Abstract
This study provides empirical information on intrapopulation gene dispersal via pollen, the size of genetic ‘neighbourhoods’, and interpopulation gene flow in a long-lived conifer, limber pine (Pinus flexilis). We used allozyme loci for a paternity analysis of 518 seeds produced in an isolated population of limber pine located in north-eastern Colorado, U.S.A., separated by 2 km from the nearest conspecific trees and nearly 100 km from populations in the Rocky Mountains. We also used indirect techniques (FST analyses) to estimate gene flow rates among subdivisions of the study population and among five widely separated populations. Within the main study population limber pine exhibited a polymorphism level of 50%, observed heterozygosity of 0.159, and 2.36 alleles per polymorphic locus. Mountain populations were slightly more variable. The main study population showed significant differentiation in allozymes among neighbouring subpopulations. The mean FST was 0.031 and the gene flow rate among subpopulations was estimated as 7.8 migrants per generation. Among widely separated populations the mean FST was 0.035 and the gene flow rate was estimated as 6.9 migrants per generation. The paternity analysis indicated a best estimate of 6.5% pollen immigration (minimum 1.1%) from populations 2 km to 100+ km away. For 4% of the seeds examined, paternity could be ascribed to a single tree in the study population. Fractional paternity and likelihood methods were used to estimate pollen dispersal distances for the remainder of the seeds. Mean pollen dispersal distance was estimated at 140 m using the fractional method, similar to results from the other techniques. This compares with a mean distance of 172 m between potential mates. These results suggest near-panmictic pollen dispersal over this population, which covers about 15 ha. The observed allozyme differences and surprisingly low estimates of among-subpopulation gene flow are ascribed to a probable restriction of gene dispersal by seeds.
Similar content being viewed by others
Introduction
Gene dispersal, whether accomplished through the movement of gametes or zygotes, exerts a controlling influence on factors such as neighbourhood size, population structure, and the differentiation of populations. Mayr (1963) expressed the view that gene flow was ultimately responsible for maintaining the genetic cohesion of species. Wright (1977) and earlier assigned to gene flow the important role of spreading new genetic combinations in his shifting balance theory of evolution. Others, however, subsequently argued that gene exchange among populations occurs too rarely to have much significance in shaping the evolutionary process (Ehrlich & Raven, 1969; Levin, 1981).
In addition to interpopulation gene flow, patterns of mating and gene dispersal within populations strongly affect genetic variation and the rate and direction of microevolutionary change. Detailed studies of breeding patterns in wild populations have improved our understanding of genetic structure in populations and allowed prediction of their future evolution (Brown, 1990; Avise, 1994).
In plants, early attempts to estimate gene exchange between mating neighbourhoods used inferences based on observations of pollen and seed movements (Levin & Kerster, 1974). ‘Indirect’ methods were subsequently developed to use the spatial distribution of genotypic frequencies to estimate past levels of gene dispersal (Slatkin, 1985a,b). Difficulties and limitations of these methods led evolutionary biologists to express the need for ‘direct’ measurements of gene dispersal in plants (Ellstrand & Marshall, 1985; Hamrick, 1987).
Ellstrand & Marshall (1985) advocated the use of paternity analysis to measure directly gene dispersal via pollen. Subsequent paternity analyses have significantly added to our understanding of realized pollen dispersal, resulting in a re-emphasis of the role of gene flow in the evolutionary process (Godt & Hamrick, 1993; Broyles et al., 1994; Kaufman et al., 1998).
Our study used both direct and indirect methods at two spatial scales to examine gene dispersal in limber pine (Pinus flexilis James). We chose to study limber pine for several reasons. First, temperate conifers share many characteristics (e.g. large size, wind pollination, outcrossed breeding system, long lifespan) expected to result in high levels of long-distance gene dispersal (evidence cited in Schuster et al., 1989). Thus a detailed study in such a species should help to define the upper bounds of gene dispersal rates and distances in nature. Secondly, the haploid endosperm of gymnosperms allows unambiguous determination of the maternal contribution to embryos, a key benefit in paternity analyses (Neale, 1983). Thirdly, limber pine produces large seeds and possesses easily extracted, highly variable electrophoretic marker enzymes (Schuster et al., 1989). Finally, its broad geographical distribution in Colorado, including discrete, isolated populations ideal for paternity analyses, provided an opportunity for accurately estimating both short- and long-distance gene dispersal.
In this report, we first use indirect methods to estimate gene flow rates among groups of limber pines at two different spatial scales (100–500 m, and 10–150 km). We then report on a paternity analysis of 518 seeds from a single isolated population to estimate directly mating patterns within and pollen flow into this population. Finally, we compare the results obtained from these methods to assess the overall dispersal of genes via seeds and pollen, both within and among populations.
Materials and methods
Study sites
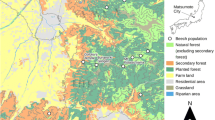
The primary study site is an isolated stand of limber pine located on the Pawnee National Grasslands in northern Colorado, U.S.A., nearly 100 km east of the Rocky Mountains (Figs 1 and 2). The trees are located in a series of adjacent north-facing canyons, separated by ridges up to 25 m high, except for a small number of trees located on the opposing, south-facing slope (Fig. 1). These trees are herein referred to as the Pawnee population, and are divided by site topography into groups termed subpopulations 1 through 5. The entire population consists of 455 trees confined to an elevational range between 1625 and 1655 m. Additional isolated populations occur along a west-facing escarpment that stretches 40 km to the north (Fig. 2; Schuster et al., 1995). The nearest population is 2 km north-east of Pawnee and the next nearest is 5 km to the west.
