Abstract
Complete D-loop sequences of 20 Mus from three localities in Turkey and seven in Iran were characterized. These countries are thought to be close to the place of origin of the subspecies Mus musculus domesticus. Five new M. m. domesticus haplotypes were added to the nine already known for the region. Four of these 14 haplotypes were very similar to the consensus D-loop sequence for western Europe defined by Nachman et al. (1994), which may represent the ancestral condition for M. m. domesticus. A divergent mtDNA lineage is found in various parts of Turkey and northern Iran; it has spread into western Europe, but other European lineages were not found in either Turkey or Iran. The other Mus D-loop sequences were of M. m. castaneus and Mus macedonicus and confirmed M. macedonicus as a monotypic species with low nucleotide diversity. The prevalence of the standard 40-chromosome complement in this region is particularly interesting with regards M. m. domesticus, as it is consistent with the in situ origin of Robertsonian karyotypic races (2n < 40) in western Europe.
Similar content being viewed by others
Introduction
The genus Mus arose within the last 4 Myr and is currently represented by nine species (Bonhomme & Guénet, 1996). The most familiar of these is the house mouse Mus musculus, which is subdivided into at least three subspecies: musculus, castaneus and domesticus (Bonhomme & Guénet, 1996). M. m. musculus occurs in eastern Europe and throughout northern Asia and M. m. castaneus is found in southern Asia. Mus musculus domesticus occupies western Europe and has colonized much of rest of the world (the Americas, Australasia, Africa) as a commensal of humans. The genus Mus in general, and M. m. domesticus in particular, have been the subject of much evolutionary study (Boursot et al., 1993; Sage et al., 1993), not only because of their instrinsic fascination as a group of rodents with interesting ecologies, genetics and colonization histories, but because of the status of the laboratory version of M. m. domesticus as the primary biomedical model for genetic analysis (Silver, 1995). Not only have the biomedical community an interest in the evolutionary history of their study organism, but they have, of course, described genetic markers which allow the evolutionary history in Mus to be more easily unravelled than in other mammals.
Recently there have been studies examining genetic variation over the whole range of Mus musculus in order to elucidate the site of origin of this species and its most important subspecies, M. m. domesticus. There are two evolutionary models relating to this (which we, for convenience, call models ‘I’ and ‘II’). For model I, Din et al. (1996) and Boursot et al. (1996) have suggested that the species Mus musculus arose in northern parts of the Indian subcontinent but that M. m. domesticus originated in the Fertile Crescent (specifically to the west of the Zagross mountains, in the vicinity of present-day Iraq) following an expansion from the species source area. The house mice currently living in northern India and the surrounding area used to be called ‘M. m. bactrianus’, but Din et al. (1996) and Boursot et al. (1996) prefer to consider these mice as an undefined ‘oriental’ form. It appears that ‘M. m. bactrianus’ is in fact a westerly extension of M. m. castaneus (Prager et al., 1998) which otherwise occurs in south-eastern Asia. Thus, model I may be restated that house mice originated in northern India and were castaneus-like, and that domesticus-like mice derived from them following spread into the Fertile Crescent.
In contrast, model II, stated by Prager et al. (1998), is that the Fertile Crescent was the source area for the whole species and that the first house mice were domesticus-like. Thus, in both models I and II, the Fertile Crescent was the site of origin of M. m. domesticus, and it is from there that the subspecies began its extensive colonization of western Europe and northern Africa, and ultimately elsewhere in the world (Boursot et al., 1996; Din et al., 1996; Prager et al., 1998).
Given that M. m. domesticus originated in south-western Asia, further genetic characterization of mice from this area is important. Such studies should help to establish more clearly the ancestral characteristics of M. m. domesticus and help interpret better the pattern of genetic variation found throughout the species range. In particular, there is much known about the mitochondrial (mt) DNA and chromosomal variation of M. m. domesticus in western Europe (Prager et al., 1993; Nachman et al., 1994; Nachman & Searle, 1995) but little information from south-western Asia. If it could be shown that some mtDNA or chromosomal variants found in western Europe are not found in the south-west Asian source area, it may suggest origin of those variants during or subsequent to the range expansion of M. m. domesticus into western Europe. Likewise, presence of certain mtDNA or chromosomal variants only in south-western Asia and not in western Europe may suggest that only part of the genetic variation present in the source area was carried during the colonization of western Europe. There is considerable interest in elucidating the genetic processes that occur during geologically recent colonizations (Hewitt, 1996).
In this paper we describe a small-scale study of M. m. domesticus mtDNA and chromosomal variation in south-western Asia, close to the putative source area for this subspecies. As in the previous extensive mtDNA studies of M. m. domesticus from western Europe (Prager et al., 1993; Nachman et al., 1994), we sequenced the D-loop (control region). In total, we describe five new D-loop haplotypes from localities in central and northern Turkey and southern Iran. This adds to nine complete or near-complete haplotypes from Georgia, southern Turkey and western Iran described by Prager et al. (1996, 1998).
