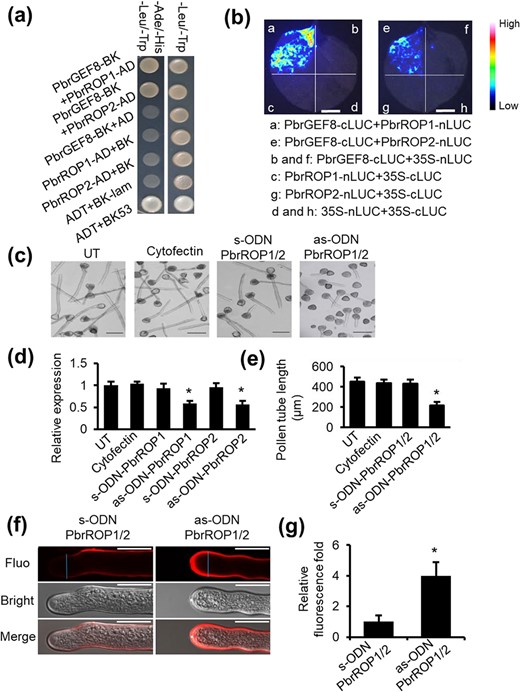
Previous studies have revealed that ROPGEFs are important for regulating ROP GTPases in Arabidopsis. To verify whether ROPGEFs bind to ROPs in pear, the physical interactions between PbrGEFs and PbrROP1 or PbrROP2 were tested with Y2H. The Y2H results showed that PbrGEF8 interacted with PbrROP1 and PbrROP2 (Fig. 5a), whereas no interaction was found in the negative controls. Next, the interactions between PbrROP1 and PbrROP2 with PbrGEF8 in the leaves of N. benthamiana were confirmed with LCI assay (Fig. 5b and S4b). The results showed that PbrGEF8 directly interacted with PbrROP1 and PbrROP2. Moreover, PbrROP1 and PbrROP2, the two closest ROP homologs in pear, share 97% identity at the amino acid level. When PbrROP1 and PbrROP2 were knocked down using the antisense oligonucleotide approach, the length of pollen tubes in the presence of as-ODNs was significantly shorter than that in the control groups (Fig. 5c–e), and the cellulose contents at the pollen tube tips were significantly increased following PbrROP1/2-as-ODN treatments compared to the contents found in the control pollen tubes (Fig. 5f and g).
PbrGEF8 interacts with PbrROP1 and PbrROP2 to regulate the pear pollen tube growth. (a) PbrGEF8 interacts with PbrROP1 and PbrROP2 in yeast. Yeast growth in the medium that lacks Trp, Leu, His, and Ade indicates protein–protein interaction. No interaction was observed in negative control groups. The experiment was repeated more than three times. (b) PbrGEF8 interacts with PbrROP1 and PbrROP2 in the leaves of N. benthamiana, as shown by LCI assays. Fluorescence on the leaf surface indicates protein–protein interaction. No fluorescence was observed in negative control groups. The experiment was repeated more than three times. The bar = 1 cm. (c) Knockdown of the expression of PbrROP1/2 using the as-ODN approach inhibits pear pollen tube growth. UT represents no-treatment controls, and cytofectin and s-ODN are used as negative controls. The bar = 200 μm. (d) The expression level of PbrROP1 and PbrROP1 after ODN treatment is shown in qRT-PCR. The value for each sample is the mean of three replicates. Differences were identified using Student’s t-test and were considered significant when P < 0.05, represented by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (e) Statistical analysis of the pollen tube length. Differences were identified using Student’s t-test and were considered significant when P < 0.05, indicated by *. More than 100 pollen tubes were measured. The value for each sample is the mean of three replicates. (f) As-ODN-PbrROP1/2 treatments increase the cellulose contents in the pear pollen tube wall. The bar = 50 μm. The value for each sample is the mean of three replicates. (g) The cellulose content was quantified by the cellulose fluorescence staining. The pollen tube marked by the blue line was used to quantify the cellulose content in the cell wall.