Limber pines were examined at four sites in addition to Pawnee, chosen to span the geographical and elevational range of limber pine in northern Colorado (Fig. 2). The Wilson/Foster stand is 13 km north-west of Pawnee (stand 6 in Schuster et al., 1995), and contains approximately 1500 limber pines in a similar ecological setting to Pawnee. Rocky Mountain juniper (Juniperus scopulorum Sarg.) is the only other tree species at both of these Great Plains sites. The Gem Lake study site is 130 km west of Pawnee in the montane zone of the Rocky Mountains (elevation 2690 m). This site contains several hundred limber pines in a south-facing basin along with less numerous Douglas-fir (Pseudotsuga menziesii (Mirb.) Franco). The Rainbow Curve study site is 14 km south-west of Gem Lake on south- and east-facing slopes near the upper treeline in Rocky Mountain National Park (elevation 3340 m). The Niwot Ridge study site is 40 km south of Rainbow Curve, also on south-facing slopes near the upper treeline, within the Niwot Ridge Long-term Ecological Research (LTER) study area (3330 m). At both of these high elevation sites, limber pines numbering in the thousands grow with numerous Engelmann spruce (Picea engelmannii (Parry) Engelm.) and subalpine fir (Abies lasiocarpa (Hook.) Nutt.).
Sample collection and preparation
Fresh needle tissue was collected for electrophoresis from all 371 trees at Pawnee that exceeded 0.6 m in height or 7 cm2 in basal area (no trees below these thresholds bore reproductive strobili over three years). Needles were also sampled from 26 of the 84 immature trees for a total of 397 trees sampled out of a population of 455. Electrophoretic analysis was used to determine the number of individuals present within multitrunk clusters (Schuster & Mitton, 1991). The entire population was mapped using aerial photographs and ground measurements.
At the four additional sites we sampled needle tissue from 50 randomly chosen trees in a 1–2 ha area. We avoided taking more than one sample from multiple-trunk clusters to minimize Wahlund effects and the oversampling of family groups (Linhart et al., 1981). Needle samples were ground in liquid nitrogen with the buffer of Mitton et al. (1979) and stored at −70°C.
For the paternity analysis, we randomly selected 10 or more cone-bearing trees from within each of the five subpopulations (total n=71). We collected four mature cones randomly from around the periphery of each tree. The sampled trees represented 27% of the trees that bore at least four mature cones that year. Cones were dried at 60°C for 24 h. All seeds were extracted, bulked by tree, then stored at 4°C until dissection.
Prior to electrophoresis, seed coats were removed and the embryos dissected away from their megagametophyte from 518 seeds. Six filled seeds per tree were dissected for most trees. However, for six of the most fecund trees (bearing >100 cones) we dissected between 12 and 29 seeds to adjust for variations in female fertility (Ennos & Dodson, 1987). Each embryo and half of each gametophyte were placed in separate, labelled microcentrifuge tubes. Tissues were immersed in 0.5 mL distilled water, homogenized with a blunt stirring rod, and then stored at −70°C.
Electrophoresis
Samples were absorbed onto filter paper wicks and electrophoresed on 12% starch gels following the methods of Mitton et al. (1979). Fifty randomly selected trees from Pawnee were initially surveyed, and five gel/electrode buffer combinations were used to resolve 17 enzyme systems. Phosphoglucoisomerase (PGI), shikimate dehydrogenase (SKDH), peroxidase (PER), isocitrate dehydrogenase (IDH) and glycerate dehydrogenase (GLYDH) were resolved using the discontinuous tris-citrate buffer of Poulik (1957). Fluorescent esterase (FE), glutamate dehydrogenase (GDH), triose-phosphate isomerase (TPI), and superoxide dismutase (SOD) were resolved using a modification of the Poulik buffer (J. Hamrick, pers. comm.). Uridine-5-diphosphoglucose pyrophosphorylase (UDP), 6-phosphogluconate dehydrogenase (6PGD), malate dehydrogenase (MDH), and aconitase (ACO) were resolved on a tris-citrate buffer system (Selander et al., 1971). Phosphoglucomutase (PGM), alcohol dehydrogenase (ADH) and mannose-6-phosphate isomerase (MPI) were resolved on the histidine II buffer system of King & Dancik (1983). Glutamic-oxaloacetic transaminase (GOT) was resolved on the Alle I buffer system of O’Malley et al. (1979). These 17 enzyme systems provided evidence of 31 putative loci, with banding patterns consistent with knowledge of enzyme structure and compartmentalization (Weeden & Wendel, 1989). Twenty-eight of these loci were consistently scorable in needle tissues. Results from the electrophoresis of gametophyte tissue confirmed Mendelian inheritance patterns for the 10 loci used in the paternity analysis.
Subsequently, the remaining trees in the Pawnee population and all samples from the four additional populations were surveyed for nine of these enzyme systems (PGM, UDP, 6PGD, MDH, PGI, IDH, SKDH, FE and PER) and scored for 12 loci. All 518 embryo and gametophyte samples were genotyped for seven of these systems (PGM, UDP, 6PGD, MDH, PGI, SKDH and FE) encoding 10 loci known to be variable at Pawnee. Multiple loci and alleles were numbered sequentially according to speed of anodal migration, with locus 1 and allele 1 migrating fastest. Identical standards were loaded on gels to ensure consistent scoring of allelomorphs.
Data analysis
Population-level polymorphism (P) at Pawnee was determined with a 99% criterion based on the random sample of 50 trees surveyed for all 28 electrophoretic loci. Mean number of alleles per locus (A) and per polymorphic locus (Ap), observed heterozygosity (HO), Nei’s (1978) unbiased estimate of gene diversity (HT; also termed expected heterozygosity) were calculated based on the 12 loci scored for all trees. These figures were converted to estimates over all 28 loci using the polymorphism level of 50%. Similar estimates of genetic variation at these 12 loci were also calculated for the other four limber pine populations.
At Pawnee, spatial genetic differentiation was assessed by comparing trees topographically divided into subpopulations 1 through 5 (Fig. 1). Similar comparisons were made among all five sampled populations, treating the 397 sampled trees at Pawnee as a single population.