Our collections from Turkey and Iran included mice with M. m. castaneus mtDNA haplotypes. The aboriginal mouse Mus macedonicus was also represented. D-loop sequences from these specimens were obtained for further comparison with data in the literature.
Materials and methods
Altogether 20 Mus were caught at three localities in Turkey and seven localities in Iran during 1996–98 (Table 1, Fig. 1). Standard external measurements were taken: head + body, tail, hindfoot and ear lengths.
Collection localities in Turkey and Iran. See Table 1 for further details.
Direct chromosome preparations were made from somatic tissue and diploid numbers obtained. Four mice from one site in Turkey (Samsun) were unavailable for measurement and chromosome preparation (Table 1), but samples were taken for DNA studies.
For DNA analysis, tail tips were preserved in 100% ethanol and maintained at 4°C. DNA was extracted by the standard phenol/chloroform procedure (Sambrook et al., 1989). The whole of the D-loop and flanking regions were amplified in two overlapping fragments by PCR: a 724-bp fragment generated with the primers L15774 and H16498 (5′-CCTGAAGTAGGAACCAGATG-3′) (Kocher et al., 1989; P. Taberlet, pers. comm.; numbering according to Anderson et al., 1981) and a 632-bp fragment produced with primers L15735 and H00072 (Prager et al., 1993; numbering according to Bibb et al., 1981).
For each primer pair, double-stranded amplifications were carried out using standard concentrations of DNA and reagents (Bilton et al., 1998). For primers L15774-H16498, 30 cycles of 2 min at 93°C, 1 min at 50°C and 2 min at 72°C were carried out. For primers L15735-H00072 there were only 25 cycles and an annealing temperature of 64°C. PCR products were purified with the QIAquick PCR purification kit (Qiagen) and sequenced using the ABI PRISM Dye Terminator Cycle Sequencing Ready Reaction Kit. For the 724-bp fragment, generally only the L15774 primer was used to generate the sequences, which were exceptionally clear in all cases (some checks were made by sequencing in the other direction as well, using the H16498 primer). The 632-bp fragment was sequenced in both directions using primers L15735 and H00072. The sequencing PCR consisted of 25 cycles of 96°C for 30 s, 50°C for 30 s and 60°C for 4 min for primers L15774 and H16498, whereas it differed in annealing temperature (53°C) for primers L15735 and H00072. The sequencing products were loaded onto an ABI PRISM 377 automated sequencer.
Sequence traces were downloaded, checked using Analysis software (ABI) and aligned using SeqEd (ABI). For all specimens we obtained sequences for the mtDNA region between positions 15283–16295 of Bibb et al. (1981), although a shorter length (15363–16295) was used in the analyses. This comprised the whole D-loop (879 bp) and a small amount of flanking tRNA sequence. Diversity estimates and phylogenetic analyses were performed using our sequences and selected sequences from the literature (Nachman et al., 1994; Prager et al., 1996, 1998). Some of the relevant sequences in Prager et al. (1998) had missing bases. These were only included if there were five or fewer missing positions; we followed the approach of Prager et al. in coding these.
Nucleotide diversities (π) were calculated according to Nei (1987) using the ARLEQUIN 1.1 package of Schneider et al. (1997).
For the phylogenetic analyses, both parsimony and distance methods were used. Maximum parsimony analyses were run using PAUP* (Swofford, 1998) with a variety of different weighting schemes. Where possible, the branch-and-bound option was used to generate the shortest trees. Otherwise we used an heuristic search with stepwise addition (10 random replicates) and the tree bisection-reconnection (TBR) setting. For each analysis a strict consensus tree was generated. Bootstrap values were produced with separate analyses, based on 1000 pseudoreplicates, using either heuristic search (same settings as above) or branch-and-bound.
Distance trees were generated using the PHYLIP package (Felsenstein, 1991). Pairwise distances between taxa were produced under the assumption of the Kimura 2-parameter model, and trees constructed by the neighbour-joining method (Saitou & Nei, 1987). Again, bootstrap values were based on 1000 pseudoreplicates.
Our sequences have been deposited in the EMBL database (accession numbers AJ286317–AJ286329).
Results
Morphology, mtDNA and karyotypes
For Iran and Turkey, previous D-loop sequences obtained by Prager et al. (1998) are readily categorized as characteristic of M. m. domesticus, M. m. castaneus and M. macedonicus. Alignments indicate that all the mtDNA sequences that we obtained clearly belong to one or other of these haplotypes.
