Abstract
Air pollution has been classified as a group 1 carcinogen in humans, but the underlying tumourigenic mechanisms remain unclear. In Xuanwei city of Yunnan Province, the lung cancer incidence is among the highest in China, owing to severe air pollution generated by the combustion of smoky coal, providing a unique opportunity to dissect lung carcinogenesis. To identify abnormal miRNAs critical for air pollution-related tumourigenesis, we performed microRNA microarray analysis in 6 Xuanwei non-small cell lung cancers (NSCLCs) and 4 NSCLCs from control regions where smoky coal was not used. We found 13 down-regulated and 2 up-regulated miRNAs in Xuanwei NSCLCs. Among them, miR-144 was one of the most significantly down-regulated miRNAs. The expanded experiments showed that miR-144 was down-regulated in 45/51 (88.2%) Xuanwei NSCLCs and 34/54 (63%) control region NSCLCs (p = 0.016). MiR-144 interacted with the oncogene Zeb1 at 2 sites in its 3′ untranslated region and a decrease in miR-144 resulted in increased Zeb1 expression and an epithelial mesenchymal transition phenotype. Ectopic expression of miR-144 suppressed NSCLCs in vitro and in vivo by targeting Zeb1. These results indicate that down-regulation of miR-144 is critical for air pollution-related lung cancer and the miR-144-Zeb1 signalling pathway could represent a potential therapeutic target.
Similar content being viewed by others
Introduction
Lung cancer is the most common cause of death from cancer worldwide and is estimated to have been responsible for nearly one in five deaths (1.59 million deaths, 19.4% of the total) in 20121. Cigarette smoke is a major cause of lung cancer2 and indoor3 and outdoor4 air pollution have been classified as group 1 carcinogens in humans by the International Agency for Research on Cancer (IARC) of the World Health Organization. Anthropogenic particulate matter (PM) smaller than 2.5 μm in diameter (PM2.5) is associated with 220,000 lung cancer mortalities annually5 and is estimated to have cause 3.7 million premature deaths worldwide in 20126. However, the carcinogenic mechanism of air pollution remains unclear.
In Xuanwei City of the Yunnan Province, the lung cancer mortality rate is among the highest in China, owing to severe air pollution produced by the combustion of smoky coal7,8,9. Residents in this city used smoky coal in unvented indoor firepits for domestic cooking and heating until the 1970s. These processes released high concentrations of PM10 and PM2.5, which contained high concentrations of polycyclic aromatic hydrocarbons (PAHs) including benzo(a)pyrene (BaP) and polar compounds that are highly mutagenic9. A reduction in lung cancer morbidity was noted in the 1990s after stove improvements were introduced in central Xuanwei, supporting the association between air pollution and lung cancer10. The IARC monograph cited the findings in Xuanwei and classified indoor emissions from household combustion of coal as “carcinogenic to humans (Group 1)”3. The population in these highly polluted regions (HPR) lends a unique opportunity to dissect carcinogenesis that is specifically related to air pollution. We utilized this opportunity by systematically analyzing the abnormalities in the cancer genomes, genome-wide DNA methylation, non-coding RNAs (miRNAs and lncRNAs) and inflammatory factors, in patients from this region. Alterations found in HPR lung cancers were tested in patients from control regions (CR) where smoky coal was not used to compare the difference in the carcinogenic mechanism between HPR and CR lung cancer. Here, we report our results in the assessment of the expression of miRNAs in HPR non-small cell lung cancers (NSCLCs).
Results
Identification of aberrant miRNAs in Xuanwei NSCLCs
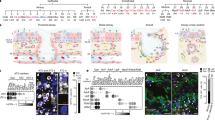
Tumour tissues and adjacent normal lung tissues were obtained with informed consent from 105 patients (51 from HPR and 54 from CR) with previously untreated lung adenocarcinoma (AD) or squamous cell carcinoma (SCC) (Table 1). Firstly, we performed miRNA microarray analysis for ten NSCLCs (6 from HPR and 4 from CR; Fig. 1A) and compared the expression of the miRNAs in tumour samples to that in their paired normal lung tissues to identify abnormal miRNA expression patterns. In total, we found 23 down-regulated and 7 up-regulated miRNAs in the 10 patients (Fig. 1A and Table 2). MiR-3195, miR-3656 and miR-144-3p (hereafter, miR-144) were the 3 most significantly down-regulated miRNAs (Fig. 1 and Table 2). We compared miRNA expression profiles between HPR and CR patients and found 21 miRNAs that were differentially expressed in HPR patients compared to CR cases. These included 13 down-regulated and 2 up-regulated miRNAs that were seen only in HPR and 5 down-regulated and 1 up-regulated miRNAs that were detected only in CR patients. Three miRNAs were down-regulated in NSCLCs from both regions (Fig. 1B).
Identification of down-regulated miR-144 in HPR and CR NSCLCs.
