-
PDF
- Split View
-
Views
-
Cite
Cite
Miquel Taron, Rafael Rosell, Enriqueta Felip, Pedro Mendez, John Souglakos, Maria Sanchez Ronco, Cristina Queralt, Joaquim Majo, Jose Miguel Sanchez, Jose Javier Sanchez, Jose Maestre, BRCA1 mRNA expression levels as an indicator of chemoresistance in lung cancer, Human Molecular Genetics, Volume 13, Issue 20, 15 October 2004, Pages 2443–2449, https://doi.org/10.1093/hmg/ddh260
Close - Share Icon Share
Abstract
Lung cancer is the most common cancer, with dismal outcome. Treatment approaches, including cisplatin-based chemotherapy and surgery, are currently based on the clinical classification of the tumor, without genetic assessment for predicting differential chemosensitivity. BRCA1 plays a central role in DNA repair, and decreased BRCA1 mRNA expression in the human breast cancer HCC1937 cell line caused cisplatin hypersensitivity, but the relation between BRCA1 and survival in lung cancer patients has never been examined. We used real-time quantitative polymerase chain reaction to determine BRCA1 mRNA levels in 55 surgically resected tumors of non-small-cell lung cancer patients who had received neoadjuvant gemcitabine/cisplatin chemotherapy, and divided the gene expression values into quartiles. When results were correlated with outcome, two cut-offs were observed; patients with levels <0.61 had better outcome, and those >2.45 had poorer outcome. Median survival was not reached for the 15 patients in the bottom quartile, whereas for the 28 in the two middle quartiles, it was 37.8 months (95% CI, 10.6–65), and for the 12 patients in the top quartile, it was 12.7 months (95% CI, 0.28–28.8) (P=0.01). Moreover, when patients were stratified by pathologic stage, those in the bottom quartile had a decreased risk of death (HR=0.206; 95% CI, 0.05–0.83; P=0.026) compared with those in the top quartile, and those in the two middle quartiles also had a decreased risk of death (HR=0.294; 95% CI, 0.10–0.83; P=0.020) compared with those in the top quartile. BRCA1 expression is potentially an important tool for use in cancer management and should be assessed for predicting differential chemosensitivity and tailoring chemotherapy in lung cancer.
INTRODUCTION
Breast cancer 1 (BRCA1) plays a crucial role in DNA repair, and decreased BRCA1 mRNA expression has been observed in both sporadic and hereditary breast cancers (1); however, its potential effect in lung cancer has never been examined. BRCA1 is implicated in transcription-coupled nucleotide excision repair (TC-NER), and modulation of its expression leads to modification of TC-NER and hence to radio- and chemoresistance. Upregulation of BRCA1 expression led to increased cisplatin resistance in the SKOV-3 human ovarian cancer cell line (2), and restoration of BRCA1 in the BRCA1-negative HCC1937 human breast cancer cell line restored radioresistance (3). BRCA1 is also involved in homologous recombination repair (HRR) and non-homologous end joining, in response to DNA damage (4). In addition, it is a component of a large DNA repair complex termed the BRCA1-associated genome surveillance complex, which contains a number of mismatch repair proteins, indicating a potential role for BRCA1 in mismatch repair (1,4). BRCA1 may also be a regulator of mitotic spindle assembly, as BRCA1 and β-tubulin colocalize to the microtubules of the mitotic spindle and to the centrosomes (5). Finally, enhanced BRCA1 expression has been linked to apoptosis through the c-Jun N-terminal kinase pathway (6), which is activated by cisplatin-induced DNA damage; inhibition of this pathway increased cisplatin sensitivity in cell lines (7). Decreased BRCA1 mRNA expression in a breast cancer cell line, as determined by real-time quantitative polymerase chain reaction (RT-QPCR), led to greater sensitivity to cisplatin and etoposide, but to greater resistance to the microtubule-interfering agents paclitaxel and vincristine (8). Recently, furthermore, reconstitution of wild-type BRCA1 into the BRCA1-negative HCC1937 breast cancer cell line (9) resulted in a 20-fold increase in cisplatin resistance and, in contrast, in a 1000–10 000-fold increase in sensitivity to antimicrotubule drugs (paclitaxel and vinorelbine) (4,10). Mouse models carrying conditional disruption of BRCA1 were highly sensitive to doxorubicin and gamma irradiation but resistant to tamoxifen, providing additional evidence for differential chemosensitivity linked to BRCA1 expression (11). When BRCA1 expression was examined by semi-quantitative PCR in women with sporadic breast cancer, low BRCA1 mRNA levels (bottom quartile) were associated with a higher frequency of distant metastases (12).
Despite the wealth of data in cell lines and mouse models, only one small study has examined the correlation of BRCA1 and BRCA2 mRNA expression with response to chemotherapy in the clinical setting. Among 25 women with docetaxel-treated locally advanced or metastatic breast cancer (13), only BRCA2 mRNA levels were significantly lower in responders than in non-responders, though a slight difference was also observed for BRCA1. Non-small-cell lung cancer (NSCLC) accounts for ∼80% of all lung cancers, with 1.2 million new cases worldwide each year. NSCLC resulted in more than 1 million deaths worldwide in 2001, and is the leading cause of cancer-related mortality in both men and women (31 and 25%, respectively) (14). The overall 5-year survival of patients with NSCLC has remained at <15% for the past 20 years. Stage grouping of TNM subsets (T, primary tumor; N, regional lymph nodes; M, distant metastases) permits the identification of patient groups with similar prognosis and treatment options. Five-year survival is around 25% for pathologic stage IIB (T1-2N1M0, T3N0M0), 13% for stage IIIA (T3N1M0, T1-2-3N2M0) and a low 7% for stage IIIB (T4N0-1-2M0) (15). Small randomized studies of cisplatin-based chemotherapy followed by surgery in clinical stage IIIA (16) or stage IIB–IIIB (17) showed remarkable improvement in survival over patients treated either with surgery alone or with surgery followed by radiotherapy. Event-free survival was similar in the two studies (12.7 (16) and 20 (17) months in the neoadjuvant chemotherapy arm and 5.8 (16) and 5 (17) months in the surgery arm). In general, neoadjuvant chemotherapy induces tumor shrinkage and sterilizes metastatic lymph nodes, leading to pathologic downstaging in ∼33% and complete pathologic remission in up to 14% of patients (18). Although a wealth of data indicates that changes in the level of several gene transcripts can modulate differential chemosensitivity between patients with the same TNM subset, at present no predictive genetic markers of chemotherapy response are used for tailoring treatment.