All long-tailed mice (tails >71 mm) had a M. m. domesticus or M. m. castaneus D-loop haplotype, whereas all short-tailed mice (tails <70 mm) had M. macedonicus haplotypes (Table 1). These tail lengths fit well with Prager et al.’s (1998) assessment of ‘short-tailed’ and ‘long-tailed’ mice. Mus macedonicus is well known to have a shorter tail than M. m. domesticus and ‘M. m. bactrianus’ (which we consider here to be within M. m. castaneus) (Marshall & Sage, 1981; Auffray et al., 1990; Chondropoulos et al., 1995).
We could not separate mice with M. m. domesticus and M. m. castaneus haplotypes on the basis of tail length or other external measurements (data not shown).
The standard Mus diploid number of 40 was found in all mice examined, i.e. seven mice with M. m. domesticus haplotypes from five localities in Turkey and Iran, five mice with M. macedonicus haplotypes from three localities in Turkey and Iran, and four mice with M. m. castaneus haplotypes from three localities in Iran.
Mus musculus domesticus D-loop haplotypes
D-loop haplotypes clearly recognizable as M. m. domesticus type (Fig. 2) were found in mice from five localities in Turkey and Iran, although the precise haplotypes differ among the localities (Table 1). All the haplotypes show differences from the laboratory mouse sequence of Bibb et al. (1981) at nucleotide positions 15493, 15823, 15912, 16119, 16241 and 16268, as is normal for wild M. m. domesticus. Four out of five haplotypes also have a T instead of a C at position 15597. Although a mouse with only these changes from Bibb et al. has, to our knowledge, not been found, this is the consensus sequence for M. m. domesticus from western Europe [Fig. 2, i.e. these are the most common bases at each nucleotide position that Nachman et al. (1994) found among 56 specimens examined across western Europe]. The two Iranian haplotypes, domIran.1 and 2, are very close to this consensus, differing only by three and two substitutions, respectively (Fig. 2). The south Turkish haplotype 107 and Iranian haplotype 110 described by Prager et al. (1998) also differ from the consensus by only two substitutions and two substitutions plus two insertions, respectively.
Mus mtDNA sequences from Turkey and Iran. All variable positions in the M. m. domesticus (dom), M. m. castaneus (cas) and M. macedonicus (mac) haplotypes relative to the laboratory mouse sequence of Bibb et al. (1981) are shown. The total sequence screened ranged between nucleotide positions 15283 and 16295, although a shorter length (15363–16295) was used for the diversity estimates and phylogenetic analyses to provide alignment with published sequences. Nucleotide substitutions are shown with reference to the numbering system of Bibb et al. (1981), and a dot indicates identity to the sequence of Bibb et al. Each insertion or deletion relative to the sequence of Bibb et al. is indicated by an ‘i’ or a ‘d’, respectively. For a deletion, a dash is shown in the sequence concerned at the relevant nucleotide position. For an insertion, a dash is shown in the Bibb et al. sequence; the insertion occurs after the nucleotide position indicated. Note that the insertions of Cs after positions 160087 and 160093 are positioned arbitrarily; they add to a string of Cs and in the analyses reported here, the addition of the two Cs is taken to be two insertion events. The plus at position 15537 for two M. m. castaneus sequences indicates a tandem repeat from 15538 to 15615 (76 bp); the 3′ variant is shown on a separate line. The 5′ variant was used in phylogenetic analysis, following Prager et al. (1998). The consensus sequence of Nachman et al. (1994) for the region between nucleotide positions 15363 and 16295 is also shown; see text for further details. Note that in their list of sequences (Table 3), Nachman et al. (1994) did not record the difference between their consensus sequence and the sequence of Bibb et al. (1981) at position 16119.
Our three Turkish haplotypes (domTurkey.1, 2 and 3) are very similar to each other, even though they derived from two well-separated parts of the country (Fig. 1). In contrast to our Iranian haplotypes, they differ substantially from the consensus of Nachman et al. (1994). The dom Turkey.1, 2 and 3 haplotypes share substitutions and indels relative to the consensus at positions 15550–1, 15718, 16009, 16250 and 16252, as do single Turkish and Iranian haplotypes of Prager et al. (1998), the Georgian haplotypes of Prager et al. (1996), and haplotypes in Greece, Portugal, Spain and Switzerland (Prager et al., 1993; Nachman et al., 1994). The dom Turkey.1, 2 and 3 haplotypes differ from each other and the Nachman et al. consensus by six, two and three further substitutions, respectively (Fig. 2).