(A) MiRNA expression profiles of tumour and adjacent normal lung tissues from 10 NSCLCs. Left, data were organized according to the expression levels of individual miRNAs. Right, heat map showing 7 representative miRNAs down-regulated in the tumour samples. (B) Differentially expressed miRNAs in HPR compared to CR NSCLCs. (C) The expression of miR-3195, −3656, −1915-3p and −451a in tumour samples and their paired normal lung tissues. The numbers of patients tested are shown in parentheses. (D, E) The expression of miR-144 in tumour samples and their counterpart normal lung tissues (D). MiR-144 expression was frequently decreased in cancer tissues (E). A log2 fold change greater than +2 or less than −2 was considered to be a significant up-regulation or down-regulation, respectively. (F) The expression of miR-144 in tumour samples and paired normal lung tissues from HPR and CR NSCLCs.
To verify the results of the miRNA microarrays, we tested the expression of miR-3195, miR-3656, miR-144, miR-1915 and miR-451a in additional NSCLCs from HPR and CR. We found that the expression of miR-3195 in 18 NSCLC tumor samples did not differ from their paired normal lung tissues (Fig. 1C). The expression of miR-3656, miR-1915 and miR-451a was tested in 44 to 55 NSCLCs and the results showed that these miRNAs were down-regulated in tumour samples compared to paired normal lung tissues (Fig. 1C). Notably, the expression of miR-144 was decreased by at least 4-fold in 68/105 (64.8%) NSCLCs compared to matched adjacent normal lung samples (Fig. 1D, E).
Down-regulation of miR-144 is frequently seen in NSCLCs from HPR
We found that the expression of miR-144 was much lower in tumour samples than paired adjacent normal lung tissues in 45/51 (88.2%) and 34/54 (63%) NSCLCs from HPR and CR (Fig. 1F), respectively. The down-regulation of miR-144 in tumor samples was more frequently seen in HPR than CR NSCLCs (p = 0.016), indicating the association between air pollution and low miR-144 expression (Table 1).
To test whether air pollutants could down-regulate miR-144, several PAH compounds including benzo(a)pyrene (BaP), benzo(a)pyrene diol epoxide (BPDE), dibenzo[a, h]anthracene (DBA) and benzo[g, h, i]perylene (BzP) as well as the tobacco-specific carcinogen nicotine-derived nitrosamine ketone (NNK) were used to treat normal human bronchial epithelial 16HBE cells11 for 2 (Fig. S1A) or 15 (Fig. S1B) days. We showed that treatment of the cells with these compounds at concentrations of 0.25 and 0.5 μM for up to 15 days did not perturb the expression of miR-144. Therefore, isolation of other carcinogens and the assessment of the effects of these compounds on the expression of miR-144 warrant further investigation.
Effects of miR-144 on NSCLC cell migrations
We analysed the expression of miR-144 in human lung cancer cell lines by quantitative RT-PCR and found that the expression of this miRNA was low in the lung cancer cell lines 95D, H1975, EPLC and A549, whereas 16HBE cells had a relatively high expression level of miR-144 (Fig. 2A). To determine whether miR-144 was a putative tumour suppressor, an miR-144 mimic was transfected into A549 and 95D cells by using Lipofectamine or into A549-luciferase cells by using lentivirus-mediated cell transfection (Fig. 2B) and cell migration assays were performed. By transwell and wound healing assays, we found that miR-144 dramatically suppressed the migration abilities of the cells (Fig. 2C, D). In contrast, suppression of miR-144 by using an miRNA inhibitor enhanced cell migration in both of the lung cancer cell lines (Fig. 2E, F).
MiR-144 suppresses cell migration in vitro.
(A) miR-144 expression is reduced in lung cancer cell lines compared to the immortalized normal human bronchial epithelial cell line 16HBE. (B) MiR-144 was increased in A549 and 95D cells after transfection of a miR-144 mimic. MiR-144 expression was determined by qRT-PCR 48 h after transfection and was calculated as the fold change relative to the miR-control. U6 served as an internal control. (C) MiR-144 overexpression decreased the number of migrated cells in the transwell assay. Cells were seeded into transwell chambers. After 24 h, migrated cells stained with 0.1% crystal violet were counted. Data represent the means ± SD from three independent experiments. (D) Inhibition of cell migration by the miR-144 mimic was confirmed by the wounding healing assay. Data represent the means ± SD from three independent experiments. (E) Inhibition of miR-144 by anti-miR-144 enhanced cell migration in the Transwell assay. *, p<0.05. (F) Anti-miR-144 treatment enhanced cell migration in the wounding healing assay.