On the basis of the evidence for the role of BRCA1 in breast and ovarian cancers, we reasoned that BRCA1 mRNA expression could also play an important role in predicting differential chemotherapy sensitivity in NSCLC. We examined the potential predictive value of BRCA1 mRNA expression in resected specimens from stage IIB, IIIA and IIIB NSCLC patients treated with neoadjuvant gemcitabine/cisplatin followed by surgery.
RESULTS
Median survival was 37.8 months (95% CI, 27–48.5 months) for all patients, 51.9 months (95% CI, 31.6–72.4 months) for patients who underwent lobectomy and 25.8 months (95% CI, 12.7–38.8 months) for those who underwent pneumonectomy. BRCA1 was detected in all tumors, although there was considerable variation in its level of expression, with values relative to the β-actin internal control ranging ∼37-fold, from 0.28 to 10.43. Amplification plots obtained for the genes BRCA1 and β-actin are shown in Figure 1. Values ranged from 0.28 to 0.61 [interpatient coefficient of variation (ICV), 30.7%] for the 15 patients in the bottom quartile, from 0.65 to 1.20 (ICV, 17.4%) for the 14 patients in the second quartile, from 1.23 to 2.37 (ICV, 17.7%) for the 14 patients in the third quartile, and from 2.45 to 10.43 (ICV, 54.7%) for the 12 patients in the top quartile. Owing to the similar values and ICVs observed in the second and third quartiles, these two groups were merged for statistical analyses.
No differences in clinical characteristics were observed according to quartiles of BRCA1 mRNA expression levels (Table 1). However, for patients in the bottom quartile, radiographic response tended to be higher than for those in the middle or top quartiles (66.7, 57.1 and 58.3%, respectively), complete resection was attained more often (93.3, 78.6 and 83.3%, respectively), and a lobectomy was performed more often [73.3, 32.1 (P=0.005) and 58.3% (P=0.2), respectively] (Table 1). Median survival was not reached for the 15 patients in the bottom quartile, whereas for the 28 patients in the two middle quartiles, it was 37.8 months (95% CI, 10.6–65), and for the 12 patients in the top quartile, it was 12.7 months (95% CI, 0.28–28.8) (P=0.01) (Fig. 2). Five patients who attained a complete pathologic response (T0N0) were all in the bottom quartile of BRCA1 levels (Table 2). Conversely, in the majority of patients with high BRCA1 levels, no clinical or pathologic downstaging was observed following chemotherapy and surgery (Table 3). When patients were stratified by pathologic stage, those in the bottom quartile had a decreased risk of death (HR=0.206; 95% CI, 0.05–0.83; P=0.026) compared with those in the top quartile, and those in the two middle quartiles also had a decreased risk of death (HR=0.294; 95% CI, 0.10–0.83; P=0.020) compared with those in the top quartile. When patients were stratified by clinical stage, a similar pattern was observed. Those in the bottom quartile had a decreased risk of death (HR=0.220; 95% CI, 0.06–0.77; P=0.018) compared with those in the top quartile, and those in the two middle quartiles also had a decreased risk of death (HR=0.430; 95% CI, 0.17–1.1; P=0.078) compared with those in the top quartile.
DISCUSSION
Resistance to cytotoxic drugs is the major impediment to the successful treatment of many tumor types, especially in lung cancer. The elucidation of the mechanisms of this resistance is crucial for improving treatment outcome and for selecting and customizing chemotherapy. Upregulation of DNA repair genes has been related to resistance to cisplatin and radiotherapy. The repair of cisplatin DNA damage occurs via the activity of the nucleotide excision repair endonuclease (ERCC1/XPF) and Rad51-related HRR proteins (19,20). We had previously used RT-QPCR to assess mRNA levels of ERCC1 and RRM1, genes related to global genome NER but not directly to TC-NER (20), and found that overexpression of either of these genes influenced survival in gemcitabine/cisplatin-treated stage IV NSCLC patients (21–23). However, unlike ERCC1, BRCA1 is involved in TC-NER (3,24), and may thus be a better predictive marker of cisplatin response.