To investigate further the relationship of our new M. m. domesticus sequences with those in the literature, we conducted phylogenetic analyses including all D-loop haplotypes previously described in Iran, Turkey and neighbouring countries (Prager et al., 1996, 1998) together with representatives of European mtDNA lineages (clades I–VI of Nachman et al., 1994). After alignment of all the sequences, the relative frequencies of transitions, transversions and indels were calculated as 1:0.32:0.11. Given that transversions and indels are relatively rare types of mutation, it can be argued that they are more informative than transitions and should therefore be weighted more highly in phylogenetic analysis (see Nachman et al., 1994). However, although insertion–deletion events occur at rather few nucleotide positions in the house mouse, there is evidence that they occur repeatedly in these positions (Prager et al., 1993, 1998; Nachman et al., 1994; Boissinot & Boursot, 1997).
Figure 3 shows a neighbour-joining tree generated with transversions weighted at the inverse of their frequency relative to transitions (to the nearest integer) and indels zero-weighted (which may be most appropriate given the problem of homoplasy), i.e. a relative weighting of 1:3:0 (transitions:transversions:indels). The tree shows the representatives of the six clades defined by Nachman et al. (1994). Based on the bootstrap values, most of these clades are retained with low (54–60%: II, III, VI), moderate (72%: V) or high (93%: I) support, but without any south-west Asian haplotypes within the clades. Many of the Turkish and Iranian haplotypes are scattered basal to clades II, III and V, but with uncertain affinity.
Neighbour-joining phylogenetic tree for the Mus musculus domesticus D-loop haplotypes that we found in Turkey (Turkey.1–3) and Iran (Iran.1–2). Also included are all other complete or near-complete D-loop haplotypes found in M. m. domesticus from Iran, Turkey and Georgia by Prager et al. (1996), (1998), using their numbering system (68, 69, 102, 104–107, 109, 110). In order to establish how these south-western Asian haplotypes relate to those found in M. m. domesticus from Europe, two divergent haplotypes from each of the six clades identified by Nachman et al. (1994) are included in the phylogenies. Their four-digit numbering system and country of origin are used: 1328 and 1334 (from their clade I), 1036 and 1262 (II), 1272 and 1083 (III), 1173 and 1369 (IV), 1316 and 1176 (V) and 1008 and 1199 (VI). A M. m. musculus sequence (Nachman et al., 1994) is included as an outgroup. The tree was generated using a transition:tranversion:indel weighting of 1:3:0 (see text). All bootstrap values over 50% are shown.
Clade IV of Nachman et al. is not retained in the tree in Fig. 3. However, one representative of this clade (1173) branches together with various haplotypes from south-western Asia into a ‘new’ clade with low bootstrap support (58%). These are the haplotypes that were described above as distinctly different from the Nachman et al. consensus.
Although weakly supported in the neighbour-joining tree with a weighting of 1:3:0, the new clade is particularly strongly supported (90% bootstrap) in a parsimony tree that we generated with the same weighting scheme. In other respects this parsimony tree shows less structure than Fig. 3, although clades I, V and VI of Nachman et al. are retained with moderate or strong bootstrap support. Parsimony trees were also generated with the weightings: 1:1:0, 1:6:0, 1:1:1, 1:3:1, 1:6:1, 1:3:3, 1:3:5, 1:3:9, 1:6:9, and the new clade as well as clade I of Nachman et al. are found in all these trees, except for those with the highest weightings for indels. Clades II, III, V and VI of Nachman et al. are less frequently retained under the various weighting schemes. These schemes failed to produce any further, well-supported groupings of south-west Asian haplotypes.
Mus musculus castaneus and Mus macedonicus D-loop haplotypes
In the phylogenetic analysis of M. m. castaneus and M. macedonicus D-loop haplotypes, we combined our data with all other complete or near-complete M. macedonicus sequences and other sequences of M. m. castaneus from Iran (Prager et al., 1996, 1998). Representative sequences of M. m. musculus, M. m. domesticus and south-east Asian M. m. castaneus were also included in the analysis, and a M. spretus sequence was used as an outgroup. As with the phylogenetic analysis of M. m. domesticus, we used a variety of weighting schemes. Figure 4 shows a parsimony tree where transversions are weighted by the inverse of their frequency relative to transitions, and indels are zero-weighted, i.e. a relative weighting of 1:3:0 (transitions:transversions:indels). Very similar trees were generated after parsimony analysis with weightings of 1:1:0, 1:2:0, 1:6:0, 1:1:1, 1:3:1, 1:3:5 and after neighbour-joining analysis with a weighting of 1:3:0. The branching order of M. macedonicus, M. m. domesticus, M. m. musculus and M. m. castaneus is identical to that in Fig. 4 in all these trees. There are also always two major clades within M. m. castaneus (one clade formed by casIran.6 and 7 and cas17Iran; the remaining haplotypes forming the other), as in Fig. 4. The relationship between M. macedonicus haplotypes varies somewhat between trees, but macIran.3 and 4 always form a well-supported clade.
