Abstract
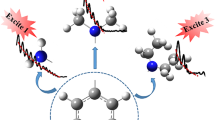
The structural properties and ultrafast electronic deactivation dynamics of the inosine dimer in CHCl3 have been investigated by two-dimensional 1H NMR and static FTIR spectroscopy and by femtosecond time-resolved transient absorption spectroscopy, respectively. The 1H NMR and IR spectra show the formation of a well-defined, symmetric dimer with an association equilibrium constant of KI·I = 690 ± 100 M-1. The excited-state dynamics after photoexcitation at λpump = 260 nm monitored by ultrafast absorption spectroscopy show great similarity with those of the monomer inosine in an aqueous solution and are governed by a decay time of t = 90 ± 10 fs, which is one of the shortest electronic lifetimes of all nucleobases and nucleobase dimers studied so far. On the basis of these observations, the inosine dimer is expected to follow a similar relaxation pathway as the monomer, involving an out-of-plane deformation of the six-membered ring. The importance of the C(2) position for the electronic deactivation of hypoxanthine and guanine is discussed. The obtained well-determined structure and straightforward dynamics qualify the inosine dimer as an excellent reference case for more complicated systems such as the G·G dimer and the G·C and A·T Watson-Crick pairs.
Similar content being viewed by others
Notes and references
M. Hill-Perkins, M. D. Jones, P. Karran, Site-specific mutagenesis in vivo by single methylated or deaminated purine bases, Mutat. Res., 1986, 162, 153–163.
K. A. Schouten, B. Weiss, Endonuclease V protects Escherichia coli against specific mutations caused by nitrous acid, Mutat. Res., 1999, 435, 245–254.
Y. Kyogoku, R. C. Lord, A. Rich, An infrared study of the hydrogen-bonding specificity of hypoxanthine and other nucleic acid derivatives, Biochim. Biophys. Acta, 1969, 179, 10–17.
F. H. Martin, M. M. Castro, F. Aboul-ela, I. Tinoco, Base pairing involving deoxyinosine: implications for probe design, Nucleic Acids Res., 1985, 13, 8927–8938.
E. Ohtsuka, S. Matsuki, M. Ikehara, Y. Takahashi, K. Matsubara, An alternative approach to deoxyoligonucleotides as hybridization probes by insertion of deoxyinosine at ambiguous codon positions, J. Biol. Chem., 1985, 260, 2605–2608.
M. R. Valentine, J. Termini, Kinetics of formation of hypoxanthine containing base pairs by HIV-RT: RNA template effects on the base substitution frequencies, Nucleic Acids Res., 2001, 29, 1191–1199.
Y. W. Kow, Repair of deaminated bases in DNA, Free Radical Biol. Med., 2002, 33, 886–893.
G. Paragi, I. Pálinkó, C. Van Alsenoy, I. K. Gyémánt, B. Penke, Z. Timár, Ab initio studies on the H-bonding of hypoxanthine and DNA bases, New J. Chem., 2002, 26, 1503–1506.
N. E. Watkins, J. SantaLucia, Nearest-neighbor thermodynamics of deoxyinosine pairs in DNA duplexes, Nucleic Acids Res., 2005, 33, 6258–6267.
P. Karran, T. Lindahl, Hypoxanthine in deoxyribonucleic acid: generation by heat-induced hydrolysis of adenine residues and release in free form by a deoxyribonucleic acid glycosylase from calf thymus, Biochemistry, 1980, 19, 6005–6011.
T. Nguyen, D. Brunson, C. L. Crespi, B. W. Penman, J. S. Wishnok, S. R. Tannenbaum, DNA damage and mutation in human cells exposed to nitric oxide in vitro, Proc. Natl. Acad. Sci. U. S. A., 1992, 89, 3030–3034.
J. P. E. Spencer, A. Jenner, K. Chimel, O. I. Aruoma, C. E. Cross, R. Wu, B. Halliwell, DNA damage in human respiratory tract epithelial cells: damage by gas phase cigarette smoke apparently involves attack by reactive nitrogen species in addition to oxygen radicals, FEBS Lett., 1995, 375, 179–182.
B. Pang, J. L. McFaline, N. E. Burgis, M. Dong, K. Taghizadeh, M. R. Sullivan, C. E. Elmquist, R. P. Cunningham, P. C. Dedon, Defects in purine nucleotide metabolism lead to substantial incorporation of xanthine and hypoxanthine into DNA and RNA, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 2319–2324.
E. Ohtsuka, Non-canonical hydrogen bonds in genetic information flow, Proc. Jpn. Acad., Ser. B, 2005, 81, 393–402.
N. A. Siegfried, S. L. Metzger, P. C. Bevilacqua, Folding cooperativity in RNA and DNA is dependent on position in the helix, Biochemistry, 2007, 46, 172–181.
X. Sun, J. K. Lee, Stability of DNA duplexes containing hypoxanthine (inosine): gas versus solution phase and biological implications, J. Org. Chem., 2010, 75, 1848–1854.