The availability of fresh tumor tissue in the clinical setting is not yet common, and the recovery of mRNA from paraffin-embedded tissue has therefore become very important. mRNA real-time assays permit quantitative and accurate measurement of gene expression (25). In the present study, we used RT-QPCR to quantitatively analyze BRCA1 mRNA expression in processed formalin-fixed, paraffin-embedded tissues from resected lung cancer patients and demonstrated that BRCA1 expression can be accurately assessed. BRCA1 gene expression was detectable in all 55 samples analyzed in this study. Patients in the bottom quartile of BRCA1 mRNA levels (<0.61) obtained the maximum benefit of neoadjuvant gemcitabine/cisplatin chemotherapy, whereas those in the top quartile (>2.45) had the poorest outcome. These findings support the hypothesis that BRCA1 mRNA expression levels could be an indicator of differential cisplatin sensitivity in NSCLC, which is consistent with findings in pre-clinical models in breast cancer (2–4,8,10,11). The HCC1937 cell line (9), from a primary breast carcinoma with a germline BRCA1 mutation, was transfected with either wild-type BRCA1 or an empty vector to test response to antimicrotubule drugs (paclitaxel and vinorelbine) and DNA-damaging drugs (cisplatin, bleomycin and etoposide). Reconstitution of wild-type BRCA1 function into HCC1937 resulted in a 1000-fold increase in sensitivity to paclitaxel and a 10 000-fold increase in sensitivity to vinorelbine. Conversely, it resulted in a 2-fold increase in resistance to bleomycin, a 20-fold increase in resistance to cisplatin and a >100-fold increase in resistance to etoposide (10). Interestingly, BRCA1 failed to modulate resistance or sensitivity to the antimetabolite 5-fluorouracil, perhaps reflecting the distinct mode of action of antimetabolites (10).
BRCA1 mRNA is reduced in sporadic breast cancer cells despite a lack of mutations. Aberrant cytosine methylation of the BRCA1 CpG island promoter may be a partial mechanism of BRCA1 repression in sporadic breast cancer (26,27). Along the same lines, it has been shown that the Fanconi anemia (FANC)-BRCA pathway (28) regulates cisplatin sensitivity, with the clinical finding that methylation of FANCF confers increased cisplatin sensitivity in ovarian cancer (29). FANC genes interact with those involved in DNA repair pathways, including BRCA1, Rad-51, ATM and NBS1 (28).
Cigarette smoking remains the principal cause of lung cancer, with 85–90% of all lung cancer patients having smoked cigarettes at some time. The profound role of cigarette smoking in lung cancer development and DNA damage could also contribute to the dismal outcome and the limited effect of chemotherapy as DNA repair capacity is stimulated in response to DNA damage caused by tobacco carcinogens. Among heavy smokers, both lung cancer patients and controls have more proficient DNA repair capacity (measured by host-cell reactivation assay) in lymphocytes than non- or light smokers (30). Elevated DNA repair capacity has been associated with cisplatin resistance both in NSCLC cell lines (31) and in lung cancer patients (32). The expression levels of DNA repair genes, including BRCA1, can be expected to be elevated in lung cancer patients, particularly those who are heavy smokers.
Several cisplatin-based doublets demonstrated similar survival in a randomized study of more than 1000 metastatic NSCLC patients (33); furthermore, other studies have found no survival differences between cisplatin alone and cisplatin/paclitaxel (34), or between docetaxel alone and docetaxel/cisplatin (35). On the basis of our results and of pre-clinical data (10), we can speculate that patients with low BRCA1 mRNA levels can benefit from single-agent cisplatin, whereas those with high levels would benefit from single-agent docetaxel or paclitaxel. In contrast, high BRCA1 levels may diminish the synergism between taxanes and cisplatin or carboplatin. Although sensitivity to antimetabolites, such as gemcitabine, may not be affected by BRCA1 levels, gemcitabine/cisplatin synergism may be partially abrogated in tumors with high BRCA1 mRNA levels; on the other hand, these tumors may benefit from the synergism observed between taxanes and gemcitabine. To date, no other clinical study has assessed BRCA1 mRNA expression as a predictive marker of chemotherapy response in lung cancer. If further research validates our findings, BRCA1 mRNA assessment will provide an important tool for customizing NSCLC chemotherapy in order to improve survival in this very common and fatal disease.
MATERIALS AND METHODS
Patients
In all patients, neoadjuvant chemotherapy was indicated after evaluation by a thoracic surgeon, a radiologist, a medical oncologist and a radiation oncologist. Patients received three cycles of neoadjuvant chemotherapy; 51 received cisplatin 100 mg/m2 day 1 plus gemcitabine 1250 mg/m2 days 1 and 8 every 21 days, and four received carboplatin AUC=5 day 1 plus gemcitabine 1000 mg/m2 days 1 and 8 every 21 days. A thoracotomy was performed within 4–5 weeks after the last chemotherapy cycle; the surgical procedure was based on the extent of tumor at the time of the initial presentation.
BRCA1 gene expression analysis by RT-QPCR
We examined BRCA1 gene expression in formalin-fixed, paraffin-embedded surgical resected specimens from the 55 patients as previously described (36,37). After standard tissue sample deparaffinization using xylene and alcohols, samples were lyzed in a Tris–chloride, EDTA, sodium dodecyl sulfate (SDS) and proteinase K containing buffer. RNA was then extracted with phenol–chloroform–isoamyl alcohol followed by precipitation with isopropanol in the presence of glycogen and sodium acetate. RNA was resuspended in RNA storage solution (Ambion Inc., Austin TX, USA) and treated with DNase I to avoid DNA contamination. cDNA was synthesized using M-MLV retrotranscriptase enzyme. Template cDNA was added to TaqMan Universal Master Mix (AB; Applied Biosystems, Foster City, CA, USA) in a 12.5µl reaction with specific primers and probe for each gene. The primer and probe sets were designed using Primer Express 2.0 Software (AB). Quantification of gene expression was performed using the ABI Prism 7900HT Sequence Detection System (AB). Primers and probe for BRCA1 mRNA expression analysis were designed according to the Ref Seq NM_007294 (http://www.ncbi.nlm.nih.gov/LocusLink). Forward primer is located in exon 8 (position 4292–4317 bp), reverse primer in exon 9 (position 4336–4360 bp) and probe in the exon 8/9 junction (position 4313 bp–4333 bp). The PCR product size generated with these primers was 69 bp. The primers and 5′labeled fluorescent reporter dye (6FAM) probe were as follows: β-actin: forward 5′-TGA GCG CGG CTA CAG CTT-3′, reverse 5′-TCC TTA ATG TCA CGC ACG ATT T-3′, probe 5′-ACC ACC ACG GCC GAG CGG-3′; BRCA1: forward 5′-GGC TAT CCT CTC AGA GTG ACA TTT TA-3′, reverse 5′-GCT TTA TCA GGT TAT GTT GCA TGG T-3′, probe 5′-CCA CTC AGC AGA GGG-3′.
