Abstract
Genetic variation was studied in unimproved grassland populations of two contrasting outbreeding perennial grass species. A total of 27 populations of Lolium perenne (perennial ryegrass) and 30 populations of Agrostis curtisii (bristle-leaved bent), sampled from seven and five regions spread across southern Britain, were assessed at three and four isozyme loci, respectively. The extent of genetic structure within and among populations was estimated using unbiased F-statistics. In A. curtisii, a nonagricultural species, populations from adjacent regions were found to be more genetically similar than those separated by greater distance. The reverse situation was observed within L. perenne, a species of major agricultural importance. It is suggested that the absence from L. perenne of the pattern of genetic variation found in A. curtisii is consistent with the occurrence of large-scale human-mediated gene flow via ‘improved’ ryegrass cultivars. If this is the case, then the disruption of natural patterns of genetic variation by the introduction of nonlocal genotypes may occur without apparent major ecological consequences.
Similar content being viewed by others
Introduction
Genetic structure and morphological diversity within species can result from restricted gene flow (Wright, 1969; Slatkin, 1985, 1993). Direct methods of estimating gene flow are problematic, and can only give estimates for a relatively restricted area and over a short period of time. Over the past 30 years, isozyme markers have been used extensively in the study of genetic structure in plant populations. The forage grasses, which dominate many temperate agricultural and seminatural areas, have been no exception (Hayward et al., 1978; McNeilly & Roose, 1984; Ennos, 1985). Population genetic studies such as these have established indirect estimates of gene flow through the analysis of differences in the frequencies of alleles at various loci among two or more populations.
Although gene flow in flowering plants can occur via both pollen and seed, wind dispersal of pollen is considered to be more important in outbreeding grasses (Griffiths, 1950; Gleaves, 1973). However, this is likely to vary greatly between agriculturally important grasses, which experience considerable amounts of seed movement, and species less interfered with by man. The majority of studies of population-level genetic variation within grasses have looked at those species important to agriculture which are widely sown as forage, such as perennial ryegrass (Lolium perenne) (Hayward et al., 1978; McNeilly & Roose, 1984; Balfourier & Charmet, 1994). Knowledge of gene flow in grasses is dominated by studies of agricultural situations (Copeland & Hardin, 1970) or from experimental blocks (Griffiths, 1950; Gleaves, 1973; Giddings et al., 1997). Surprisingly little is known about gene flow and genetic structure in nonagricultural grass species. As a result of this lack of information, it has been difficult to infer how much of the genetic structure observed in agriculturally important species has resulted from human-mediated gene flow and what remains of natural patterns of variation. In this study we compare the genetic structure of two diploid species of outbreeding perennial grass, both sampled from apparently seminatural, agriculturally unimproved sites. The first of these, L. perenne, is probably the most important forage grass species of northern Europe and, as a consequence, it has experienced massive amounts of gene flow through the sowing of selected cultivars. In contrast, the second species, bristle-leaved bent (Agrostis curtisii) is a nonagricultural species, in the UK occurring in southern and south-western counties of England on well-drained acid soils of lowland heaths (Hubbard, 1984).
Materials and methods
Study populations
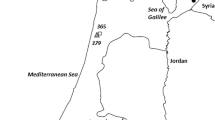
Twenty-seven populations of L. perenne were sampled from across Britain from old unimproved sites. The sampling occurred during the summer of 1976, when unimproved pastures were more common. The populations were chosen from within seven widely spaced regions whose locations can be seen in Fig. 1. Approximately 100 plants were collected from each population as isolated tillers spaced at least 1 m apart, taken from areas containing no obvious ecological differences or gradients. The tillers were established in pots in the glasshouse prior to sampling for electrophoresis.
During 1979 seeds of A. curtisii were collected from five sites in each of six regions in southern Britain, whose locations are shown in Fig. 2. At each site, seed was collected from 100 randomly chosen individuals within an area of 100 m2. Seeds from each plant were sown into lime-free potting compost, with one randomly selected seedling per parent being grown on in a 13 cm pot of the same compost until sampling for electrophoresis.