Allelic frequencies were compared for each locus and over all loci using the methods of Workman & Niswander (1970). Nei’s genetic distance (D) and identity (I) were calculated among all pairs of populations or subpopulations (Nei, 1972). Wright’s F-statistics (FIS, FIT and FST) were calculated both within the Pawnee population and for the five sampled populations (Wright, 1965). We employed Nei’s method (1977) for calculating average F coefficients over multiple loci, weighting these averages over alleles. Significance of these F-values was tested using a chi-squared test. The resulting FST values were then used to generate indirect estimates of gene flow among populations. The number of migrants exchanged per generation, Nm, was estimated as (1/FST − 1)/4 (Wright, 1965).
A total of 277 potential male parents were identified at Pawnee by surveying male strobilus production in the spring of the year prior to cone collection. Pollen immigration from outside the Pawnee population was restricted to trees growing below about 2700 m, because of the strong relationship between elevation and date of pollen release (Schuster et al., 1989). The distance to other limber pines shedding pollen during this period was 2–40 km for other Plains populations or 100+ km for mountain populations below 2700 m.
For each seed, all possible male parents in the Pawnee stand were identified by the genetic exclusion method (Ellstrand & Marshall, 1985). Electrophoretic analysis of the embryo and haploid maternal gametophyte tissue in each seed allowed direct determination of the paternal contribution to the embryo for each of the 10 surveyed loci, and trees that could not have made such contributions were excluded. The average exclusion probability, representing the probability that any potential male parent could be excluded, was calculated as in Brown et al. (1989) based on population allele frequencies.
In cases where only one tree in the population could have provided the paternal complement of alleles, that tree was assigned paternity and the distance of realized gene dispersal (d ) was recorded as the distance between mates. In cases where none of the trees at Pawnee could have provided the paternal complement, paternity was assigned to an unknown source outside the population. A minimum distance of 2000 m was assigned to all such mating events. Mean effective pollen dispersal distance (d) was estimated using those cases where paternity was assigned either within or outside the population. The distribution of realized mating distances was used to estimate the neighbourhood size using the formula of Wright (1946); N=4πD(∑pi2/2Np), where D=density of adult plants, pi=axial distance of realized pollen dispersal, and Np=the number of pollen grains sampled (Crawford, 1984). These neighbourhood size estimates thus discount any contribution from seed dispersal.
Because, as expected (Chakraborty et al., 1988), paternity could be unambiguously assigned only in a small proportion of cases, two other methods were employed to assign paternity. The first was the likelihood method (Meagher, 1986; Meagher & Thompson, 1987), in which the likelihood of each potential male parent is calculated based on segregation probabilities. We assigned paternity only when the likelihood that a tree sired a seed was at least twice as great as for any other tree. We also employed the fractional method (Devlin et al., 1988), in which paternity for each seed is assigned fractionally to all possible male parents, based on their likelihood (i.e. segregation probability) of having sired that seed. Both d and N were estimated as above for each of these methods.
The simple exclusion method provided a minimum estimate for the total rate of gene migration via pollen entering the Pawnee population, because only pollen grains with a multilocus contribution not present at Pawnee could be identified as unambiguous immigrants. We also used a multilocus estimation method (Adams & Birkes, 1991) to estimate total immigration into the seed pool, including the fraction of ‘cryptic’ pollen which could not be discriminated from local pollen. The combined allelic frequencies from the Wilson/Foster and Gem Lake sites were used to approximate the genetic makeup of background pollen.
Results
Genetic variation
Of the 28 loci observed in samples from Pawnee, 14 were variable and 14 were fixed, producing a polymorphism estimate of 50% (Table 1). A total of 47 alleles were observed with a mean of 1.68 alleles per locus or 2.36 alleles per polymorphic locus. Mean observed heterozygosity (HO) over the 28 scorable loci was 0.159, and expected heterozygosity (HT) was 0.162.
Fifty trees randomly selected from the Pawnee population were compared for variation at 12 of these loci to trees sampled from each of four other populations (Table 1). Populations from the Great Plains (Pawnee and Wilson/Foster) were slightly less variable at these loci compared to mountain populations. Two loci, PGM-1 and IDH, were polymorphic in the populations in the mountains but were monomorphic at Pawnee.
Genetic differentiation and indirect estimates of gene flow among subpopulations
Allelic frequencies differed significantly among Pawnee subpopulations for five of the 10 variable loci surveyed (Table 2). The summed chi-squared value indicated significant spatial differentiation over all loci (P < 0.001). Although most alleles were observed in all subpopulations, five of the less common alleles were absent in one or two subpopulations. Frequencies of individual alleles differed among subpopulations by as much as 35%. Genetic distances among subpopulations ranged from a minimum of 0.003 between subpopulations 3 and 4 to a maximum of 0.031 between subpopulations 1 and 5. The results indicated that subpopulation 3 was most genetically similar to the other four subpopulations whereas subpopulations 1 and 5 were most genetically distinct.
Analysis of Wright’s F-statistics provided further information on population structure (Table 3). FIS values indicated no significant deviations from Hardy–Weinberg expectations within subpopulations. FST values ranged from 0.010 to 0.062 among loci, with an average of 0.031 over all alleles and loci (P < 0.001). These FST values produced estimates of gene flow among subpopulations (Nm) ranging from 3.8 (6PGD-1) to 24.8 (UDP-1), with a weighted mean of 7.8 migrants per generation.
Further information was obtained by repeating the FST analyses five times, removing one of the subpopulations at each iteration (Slatkin, 1985b). Removing subpopulation 3 resulted in a slight decrease of Nm to 6.3, suggesting that this subpopulation may have a higher rate of gene exchange with other subpopulations. In contrast, removing either subpopulation 1 or 5 resulted in a nearly 50% increase in Nm, suggesting that these subpopulations may experience restricted gene exchange.