Relative gene expression quantification was calculated according to the comparative Ct method using β-actin as an endogenous control and commercial RNA controls (Stratagene, La Jolla, CA) as calibrators. Final results, were determined as follows: 2−(ΔCt sample−ΔCt calibrator), where ΔCt values of the calibrator and sample are determined by subtracting the Ct value of the target gene from the value of the β-actin gene. In all experiments, only triplicates with a SD of the Ct value <0.20 were accepted. In addition, for each sample analyzed, a retrotranscriptase minus control was run in the same plate to assure lack of genomic DNA contamination (Fig. 1).
Statistical methods
In order to provide an easily interpretable evaluation of the effect of BRCA1 mRNA expression, gene expression values were divided into quartiles. ICVs were calculated to assess similarities between quartiles. Hazard ratios were calculated with the univariate Cox model, stratifying by pathologic and clinical stage, and comparison between Kaplan–Meier survival curves was performed with the log-rank test. All tests of statistical significance were two-sided, with a statistical power of 80%, and significance was set at 0.05 except in multiple comparisons, where it was set at 0.017 in accordance with the Bonferroni correction.
ACKNOWLEDGEMENTS
The authors thank Renée O'Brate for assistance with the manuscript. This study was supported by Spanish Ministry of Health research grants provided through Red Temática de Investigación Cooperativa de Centros de Cáncer (CO-010) and through Ayuda Carlos III (RCESP 03/09) and by funding from La Fundació Badalona Contra El Càncer.
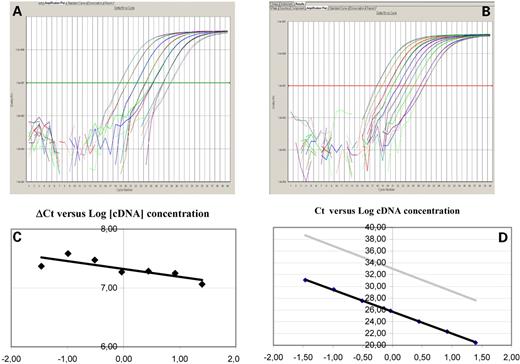
Figure 1. Example of the amplification plots (ΔRn versus cycle number) of (A) β-actin and (B) BRCA1 cDNAs. Both figures correspond to serial dilutions of cDNA obtained from one of the samples. (C) and (D) Examples of the validation curves for relative quantification. Different primers and probe concentrations were assayed for β-actin and BRCA1 gene expression analysis to obtain the optimal PCR efficiency. In order for the relative quantification to be valid, the amplification efficiency of the target (BRCA1) and the reference (β-actin) amplification must be approximately equal. A sensitive method for assessing whether two amplicons have the same amplification efficiency is to see how ΔCt varies when using a serial dilution of a control cDNA. We performed two validations: one using control cDNA, another using cDNA from paraffin-embedded samples. (C) Several runs with serial dilutions were performed to confirm that the slope <0.1 in the plot ΔCt value versus log10 input amount cDNA, defined as Ct BRCA1 in each dilution minus Ct β-actin in the same dilution. (D) For primers and probe sets, the slope of the plot Ct versus log10 input amount cDNA needed to be between −3.25 and −3.45, since a slope of −3.33 represents 100% efficiency. The slopes in our assays were −3.36 for β-actin and −3.32 for BRCA1, with a correlation coefficient (R2) >0.98.
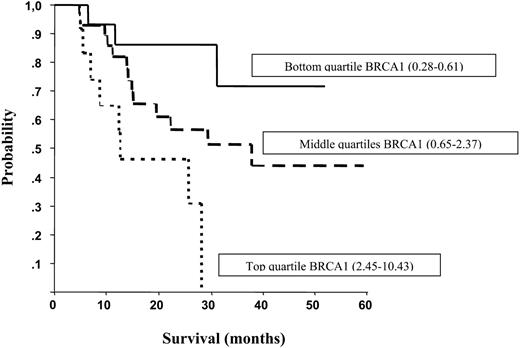
Figure 2. Median survival according to quartiles of BRCA1 mRNA expression levels. Median survival was not reached for those in the bottom quartile, whereas it was 37.8 months for those in the middle quartiles, and 12.7 months for those in the top quartile.