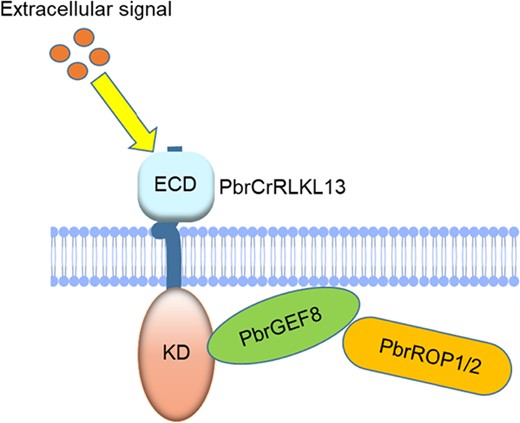
Taken together, these results show that after receiving extracellular signals, PbrCrRLK1L13 regulates the downstream ROP GTPase and affects the cellulose content balance of the pollen tube cell wall, inhibiting pear pollen tubes growth (Fig. 6).
The model for CrRLK1L-ROP signaling in regulating pear pollen tube growth. PbrCrRLK1L13 binds to the C-terminal region of PbrGEF8, with PbrROP1 and PbrROP2 forming the complex, which affects the balance of cellulose content and regulates pear pollen tube growth.
Discussion
Recent studies have shown the signal model of the CrRLK1L subfamily controlling pollen tube growth. For example, FERONIA plays a critical role in female fertility and controlling pollen tube–ovule interaction [4, 7, 25, 33, 34]. In addition, ANX and BUPS as the receptor mediate RALF signaling to maintain pollen tube integrity [35]. The pollen tube precociously ruptured in bups1 bups2 and anx1 anx2 double mutants [8, 9, 36]. RUPO is known to facilitate male transmission by controlling the potassium concentration in the pollen tube [32]. Here, we identified a CrRLK1L protein in pear, PbrCrRLK1L13, which is critical for the growth of pear pollen tubes. Although PbrCrRLK1L13 is the ortholog of RUPO from rice (Oryza sativa) [32], and the ortholog of BUPS1 (At4g39110) and BUPS2 (At2g21480) [5], RUPO, BUPS1, and BUPS2 have pollen-specific expression in rice and Arabidopsis, whereas PbrCrRLK1L13 was expressed in multiple tissues in pear (previous data), indicating functional differences in PbrCrRLK1L13 and BUPS and RUPO. When PbrCrRLK1L13 was knocked down, the pollen tubes grew faster (Fig. 1b–d), indicating that PbrCrRLK1L13 acted as a negative regulator in pear pollen tube growth. Although BUPS in Arabidopsis and RUPO in rice have been shown to play important roles in pear pollen tube growth, the downstream signaling pathway in pollen tube growth is poorly understood. In the present study, we identified PbrCrRLK1L13, which acts as a surface regulator for the downstream signaling pathway of the PbrROP1/2 signal to affect the balance of cellulose contents and regulate pollen tube growth.
Membrane receptor proteins act as a bridge connecting extracellular signals and cytoplasmic proteins. Previous reports have revealed that the ROP signaling pathway is regulated by receptor kinase. For example, AtPRK2 activates ROP1 signaling to control the polarized pollen tube growth [37, 38]. FERONIA RLK mediates Rop GTPase signaling to regulate root hair development [25, 39, 40]. Therefore, we posed a question of whether the ROP signaling pathway is regulated by PbrCrRLK1L13 in pear, as previous reports of the RLK-ROP can mediate pollen tube growth. Here, we provide evidence that PbrGEF8 is a downstream target of PbrCrRLK1L13. When we knock down the expression of PbrGEF8, the pear pollen tube will grow slowly (Fig. 3a–c). In addition, according to Y2H (Fig. 2a) and LCI assays (Fig. 2c and S4a), PbrCrRLK1L13 interacts with PbrGEF8 in vitro and in vivo, indicating that PbrCrRLK1L13 can act as an upstream regulator of PbrGEF8; PbrGEF8 interacts with the KD of PbrCrRLK1L13 via the CT (Fig. 2b), which is essential for GEF activity in an auto-inhibitory manner, similar to the region where AtROPGEF1 physically interacts with AtPRK2 [37]. Besides, the phosphorylation of PbrGEF8 can be directly promoted by PbrCrRLK1L13 (Fig. 2d), indicating that the interaction between PbrCrRLK1L13 and PbrGEF8 is functionally relevant. Such phosphorylation of RopGEFs regulated by RLK proteins has been observed in multiple papers. For example, AGC1.5 Kinase mediates the phosphorylation of RopGEFs to control pollen tube growth in Arabidopsis [41]. The phosphorylation of RopGEFs by AtPRK2 is essential for the polarized pollen tube growth [37]. Based on the results, phosphorylation of RopGEFs by RLKs may represent a general mechanism for ROP activity to control growth and development.