Genetic differentiation and indirect estimates of gene flow among populations
Allelic frequencies differed significantly among the five sampled populations for nine of the 12 loci surveyed (Table 4). The summed chi-squared value indicated significant spatial differentiation of allele frequencies (P < 0.001). All loci that were highly differentiated among subpopulations within the Pawnee population (UDP-2, 6PGD-1, MDH-2 and PGI) were also highly differentiated among the five populations. In a similar fashion, most of the loci that were not spatially differentiated within the Pawnee population (6PGD-2, SKDH, FE-2 and PER-2) were weakly or not at all differentiated among populations. However, two other loci that showed little or no spatial differentiation at Pawnee were highly differentiated among populations (PGM-2 and UDP-1). No ‘private alleles’, sensu Slatkin (1985a), were observed, although six alleles (three at the SKDH locus) were observed in only two or three populations. The maximum allelic frequency difference between any pair of populations was 30%. Genetic distances between populations ranged from a minimum of 0.010 between the mid-elevation Gem Lake population and each of the upper treeline populations, to a maximum of 0.034 between the Pawnee and Wilson/Foster populations.
Values for Wright’s FST were less than 0.030 for nine of the 12 loci (Table 5). PGM-2 was the locus most strongly differentiated among populations, with an FST value (0.125) several times higher than that for most other loci. The FST values produced estimates of Nm ranging from 1.8 to 19.0, with a weighted mean estimate of 6.9 migrants per generation. Repeating this analysis removing the plains populations resulted in an increase in mean Nm to 8.4 and 9.0 when Wilson/Foster and Pawnee, respectively, were removed, suggesting that these populations may experience more restricted gene flow compared to mountain populations.
Paternity analysis
The average exclusion probability for our paternity analysis, using the 10 loci for which all trees, embryos and gametophytes were scored, was 0.88. Of the 518 embryo/gametophyte pairs examined, 44 (8.5%) produced few or no bands upon staining, and were apparently inviable. For the remaining 474 embryos, the number of nonexcluded potential male parents among the pollen-producing trees in the Pawnee population ranged from 0 to 125, with a mean of 47.6. For five of the seeds examined, no trees in the Pawnee population could have produced the pollen contribution inferred, indicating a 1.1% minimum estimate for pollen immigration. For 19 seeds (4% of the total), only a single tree in the population could have produced the observed pollen contribution. These matings are indicated graphically in Fig. 3 along with the trees that captured immigrant pollen. The remaining 450 seeds each had two or more possible male parents in the Pawnee population.
Map of the Pawnee limber pine population showing trees identified as having captured immigrant pollen (circled) and matings indicated by the paternity analysis (male and female parents connected by solid lines). The tree above the asterisk symbol captured two immigrant pollen grains and the tree above the letter ‘s’ produced one selfed progeny.
Mean pollen dispersal distance (d ) for the 19 seeds for which paternity was assigned using the simple exclusion method was 174.3 m (Table 6a). Slightly lower mean pollen dispersal estimates of 149.8 m and 160.5 m were obtained using the most likely and fractional methods, respectively. However, mean pollen dispersal distances varied widely among subpopulations, from as little as 68.5 m for centrally located subpopulation 3 to as much as 300 m for subpopulation 2. Mean dispersal distances estimated over all seeds examined were somewhat upwardly biased because of our sampling design; more seeds were examined per cone produced (0.26) in peripherally located subpopulation 1, compared to centrally located subpopulation 3 (0.05 seeds examined per cone produced). We attempted to eliminate this bias by weighting the mean pollen dispersal distance for each subpopulation by these ratios. The weighted mean dispersal distance estimates agreed well among methods, indicating mean gene dispersal distance via pollen within the Pawnee population between 133 m and 140 m.
However, these values do not reflect the distances traversed by immigrant pollen. Our mean, weighted dispersal distance estimates increased to 155–265 m when we used a minimum estimate (2 km) as the mating distance for all immigrant pollen (Table 6b).
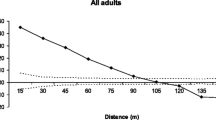
The fractional paternity analysis produced a frequency distribution of mating distances with a pronounced peak between 50 m and 200 m and a median of 133 m (Fig. 4). Matings at distances less than 50 m were much less common. A substantial number of matings were attributed to distances between 200 m and 400 m. The tail of the distribution was represented by the small number of matings ascribed to distances of more than 500 m.
Selfing events (assigned a distance of 0) were apparently infrequent. We observed only one unambiguously selfed seed among 474 total seeds, although 14 seeds (3%) were at least twice as likely to be selfed as opposed to outcrossed seeds. The overall outcrossing rate as estimated by the fractional paternity method was nearly 98%, with variations among maternal families but no significant differences among subpopulations.
Estimates of neighbourhood size based solely on pollen dispersal estimates ranged from 143 to 290 trees depending on the paternity analysis method used (Table 7). These are minimum estimates because some gene dispersal undoubtedly occurs via seeds. These neighbourhood size estimates also discount the contribution of immigrant pollen. Including these long-distance mating events, even at the minimum distance of 2 km, resulted in neighbourhood size estimates far in excess of the size of the Pawnee population.
Our minimum estimate of 1.1% pollen immigration also neglects any contribution from “cryptic” immigrant pollen. The multilocus estimation method of Adams & Birkes (1991) indicated that this ‘cryptic’ pollen was several fold more important than genetically distinguishable immigrant pollen, estimating the total rate of pollen immigration into the Pawnee population at 6.5%.
Discussion
Direct measurement of gene dispersal by pollen
The relatively high levels of genetic variation we discovered in limber pine are consistent with previous results for populations of forest trees, especially conifers (Hamrick & Godt, 1990). The marginal and isolated Pawnee population maintains nearly as high a level of genetic diversity as populations in the Rocky Mountains where limber pine is much more widespread. Only a few rare alleles seem to be absent, similar to studies of other tree species that have compared marginal to centrally located populations (Guries & Ledig, 1982; Betancourt et al., 1991).