Patient characteristics according to BRCA1 mRNA expression levels (bottom quartile versus two middle quartiles versus top quartile)
| . | Bottom quartile of BRCA1 levels (0.28–0.61) N . | Middle quartiles of BRCA1 levels (0.65–2.37) N . | Top quartile of BRCA1 levels (2.45–10.43) N . |
|---|---|---|---|
| Sex | |||
| Female | 3 (20%) | 3 (10.7%) | 0 |
| Male | 12 (80%) | 25 (89.3%) | 12 (100%) |
| Age | |||
| Median, range | 60 (49–74%) | 65 (51–76%) | 61 (45–71%) |
| Histology | |||
| Squamous cell carcinoma | 5 (33.3%) | 16 (57.1%) | 5 (41.7%) |
| Adenocarcinoma | 7 (46.7%) | 11 (39.3%) | 2 (16.7%) |
| Large cell carcinoma | 3 (20%) | 1 (3.6%) | 5 (41.7%) |
| Initial staging | |||
| IIB | |||
| T3N0 | 2 (13.3%) | 3 (10.7%) | 1 (8.3%) |
| IIIA | |||
| T3N1 | 0 | 1 (3.6%) | 3 (25%) |
| T1N2 | 0 | 0 | 0 |
| T2N2 | 1 (6.7%) | 6 (21.4%) | 1 (8.3%) |
| T3N2 | 3 (20%) | 7 (25%) | 2 (16.7%) |
| IIIB | |||
| T4N0 | 6 (40%) | 8 (28.6%) | 3 (25%) |
| T4N1 | 1 (6.7%) | 2 (7.1%) | 1 (8.3%) |
| T4N2 | 2 (13.3%) | 1 (3.6%) | 1 (8.3%) |
| Chemotherapy regimen | |||
| Gemcitabine/cisplatin | 15 (100%) | 26 (92.9%) | 10 (83.3%) |
| Gemcitabine/carboplatin | 0 | 2 (7.1%) | 2 (16.7%) |
| Radiographic response | |||
| Partial response | 10 (66.7%) | 16 (57.1%) | 7 (58.3%) |
| Stable disease | 5 (33.3%) | 10 (35.7%) | 4 (33.3%) |
| Progressive disease | 0 | 2 (7.1%) | 1 (8.3%) |
| Surgical results | |||
| Complete resection | 14 (93.3%) | 22 (78.6%) | 10 (83.3%) |
| Incomplete resection | 1 (6.7%) | 5 (17.9%) | 2 (16.7%) |
| Unresectable | 0 | 1 (3.6%) | 0 |
| Surgical procedures | |||
| Lobectomy | 11 (73.3%) | 9 (32.1%) | 7 (58.3%) |
| Pneumonectomy | 4 (26.7%) | 14 (50%) | 5 (41.7%) |
| Bilobectomy | 0 | 4 (14.3%) | 0 |
| Unresectable | 0 | 1 (3.6%) | 0 |
| . | Bottom quartile of BRCA1 levels (0.28–0.61) N . | Middle quartiles of BRCA1 levels (0.65–2.37) N . | Top quartile of BRCA1 levels (2.45–10.43) N . |
|---|---|---|---|
| Sex | |||
| Female | 3 (20%) | 3 (10.7%) | 0 |
| Male | 12 (80%) | 25 (89.3%) | 12 (100%) |
| Age | |||
| Median, range | 60 (49–74%) | 65 (51–76%) | 61 (45–71%) |
| Histology | |||
| Squamous cell carcinoma | 5 (33.3%) | 16 (57.1%) | 5 (41.7%) |
| Adenocarcinoma | 7 (46.7%) | 11 (39.3%) | 2 (16.7%) |
| Large cell carcinoma | 3 (20%) | 1 (3.6%) | 5 (41.7%) |
| Initial staging | |||
| IIB | |||
| T3N0 | 2 (13.3%) | 3 (10.7%) | 1 (8.3%) |
| IIIA | |||
| T3N1 | 0 | 1 (3.6%) | 3 (25%) |
| T1N2 | 0 | 0 | 0 |
| T2N2 | 1 (6.7%) | 6 (21.4%) | 1 (8.3%) |
| T3N2 | 3 (20%) | 7 (25%) | 2 (16.7%) |
| IIIB | |||
| T4N0 | 6 (40%) | 8 (28.6%) | 3 (25%) |
| T4N1 | 1 (6.7%) | 2 (7.1%) | 1 (8.3%) |
| T4N2 | 2 (13.3%) | 1 (3.6%) | 1 (8.3%) |
| Chemotherapy regimen | |||
| Gemcitabine/cisplatin | 15 (100%) | 26 (92.9%) | 10 (83.3%) |
| Gemcitabine/carboplatin | 0 | 2 (7.1%) | 2 (16.7%) |
| Radiographic response | |||
| Partial response | 10 (66.7%) | 16 (57.1%) | 7 (58.3%) |
| Stable disease | 5 (33.3%) | 10 (35.7%) | 4 (33.3%) |
| Progressive disease | 0 | 2 (7.1%) | 1 (8.3%) |
| Surgical results | |||
| Complete resection | 14 (93.3%) | 22 (78.6%) | 10 (83.3%) |
| Incomplete resection | 1 (6.7%) | 5 (17.9%) | 2 (16.7%) |
| Unresectable | 0 | 1 (3.6%) | 0 |
| Surgical procedures | |||
| Lobectomy | 11 (73.3%) | 9 (32.1%) | 7 (58.3%) |
| Pneumonectomy | 4 (26.7%) | 14 (50%) | 5 (41.7%) |
| Bilobectomy | 0 | 4 (14.3%) | 0 |
| Unresectable | 0 | 1 (3.6%) | 0 |