MiR-144 directly targets Zeb1 by interacting with its 3′-untranslated regions (UTRs)
Next, we investigated the targets of miR-144. Based the bioinformatics analysis using TargetScan (http://www.targetscan.org/), Miranda (http://www.microrna.org/microrna/home.do) and microcosm (http://www.ebi.ac.uk/enright-srv/microcosm/htdocs/targets/v5/), four genes (Zeb1, Zeb2, Rgs17 and Rock1) were predicted to be targeted by miR-144. Zeb1 and Rock1 had 2 potential interacting sites, while Zeb2 and Rgs1 had 1 potential interacting site with miR-144 (Fig. 3A). Then, we performed firefly luciferase reporter assays using the 3′-UTRs of Zeb1 (Zeb1-3UTR), Zeb2 (Zeb2-3UTR), Rgs17 (Rgs17-3UTR) and Rock1 (Rock1-3UTR) and found that the relative luciferase activity of the reporter that contained the Zeb1-3UTR was decreased by 47.0% in HEK293T cells and 47.5% in A549 cells after co-transfection of the miR-144 mimic (Fig. 3B). Conversely, co-transfection of the miR-144 mimic did not perturb Zeb2, Rgs17 or Rock1 reporter activity (Fig. 3B). Furthermore, mutations in potential interaction sites 1 and 2 of Zeb1 (Fig. 3C) abrogated miR-144-induced suppression of Zeb1-luciferase in the 293T (Fig. 3D) and A549 cells (Fig. 3E).
MiR-144 directly targets Zeb1.
(A) Zeb1, Zeb2, Rock1 and Rgs17 are targets of miR-144. Predicted duplex formation between Zeb1-3′-UTR, Zeb2-3′-UTR, Rock1-3′UTR, Rgs17-3′UTR and miR-144. (B) Luciferase assays in 293T and A549 cells transfected with a luciferase reporter controlled by the indicated elements. (C) Sequences of wild-type and mutant target sites for miR-144 in the Zeb1-3′ UTR. Two point mutations (underlined) predicted to abolish miRNA-mRNA binding were introduced into the miR-144 recognition region. (D, E) Luciferase reporter assays were performed in 293T (D) and A549 (E) cells co-transfected with the miR-control or miR-144 mimic (miR-144) together with the luciferase gene driven by the wild-type or mutant Zeb1-3′UTR sequences. The normalized luciferase activity in the control group was set as the relative luciferase activity. Each bar represents the mean ± S.D. for triplicate experiments. P values were determined by Student’s t-test. *p < 0.05; **p < 0.01. (F, G) The cells were transfected with the miR-control or miR-144 mimic (F) or anti-miR-144 (G) at a final concentration of 100 nM. The expression of the indicated proteins was determined by immunoblotting. (H) The expression of Zeb1 in tumour samples and their paired normal lung tissues from the 105 NSCLCs. (I) The correlation between Zeb1 and miR-144 expression was evaluated by Pearson correlation analysis. R and p-values are shown.
To further determine whether miR-144 affected endogenous Zeb1 expression, A549 and 95D cells were transfected with a miR-control or miR-144 mimic. In cells transfected with the miR-144 mimic, Zeb1 expression was down-regulated (Fig. 3F) at the protein level, whereas its target E-cadherin12 was up-regulated and Vimentin was down-regulated (Fig. 3F). In contrast, transfection of anti-miR-144 into the cells resulted in up-regulation of the Zeb1 protein (Fig. 3G), suggesting that Zeb1 could be a target gene of miR-144. However, the expression of Zeb2 was not affected by miR-144 overexpression or silencing (Fig. 3B, F, G). Moreover, the transcription factors Snail and Slug which play important roles in the epithelial mesenchymal transition (EMT) through transcriptional activation of Zeb113,14, were not affected by miR-144 overexpression or silencing (Fig. 3F, G), confirming the results that Zeb1 has no effect on the expression of Snail and Slug15.
The expression of Zeb1 in HPR NSCLCs
We tested the expression of Zeb1 in the 105 NSCLCs and found that Zeb1 expression in tumour samples was much higher than in paired normal lung tissues (Fig. 3H). The expression of Zeb2 was also elevated in the patients’ tumour samples (Fig. S2), which was consistent with previous reports16. Using Pearson correlation analysis, we evaluated the potential correlation between miR-144 and Zeb1 expression and found that miR-144 was inversely associated with Zeb1 expression (Fig. 3I).
Zeb1 inhibition recapitulates the tumour suppressing effect of miR-144
To evaluate whether down-regulation of Zeb1 was involved in the miR-144-induced suppression of cell migration, A549 and 95D cells were transfected with a Zeb1-specific siRNA (siZeb1) that caused down-regulation of Zeb1 at the mRNA (Fig. 4A) and protein (Fig. 4B) levels. Silencing of Zeb1 led to up-regulation of E-Cadherin and down-regulation of Vimentin (Fig. 4B). Furthermore, we found that silencing of Zeb1 significantly suppressed the cell migration of A549 and 95D cells in the transwell (Fig. 4C) and wound-healing (Fig. 4D) assays. Whereas transfection of miR-144 suppressed the migration of cancer cells, co-transfection of Zeb1 attenuated this effect (Fig. 4E). Consistent with this, transfection of miR-144 into the cells caused down-regulation of Zeb1 and Vimentin and up-regulation of E-Cadherin, whereas ectopic expression of Zeb1 partially reversed these effects (Fig. 4F).
Inhibition of Zeb1 mediates the tumour suppressor functions of miR-144.