This variation, combined with the advantage of being able to deduce paternal alleles in conifer embryos by comparison with gametophyte tissue, allowed us to use paternity analysis to study gene dispersal by pollen. Our results indicate that limber pine pollen effects gene dispersal over moderate (100 m to 200 m) to long (>2 km) distances at measurable frequencies. The data indicate low selfing rates (2–3%), although it is possible that the 8.5% of seeds which exhibited no enzyme activity could have been inviable products from self-pollinations. The results also indicate infrequent nearest-neighbour mating, because fewer than 10% of matings apparently occurred at distances less than 50 m (all trees had mature neighbours within this distance). Estimates of mean pollen dispersal distance within the population ranged from 133 m to 140 m and matings across more than 500 m were detected. Ridges up to 25 m high had little apparent effect on effective pollen dispersal. The distribution of pollen flow distances estimated by the fractional paternity method resembled a random mating pattern, with the exceptions of more matings near the mean (100 m to 150 m) and upper end (>500 m) of the distribution.
Results from several paternity analyses have reported an excess of near-neighbour matings as compared to random mating expectations (Meagher, 1986; Devlin & Ellstrand, 1990; Hamrick & Murawski, 1990; Godt & Hamrick, 1993; Schnabel & Hamrick, 1995). Notably these studies were all accomplished on insect-pollinated species (but see Broyles & Wyatt, 1991). In contrast, two studies of the wind-pollinated conifer Pseudotsuga menziesii (Neale, 1983; Adams & Birkes, 1991) and one examining Pinus ponderosa (Latta et al., 1998) indicated no excess of near-neighbour matings, similar to our findings in limber pine. Our mean within-population pollen dispersal distances are an order of magnitude higher than reported in most previous paternity analyses. This is partly because of the dispersed nature of this population (fewer than 20 mature trees per hectare), but also indicates near panmixia within the area.
Our estimates of genetic neighbourhood area (minimum 6 ha to 12 ha) are the highest we have seen documented, again partly caused by the dispersed nature of our study population. Our neighbourhood size estimates range from 143 to 290 individuals depending on the method used. These estimates do not include interpopulation gene flow by pollen nor any contribution from seed dispersal. Such seed dispersal must occur, although it may be restricted as indicated by the genetic structure analysis (see below). Because there are fewer than 300 reproductive trees in the study population, our paternity analysis indicates a genetic neighbourhood of the order of the actual size of the Pawnee population (15 ha).
We detected long-distance gene flow by pollen occurring across a minimum of 2 km of grasslands. Because of the prevailing westerly winds, this pollen probably travelled at least 5 km and may have traversed the distance from populations in the mountains (>100 km). A minimum of 1.1% of the seeds we examined (five out of 474) were sired by immigrant pollen. However, we estimated total immigrant pollen flow, including ‘cryptic’ pollen, as 6.5% (3.25% gene flow given the haploid nature of pollen). Including this interpopulation pollen flow increased our estimate of mean effective pollen dispersal distance to at least 155 m.
Previous paternity analyses have often reported significant interpopulation gene flow, even in studies of insect-pollinated herbs. Ellstrand & Marshall (1985) and Devlin & Ellstrand (1990) measured interpopulation gene flow by pollen in wild radish, Raphanus sativus. They reported 6% to 18% of seed fertilized by immigrant pollen travelling distances of 100 m or more and 8% to 9% in populations isolated by 600 m to 1 km. Godt & Hamrick (1993) estimated pollen-mediated gene flow in the perennial vine Lathyrus latifolius at 16% to 46% into sites separated by minimum distances of 20 m to 70 m from surrounding populations. Broyles et al. (1994) estimated 29% to 50% pollen immigration rate into natural populations of milkweed (Asclepias exaltata) over distances ranging from 50 m to 1 km.
Previous studies of wind-pollinated trees have indicated pollen flow of 30% to 50% or more across minimum distances of 100 m (Neale, 1983; Friedman & Adams, 1985; Adams & Birkes, 1991). Pollen flow was measured at 17% to 30% in two populations of the insect-pollinated tree Gleditsia triacanthos, isolated by a minimum distance of 85 m (Schnabel & Hamrick, 1995). Even longer distances of pollen flow, from 500 m to as much as 14 km, have been indicated for some widely spaced tropical trees (Hamrick & Murawski, 1990; Kaufman et al., 1998; Loveless et al., 1998).
Our estimate of 6.5% pollen flow is less than many of these previous estimates, but our minimum pollen immigration distance of 2 km is among the longest documented in the literature to date. Pollen from conifers has previously been captured 100 km or more from its probable source (Levin & Kerster, 1974). Our results suggest that gene flow in conifers may actually occur across distances of many kilometres, although pollen viability will certainly decrease with time spent in transport.
Our results are concordant with the growing body of evidence indicating that gene flow occurs in nature at a sufficient magnitude to be a substantial evolutionary force. Predictions that migration rates in nature are usually less than 1% (Levin & Kerster, 1974), that gene flow occurs at a scale of metres, not kilometres (Bradshaw, 1972), and that populations more than a few kilometres apart should be completely isolated (Levin, 1981) appear not to be supported by the data for limber pine and some other tree species. Our data on long-distance gene flow and a large area of panmictic pollen dispersal should help to circumscribe upper bounds for these parameters in nature. These levels of gene flow should be high enough to override the diversifying effects of genetic drift (Wright, 1951; Maruyama, 1970). We expect that natural selection would only be able to produce geographical differentiation under these circumstances if its strength is very high, exceeding the rate of gene flow.
Genetic structure at different spatial scales
Genetic substructure persists in the Pawnee limber pine population in the presence of apparently extensive gene dispersal by pollen. Our findings are reminiscent of those reported by Linhart et al. (1981) in which a family structure was documented in a population of a widely distributed tree (ponderosa pine) with few apparent barriers to gene dispersal. Our FST value of 0.031 and our Nm estimate of 6.9 among subpopulations only 100 m to 500 m apart are similar to previous results for conifer populations separated by many kilometres (Hamrick, 1987). Slatkin & Barton (1989) have shown that Nei’s FST methods slightly underestimate Nm, so ours must be considered conservative gene dispersal estimates. Whitlock & McCauley (1999) have argued that FST and similar measures provide at best only rough estimates of gene dispersal because of frequent violation of the assumptions underlying the models. Nevertheless, significant spatial genetic structure remains evident at Pawnee: in particular, the most peripheral subpopulations (1 and 5) show moderate differentiation at several loci.