Patient characteristics according to BRCA1 mRNA expression levels (bottom quartile versus two middle quartiles versus top quartile)
| . | Bottom quartile of BRCA1 levels (0.28–0.61) N . | Middle quartiles of BRCA1 levels (0.65–2.37) N . | Top quartile of BRCA1 levels (2.45–10.43) N . |
|---|---|---|---|
| Sex | |||
| Female | 3 (20%) | 3 (10.7%) | 0 |
| Male | 12 (80%) | 25 (89.3%) | 12 (100%) |
| Age | |||
| Median, range | 60 (49–74%) | 65 (51–76%) | 61 (45–71%) |
| Histology | |||
| Squamous cell carcinoma | 5 (33.3%) | 16 (57.1%) | 5 (41.7%) |
| Adenocarcinoma | 7 (46.7%) | 11 (39.3%) | 2 (16.7%) |
| Large cell carcinoma | 3 (20%) | 1 (3.6%) | 5 (41.7%) |
| Initial staging | |||
| IIB | |||
| T3N0 | 2 (13.3%) | 3 (10.7%) | 1 (8.3%) |
| IIIA | |||
| T3N1 | 0 | 1 (3.6%) | 3 (25%) |
| T1N2 | 0 | 0 | 0 |
| T2N2 | 1 (6.7%) | 6 (21.4%) | 1 (8.3%) |
| T3N2 | 3 (20%) | 7 (25%) | 2 (16.7%) |
| IIIB | |||
| T4N0 | 6 (40%) | 8 (28.6%) | 3 (25%) |
| T4N1 | 1 (6.7%) | 2 (7.1%) | 1 (8.3%) |
| T4N2 | 2 (13.3%) | 1 (3.6%) | 1 (8.3%) |
| Chemotherapy regimen | |||
| Gemcitabine/cisplatin | 15 (100%) | 26 (92.9%) | 10 (83.3%) |
| Gemcitabine/carboplatin | 0 | 2 (7.1%) | 2 (16.7%) |
| Radiographic response | |||
| Partial response | 10 (66.7%) | 16 (57.1%) | 7 (58.3%) |
| Stable disease | 5 (33.3%) | 10 (35.7%) | 4 (33.3%) |
| Progressive disease | 0 | 2 (7.1%) | 1 (8.3%) |
| Surgical results | |||
| Complete resection | 14 (93.3%) | 22 (78.6%) | 10 (83.3%) |
| Incomplete resection | 1 (6.7%) | 5 (17.9%) | 2 (16.7%) |
| Unresectable | 0 | 1 (3.6%) | 0 |
| Surgical procedures | |||
| Lobectomy | 11 (73.3%) | 9 (32.1%) | 7 (58.3%) |
| Pneumonectomy | 4 (26.7%) | 14 (50%) | 5 (41.7%) |
| Bilobectomy | 0 | 4 (14.3%) | 0 |
| Unresectable | 0 | 1 (3.6%) | 0 |
| . | Bottom quartile of BRCA1 levels (0.28–0.61) N . | Middle quartiles of BRCA1 levels (0.65–2.37) N . | Top quartile of BRCA1 levels (2.45–10.43) N . |
|---|---|---|---|
| Sex | |||
| Female | 3 (20%) | 3 (10.7%) | 0 |
| Male | 12 (80%) | 25 (89.3%) | 12 (100%) |
| Age | |||
| Median, range | 60 (49–74%) | 65 (51–76%) | 61 (45–71%) |
| Histology | |||
| Squamous cell carcinoma | 5 (33.3%) | 16 (57.1%) | 5 (41.7%) |
| Adenocarcinoma | 7 (46.7%) | 11 (39.3%) | 2 (16.7%) |
| Large cell carcinoma | 3 (20%) | 1 (3.6%) | 5 (41.7%) |
| Initial staging | |||
| IIB | |||
| T3N0 | 2 (13.3%) | 3 (10.7%) | 1 (8.3%) |
| IIIA | |||
| T3N1 | 0 | 1 (3.6%) | 3 (25%) |
| T1N2 | 0 | 0 | 0 |
| T2N2 | 1 (6.7%) | 6 (21.4%) | 1 (8.3%) |
| T3N2 | 3 (20%) | 7 (25%) | 2 (16.7%) |
| IIIB | |||
| T4N0 | 6 (40%) | 8 (28.6%) | 3 (25%) |
| T4N1 | 1 (6.7%) | 2 (7.1%) | 1 (8.3%) |
| T4N2 | 2 (13.3%) | 1 (3.6%) | 1 (8.3%) |
| Chemotherapy regimen | |||
| Gemcitabine/cisplatin | 15 (100%) | 26 (92.9%) | 10 (83.3%) |
| Gemcitabine/carboplatin | 0 | 2 (7.1%) | 2 (16.7%) |
| Radiographic response | |||
| Partial response | 10 (66.7%) | 16 (57.1%) | 7 (58.3%) |
| Stable disease | 5 (33.3%) | 10 (35.7%) | 4 (33.3%) |
| Progressive disease | 0 | 2 (7.1%) | 1 (8.3%) |
| Surgical results | |||
| Complete resection | 14 (93.3%) | 22 (78.6%) | 10 (83.3%) |
| Incomplete resection | 1 (6.7%) | 5 (17.9%) | 2 (16.7%) |
| Unresectable | 0 | 1 (3.6%) | 0 |
| Surgical procedures | |||
| Lobectomy | 11 (73.3%) | 9 (32.1%) | 7 (58.3%) |
| Pneumonectomy | 4 (26.7%) | 14 (50%) | 5 (41.7%) |
| Bilobectomy | 0 | 4 (14.3%) | 0 |
| Unresectable | 0 | 1 (3.6%) | 0 |
BRCA1 mRNA levels and clinical stage in patients who attained complete pathologic response after neoadjuvant chemotherapy followed by surgery