(A, B) Endogenous Zeb1 expression was efficiently repressed by siRNA. A549 and 95D cells were transfected with non-specific siRNA control (sicontrol) or Zeb1-specific siRNA (siZeb1) for 48 h. The expression of Zeb1 were measured by qRT-PCR (A) and Western blotting (B). Inhibition of Zeb1 expression led to up-regulation of E-cadherin and down-regulation of Vimentin (B). (C) Inhibition of Zeb1 expression reduced the number of migrated cells in the Transwell assay. (D) Inhibition of cell migration by siZeb1 was confirmed by the wounding healing assay. Data represent the means ± SD from three independent experiments. (E) Ectopic expression of Zeb1 rescued the inhibition of cell migration associated with the re-expression of miR-144 in A549, 95D and A549-luciferase-miR-144 cells. Transwell assays were performed after the co-transfection of the miR-144 mimic and the plasmid containing Zeb1. (F) Zeb1 re-expression abrogated miR-144-induced up-regulation of E-cadherin and down-regulation of Vimentin. (G–J) Six-week-old SCID Beige female mice were administered with 1 × 106 A549-luciferase cells stably expressing miR-144 or the miR-control via the tail veins. (G) One month later, tumours in the lungs were surveyed with the IVIS Spectrum Imaging System. (H) Body weights of the mice were measured. (I) Immunohistochemistry (IHC) staining of Zeb1 in lung tissues. (J) The expression of Zeb1, E-cadherin and Vimentin was detected by Western blotting.
MiR-144 inhibits cancer progression in vivo
To test the in vivo tumour suppressing effect of miR-144, either the miR-control or miR-144 was stably transfected into A549-luciferase cells, which were then injected into the tail veins of SCID Beige mice. We showed that, compared to mice injected with miR-control-transfected cells, mice injected with miR-144-A549-luciferase cells showed a three-fold reduction in lung tumour volume detected by the IVIS Spectrum Imaging System (Fig. 4G). The body weights of the mice harboring miR-control-A549-luciferase cells decreased rapidly, whereas the body weights of mice bearing miR-144-A549-luciferase cells decreased gradually (Fig. 4H), reflecting the tumour burden and progression. The immunohistochemistry (Fig. 4I) and Western blot (Fig. 4J) assays showed that both Zeb1 and Vimentin were down-regulated and E-Cadherin was up-regulated in tumours harvested from mice inoculated with the miR-144-A549-luciferase cells.
Discussion
MiRNAs are small non-coding RNAs that post-transcriptionally mediate the expression of their target genes by perfect or imperfect binding to the 3′-UTR17, the 5′-UTR18, or the open reading frame of the target mRNAs19, thereby causing mRNA degradation or inhibition or enhancement of mRNA translation. It has been estimated that miRNAs regulate more than 60% of human genes20. MiRNAs participate in diverse biological processes, including cell proliferation, apoptosis, migration and invasion21. Some miRNAs have been shown to be involved in the tumourigenesis of lung cancer and miRNA expression profiles are diagnostic and prognostic markers of lung cancer22. For instance, microRNA-135b promotes lung cancer metastasis by targeting tumour suppressor LZTS1 and multiple key components in the Hippo pathway such as LATS2, β-TrCP and NDR223. MicroRNA-193a-3p and −5p function as tumour suppressors and inhibit the metastasis of lung cancer by down-regulating the ERBB4/PIK3R3/mTOR/S6K2 signalling pathway24. MiR-2125, miR-17-9226 and miR-196a27 are up-regulated and function as oncogenes, whereas let-728, miR-19429 and miR-12630 suppress carcinogenesis. However, abnormalities in miRNAs in air pollution-related lung cancer have not been investigated.
PAHs, the main carcinogens in HPR9, are ubiquitous environmental pollutants that are generated primarily through incomplete combustion of carbon-containing materials. In cellular or animal models, PAHs down-regulate miR-34c31, miR-21, miR-221, miR-222 and miR-42932 and up-regulate miR-34a33,34, miR-181a, miR-181b and miR-181d35. Deng et al.36 performed a study in healthy male coke oven workers to identify miRNAs associated with PAH exposure and found that urinary 4-hydroxyphenanthrene and/or plasma BPDE–Alb adducts were associated with lower miR-24-3p, miR-27a-3p, miR-142-5p and miR-28-5p expression. Urinary 1-hydroxynaphthalene, 2-hydroxynaphthalene, 2-hydroxyphenanthrene and the sum of monohydroxy-PAHs were associated with higher miR-150-5p expression. Here, we performed miRNA microarrays to screen for abnormal miRNAs in air pollution-related lung cancer, using samples from a unique HPR and found that miR-144 and its family member miR-45137 were significantly down-regulated in the patients. We expanded this observation and reported that miR-144 was down-regulated in 45/51 (88.2%) HPR NSCLCs, whereas in CR patients miR-144 was suppressed in 34/54 (63%) tumour samples (p = 0.016). Our results provided the first evidence of association between a miRNA and air pollution-related lung cancer. However, treatment of 16HBE cells with the carcinogens BaP, BPDE, BzP, DBA and NNK for up to 15 days did not result in the down-regulation of miR-144, suggesting that other carcinogens may be responsible for the down-regulation of this miRNA; therefore, investigations are warranted to uncover the carcinogen(s).