Strong natural selection appears a possible but unlikely culprit for this pattern, as most of the subpopulations appear ecologically identical. Although the 18 tree clusters in the population appear to contain related individuals (Schuster & Mitton, 1991), significant genetic differentiation was evident even when these clusters were eliminated from the analysis.
Spatial genetic structure in plant populations has usually been ascribed to limited seed and/or pollen dispersal (Levin & Kerster, 1974; Loveless & Hamrick, 1984). In light of the extensive pollen dispersal revealed by our paternity analysis, we hypothesize that this genetic structure could persist only if gene dispersal by seeds is restricted. Limber pine seeds are heavy, wingless, and adapted for dispersal by birds, in particular the Clark’s nutcracker (Nucifraga columbiana; Tomback, 1977). In three years of study we never observed a Clark’s nutcracker at Pawnee despite good cone crops. This site is peripheral to the main range of both the nutcracker and limber pine, and the birds may visit these stands only when seed crops fail in mountain populations. In the absence of its primary disperser, gene movement via seeds would be expected to be highly restricted. Our implication of restricted seed dispersal resulting in genetic structure despite extensive pollen dispersal is similar to the findings of Latta et al. (1998) in the aforementioned population of ponderosa pine.
The larger-scale patterns we discovered in limber pine are consistent with previous results for conifer trees exhibiting high levels of genetic variation residing mostly within rather than among populations (Loveless & Hamrick, 1984). The five populations of limber pine that we examined, separated by as much as 150 km, were only slightly more differentiated than the subpopulations at Pawnee (mean genetic distance of 0.020 vs. 0.015). The three populations in the mountains, where limber pine is more widespread, are highly similar (mean genetic identity of 0.989). This is consistent with our previous findings (Schuster et al., 1989), suggesting significant gene flow among mountain populations. Latta & Mitton (1997) also found low levels of among-population variation in allozymes (and cpDNA) for seven populations of limber pine in the Rocky Mountains. The two populations we examined on the Great Plains (Pawnee and Wilson/Foster) were more differentiated and lacked some rare alleles present in the mountain populations. Yet the indirect analyses mainly indicated high genetic similarity and rates of gene flow among populations, consistent with the results from the paternity analysis.
Utility and limitations of the methods
Inferences from our paternity analysis must be limited by the fact that it was accomplished in only one year, because previous studies have revealed significant year-to-year variation in gene flow frequencies among plant populations (Hamrick & Murawski, 1990; Schnabel & Hamrick, 1995). Also, despite high genetic variation, the electrophoretic markers available were insufficient to exclude a large proportion of potential male parents for many of the seeds. We found the fractional paternity method to be superior to the simple exclusion and likelihood methods in gleaning gene dispersal data from a larger proportion of the seeds.
We consider our best estimate of mean intrapopulation pollen flow distance to be the one produced using the fractional paternity method (i.e. 140 m). However, Adams et al. (1992) have warned that estimates from fractional paternity analyses may be upwardly biased if male mating success decreases with distance. This may not be the case in limber pine, and thus adding mating probability estimates based on distance could lead to erroneous results. The fractional method also contains a bias towards more uniform fertility and random mating than may actually exist. Despite these shortcomings, we are encouraged by the substantial agreement of the results of the fractional method with those from the simple exclusion and likelihood methods.
Paternity analyses provide no information on which seeds sired will result in mature trees, no information on seed dispersal, and are usually only accomplished over one or a few years. ‘Indirect’ methods provide no means to discriminate between pollen and seed flow, and they estimate historical gene flow levels rather than current rates. Their accuracy may also be affected by the validity of assumptions such as genetic equilibrium and the absence of strong natural selection (Slatkin, 1985b). Slatkin & Barton (1989) have advised that indirect estimates should be considered rough indicators of gene flow, with their accuracy dependent on departures of the data from underlying models.
The simultaneous use of both direct and indirect methods can provide more information than either method alone, overcoming some of the above limitations (Kaufman et al., 1998; Loveless et al., 1998). Whitlock & McCauley (1999) suggested that divergent results between the methods could indicate important ecological deviations from typical island model assumptions. In our case, our conclusion of extensive interpopulation gene flow in limber pine is substantiated by general agreement between the results of both methods. However, the fairly low estimate of gene dispersal within the Pawnee population produced by the indirect method may reflect the poor fit of this population to an island model.
Comparing the results of both methods also permits inferences about the magnitude of gene flow via seeds vs. pollen. At the intrapopulation level, contrasting results between the methods led us to conclude that restricted seed dispersal is the reason for persistent genetic substructure. Uniparentally inherited organellar DNA markers have provided a powerful alternative means to discriminate between pollen and seed flow, especially in conifers in which cpDNA is paternally inherited whereas mtDNA is inherited maternally (Dong & Wagner, 1994; Latta & Mitton, 1997). In limber pine, the spatial distribution of cpDNA and mtDNA haplotypes strongly suggests that pollen travels more readily among populations than do seeds (Latta & Mitton, 1997; Mitton et al., 1999). Our empirical results confirm both high levels and long distances of interpopulation gene flow via pollen in this species. Although the dispersal of diploid seeds can potentially be more important than pollen dispersal to overall gene flow, its relative importance will vary with the effectiveness of dispersal over different scales of space and time.
References
Adams, W. T. and Birkes, D. S. (1991). Estimating mating patterns in forest tree populations. In: Fineschi, S., Malvolti, M. E., Cannata, F. and Hattemer, H. H. (eds) Biochemical Markers in the Population Genetics of Forest Trees, pp. 157–172. SPB Academic Publishing, The Hague.