A parsimony phylogenetic tree for the Mus musculus castaneus and M. macedonicus D-loop haplotypes that we found in Turkey (Turkey.4–6) and Iran (Iran.3–7). Also included are all the other complete M. macedonicus haplotypes available in the literature (Prager et al., 1996, 1998) and all the complete and near-complete M. m. castaneus haplotypes found previously in Iran (Prager et al., 1998). In addition, sequences for M. m. castaneus from Thailand (Prager et al., 1996), M. m. musculus from the Czech Republic (Nachman et al., 1994) and M. m. domesticus from Iran (Iran.2) are included for comparison and Mus spretus is included as an outgroup following Prager et al. (1996), whose sequence we use. The tree was generated using a transition:tranversion:indel weighting of 1:3:0 (see text) and shows all bootstrap values over 50%. It is a strict consensus of the nine shortest trees obtained by the branch-and-bound method and has a length of 175 steps, a consistency index of 0.771 and a homoplasy index of 0.229.
Considering the specific sequences that we obtained (Fig. 2): the M. m. castaneus haplotypes that we observed in Iranian mice are new, but similar to those obtained previously in the same country by Prager et al. (1998). In M. m. castaneus the D-loop in some haplotypes includes a 76-bp repeat (Prager et al., 1998). We observed haplotypes with and without the repeat in Iran, as found previously by Prager et al. (1998). As described above, these sequences always form two well-supported clades in the phylogenetic analyses; casIran.6 and 7 and cas17Iran have the repeat and form one clade, the other Iranian haplotypes and that from Thailand (cas Thailand from Prager et al., 1996) lack the repeat and form a second clade (Fig. 4).
In our phylogenies, the M. m. castaneus haplotypes are clearly grouped close to the M. m. musculus and M. m. domesticus haplotypes, whereas the Mus macedonicus haplotypes are well separated from the Mus musculus haplotypes (Fig. 4). The distinctiveness of M. macedonicus is also apparent from the nucleotide sequences (Fig. 2). The nucleotide diversity (π=0.0080) among the 11 specimens of M. macedonicus from six localities dispersed over the species range (Macedonia and south-western Asia) is lower than that recorded in M. m. domesticus (19 animals, 13 localities) and M. m. castaneus (seven animals, six localities) from south-western Asia (0.0100 and 0.0153, respectively).
Discussion
Karyotypes
A diploid complement of 40 chromosomes is considered to be the norm for M. macedonicus, M. m. domesticus and M. m. castaneus (both in mice from south-eastern Asia and those in south-western Asia previously described as ‘M. m. bactrianus’) (Silver, 1995). ‘Mus musculus bactrianus’ with 40 chromosomes have been found in Pakistan, Afghanistan, Turkmenistan, Tadjikistan and Kyrgyzstan by Moriwaki et al. (1986) and Zima et al. (1990). Clearly, our results for Iranian mice with a M. m. castaneus haplotype are consistent with these earlier findings. Mus macedonicus have been karyotyped in Macedonia (Zima et al., 1990; Giagia-Athanasopoulou et al., 1994), Israel (Ivanitskaya et al., 1996), Armenia and Azerbaijan (Bulatova et al., 1991); all individuals examined also had the standard number of chromosomes. Our data from the south-east of the distribution of M. macedonicus (Turkey and Iran) again confirm a 40-chromosome complement.
Although the standard diploid number for M. m. domesticus is 40, numerous karyotypic races characterized by Robertsonian fusions and a reduced chromosome number have been described in western Europe and northern Africa (Nachman & Searle, 1995). It is generally assumed that these Robertsonian fusions arose in situ, after colonization by 40-chromosome mice (Britton-Davidian et al., 1989). These colonists would have derived ultimately from south-western Asia, so it is of interest that we were able to confirm a standard karyotype in mice with a M. m. domesticus haplotype from five localities within that region. Mus musculus domesticus with 40 chromosomes have also been described in Israel (Auffray, 1993).
Mus musculus D-loop haplotypes
As in the previous study by Prager et al. (1998), we found two major lineages of mtDNA in Mus musculus from Turkey and Iran. One is the M. m. domesticus mtDNA lineage. The other is the ‘oriental’ lineage of Boursot et al. (1996), but which Prager et al. (1998) showed could readily be incorporated into the M. m. castaneus lineage; we follow Prager et al.’s nomenclature here.
The three M. m. castaneus haplotypes that we found are new and double the number of complete D-loop sequences available for Iran. However, our three haplotypes belonged to two clades already identified in M. m. castaneus by Prager et al. (1998), one characterized by a 76-bp repeat (casIran.6 and 7, cas17Iran: Fig. 4) and the other not. Prager et al. (1998) analysed M. m. castaneus from throughout southern Asia and the reader is referred to their detailed account on this subspecies.