MiR-144 can modulate TRAIL-induced apoptosis by targeting caspase-338. Down-regulation of miR-144 is associated with colorectal cancer progression via activation of the mTOR signalling pathway39. MiR-144 inhibits NSCLC cell growth and induces apoptosis by down-regulating ZFX40 and re-establishing miR-144 in gastric cancer restores the chemosensitivity41. We showed that Zeb1, a transcriptional repressor42 that inhibits E-cadherin and promotes EMT and metastasis12,43, is a target of miR-144. MiR-144 interacts with Zeb1 via two sites in Zeb1’s 3′-UTR and mutations in these sites abrogate miR-144’s repression of Zeb1 functions (Fig. 3), a result in agreement with previous reports44,45. In lung cancer, the expression of Zeb1 is up-regulated46, whereas knockdown of Zeb1 results in dramatic growth inhibition47. MiR-144 could also bind Zeb2 at positions 1003-1023 in its 3′-UTR (Fig. 3A), but in NSCLC cells overexpression or knockdown of miR-144 had no effect on the expression of Zeb2 at the mRNA and protein levels (Fig. 3B, F, G). In NSCLCs, miR-200b and miR-200c, which are able to bind Zeb2 at positions 1015-1036 of its 3′-UTR, were up-regulated48,49 and may therefore antagonize the effects of miR-144 on Zeb2. Hence, our results indicated that miR-144 is an important tumour suppressor that is inactivated during malignant transformation and the miR-144-Zeb1 signal pathway could represent a rational therapeutic target.
Methods
Patients and tissue samples
Use of the samples was approved by the Institutional Review Board of the Institute of Zoology, Chinese Academy of Sciences and the local research ethics committees of all participating hospitals. The methods were performed in accordance with the approved guidelines. Tumour tissues and adjacent normal lung tissues were obtained with informed consent from 105 patients with previously untreated lung adenocarcinoma (AD) or squamous cell carcinoma (SCC) at local hospitals. The diagnosis of lung cancer was confirmed by at least 3 pathologists and the HPR patients enrolled met the following criteria: (1) patients were residents of Xuanwei where the smoky coal was used; (2) patients resided in their communities and never stayed in other regions for a long period of time (6 months or more); (3) patients had previously untreated primary lung cancer; and (4) patients’ tissue samples were taken at the time of surgery and quickly frozen in liquid nitrogen. The tumour samples contained a tumour cellularity of greater than 60% and the matched control samples had no tumour content. The clinical and pathological data for these patients are shown in Table 1 and Fig. 1A.
Cell culture
The NSCLC cell lines A549 and 95D, human bronchial epithelial cell line 16HBE and a human embryonic kidney cell line HEK293T were cultured in DMEM (Hyclone, Logan, UT, USA) medium supplemented with 10% foetal bovine serum (Hyclone). The cells were treated with PAHs (BaP, BPDE, DBA and BzP) or the tobacco specific carcinogen NNK. Cell viability was estimated by trypan blue dye exclusion analysis and cell proliferation was measured by the MTT assay50.
MiRNA microarray and quantitative RT-PCR
Total RNA was isolated using TRIzol (Invitrogen, Frederick, MD, USA) and the miRNeasy mini kit (Qiagen, Hilden, Germany), quantitated by using the NanoDrop 1000 (Thermo Scientific, Wilmington, DE, USA), labelled by using the miRCURY™ Hy3™/Hy5™ Power labelling kit (Exiqon, Vedbaek, Denmark) and hybridized onto the miRCURY™ LNA Array (v.16.0; Exiqon). Following the washing steps, the slides were scanned using the Axon GenePix 4000B microarray scanner (Axon Instruments, Foster City, CA, USA) and images were imported into the GenePix Pro 6.0 software (Axon Instruments) for grid alignment and data extraction. All expressed data were normalized using the Median normalization method51 and significantly differentially expressed miRNAs were identified through Volcano Plot filtering. Quantitative RT-PCR analysis for miR-3195, −3656, −144, −451a and −1915-3p was performed with a miScript SYBR Green PCR Kit (Qiagen). The expression of related genes was measured by qPCR with SYBR® Green Real time PCR Master Mix (Takara Biotechnology, Dalian, China). The fold changes in mRNA expression were calculated using the 2−ΔΔCt method52. The primers used for quantitative RT-PCR are as follows: Zeb1, forward, 5′-ATGCACAACCAAGTGCAGAAGA-3′ and reverse, 5′-TTGCCTGGTTCAGGAGAAGATG-3′; Zeb2, forward, 5′- CAAGAGGCGCAAACAAGC-3′ and reverse, 5′- GGTTGGCAATACCGTCAT-3′; E-cadherin, forward, 5′-TGCCCAGAAAATGAAAAAGG-3′ and reverse, 5′-GTGTATGTGGCAATGCGTTC-3′; Vimentin, forward, 5′-GAGAACTTTGCCGTTGAAGC-3′ and reverse, 5′-GCTTCCTGTAGGTGGCAATC-3′; and GAPDH, forward, 5′-GAGTCAACGGATTTGGTCGT-3′ and reverse, 5′-GACAAGCTTCCCGTTCTCAG-3′.