Adams, W. T., Griffin, A. R. and Moran, G. R. (1992). Using paternity analysis to measure effective pollen dispersal in plant populations. Am Nat, 140: 762–780.
Avise, J. C. (1994). Molecular Markers, Natural History, and Evolution. Chapman & Hall, New York.
Betancourt, J. L., Schuster, W. S., Mitton, J. B. and Anderson, R. S. (1991). Fossil and genetic history of a pinyon pine (Pinus edulis) isolate. Ecology, 72: 1685–1697.
Bradshaw, A. D. (1972). Some of the evolutionary consequences of being a plant. Evol Biol, 5: 25–47.
Brown, A. H. D. (1990). Genetic characterization of plant mating systems. In: Brown, A. H. D., Clegg, M. T., Kahler, A. L. and Weir, B. S. (eds) Plant Population Genetics, Breeding, and Genetic Resources, pp. 145–162. Sinauer Associates, Sunderland, MA.
Brown, A. H. D., Burdon, J. J. and Jarosz, A. M. (1989). Isozyme analysis of plant mating systems. In: Soltis, D. E. and Soltis, P. S. (eds) Isozymes in Plant Biology, pp. 73–86. Dioscorides Press, Portland, OR.
Broyles, S. B. and Wyatt, R. (1991). Effective pollen dispersal in a natural population of Asclepias exaltata: the influence of pollinator behavior, genetic similarity, and mating success. Am Nat, 138: 1239–1249.
Broyles, S. B., Schnabel, A. and Wyatt, R. (1994). Evidence for long-distance pollen dispersal in milkweeds (Asclepias exaltata). Evolution, 48: 1032–1040.
Chakraborty, R., Meagher, T. R. and Smouse, P. E. (1988). Parentage analysis with genetic markers in natural populations. I. The expected proportion of offspring with unambiguous paternity. Genetics, 118: 527–536.
Crawford, T. J. (1984). The estimation of neighbourhood parameters for plant populations. Heredity, 52: 273–283.
Devlin, B. and Ellstrand, N. C. (1990). Male and female fertility variation in wild radish, a hermaphrodite. Am Nat, 136: 87–107.
Devlin, B., Roeder, K. and Ellstrand, N. C. (1988). Fractional paternity assignment: theoretical development and comparison to other methods. Theor Appl Genet, 76: 369–380.
Dong, J. and Wagner, D. B. (1994). Paternally inherited chloroplast polymorphism in Pinus: estimation of diversity and population subdivision, and tests of disequilibrium with a maternally inherited mitochondrial polymorphism. Genetics, 136: 1187–1194.
Ehrlich, P. R. and Raven, P. H. (1969). Differentiation of populations. Science, 165: 1228–1232.
Ellstrand, N. C. and Marshall, D. L. (1985). Interpopulation gene flow by pollen in wild radish, Raphanus sativus. Am Nat, 126: 606–616.
Ennos, R. A. and Dodson, R. K. (1987). Pollen success, functional gender and assortative mating in an experimental plant population. Heredity, 58: 119–126.
Friedman, S. T. and Adams, W. T. (1985). Estimation of gene flow into two seed orchards of loblolly pine (Pinus taeda L.). Theor Appl Genet, 69: 609–615.
Godt, M. J. and Hamrick, J. L. (1993). Patterns and levels of pollen-mediated gene flow in Lathyrus latifolius. Evolution, 47: 98–110.
Guries, R. P. and Ledig, F. T. (1982). Genetic diversity and population structure in pitch pine (Pinus rigida Mill.). Evolution, 36: 387–399.
Hamrick, J. L. (1987). Gene flow and distribution of genetic variation in plant populations. In: Urbanska, K. M. (ed.) Differentiation Patterns in Higher Plants, pp. 53–67. Academic Press, New York.
Hamrick, J. L. and Godt, M. J. (1990). Allozyme diversity in plant species. In: Brown, A. H. D., Clegg, M. T., Kahler, A. L. and Weir, B. S. (eds) Plant Population Genetics, Breeding, and Genetic Resources, pp.43–63. Sinauer Associates, Sunderland, MA.
Hamrick, J. L. and Murawski, D. (1990). The breeding structure of tropical tree populations. Pl Sp Biol, 5: 157–165.
Kaufman, S. R., Smouse, P. E. and Alvarez-Buylla, E. R. (1998). Pollen-mediated gene flow and differential male reproductive success in a tropical pioneer tree, Cecropia obtusifolia Bertol. (Moraceae): a paternity analysis. Heredity, 81: 164–173.
King, J. N. and Dancik, B. P. (1983). Inheritance and linkage of isozymes in white spruce (Picea glauca). Can J Genet Cytol, 25: 430–436.
Latta, R. G. and Mitton, J. B. (1997). A comparison of population differentiation across four classes of gene markers in limber pine (Pinus flexilis James). Genetics, 146: 1153–1163.
Latta, R. G., Linhart, Y. B., Fleck, D. and Elliot, M. (1998). Direct and indirect estimates of seed versus pollen movement within a population of ponderosa pine. Evolution, 52: 61–67.
Levin, D. A. (1981). Dispersal versus gene flow in plants. Ann Mo Bot Gard, 68: 233–253.
Levin, D. A. and Kerster, H. W. (1974). Gene flow in seed plants. Evol Biol, 7: 139–220.
Linhart, Y. B., Mitton, J. B., Sturgeon, K. B. and Davis, M. L. (1981). Genetic variation in space and time in a population of ponderosa pine. Heredity, 46: 407–426.
Loveless, M. D. and Hamrick, J. L. (1984). Ecological determinants of genetic structure in plant populations. Ann Rev Ecol Syst, 15: 65–95.