ROP GTPases act as switches cycling between GDP and GTP, which are important for plant growth and development [13, 16, 18]. Although ROP GTPases have been identified in many plants, their biological role in pear remains uncharacterized. In this study, we identified 112 small GTPase genes in the pear genome (Table S1). Phylogenetic tree analysis of pear and Arabidopsis shows that the small GTPase gene family includes four subgroups: Rab, Rop, Ran, and Arf/Sar (Fig. S2). The Rop subfamily contains the fewest members in pear, including an RHO domain with proved function in pollen tube growth in Arabidopsis [11, 15, 37, 42]. Based on the transcriptome data from pear pollen tube growth and RT-PCR in the tissue-specific analysis, PbrROP1 and PbrROP2 are highly expressed in pear pollen tube growth (Fig. 4 and S3), which indicates that PbrROP1 and PbrROP2 play important roles in pear pollen tube growth. When PbrROP1 and PbrROP2 are knocked down in pear pollen, the pear pollen tubes will grow more slowly (Fig. 5c–e), demonstrating that PbrROP1 and PbrROP2 could regulate pear pollen tube growth.
ROPGEFs have been identified as activators of ROP GTPases in Arabidopsis [20, 37, 43]. However, the connection between ROPGEFs and ROP in pear has not been revealed. Here, interaction studies reveal that PbrGEF8 can bind to PbrROP1 and PbrROP2 (Fig. 5a, 5b and S4b), demonstrating that PbrGEF8 can be considered upstream regulators of PbrROP1 and PbrROP2. In addition, the phenotype of PbrROP/2 is consistent with that of PbrGEF8 in pear pollen tube growth, indicating that PbrGEF8 can positively mediate the PbrROP1/2 signal to regulate pollen tube growth. Therefore, PbrCrRLK1L13 regulates the ROP GTPase activity via RopGEFs. The phenotypes of PbrCrRLK1L13 and PbrROP1/2 in pollen tube growth rate are opposite, suggesting a negative regulation between PbrCrRLK1L13 and ROP activity.
The pollen tube is a fast tip-growing cell, and the cell wall is important for pollen tube growth. The calcium gradient, cytoskeleton dynamics, and cellulose distribution are essential to maintaining the cell wall integrity [44, 45]. Cellulose, a major component of the cell wall, is critical for pollen tube growth. However, as yet, little is known of the molecular mechanism that controls cellulose deposition in pollen tubes. Here, functional characterization of the cellulose deposition pathway is studied. In this study, via the as-ODN approach, the cellulose content in the pollen tube wall becomes imbalanced in as-ODN-PbrCrRLK1L13 (Fig. 1d), as-ODN-PbrGEF8 (Fig. 3e), and as-ODN-PbrROP1/2 (Fig. 5f). Thus, RLK-ROP signaling mediates cellulose synthesis and deposition, disrupts the stability of the pollen tube, and further affects pollen tube growth. PbrCrRLK1L13 is identified as a surface regulator of ROP signaling for cellulose deposition-mediated elongation of the pear pollen tube, which was puzzling initially but provided a novel mechanism for the RLK-mediated cell wall to regulate pollen tube growth. However, the connection between Rop GTPase and cellulose signaling remains elusive, so future work needs to unravel the underlying mechanisms.