The most interesting specimens carrying a M. m. domesticus haplotype were those from south-eastern Iran (localities I and J; Fig. 1). In both models I and II for the origin of M. m. domesticus (see Introduction), it is believed that the subspecies arose in the Fertile Crescent and expanded westwards. The discovery of M. m. domesticus haplotypes to the east of the Fertile Crescent is therefore something of a surprise, although in model I the progenitors of M. m. domesticus are thought to have migrated through southern Iran (Boursot et al., 1996). The M. m. domesticus mtDNA in Iran may be a relict of this migration, or it may be a consequence of a later human-mediated transport of M. m. domesticus after its formation in the Fertile Crescent (localities I and J are on the coast and mice may have been brought there accidentally, by boat). In either case, the haplotypes from Iran are of interest as they provide insight into the ancestral D-loop sequence of M. m. domesticus.
The two M. m. domesticus haplotypes that we obtained from south-eastern Iran were very similar to the consensus for the D-loop sequence described for western Europe by Nachman et al. (1994). Two haplotypes from northern Iran/southern Turkey described by Prager et al. (1998) are also extremely close to this consensus. Therefore, one possibility is that the consensus sequence of Nachman et al. represents the ancestral condition for M. m. domesticus, perhaps becoming fixed within or en route to the Fertile Crescent. A consensus mtDNA sequence based on haplotypes collected widely over the range of M. m. domesticus is likely to be similar to the ancestral sequence if the subspecies spread rapidly over that range from a small source population characterized by the ancestral mtDNA haplotype. Such expansion leads to a star phylogeny (Slatkin & Hudson, 1991), as well-demonstrated for humans colonizing western Europe (Richards et al., 1998). It is possible that M. m. domesticus, as a commensal of humans, has a similar true phylogeny.
Whereas inspection of haplotypes suggests one type of ancestral mtDNA sequence for M. m. domesticus, the phylogenetic trees that we produced by the outgroup method of rooting, suggest another. Clade VI of Nachman et al. (1994) tended to be basal within the phylogenetic trees that we generated, using M. m. musculus as an outgroup (it is also basal in the trees that Nachman et al. produced). However, this clade was defined by haplotypes of M. m. domesticus collected in western Europe by Nachman et al. Not one of the 14 D-loop haplotypes that we considered from south-western Asia belongs to this clade.
Clearly, there is a need for further sampling in south-western Asia, and particularly the Fertile Crescent itself, to resolve the ancestral D-loop sequence of M. m. domesticus and to gain an understanding of the evolution of mtDNA clades found in western Europe and northern Africa. Surprisingly, none of the Iranian and Turkish D-loop haplotypes so far typed belongs to the west European clades I, II, III, V and VI identified by Nachman et al. (1994). Whether this reflects an inadequacy of sampling in south-western Asia or evidence of evolution of these lineages in the Mediterranean area itself, requires further sampling.
However, one lineage not clearly identified by Nachman et al. (1994) is found in both south-western Asia and Europe. This is our strongly supported new clade of south-west Asian and Greek haplotypes which, on the basis of sequence similarity, is also probably found in Portugal, Spain and Switzerland. Turkey or northern Iran may therefore be the source area for this clade, which appears to have spread widely across Europe.
Taxonomic status of the Mus musculus studied in south-western Asia
In considering both the chromosomal and mtDNA data, it is important to be aware of the problems with matching the mtDNA haplotype of mice to their taxonomic status, defined by morphology or nuclear markers. This is particularly likely to be the case for samples from Iran. Our data, and those of Prager et al. (1998), imply that although M. m. domesticus does occur in the west and south of Iran, it is M. m. castaneus haplotypes that are particularly widespread. It is tempting to suggest that these match with a morphology that used to be called ‘M. m. bactrianus’. However, the correspondence between domesticus mtDNA haplotype and domesticus morphology, and castaneus mtDNA and ‘bactrianus’ morphology, is far from perfect in Iran (see data in Tables 1 and 2 of Prager et al., 1998). There is also the complication of M. m. musculus haplotypes in the north-east of the country which are not definitely associated with a musculus morphology (Boissinot & Boursot, 1997; Prager et al., 1998). Thus, any model of evolution based purely on mtDNA sequences can only be considered a working hypothesis of the mode of evolution of M. musculus as a taxonomic entity. Studies involving morphology and nuclear markers of house mice in south-western Asia, initiated by Din et al. (1996) and Prager et al. (1998), need to be continued for a complete picture of house mice in this region.