Loveless, M. D., Hamrick, J. L. and Foster, R. B. (1998). Population structure and mating system in Tachigali versicolor, a monocarpic neotropical tree. Heredity, 81: 134–143.
Maruyama, T. (1970). On the rate of decrease of heterogeneity in circular stepping stone models of populations. Theor Pop Biol, 1: 101–119.
Mayr, E. (1963). Animal Species and Evolution. Harvard University Press, Cambridge, MA.
Meagher, T. R. (1986). Analysis of paternity within a natural population of Chamaelirium luteum. I. Identification of most likely male parents. Am Nat, 128: 199–215.
Meagher, T. R. and Thompson, E. (1987). Analysis of parentage for naturally established seedlings of Chamaelirium luteum (Liliaceae). Ecology, 68: 803–812.
Mitton, J. B., Linhart, Y. B., Sturgeon, K. B. and Hamrick, J. L. (1979). Allozyme polymorphisms detected in mature needle tissue of ponderosa pine. J Hered, 70: 86–89.
Mitton, J. B., Kreiser, B. R. and Latta, R. G. (1999). Glacial refugia of limber pine (Pinus flexilis James) inferred from the populations structure of mitochondrial DNA. Genetics, in press.
Neale, D. B. (1983). Population Genetic Structure of the Shelterwood Regeneration System in Southwest Oregon. PhD Thesis, Oregon State University, Corvallis.
Nei, M. (1972). Genetic distance between populations. Am Nat, 106: 283–292.
Nei, M. (1977). F-statistics and analysis of gene diversity in subdivided populations. Ann Hum Genet, 41: 225–233.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
O’malley, D. M., Allendorf, F. W. and Blake, G. M. (1979). Inheritance of isozyme variation and heterozygosity in Pinus ponderosa. Biochem Genet, 17: 233–250.
Poulik, M. D. (1957). Starch gel electrophoresis in a discontinuous system of buffers. Nature, 180: 1477–1479.
Schnabel, A. and Hamrick, J. L. (1995). Understanding the population genetic structure of Gleditsia triacanthos L.: The scale and pattern of pollen gene flow. Evolution, 49: 921–931.
Schuster, W. S. F. and Mitton, J. B. (1991). Relatedness within clusters of a bird-dispersed pine and the potential for kin interactions. Heredity, 67: 41–48.
Schuster, W. S., Alles, D. L. and Mitton, J. B. (1989). Gene flow in limber pine: evidence from pollination phenology and genetic differentiation along an elevational transect. Am J Bot, 76: 1395–1403.
Schuster, W. S. F., Mitton, J. B., Yamaguchi, D. K. and Woodhouse, C. A. (1995). A comparison of limber pine (Pinus flexilis) ages at lower and upper treeline sites east of the Continental Divide in Colorado. Am Midl Nat, 133: 101–111.
Selander, R. K., Smith, M. H., Yang, S. Y. and Gentry, J. B. (1971). Biochemical polymorphism and systematics in the genus Peromyscus. I. Variation in the old-field mouse. Studies in Genetics VI. University Texas Publication, 7103: 49–90.
Slatkin, M. (1985a). Rare alleles as indicators of gene flow. Evolution, 39: 53–65.
Slatkin, M. (1985b). Gene flow in natural populations. Ann Rev Ecol Syst, 16: 393–430.
Slatkin, M. and Barton, N. H. (1989). A comparison of three indirect methods for estimating average levels of gene flow. Evolution, 43: 1349–1368.
Tomback, D. F. (1977). Foraging strategies of Clark’s Nutcracker. Living Bird, 16: 123–161.
Weeden, N. F. and Wendel, J. F. (1989). Genetics of plant isozymes. In: Soltis, D. E. and Soltis, P. S. (eds) Isozymes in Plant Biology, pp. 46–72. Dioscorides Press, Portland, OR.
Whitlock, M. C. and Mccauley, D. E. (1999). Indirect measures of gene flow and migration: FST ≠ 1/ (4Nm + 1). Heredity, 82: 117–125.
Workman, P. L. and Niswander, J. D. (1970). Population studies on southwestern Indian Tribes. II. Local genetic differentiation in the Papago. Am J Hum Genet, 22: 24–29.
Wright, S. (1946). Isolation by distance under diverse mating systems. Genetics, 31: 39–59.
Wright, S. (1951). The genetical structure of populations. Ann Eugen, 15: 323–354.
Wright, S. (1965). The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution, 19: 395–420.
Wright, S. (1977). Evolution and the Genetics of Populations, vol. 3, Experimental Results and Evolutionary Deductions. University of Chicago Press, Chicago, IL.
Acknowledgements
This research was funded by NSF grant BSR-8614937. We would like to thank the Pawnee National Grasslands of the USDA Forest Service for allowing us to work at the Pawnee site. Eric Stark, Steve Levin, Wendy Sell, David Alles, Pam Miller, Kristen Cottle and Gwen Hunt provided much assistance with field and laboratory work. Yan Linhart and Mike Grant provided helpful comments and advice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schuster, W., Mitton, J. Paternity and gene dispersal in limber pine (Pinus flexilis James). Heredity 84, 348–361 (2000). https://doi.org/10.1046/j.1365-2540.2000.00684.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00684.x
- Springer Nature Switzerland AG
Keywords
This article is cited by
-
Reduced within-population quantitative genetic variation is associated with climate harshness in maritime pine
Heredity (2023)
-
SNP-based analysis reveals unexpected features of genetic diversity, parental contributions and pollen contamination in a white spruce breeding program
Scientific Reports (2021)
-
RNA-Seq reveals adaptive genetic potential of the rare Torrey pine (Pinus torreyana) in the face of Ips bark beetle outbreaks
Conservation Genetics (2021)
-
Wind pollination over 70 years reduces the negative genetic effects of severe forest fragmentation in the tropical oak Quercus bambusifolia
Heredity (2020)
-
Conserving genetic diversity in ecological restoration: a case study with ponderosa pine in northern Arizona, USA
New Forests (2017)