In summary, based on the findings, we propose a model of signaling events in which the interaction of receptor kinase and cytoplasmic protein can contribute to pear pollen tube growth (Fig. 6). During pear pollen tube elongation, PbrCrRLK1L13 binds to the CT region of PbrGEF8, and phosphorylated PbrGEF8 mediates PbrROP1 and PbrROP2, thus leading to the imbalance of cellulose deposition and regulating pear pollen tube growth.
Materials and methods
Sequence collection and identification of small GTPase genes in Chinese white pear
To identify potential members of the small GTPase gene family in pear, we performed multiple database searches. The small GTP-binding proteins from Arabidopsis were used as query sequences in a BLASTP search for predicted pear proteins. The small GTP-binding protein sequences in pear were downloaded from the Pear Genome Project database (http://peargenome.njau.edu.cn/). Small GTPase genes with expected E values <0.001 were collected. After removing redundant sequences, a total of 112 small GTPase genes were identified in pear genome.
Phylogenetic analysis
The complete small GTPase protein sequences from pear and Arabidopsis were used for multiple sequence alignments with ClustalX (http://www.clustal.org/). Phylogenetic trees were constructed with the full-length amino acid sequences of small GTPase proteins using the neighbor-joining method in MEGA6.
Analysis of small GTPase gene expression based on transcriptome data from pear pollen tube growth
Pollen samples were obtained from the Chinese white pear “Dangshansuli”. Based on the previous transcriptome sequencing data and the expression data from pollen, we measured the expression levels of small GTPase genes. The RNA-seq data was downloaded from the website (http://peargenome.njau.edu.cn/) of our center. During the four development stages of pear pollen [mature pollen grains (MP), hydrated pollen grains (HP), growing pollen tubes (PT), and stopped-growth pollen tubes (SPT)], the RPKM (reads per kilobase per million) values, were used to measure the gene expression level. The log2-transformed RPKM values were adopted to measure the expression of the small GTPase genes and generate heat maps. RPKM values of the small GTPase genes expressed in pollen are shown in Table S2.
Pollen culture
Pollen of Pyrus bretschneideri Rehd. cv. Danshansuli was cultured at 25°C in darkness in the basal medium, which contained (w/v): 0.01% KNO3, 0.07% Ca(NO3)2·4H2O, 15% polyethylene glycol 4000, 10% sucrose, 0.01% H3BO3, and 0.02% MgSO4·7H2O, and pH of the medium was adjusted to 6.0~6.5 with Tris.
Quantitative RT-PCR
The purified total RNA, obtained from different tissues in pear, which was used for reverse transcription into cDNA for quantitative PCR (qRT-PCR). Subsequently, qRT-PCR was performed via SYBR green (Takara, Dalian, China) on an IQ5 real-time PCR machine (Bio-Rad, Hercules, CA, USA). The pear TUB gene acted as an internal control, and relative expression levels were calculated by the 2−△△CT method described previously [46]. Primers were designed based on the target genes via Primer5 software, as shown in Table S3.
Antisense oligodeoxynucleotides
Antisense oligodeoxynucleotides (ODN) assay was conducted as described previously [47]. The ODN sequences of PbrCrRLK1L13, PbrROP1, PbrROP2, and PbrGEF8 were constructed via the RNAfold Web Server (http://rna.tbi.univie.ac.at/cgi-bin/RNAfold.cgi). The ODN sequences were phosphorothioated and purified by high-performance liquid chromatography. Pollen was pre-cultured in a liquid medium at 25°C and 120 rpm for 45 min. The ODN primers mixed with the transfection reagent (Lipofectamine® 2000, Thermo Fisher Scientific, Shanghai) were added to the pre-cultured pollen, and the pollen tubes were retained in the culture for 1.5 h. The pollen tubes were imaged with a microscope and measured with IPWin32 software. Finally, the pollen samples were centrifuged to remove the medium and retain the precipitate. Samples were preserved frozen in liquid nitrogen at −80°C. Total RNA was harvested with these materials for qRT-PCR. The primers of ODN are listed in Table S4.