Mus macedonicus D-loop haplotypes
It is safe to consider the M. macedonicus D-loop haplotypes that we found as truly representing the morphological and biological species M. macedonicus, as it is unlikely that this form currently hybridizes with or has recently hybridized with any of the M. musculus subspecies (Bonhomme & Guénet, 1996). By combining our data with those of Prager et al. (1998), we have been able to examine the mtDNA variation over the broad range of M. macedonicus, from Macedonia in the extreme west to Turkey in the centre and Iran in the extreme east. On the basis of low nucleotide variation and tight phylogenetic clustering, M. macedonicus appears to be a monotypic species that has recently expanded from a rather small source, perhaps in relation to the last glaciation. Obviously larger sample sizes would be desirable to confirm this result.
References
Anderson, S., Bankier, A. T., Barrell, B. G., de Bruijn, M. H. L., Coulson, A. R. and Drouin, J. et al.1981). Sequence and organisation of the human mitochondrial genome. Nature. 290: 457–465.
Auffray, J. -C. (1993). Chromosomal divergence in the house mouse in the light of palaeontology: a colonization-related event. In: Chaline, J. and Werdelin, L. (eds) Modes and Tempos of Evolution in the Quaternary, pp. 21–25. Pergamon Press, Oxford.
Auffray, J. -C., Tchernov, E., Bonhomme, F., Heth, G., Simson, S. and Nevo, E. (1990). Presence and ecological distribution of Mus “spretoides” and Mus musculus domesticus in Israel. Circum-Mediterranean vicariance in the genus Mus. Z Säugetierk. 55: 1–10.
Bibb, M. J., van Etten, R. A., Wright, C. T., Walberg, M. W. and Clayton, D. A. (1981). Sequence and gene organization of mouse mitochondrial DNA. Cell. 26: 167–180.
Bilton, D. T., Mirol, P. M., Mascheretti, S., Fredga, K., Zima, J. and Searle, J. B. (1998). Mediterranean Europe as an area of endemism for small mammals rather than a source for northwards postglacial colonisation. Proc R Soc B. 265: 1219–1226.
Boissinot, S. and Boursot, P. (1997). Discordant phylogeographic patterns between the Y chromosome and mitochondrial DNA in the house mouse: selection on the Y chromosome? Genetics. 146: 1019–1034.
Bonhomme, F. and Guénet, J. -L. (1996). The laboratory mouse and its wild relatives. In: Lyon, M. F., Rastan, S. and Brown, S. D. M. (eds) Genetic Variants and Strains of the Laboratory Mouse, 3rd edn, pp. 1577–1596. Oxford University Press, Oxford.
Boursot, P., Auffray, J.-C., Britton-Davidian, J. and Bonhomme, F. (1993). The evolution of house mice. Ann Rev Ecol Syst. 24: 119–152.
Boursot, P., Din, W., Anand, R., Darviche, D., Dod, B. and von Deimling, F. Et Al.1996). Origin and radiation of the house mouse: mitochondrial DNA phylogeny. J Evol Biol. 9: 391–415.
Britton-Davidian, J., Nadeau, J. H., Croset, H. and Thaler, L. (1989). Genic differentiation and origin of Robertsonian populations of the house mouse (Mus musculus domesticus Rutty). Genet Res. 53: 29–44.
Bulatova, N. S., Nadjafova, R. S. and Kozlovsky, A. I. (1991). Cytotaxonomic analysis of species of the genera Mus, Apodemus and Rattus in Azerbaijan. Z zool Syst Evol-Forsch. 29: 139–153.
Chondropoulos, B. P., Markakis, G. and Fraguedakis-Tsolis, S. E. (1995). Morphometric and immunological relationships among some Greek Mus L. populations (Mammalia, Rodentia, Muridae). Z Säugetierk. 60: 361–372.
Din, W., Anand, R., Boursot, P., Darviche, D., Dod, B. and Jouvin-Marche, E. et al.1996). Origin and radiation of the house mouse: clues from nuclear genes. J Evol Biol. 9: 519–539.
Felsenstein, J. (1991) Phylogeny Inference Package (Phylip), Version 3.4. University of Washington, Seattle, WA.
Giagia-Athanasopoulou, E. B., Fraguedakis-Tsolis, S. E. and Chondropoulos, B. P. (1994). Chromosomal variation and geographical distribution of some wild mice populations of the genus Mus (Mammalia, Rodentia) in Greece. Biol Gallo-Hell. 22: 193–202.
Hewitt, G. M. (1996). Some genetic consequences of the ice ages, and their role in divergence and speciation. Biol J Linn Soc. 58: 247–276.
Ivanitskaya, E., Gorlov, I., Gorlova, O. and Nevo, E. (1996). Chromosome markers for Mus macedonicus (Rodentia, Muridae) from Israel. Hereditas. 124: 145–150.