Isozyme electrophoresis
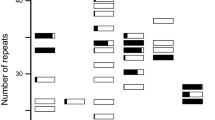
During 1977/78, leaf material was sampled from each L. perenne plant from all 27 populations. This material was crushed in 0.1 M Tris-HCL, pH 7.2 extraction buffer with 0.1% mercaptoethanol, and starch gel electrophoresis was performed following the method and staining procedures of Hayward & McAdam (1977). The gels were scored for the multiallelic enzyme loci: phosphoglucoisomerase (Pgi-2, EC 5.3.1.9), acid phosphatase (Acp-2, EC 3.1.3.2) and glutamic-oxaloacetic transaminase (Got-3, EC 2.6.1.1). These loci show regular Mendelian inheritance and their enzyme products are dimeric.
During 1980 leaf samples of A. curtisii were taken from plants of a uniform age and homogenized in 0.1 M Tris-HCL, pH 7.0 with 0.1% mercaptoethanol. Electrophoresis, on 13% starch gels, used the buffer system and staining methods of Shaw & Prasad (1970). The following enzyme loci were scored: glutamic-oxaloacetic transaminase (Got-3) and phosphoglucoisomerase (Pgi-2), whose products are dimeric, and esterase (Est-2, EC 3.1.1.1) and peroxidase (Per-1, EC 1.11.1.7), which code for monomeric enzymes.
Data analysis
The analysis of genetic structure was the same for both species. The extent of genetic structure within populations (FIS) and among populations (FST) was estimated using the program FSTAT (Goudet, 1995), which calculates Weir & Cockerham's (1984) unbiased estimators of F-statistics. FST values were obtained between all pairs of populations and classified into those between populations in the same region (coded 1 in a variable termed ‘region membership’) and those between populations in different regions (coded 0) (Raybould et al., 1996, 1997). The effects of distance and regional grouping were examined using partial regressions and partial matrix correspondence tests (e.g. Manly, 1991; Thorpe & Baez, 1993) of pairwise genetic distances (FST) and geographical distance, region membership and the interaction between distance and region membership (distance multiplied by the region membership variable). The interaction tests for the uniformity of the effect of distance within and between regions (Raybould et al., 1997).
Results
Isozymes detected significant nonrandom mating in several Lolium populations (Table 1), and in only one population (Romney Marshes 4) was there a slight, but nonsignificant, heterozygote excess. The over all populations mean FIS was significantly greater than zero, both over all loci and at each locus individually. There was some variation among loci; heterozygote deficits occurred frequently at Acp, but rarely at Got and Pgi.
Compared with Lolium, a lower proportion of Agrostis populations showed significant heterozygote deficit, and a number showed significant heterozygote excess (Table 2). The mean over all populations FIS over all loci was slightly negative. There was more variation among loci than for Lolium. Most populations had a significant excess of heterozygotes at Pgi and a significant deficit of heterozygotes at Est. At Got and Per, FIS was not significantly different from zero in most populations.
In both species, FST was significant at all loci (Table 3). The over all loci estimate of FST was higher in Lolium, but this relates specifically to the high value for Acp. Estimates of FIT were significantly greater than zero for all loci in Lolium, and for Est, Got and Per in Agrostis. There was a significant heterozygote excess in the whole sample at Pgi in Agrostis, which gave a much lower value of FIT over all loci than for Lolium.
The proportion of variation in pairwise FST values that was explained by distance or region membership was low in both species (r2<0.07, Table 4). However, significant effects and differences between the species were detected. In Agrostis there was a significant positive relationship between distance and FST, whereas in Lolium there was a significant negative relationship between distance and FST (Table 4). In Lolium there was no significant region membership effect, but region did have a negative effect in Agrostis.
In both species there was a significant distance effect when the region effect was removed from the analysis. When the over-regions negative distance effect was removed in Lolium, a significant effect of distance was detected within regions. Populations of Lolium within regions were more similar to each other than were populations in different regions, but apart from this effect, populations generally were more similar at greater distances apart. In Agrostis, the significant region effect was not detectable when the effect of distance was removed. In other words, there was no greater similarity of populations within regions than would be expected given their geographical proximity. In neither species was there a significant interaction between distance and region membership (Table 4).