Detection of cellulose in cell walls of pollen tubes
The pear pollen was pre-cultured and treated with the ODN, the samples were incubated for 2 h. After the pollen tubes were cultured, 1 mg/mL of cellulose fluorescent dye (fluorescent whitening agent; Sigma-Aldrich) was added for 5 min incubation. The pollen tubes were then washed three times with pollen medium. The pollen tubes were observed under a Zeiss LSM700 laser confocal microscope and photographed. ImageJ software was used to calculate the pixel intensity of formazan precipitation in the pollen tube wall. The average value of fluorescence intensity on both sides of the pollen tube cell wall was taken as the fluorescence intensity of each pollen tube.
Yeast two-hybrid assay
The Y2H assay was analyzed as described previously [48]. PbrGEF8 was cloned into the pGADT7 vector, while PbrCrRLK1L13 kinase domain (aa, 441–770) and PbrROP1/2 were cloned into the pGBKT7 vector. The constructed vectors were co-transformed into the yeast strain AH109. Transformants were selected on SD/−Leu-Trp and verified on SD/−Leu-Trp-His-Ade. The primers used for Y2H assay are listed in Table S5.
Luciferase complementation imaging assay
A LCI assay was conducted as described previously [49]. The full-length coding sequences of PbrCrRLK1L13, PbrROP1, and PbrROP2 were cloned into the vector cLUC, while the full-length coding sequences of PbrGEF8 were cloned into the vector nLUC. Paired constructs of PbrGEF8-nLUC, PbrCrRLK1L13-cLUC, PbrROP1-cLUC, and PbrROP2-cLUC were transiently co-expressed in the leaves of N. benthamiana through Agrobacteria-mediated co-infiltration. The primers used for LCI assay are listed in Table S6.
Expression and purification of the His–PbrGEF8 fusion protein
The mature coding sequence for the PbrGEF8 gene was amplified via PCR with BamHI and XbaI sites at the 5′- and 3′-ends. The primers for His-PbrGEF8 were: forward, ctcggtaccctcgagggatccATGGTTCGGGCGCTCG, and reverse, agcagagattacctatctagaTTAGTGGCGTGCCGTTGG. The amplified PCR products were ligated into the pCold-TF DNA vector (Takara Bio, Dalian, China). The recombinant plasmids were transferred into E. coli BL21 (DE3) cells. When the OD600 of the culture reached 0.6 at 37°C, we added isopropylthio-β-D-1-galactopyranoside (IPTG, 0.5 mM) to express the protein (His-PbrGEF8) at 15°C. SDS-PAGE and western blot were used to confirm expression of proteins. The His-PbrGEF8 protein was purified using Ni-NTA His Bind Resin (EMD Millipore, MA, USA). The purified buffers contained the bind buffer (20 mM Tris–HCl, 500 mM NaCl, 10 mM imidazole, pH 7.9), wash buffer (20 mM Tris–HCl, 500 mM NaCl, 100 mM imidazole, pH 7.9), and elution buffer (20 mM Tris–HCl, 500 mM NaCl, 600 mM imidazole, pH 7.9). The protein quality was assessed by SDS/PAGE.
Expression and purification of GST-PbrCrRLK1L13-ECD
The encoding sequence of the IKD of PbrCrRLK1L13 was cloned into the pGEX-4 T-1 vector. The corresponding primers for GST-PbrCrRLK1L13-ECD were: forward, 5′- GATCTGGTTCCGCGTGGATCCATGAAATGGCGAAAGAGACCT −3′, and reverse, 5′- CTCGAGTCGACCCGGGAATTCCCTACCATTTAGATTGGAAAATTG −3′. The expression vectors were transformed into E. coli BL21 (DE3). When the OD600 of the subcultured medium reached 0.6, IPTG (final concentration, 0.5 mM) was added to induce protein expression at 26°C for 8 h. The GST-PbrCrRLK1L13-ECD protein was purified using GST agarose resin (Thermo Fisher Scientific, USA) with 1 mL binding buffer (25 mM Tris•HCl, pH 7.5, 100 mM NaCl, and 1 mM DTT) three times. The proteins were confirmed using SDS-PAGE and western blot.