Kocher, T. D., Thomas, W. K., Meyer, A., Edwards, S. V., Pääbo, S., Villablanca, F. X. and Wilson, A. C. (1989). Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci USA. 86: 6196–6200.
Marshall, J. T. and Sage, R. D. (1981). Taxonomy of the house mouse. Symp zool Soc Lond. 47: 15–25.
Moriwaki, K., Miyashita, N., Suzuki, H., Kurihara, Y. and Yonekawa, H. (1986). Genetic features of major geographical isolates of Mus musculus. Curr Topics Microbiol Immunol. 127: 55–61.
Nachman, M. W. and Searle, J. B. (1995). Why is the house mouse karyotype so variable? Trends Ecol Evol. 10: 397–402.
Nachman, M. W., Boyer, S. N., Searle, J. B. and Aquadro, C. F. (1994). Mitochondrial DNA variation and the evolution of Robertsonian chromosomal races of house mice, Mus domesticus. Genetics. 136: 1105–1120.
Nei, M. (1987) Molecular Evolutionary Genetics. Colombia University Press, New York.
Prager, E. M., Sage, R. D., Gyllensten, U., Thomas, W. K., Hübner, R. and Jones, C. S. et al.1993). Mitochondrial DNA sequence diversity and the colonization of Scandinavia by house mice from East Holstein. Biol J Linn Soc. 50: 85–122.
Prager, E. M., Tichy, H. and Sage, R. D. (1996). Mitochondrial DNA sequence variation in the eastern house mouse, Mus musculus: comparison with other house mice and report of a 75-bp tandem repeat. Genetics. 143: 427–446.
Prager, E. M., Orrego, C. and Sage, R. D. (1998). Genetic variation and phylogeography of central Asian and other house mice, including a major new mitochondrial lineage in Yemen. Genetics. 150: 835–861.
Richards, M. B., Macaulay, V. A., Bandelt, H. J. and Sykes, B. C. (1998). Phylogeography of mitochondrial DNA in western Europe. Ann Hum Genet. 62: 241–260.
Sage, R. D., Atchley, W. R. and Capanna, E. (1993). House mice as models in systematic biology. Syst Biol. 42: 523–561.
Saitou, N. and Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 4: 406–425.
Sambrook, J., Fritsch, E. F. and Maniatis, T. (1989) Molecular Cloning: a Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Schneider, S., Kueffer, J. -M., Roessli, D. and Excoffier, L. (1997) Arlequin, Version 1.1: a Software for Population Genetic Data Analysis. Genetics and Biometry Laboratory, University of Geneva.
Silver, L. M. (1995) Mouse Genetics: Concepts and Applications. Oxford University Press, New York.
Slatkin, M. and Hudson, R. R. (1991). Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations. Genetics. 129: 555–562.
Swofford, D. L. (1998) PAUP*: Phylogenetic Analysis Using Parsimony and Other Methods. Sinauer, Sunderland, MA.
Zima, J., Gaichenko, V. A., Macholán, M., Radjabli, S. I., Sablina, O. V. and Wójcik, J. M. (1990). Are Robertsonian variations a frequent phenomenon in mouse populations in Eurasia? Biol J Linn Soc. 41: 229–233.
Acknowledgements
We are grateful for discussions with Drs Milos Macholán and Jean-Christophe Auffray. We thank Dr Haluk Kefelioğlu for help in collection of specimens and Dr Pierre Taberlet for details of one of the primers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gündüz, İ., Tez, C., Malikov, V. et al. Mitochondrial DNA and chromosomal studies of wild mice (Mus) from Turkey and Iran. Heredity 84, 458–467 (2000). https://doi.org/10.1046/j.1365-2540.2000.00694.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00694.x
- Springer Nature Switzerland AG
Keywords
This article is cited by
-
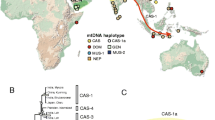
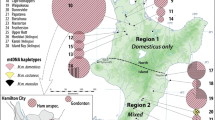
Genetic structure and invasion history of the house mouse (Mus musculus domesticus) in Senegal, West Africa: a legacy of colonial and contemporary times
Heredity (2017)
-
Multiple origins of the western European house mouse in the Aeolian Archipelago: clues from mtDNA and chromosomes
Biological Invasions (2013)
-
A chromosomal study on Greek populations of the genusApodemus (Rodentia, Murinae) reveals new data on B chromosome distribution
Acta Theriologica (2008)
-
Genetic evidence confirms the origin of the house mouse on sub-Antarctic Marion Island
Polar Biology (2007)
-
Two deeply divergent mitochondrial clades in the wild mouse Mus macedonicus reveal multiple glacial refuges south of Caucasus
Heredity (2002)