M. Krepl, M. Otyepka, P. Banáš, J. Šponer, Effect of guanine to inosine substitution on stability of canonical DNA and RNA duplexes: molecular dynamics thermodynamics integration study, J. Phys. Chem. B, 2013, 117, 1872–1879.
K. Röttger, R. Siewertsen, F. Temps, Ultrafast electronic deactivation dynamics of the rare natural nucleobase hypoxanthine, Chem. Phys. Lett., 2012, 536, 140–146.
J. Chen, B. Kohler, Ultrafast nonradiative decay by hypoxanthine and several methylxanthines in aqueous and acetonitrile solution, Phys. Chem. Chem. Phys., 2012, 14, 10677–10682.
J. P. Villabona-Monsalve, R. Noria, S. Matsika, J. Peón, On the accessibility to conical intersections in purines: hypoxanthine and its singly protonated and deprotonated forms, J. Am. Chem. Soc., 2012, 134, 7820–7829.
A. L. Sobolewski, W. Domcke, Ab initio studies on the photophysics of the guanine-cytosine base pair, Phys. Chem. Chem. Phys., 2004, 6, 2763–2771.
A. L. Sobolewski, W. Domcke, C. Hättig, Tautomeric selectivity of the excited-state lifetime of guanine/cytosine base pairs: the role of electron-driven proton-transfer processes, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 17903–17906.
A. Abo-Riziq, L. Grace, E. Nir, M. Kabelac, P. Hobza, M. S. de Vries, Photochemical selectivity in guanine-cytosine base-pair structures, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 20–23.
A. L. Sobolewski, W. Domcke, Relevance of electron-driven proton-transfer processes for the photostability of proteins, ChemPhysChem, 2006, 7, 561–564.
N. K. Schwalb, F. Temps, Ultrafast electronic relaxation in guanosine is promoted by hydrogen bonding with cytidine, J. Am. Chem. Soc., 2007, 129, 9272–9273.
G. Groenhof, L. V. Schäfer, M. Boggio-Pasqua, M. Goette, H. Grubmüller, M. A. Robb, Ultrafast deactivation of an excited cytosine-guanine base pair in DNA, J. Am. Chem. Soc., 2007, 129, 6812–6819.
N. K. Schwalb, T. Michalak, F. Temps, Ultrashort fluorescence lifetimes of hydrogen-bonded base pairs of guanosine and cytidine in solution, J. Phys. Chem. B, 2009, 113, 16365–16376.
L. Biemann, S. A. Kovalenko, K. Kleinermanns, R. Mahrwald, M. Markert, R. Improta, Excited state proton transfer is not involved in the ultrafast deactivation of guanine-cytosine pair in solution, J. Am. Chem. Soc., 2011, 133, 19664–19667.
S. Yamazaki, T. Taketsugu, Photoreaction channels of the guanine-cytosine base pair explored by long-range corrected TDDFT calculations, Phys. Chem. Chem. Phys., 2012, 14, 8866–8877.
K. K. Ogilvie, The tert-butyldimethylsilyl group as a protecting group in deoxynucleosides, Can. J. Chem., 1973, 51, 3799–3807.
T. Hupp, C. Sturm, E. M. Basílio Janke, M. Pérez Cabre, K. Weisz, B. Engels, A combined computational and experimental study of the hydrogen-bonded dimers of xanthine and hypoxanthine, J. Phys. Chem. A, 2005, 109, 1703–1712.
M. Yang, L. Szyc, K. Röttger, H. Fidder, E. T. J. Nibbering, T. Elsaesser, F. Temps, Dynamics and couplings of N-H stretching excitations of guanosine-cytidine base pairs in solution, J. Phys. Chem. B, 2011, 115, 5484–5492.
H. Fidder, M. Yang, E. T. J. Nibbering, T. Elsaesser, K. Röttger, F. Temps, N-H stretching vibrations of guanosine-cytidine base pairs in solution: ultrafast dynamics, couplings, and line shapes, J. Phys. Chem. A, 2013, 117, 845–854.
L. Biemann, T. Häber, D. Maydt, K. Schaper, K. Kleinermanns, Structural assignment of adenine aggregates in CDCl3, J. Chem. Phys., 2008, 128, 195103.
C. Greve, N. K. Preketes, H. Fidder, R. Costard, B. Koeppe, I. A. Heisler, S. Mukamel, F. Temps, E. T. J. Nibbering, T. Elsaesser, N-H stretching excitations in adenosine-thymidine base pairs in solution: pair geometries, infrared line shapes, and ultrafast vibrational dynamics, J. Phys. Chem. A, 2013, 117, 594–606.
K. Röttger, N. K. Schwalb, F. Temps, Electronic deactivation of guanosine in extended hydrogen-bonded self-assemblies, J. Phys. Chem. A, 2013, 117, 2469–2478.