Wound-healing assay and in vitro migration assay
For the wound-healing assay, the cells (4 × 105/wells) were seeded into six-well plates and transfected the next day with a miR-144 mimic or a miR-control at a final concentration of 100 nM. Twenty-four hours after transfection, wounds were created in the cell monolayer using a p200 micropipette tip. The healing process was followed for the next 48 hours. For the transwell assay, transwell inserts (6.5 mm diameter and 8 μm pore size; Corning Inc., Corning, NY, USA) were rehydrated by adding serum-free medium for at least 1 hour. The cells transfected with the miR-144 mimic in serum-free medium were seeded (2 × 104 cells) into the inserts (the upper chamber). Complete medium was used as a chemoattractant in the bottom chamber. After 24 hours of migration, the cells in the upper surface of the insert membrane were removed by wiping with a cotton swab and cells in the lower surface were fixed with methanol, stained with crystal violet and counted by microscopy.
Luciferase report assays
The miR-144 binding site-containing 3′ UTR fragment of Zeb1 was amplified and cloned into the modified pGL3-luciferase vector. Mutations in the Zeb1 3′-UTR were generated by using the QuikChange Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA). The miR-144 mimic (5′-UACAGUAUAGAUGAUGUACU-3′), miR-control (5′-UUCUCCGAACGUGUCACGUTT-3′), anti-miR-144 (5′-AGUACAUCAUCUAUACUGUA), anti-miR-control (CAGUACUUUUGUGUAGUACAA), siZeb1 (5′-UGAUCAGCCUCAAUCUGCATT-3′) and si-control (UUCUCCGAACGUGUCACGUTT) were synthesized by GenePharma (Shanghai, China). The cells were transfected with 0.5 μg of pGL3-luciferase vector and 50 nM of the miR-144 mimic or miR-control together with a Renilla plasmid using Lipofectamine 2000 (Invitrogen) according to the manufacturer′s instructions. Luciferase activities were measured using the Dual luciferase reporter assay system (Promega, Madison, WI, USA).
Lentivirus-mediated cell transfection and transduction
Pre-miR-144 DNA sequences were amplified from human genomic DNA, subcloned into the EcoR I and BamH I sites downstream of the CMV promoter in the pCDH vector (kindly provided by Dr. Wanzhu Jin at the Institute of Zoology, Chinese Academy of Sciences) and verified by DNA sequencing. The primers used for the genomic PCR amplification of miR-144 were as follows: 5′-GCGCGAATTCGAGATCTTAACAGACCCTAGCTC-3′ (forward primer) and 5′-GCGCGGATCCGTGCCCTGGCAGTCAGTAGG-3′ (reverse primer). Infectious virus particles were harvested 48 hours after co-transfection of pCDH-miR-144 or pCDH with the lentivirus packing vector (psPAX2 and pMD2G) into HEK293FT cells. A549 cells were infected with lentiviruses in medium containing polybrene (8 mg/ml). One week after infection, the cells were sorted by flow cytometry.
Western Blotting
Cells were lysed in RIPA buffer supplemented with a protease inhibitors cocktail (Sigma, St. Louis, MO, USA). Proteins (20 μg) were subjected to 10–15% SDS-PAGE, electrophoresed and transferred on to a nitrocellulose membrane. After blocking with 5% non-fat milk in Tris-buffered saline, the membrane was washed and incubated with the indicated primary and secondary antibodies and detected using the Luminescent Image Analyser LSA 4000 (GE, Fairfield, CO, USA).
Animal studies
The animal studies were approved by the Institutional Review Board of the Institute of Zoology, Chinese Academy of Sciences. The methods were performed in accordance with the approved guidelines. Six-week-old SCID Beige mice were maintained under specific pathogen-free (SPF) conditions. A549-luciferase cells (1 × 106) stably expressing miR-144 or the miR-control were injected into the lateral tail veins of the mice (6 per group). After 30 days, the tumours were monitored with the IVIS Spectrum Imaging System (Caliper Life Sciences; Hopkinton, MA, USA). For the immunohistochemistry (IHC) assay, sections were fixed in formalin and embedded in paraffin, incubated with primary antibodies overnight and then incubated with anti-rabbit IgG secondary antibodies. Detection was conducted using 3, 3′-diaminobenzidine (DAB, Zhongshan Golden Bridge Biotechnology Co., Ltd, Beijing, China) and haematoxylin.
Statistical analysis
Experimental data were presented as the mean ± SD of three independent experiments. The differences between groups were estimated using an independent two-tailed Student’s t-test (normal distribution data) or Wilcoxon rank sum test (non-normal distribution data) and the association between miR-144 expression and the Zeb1 level was analysed by Pearson correlation analysis. P values less than 0.05 were considered statistically significant in all cases.