| Patient . | BRCA1 mRNA levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 0.31 | T3N2 | T2N0 | T0N0 |
| 2 | 0.28 | T2N2 | T1N0 | T0N0 |
| 3 | 0.30 | T4N2 | T2N1 | T0N0 |
| 4 | 0.33 | T4N2 | T2N0 | T0N0 |
| 5 | 0.34 | T4N1 | T4N1 | T0N0 |
| Patient . | BRCA1 mRNA levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 0.31 | T3N2 | T2N0 | T0N0 |
| 2 | 0.28 | T2N2 | T1N0 | T0N0 |
| 3 | 0.30 | T4N2 | T2N1 | T0N0 |
| 4 | 0.33 | T4N2 | T2N0 | T0N0 |
| 5 | 0.34 | T4N1 | T4N1 | T0N0 |
BRCA1 mRNA levels and clinical stage in patients who attained complete pathologic response after neoadjuvant chemotherapy followed by surgery
| Patient . | BRCA1 mRNA levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 0.31 | T3N2 | T2N0 | T0N0 |
| 2 | 0.28 | T2N2 | T1N0 | T0N0 |
| 3 | 0.30 | T4N2 | T2N1 | T0N0 |
| 4 | 0.33 | T4N2 | T2N0 | T0N0 |
| 5 | 0.34 | T4N1 | T4N1 | T0N0 |
| Patient . | BRCA1 mRNA levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 0.31 | T3N2 | T2N0 | T0N0 |
| 2 | 0.28 | T2N2 | T1N0 | T0N0 |
| 3 | 0.30 | T4N2 | T2N1 | T0N0 |
| 4 | 0.33 | T4N2 | T2N0 | T0N0 |
| 5 | 0.34 | T4N1 | T4N1 | T0N0 |
Correlation of clinical and pathologic stage in patients in the top quartile of BRCA1 mRNA expression
| Patient . | mRNA BRCA1 levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 2.8 | T3N2 | T3N2 | T2N2 |
| 2 | 5.5 | T2N2 | —a | T2N0 |
| 3 | 10.43 | T3N1 | T2N0 | T2N0 |
| 4 | 2.45 | T3N0 | T3N0 | T3N0 |
| 5 | 4.12 | T4N0 | T1N0 | T4N0 |
| 6 | 6.93 | T4N2 | T3N0 | T2N0 |
| 7 | 2.81 | T4N1 | T4N1 | T3N1 |
| 8 | 3.09 | T3N1 | T2N0 | T2N0 |
| 9 | 5.61 | T3N1 | T2N0 | T2N0 |
| 10 | 3.36 | T3N2 | T3N2 | T2N2 |
| 11 | 2.8 | T4N0 | T3N0 | T3N0 |
| 12 | 2.62 | T4N0 | T4N0 | T2N0 |
| Patient . | mRNA BRCA1 levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 2.8 | T3N2 | T3N2 | T2N2 |
| 2 | 5.5 | T2N2 | —a | T2N0 |
| 3 | 10.43 | T3N1 | T2N0 | T2N0 |
| 4 | 2.45 | T3N0 | T3N0 | T3N0 |
| 5 | 4.12 | T4N0 | T1N0 | T4N0 |
| 6 | 6.93 | T4N2 | T3N0 | T2N0 |
| 7 | 2.81 | T4N1 | T4N1 | T3N1 |
| 8 | 3.09 | T3N1 | T2N0 | T2N0 |
| 9 | 5.61 | T3N1 | T2N0 | T2N0 |
| 10 | 3.36 | T3N2 | T3N2 | T2N2 |
| 11 | 2.8 | T4N0 | T3N0 | T3N0 |
| 12 | 2.62 | T4N0 | T4N0 | T2N0 |
aData not available.
Correlation of clinical and pathologic stage in patients in the top quartile of BRCA1 mRNA expression
| Patient . | mRNA BRCA1 levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 2.8 | T3N2 | T3N2 | T2N2 |
| 2 | 5.5 | T2N2 | —a | T2N0 |
| 3 | 10.43 | T3N1 | T2N0 | T2N0 |
| 4 | 2.45 | T3N0 | T3N0 | T3N0 |
| 5 | 4.12 | T4N0 | T1N0 | T4N0 |
| 6 | 6.93 | T4N2 | T3N0 | T2N0 |
| 7 | 2.81 | T4N1 | T4N1 | T3N1 |
| 8 | 3.09 | T3N1 | T2N0 | T2N0 |
| 9 | 5.61 | T3N1 | T2N0 | T2N0 |
| 10 | 3.36 | T3N2 | T3N2 | T2N2 |
| 11 | 2.8 | T4N0 | T3N0 | T3N0 |
| 12 | 2.62 | T4N0 | T4N0 | T2N0 |
| Patient . | mRNA BRCA1 levels . | Pre-treatment clinical stage . | Post-treatment clinical stage . | Pathologic stage . |
|---|---|---|---|---|
| 1 | 2.8 | T3N2 | T3N2 | T2N2 |
| 2 | 5.5 | T2N2 | —a | T2N0 |
| 3 | 10.43 | T3N1 | T2N0 | T2N0 |
| 4 | 2.45 | T3N0 | T3N0 | T3N0 |
| 5 | 4.12 | T4N0 | T1N0 | T4N0 |
| 6 | 6.93 | T4N2 | T3N0 | T2N0 |
| 7 | 2.81 | T4N1 | T4N1 | T3N1 |
| 8 | 3.09 | T3N1 | T2N0 | T2N0 |
| 9 | 5.61 | T3N1 | T2N0 | T2N0 |
| 10 | 3.36 | T3N2 | T3N2 | T2N2 |
| 11 | 2.8 | T4N0 | T3N0 | T3N0 |
| 12 | 2.62 | T4N0 | T4N0 | T2N0 |
aData not available.