Discussion
The results obtained indicate a number of differences between the genetic structure of L. perenne and A. curtisii populations, some of which may result from the agricultural sowing of Lolium whereas other differences may relate to biological differences between the species. The significant positive FIS over all populations of Lolium and significant negative FIS over all Agrostis populations, may be indicative of a violation of the assumptions of Hardy–Weinberg in Lolium. Although the collecting of Lolium material occurred during the autumn following the drought summer of 1976, if we assume the markers to be neutral, the fact that the positive FIS was observed for all populations and all loci is difficult to reconcile with chance linkages to non-neutral loci. However, if the markers are not neutral, then the observed nonrandom mating in old pasture populations of Lolium may result from the nonrandom sampling of genotypes. McNeilly & Roose (1984), using isozyme markers, demonstrated that old unimproved ryegrass pastures can be dominated by very few genotypes, with some clones being much larger than others. If homozygotes were over-represented in the larger clones at the end of the summer of 1976, then spatially, randomly sampling such a turf would be likely to result in multiple samples being taken from homozygous individuals. The observed deficit of heterozygotes, nonrandom mating in all populations of Lolium, over all loci, could thus result from such a biased sampling strategy. In contrast, A. curtisii has a densely tufted growth form (Hubbard, 1984), and is unlikely to form large spreading clones; it is thus more likely to have been sampled in an unbiased manner. The different sampling strategies used for the two species may also influence the apparent genetic structure of their populations, particularly so if the markers are not neutral and there is strong selection occurring during seedling establishment.
At the other end of the spatial scale, L. perenne and A. curtisii were seen to differ in their genetic structure as partitioned between regions. Agrostis curtisii was seen to exhibit classical isolation by distance (Wright, 1943, 1946; Slatkin, 1993) with a significant positive relationship between distance and FST; A. curtisii populations from adjacent regions were found to be genetically more similar than those separated by greater distance. In contrast, L. perenne showed the exact opposite relationship at the regional level, with a significant negative relationship between FST and distance. This relationship was observed over all loci. Although it may be possible to postulate a selective explanation for why populations of L. perenne separated by large distances are genetically similar (i.e. they are coastal populations), this is difficult to reconcile with the observation holding true over several neutral loci. A more plausible explanation is long-scale gene flow in L. perenne associated with agricultural usage. Much of the observed relationship can be accounted for by similarity between the L. perenne populations sampled around Aberystwyth and those collected from Romney Marsh, Kent. Indeed, if either of these regions is dropped from the analysis the relationship breaks down. There is some historical evidence (Beddows, 1953) that suggests that the fertile grasslands of Romney Marsh may have been sampled as a source of germplasm for early breeding work at the Welsh Plant Breeding Station, Aberystwyth. The resulting improved varieties of L. perenne were widely planted around mid-Wales and may have been exchanging genes with local material for some time. This human-mediated long-distance gene flow may therefore be responsible for the observed genetic similarity between geographically separated populations. Similarly, Balfourier & Charmet (1994) suggested that the historical long-distance movement of L. perenne from Italy to Corsica, for agricultural use, was responsible for the current genetic similarities between these populations.
The observed difference between L. perenne and A. curtisii in genetic variation within regions may be a consequence of both natural and agriculture-related gene flow. The greater similarity of populations of L. perenne within regions than between regions may result from extensive gene flow between populations within regions, which is possible because the species is extremely abundant. This extent of gene flow may not be possible in the rarer A. curtisii. However, the same pattern of genetic similarity between populations of L. perenne within regions could also result from different regional preferences by farmers in selecting improved L. perenne varieties to sow.
Many of the patterns of genetic variation reported here within L. perenne, but not A. curtisii, are consistent with long-distance human-mediated gene flow associated with agricultural usage. This is perhaps surprising as this occurred in plants sampled from apparently old unimproved pastures and thus implies gene flow from neighbouring sown fields. This level of gene flow of nonlocal genes is relevant to current concerns regarding the release of genetically modified organisms and of the sowing of nonlocal provenance wild flower seeds. In the case of the very widespread and abundant L. perenne at least, considerable gene flow must have occurred, resulting from the agricultural sowing of nonlocal types, apparently without disrupting the ecology of the seminatural plant communities in which it also is found. However, that is not to say that more subtle changes in the ecology of grasslands containing L. perenne have not occurred but are as yet unnoticed.
References
Balfourier, F. and Charmet, G. (1994). Geographical patterns of isozyme variation in Mediterranean populations of perennial ryegrass. Heredity, 72: 55–63.
Beddows, A. R. (1953). The Rye Grasses in British Agriculture — A Survey. H series bulletin no. 17. University College of Wales. Welsh Plant Breeding Station, Cambrian News, Aberystwyth.