Additional Information
How to cite this article: Pan, H.-L. et al. Down–regulation of microRNA-144 in air pollution-related lung cancer. Sci. Rep. 5, 14331; doi: 10.1038/srep14331 (2015).
References
World Health Organization GLOBOCAN 2012. GLOBOCAN. Available at: http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx. Day of access: 24-12/2014 (2013).
Hecht, S. S. Lung carcinogenesis by tobacco smoke. Int J Cancer 131, 2724–2732 (2012).
World Health Organization International Agency for Research on Cancer Household Use of Solid Fuels and High-temperature Frying. IARC Monographs on the Evaluation of Carcinogenic Risks to HumansVolume 95, Available at: http://monographs.iarc.fr/ENG/Monographs/vol95/.-Day of access: 24/12/2014 (2010).
Loomis, D. et al. The carcinogenicity of outdoor air pollution. Lancet Oncol 14, 1262–1263 (2013).
Anenberg, S. C., Horowitz, L. W., Tong, D. Q. & West, J. J. An estimate of the global burden of anthropogenic ozone and fine particulate matter on premature human mortality using atmospheric modeling. Environ. Health Perspect 118, 1189–1195 (2010).
World Health Organization Ambient (outdoor) air quality and health. Available at: http://www.who.int/mediacentre/factsheets/fs313/en/. Day of access: 03/12/2014 (2014).
Xiao, Y., Shao, Y., Yu, X. & Zhou, G. The epidemic status and risk factors of lung cancer in Xuanwei City, Yunnan Province, China. Front Med 6, 388–394 (2012).
Cao, Y. & Gao, H. Prevalence and causes of air pollution and lung cancer in Xuanwei City and Fuyuan County, Yunnan Province, China. Front Med 6, 217–220 (2012).
Mumford, J. L. et al. Lung cancer and indoor air pollution in Xuan Wei, China. Science 235, 217–220 (1987).
Lan, Q., Chapman, R. S., Schreinemachers, D. M., Tian, L. & He, X. Household Stove Improvement and Risk of Lung Cancer in Xuanwei, China. J Natl Cancer Inst 94, 826–835 (2002).
Cozens, A. L. et al. CFTR expression and chloride secretion in polarized immortal human bronchial epithelial cells. Am J Respir Cell Mol Biol 10, 38–47 (1994).
Sanchez-Tillo, E. et al. ZEB1 represses E-cadherin and induces an EMT by recruiting the SWI/SNF chromatin-remodeling protein BRG1. Oncogene 29, 3490–3500 (2010).
Guaita, S. et al. Snail induction of epithelial to mesenchymal transition in tumor cells is accompanied by MUC1 repression and ZEB1 expression. J Biol Chem 277, 39209–39216 (2002).
Wels, C., Joshi, S., Koefinger, P., Bergler, H. & Schaider, H. Transcriptional activation of ZEB1 by Slug leads to cooperative regulation of the epithelial-mesenchymal transition-like phenotype in melanoma. J Invest Dermatol 131, 1877–1885 (2011).
Gemmill, R. M. et al. ZEB1-responsive genes in non-small cell lung cancer. Cancer Lett 300, 66–78 (2011).
Miura, N. et al. Clinicopathological significance of Sip1-associated epithelial mesenchymal transition in non-small cell lung cancer progression. Anticancer Res 29, 4099–4106 (2009).
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Orom, U. A., Nielsen, F. C. & Lund, A. H. MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell 30, 460–471 (2008).
Tay, Y., Zhang, J., Thomson, A. M., Lim, B. & Rigoutsos, I. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455, 1124–1128 (2008).
Friedman, R. C., Farh, K. K.-H., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Research 19, 92–105 (2009).
Garzon, R., Fabbri, M., Cimmino, A., Calin, G. A. & Croce, C. M. MicroRNA expression and function in cancer. Trends in Mol Med 12, 580–587 (2006).
Yanaihara, N. et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9, 189–198 (2006).
Lin, C. W. et al. MicroRNA-135b promotes lung cancer metastasis by regulating multiple targets in the Hippo pathway and LZTS1. Nat Commun 4, 1877 10.1038/ncomms2876 (2013).
Yu, T. et al. MicroRNA-193a-3p and -5p suppress the metastasis of human non-small-cell lung cancer by downregulating the ERBB4/PIK3R3/mTOR/S6K2 signaling pathway. Oncogene 34, 413–423 (2014).
Shen, H. et al. Alteration in Mir-21/PTEN Expression Modulates Gefitinib Resistance in Non-Small Cell Lung Cancer. PLoS ONE 9, e103305 10.1371/journal.pone.0103305 (2014).
Hayashita, Y. et al. A Polycistronic MicroRNA Cluster, miR-17-92, Is Overexpressed in Human Lung Cancers and Enhances Cell Proliferation. Cancer Res 65, 9628–9632 (2005).
Liu, X. H. et al. MicroRNA-196a promotes non-small cell lung cancer cell proliferation and invasion through targeting HOXA5. BMC Cancer 12, 348 10.1186/1471-2407-12-348 (2012).