References
Kennedy, R.D., Quinn, J.E., Johnston, P.G. and Harkin, D.P. (
Husain, A., He, G., Venkatraman, E.S. and Spriggs, D.R. (
Abbott, D.W., Thompson, M.E., Robinson-Benion, C., Tomlinson, G., Jensen, R.A. and Holt, J.T. (
Mullan, P.B., Quinn, J.E., Gilmore, P.M., McWilliams, S., Andrews, H., Gervin, C., McCabe, N., McKenna, S., White, P., Song, Y.H. et al. (
Lotti, L.V., Ottini, L., D'Amico, C., Gradini, R., Cama, A., Belleudi, F., Frati, L., Torrisi, M.R. and Mariani-Costantini, R. (
Harkin, D.P., Bean, J.M., Miklos, D., Song, Y.H., Truong, V.B., Englert, C., Christians, F.C., Ellisen, L.W., Maheswaran, S., Oliner, J.D. et al. (
Potapova, O., Haghighi, A., Bost, F., Liu, C., Birrer, M.J., Gjerset, R. and Mercola, D. (
Lafarge, S., Sylvain, V., Ferrara, M. and Bignon, Y.J. (
Tomlinson, G.E., Chen, T.T.L., Stastny, V.A., Virmani, A.K., Spillman, M.A., Tonk, V., Blum, J.L., Schneider, N.R., Wistuba, I.I., Shay, J.W. et al. (
Quinn, J.E., Kennedy, R.D., Mullan, P.B., Gilmore, P.M., Carty, M., Johnston, P.G. and Harkin, D.P. (
Brodie, S.G., Xu, X., Qiao, W., Li, W.M., Cao, L. and Deng, C.X. (
Seery, L.T., Knowlden, J.M., Gee, J.M.W., Robertson, J.F.R., Kenny, F.S., Ellis, I.O. and Nicholson, R.I. (
Egawa, C., Miyoshi, Y., Takamura, Y., Taguchi, T., Tamaki, Y. and Noguchi, S. (
Jemal, A., Murray, T., Samuels, A., Ghafoor, A., Ward, E. and Thun, M.J. (
Mountain, C.F. (
Pass, H.I., Pogrebniak, H.W., Steinberg, S.M., Mulshine, J. and Minna, J. (
Rosell, R., Gómez-Codina, J., Camps, C., Maestre, J., Padilla, J., Cantó, A, Mate, J.L., Li Shanrong, Roig J., Olazábal, A. et al. (
Martini, N., Kris, M.G., Flehinger, B.J., Gralla, R.J., Bains, M.S., Burt, M.E., Heelan, R., McCormack, P.M., Pisters, K.M.W., Rigas, J.R. et al. (
Aloyz, R., Xu, Z.Y., Bello, V., Bergeon, J., Han, F.Y., Yan, Y., Malapetsa, A., Alaoui-Jamali, M.A., Duncan, A.M.V. and Panasci, L. (
Furuta, T., Ueda, T., Aune, G., Sarasin, A., Kraemer, K.H. and Pommier, Y. (
Lord, R.V.N., Brabender, J., Gandara, D., Alberola, V., Camps, C., Domine, M., Cardenal, F., Sanchez, J.M., Gumerlock, P.H., Taron, M. et al. (
Rosell, R., Scagliotti, G., Danenberg, K.D., Lord, R., Bepler, G., Novello, S., Cooc, J., Crino, L., Sancehz, J.J., Taron, M. et al. (
Rosell, R., Danenberg, K.D., Alberola, V., Bepler, G., Sanchez, J.J., Camps, C., Provencio, M., Isla, D., Taron, M., Diz, P. et al. (
LePage, F., Randrianarison, V., Marot, D., Cabannes, J., Perricaudet, M., Feunteun, J. and Sarasin, A. (
Einspahr, J.G., Krouse, R.S., Yochim, J.M., Danenberg, P.V., Danenberg, K.D., Bhattacharyya, A.K., Martinez, M.E. and Alberts, D.S. (
Rice, J.C., Massey-Brown, K.S. and Futscher, B.W. (
Esteller, M., Silva, J.M., Domínguez, G., Bonilla, F., Matias-Guiu, X., Lerma, E., Bussaglia, Prat, J., Harkes, I.C., Repasky, E.A. et al. (
Hussain, S., Uit E, Huber, P.A.J., Medhurst, A.L., Ashworth, A. and Mathew, C.G. (
Taniguchi, T., Tischkowitz, M., Ameziane, N., Hodgson, S.V., Mathew, C.G., Joenje, H., Mok, S.C and D'Andrea, A.D. (
Wei, Q., Cheng, L., Amos, C.I., Wang, L.E., Guo, Z., Hong, W.K.H and Spitz, M.R. (
Zeng-Rong, N., Paterson, J., Alpert, L., Tsao, M.S., Viallet, J. and Alaoui-Jamali, M.A. (
Bosken, C.H., Wei, Q., Amos, C.I. and Spitz, M.R. (
Schiller, J.H., Harrington, D., Velan, C.P., Langer, C., Sandler, A., Krrok, J., Zhu, J. and Johnson, D.H. (
Gatzemeier, U., von Pawel, J., Gottfried, M., ten Velde, G.P.M., Mattson, K., DeMarinis, F., Harper, P., Salvati, F., Robinet, G., Lucenti, A. et al. (
Georgoulias, V., Ardavanis, A., Agelidou, A., Agelidou, M., Chandrinos, V., Tsaroucha, E., Toumbis, M., Kouroussis, C., Syrigos, K., Polyzos, A. et al. (
Specht, K., Richter, T., Müller, U., Walch, A., Werner, M. and Hofler, H. (