Phos-tag phosphorylation assay
Phos-tag SDS/PAGE was performed as described previously [50]. PbrCrRLK1L13 and PbrGEF8 phosphorylation activity assays were performed with the reaction buffer [25 mM Tris•HCl (pH 7.5), 1 mM MnSO4, 0.5 mM CaCl2, 2 mM DTT, 10 μM ATP, 1 mM DTT, and 1 mM ATP] at 30°C for 2 h and heated at 100°C for 5 min in 4 × SDS sample buffer. Denatured samples were subjected to Phos-tag SDS-PAGE. Phos-tag SDS/PAGE contains concentrated gel [4% (w/v) acrylamide, 125 mM Tris/HCl, pH 6.8, 0.1% (w/v) SDS, 0.2% (v/v) N,N,N′,N′_tetramethylethylenediamine (TEMED), and 0.08% (w/v) ammonium persulfate (APS)] and a separating gel [12% (w/v) acrylamide, 375 mM Tris/HCl, pH 8.8, 0.1% (w/v) SDS, 75 μM phos-tag acrylamide AAL-107, 150 μM MnCl2, 0.1% (v/v) TEMED, and 0.05% (w/v) APS]. Finally, we conducted western blot analysis to detect phosphorylated PbrGEF8 via the anti-6 × His antibody, the phosphatase treatment was used as a negative control.
Statistical analysis
GraphPad Prism (version 6.05) was used to conduct the statistical analysis. The asterisk represents significant statistical differences performed via Student’s t-test (P < 0.05). All experimental data represented at least three independent experiments and are expressed as the mean ± standard error (SE).
Acknowledgements
This study is supported by the National Key Research and Development Program of China (2018YFD1000107), the National Natural Science Foundation of China (32172543, 31772256, 31772276), and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Contributions
X.B.K. performed most of the experiments and wrote the manuscript; P.C. and Q.K.H. performed the protein expression assays; P.W. analyzed the data; S.L.Z. revised the manuscript; J.Y.W. conceived the research and revised the manuscript. All authors have read and approved the final manuscript.
Data availability
The authors confirmed that the data in this study are available both in the article and supplementary materials. Original data is available from the corresponding authors upon request.
Conflict of interest statement
The authors declared no competing interests.
Supplementary data
Supplementary data is available at Horticulture Research Journal online.



![The expression profile of small GTPase genes during pear pollen tube growth. (a) Heat map of RNA-Seq expression of small GTPase genes in the four stages of pollen tube growth [mature, dry pollen (MP), hydrated pollen grains (HP), pollen tubes (PT), and stopped-growth pollen tubes (SPT)]. (b) The qRT-PCR analysis of the expression of eight small GTPase genes in four stages of pear pollen tube growth. PbrTUB was used as the reference gene.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/hr/9/10.1093_hr_uhab034/9/m_uhab034f4.jpeg?Expires=1716432109&Signature=EpUeEbcPz1JMshSVk4zvGNg~oSrY5EyoTOKTL-MSduSdVMfbOpEuKZbysKQT~HE1sADhuRE-m7nkhr9IhvA9n9cqzg2EN2lU3wXs-OmSX3jU4u2JT5V-KjPoNJZMRvZjKuJGLluHtfYA369N3Sb4TEeg3VoZ7yK1fv-UDQCRmzuHJR1sJ7EPXf2Oo4-c-1ukkHllIzFz6rGzkFjq5cAJi96lomhR1CjFnatMaVyzFmAhmb4eDFf98MK5~6mSSgiH5MfVOn~gXxHJXc51WXDUo4siPMZ-TQzYSzZ289a0x8mzSNtIfYXSMQDvKT48n~V-v~VJKC7rOTMgKs-DlEW9VA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)