Copeland, L. O. and Hardin, E. E. (1970). Outcrossing in the ryegrasses (Lolium spp.) as determined by fluorescence tests. Crop Sci, 10: 254–257.
Ennos, R. A. (1985). The mating system and genetic structure in a perennial grass, Cynosurus cristatus L. Heredity, 55: 121–126.
Giddings, G. D., Hamilton, N. R. S. and Hayward, M. D. (1997). The release of genetically modified grasses. 1. Pollen dispersal to traps in Lolium perenne. Theor Appl Genet, 94: 1000–1006.
Gleaves, J. T. (1973). Gene flow mediated by wind-borne pollen. Heredity, 31: 355–366.
Goudet, J. (1995). FSTAT (version 1.2): A computer program to calculate F- statistics. J Hered, 86: 485–486.
Griffiths, D. J. (1950). The liability of seed crops of perennial ryegrass (Lolium perenne) to contamination by wind-borne pollen. J Agri Sci, 40: 19–38.
Hayward, M. D. and McAdam, N. J. (1977). Isozyme polymorphism as a measure of distinctiveness and stability in cultivars of Lolium perenne L. Z PflZücht, 79: 79–68.
Hayward, M. D., Gottlieb, L. D. and McAdam, N. J. (1978). Survival of allozyme variants in swards of Lolium perenne L. Z PflZücht, 81: 228–234.
Hubbard, C. E. (1984). Grasses, 3rd edn (revised by J. C. E. Hubbard). Penguin Books, London.
Manly, B. J. F. (1991). Randomization and Monte Carlo Methods in Biology. Chapman & Hall, New York.
McNeilly, T. and Roose, M. L. (1984). Distribution of perennial ryegrass genotypes in swards. New Phytol, 98: 503–513.
Raybould, A. F., Goudet, J., Mogg, R. J., Gliddon, C. J. and Gray, A. J. (1996). The genetic structure of linear populations of sea beet (Beta vulgaris ssp. maritima) revealed by isozyme and RFLP analysis. Heredity, 76: 111–117.
Raybould, A. F., Mogg, R. J. and Gliddon, C. J. (1997). The genetic structure of Beta vulgaris ssp. maritima (sea beet) populations. 2. Differences in gene flow estimated from RFLP and isozyme loci are habitat-specific. Heredity, 78: 532–538.
Shaw, C. R. and Prasad, R. (1970). Starch gel electrophoresis of enzymes — a compilation of recipes. Biochem Genet, 4: 297–320.
Slatkin, M. (1985). Gene flow in natural populations. Ann Rev Ecol Syst, 16: 393–430.
Slatkin, M. (1993). Isolation by distance in equilibrium and non-equilibrium populations. Evolution, 47: 264–279.
Thorpe, R. S. and Baez, M. (1993). Geographic variation in scalation of the lizard Gallotia stehlini within the islands of Grand Canaria. Biol J Linn Soc, 48: 75–87.
Weir, B. S. and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Wright, S. (1943). Isolation by distance. Genetics, 28: 114–138.
Wright, S. (1946). Isolation by distance under diverse systems of mating. Genetics, 31: 39–59.
Wright, S. (1969). Evolution and the Genetics of Populations. vol. 2: The Theory of Gene Frequencies. University of Chicago Press, Chicago.
Acknowledgements
This work was in part funded under Contract PECD/7/8/238 of the Department of the Environment's Genetically Modified Organism Research Programme. The views expressed here are those of the authors and not necessarily those of the Department of the Environment.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Warren, J., Raybould, A., Ball, T. et al. Genetic structure in the perennial grasses Lolium perenne and Agrostis curtisii. Heredity 81, 556–562 (1998). https://doi.org/10.1046/j.1365-2540.1998.00426.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00426.x
- Springer Nature Switzerland AG
Keywords
This article is cited by
-
Genetic diversity analysis of switchgrass (Panicum virgatum L.) populations using microsatellites and chloroplast sequences
Agroforestry Systems (2014)
-
Genetic diversity of natural orchardgrass (Dactylis glomerataL.) populations in three regions in Europe
BMC Genetics (2013)
-
Value of permanent grassland habitats as reservoirs of Festuca pratensis Huds. and Lolium multiflorum Lam. populations for breeding and conservation
Euphytica (2008)