CRC Handbook of Chemistry and Physics, ed. R. C. Weast, CRC Press, 67th edn, 1986-1987, pp. E58-E59.
N. K. Schwalb, F. Temps, On the structure and excited electronic state lifetimes of cytidine self-assemblies with extended hydrogen-bonding networks, J. Photochem. Photobiol., A, 2009, 208, 164–170.
V. Karunakaran, K. Kleinermanns, R. Improta, S. A. Kovalenko, Photoinduced dynamics of guanosine monophosphate in water from broad-band transient absorption spectroscopy and quantum-chemical calculations, J. Am. Chem. Soc., 2009, 131, 5839–5850.
J.-M. L. Pecourt, J. Peon, B. Kohler, DNA excited-state dynamics: ultrafast internal conversion and vibrational cooling in a series of nucleosides, J. Am. Chem. Soc., 2001, 123, 10370–10378.
W. G. Rothschild, G. J. Rosasco, R. C. Livingston, Dynamics of molecular reorientational motion and vibrational relaxation in liquids. Chloroform, J. Chem. Phys., 1975, 62, 1253–1268.
H. Graener, G. Seifert, Vibrational and orientational relaxation of monomeric water molecules in liquids, J. Chem. Phys., 1993, 98, 36–45.
H. Graener, R. Zürl, M. Hofmann, Vibrational relaxation of liquid chloroform, J. Phys. Chem. B, 1997, 101, 1745–1749.
E. L. Sibert, R. Rey, Vibrational relaxation in liquid chloroform following ultrafast excitation of the CH stretch fundamental, J. Chem. Phys., 2002, 116, 237–257.
H. Graener, R. Zürl, M. Bartel, G. Seifert, Thermalization of vibrational excess energy in liquid chloroform, J. Mol. Liq., 2000, 84, 161–168.
F.-A. Miannay, T. Gustavsson, A. Banyasz, D. Markovitsi, Excited-state dynamics of dGMP measured by steady-state and femtosecond fluorescence spectroscopy, J. Phys. Chem. A, 2010, 114, 3256–3263.
L. Serrano-Andrés, M. Merchán, A. C. Borin, A three-state model for the photophysics of guanine, J. Am. Chem. Soc., 2008, 130, 2473–2484.
S. Yamazaki, W. Domcke, Ab initio studies on the photophysics of guanine tautomers: out-of-plane deformation and NH dissociation pathways to conical intersections, J. Phys. Chem. A, 2008, 112, 7090–7097.
S. Yamazaki, W. Domcke, A. L. Sobolewski, Nonradiative decay mechanisms of the biologically relevant tautomer of guanine, J. Phys. Chem. A, 2008, 112, 11965–11968.
L. Serrano-Andrés, M. Merchán, Are the five natural DNA/RNA base monomers a good choice from natural selection? A photochemical perspective, J. Photochem. Photobiol., C, 2009, 10, 21–32.
Z. Lan, E. Fabiano, W. Thiel, Photoinduced nonadiabatic dynamics of 9 H-guanine, ChemPhysChem, 2009, 10, 1225–1229.
M. Barbatti, J. J. Szymczak, A. J. A. Aquino, D. Nachtigallová, H. Lischka, The decay mechanism of photoexcited guanine - a nonadiabatic dynamics study, J. Chem. Phys., 2011, 134, 014304.
B. Heggen, Z. Lan, W. Thiel, Nonadiabatic decay dynamics of 9 H-guanine in aqueous solution, Phys. Chem. Chem. Phys., 2012, 14, 8137–8146.
C. C.-W. Cheng, C. Ma, C. T.-L. Chan, K. Y.-F. Ho, W.-M. Kwok, The solvent effect and identification of a weakly emissive state in nonradiative dynamics of guanine nucleosides and nucleotides - a combined femtosecond broadband time-resolved fluorescence and transient absorption study, Photochem. Photobiol. Sci., 2013. 10.1039/C3PP25450J
K. Hunger, L. Buschhaus, L. Biemann, M. Braun, S. Kovalenko, R. Improta, K. Kleinermanns, UV light-induced hydrogen transfer in guanosine-guanosine aggregates, Chem.-Eur. J., 2013, 19, 5425–5431.
T. Zelený, M. Ruckenbauer, A. J. A. Aquino, T. Müller, F. Lankaš, T. Dršata, W. L. Hase, D. Nachtigallova, H. Lischka, Strikingly different effects of hydrogen bonding on the photodynamics of individual nucleobases in DNA: comparison of guanine and cytosine, J. Am. Chem. Soc., 2012, 134, 13662–13669.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Röttger, K., Sönnichsen, F.D. & Temps, F. Ultrafast electronic deactivation dynamics of the inosine dimer — a model case for H-bonded purine bases. Photochem Photobiol Sci 12, 1466–1473 (2013). https://doi.org/10.1039/c3pp50093d
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1039/c3pp50093d