Johnson, S. M. et al. RAS Is Regulated by the let-7 MicroRNA Family. Cell 120, 635–647 (2005).
Wu, X. et al. miR-194 suppresses metastasis of non-small cell lung cancer through regulating expression of BMP1 and p27kip1. Oncogene 33, 1506–1514 (2014).
Liu, B., Peng, X. C., Zheng, X. L., Wang, J. & Qin, Y. W. MiR-126 restoration down-regulate VEGF and inhibit the growth of lung cancer cell lines in vitro and in vivo. Lung Cancer 66, 169–175 (2009).
Han, Z. et al. Cell cycle changes mediated by the p53/miR-34c axis are involved in the malignant transformation of human bronchial epithelial cells by benzo[a]pyrene. Toxicology Letters 225, 275–284 (2014).
Chanyshev, M. D., Kosorotikov, N. I., Titov, S. E., Kolesnikov, N. N. & Gulyaeva, L. F. Expression of microRNAs, CYP1A1 and CYP2B1 in the livers and ovaries of female rats treated with DDT and PAHs. Life Sciences 103, 95–100 (2014).
Malik, A. I., Williams, A., Lemieux, C. L., White, P. A. & Yauk, C. L. Hepatic mRNA, microRNA and miR-34a-Target responses in mice after 28 days exposure to doses of benzo(a)pyrene that elicit DNA damage and mutation. Environ Mol Mutagen 53, 10–21 (2012).
Malik, A. I. et al. Hepatic genotoxicity and toxicogenomic responses in MutaMouse males treated with dibenz[a,h]anthracene. Mutagenesis 28, 543–554 (2013).
Song, M. K., Park, Y. K. & Ryu, J. C. Polycyclic aromatic hydrocarbon (PAH)-mediated upregulation of hepatic microRNA-181 family promotes cancer cell migration by targeting MAPK phosphatase-5, regulating the activation of p38 MAPK. Toxicol Appl Pharm 273, 130–139 (2013).
Deng, Q. et al. Plasma microRNA expression and micronuclei frequency in workers exposed to polycyclic aromatic hydrocarbons. Environ Health Perspect 122, 719–725 (2014).
Pan, X., Wang, R. & Wang, Z. X. The Potential Role of miR-451 in Cancer Diagnosis, Prognosis and Therapy. Mol Cancer Ther 12, 1153–1162 (2013).
Ovcharenko, D., Kelnar, K., Johnson, C., Leng, N. & Brown, D. Genome-Scale MicroRNA and Small Interfering RNA Screens Identify Small RNA Modulators of TRAIL-Induced Apoptosis Pathway. Cancer Res 67, 10782–10788 (2007).
Iwaya, T. et al. Downregulation of miR-144 is associated with colorectal cancer progression via activation of mTOR signaling pathway. Carcinogenesis 33, 2391–2397 (2012).
Zha, W., Cao, L., Shen, Y. & Huang, M. Roles of Mir-144-ZFX Pathway in Growth Regulation of Non-Small-Cell Lung Cancer. PLoS ONE 8, e74175 10.1371/journal.pone.0074175 (2013).
Akiyoshi, S. et al. Clinical significance of miR-144-ZFX axis in disseminated tumour cells in bone marrow in gastric cancer cases. Br J Cancer 107, 1345–1353 (2012).
Postigo, A. A. & Dean, D. C. ZEB represses transcription through interaction with the corepressor CtBP. Proc Natl Aca Sci USA 96, 6683–6688 (1999).
Spaderna, S. et al. The Transcriptional Repressor ZEB1 Promotes Metastasis and Loss of Cell Polarity in Cancer. Cancer Res 68, 537–544 (2008).
Sureban, S. et al. Nanoparticle-based delivery of siDCAMKL-1 increases microRNA-144 and inhibits colorectal cancer tumor growth via a Notch-1 dependent mechanism. J Nanobiotechnology 9, 40 10.1186/1477-3155-9-40 (2011).
Guan, H. et al. Down-regulation of miR-144 promotes thyroid cancer cell invasion by targeting ZEB1 and ZEB2. Endocrine 48, 566–574 (2015).
Ohira, T. et al. WNT7a induces E-cadherin in lung cancer cells. Proc Natl Acad Sci USA 100, 10429–10434 (2003).
Takeyama, Y. et al. Knockdown of ZEB1, a master epithelial-to-mesenchymal transition (EMT) gene, suppresses anchorage-independent cell growth of lung cancer cells. Cancer Letters 296, 216–224 (2010).
Boeri, M. et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc Natl Acad. Sci USA 108, 3713–3718 (2011).
Liu, X. G. et al. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med Oncol 29, 618–626 (2012).
Carmichael, J. et al. Chemosensitivity testing of human lung cancer cell lines using the MTT assay. Br J Cancer 57, 540–547 (1988).
Liu, C. G. et al. An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Proc Natl Acad Sci USA 101, 9740–9744 (2004).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Acknowledgements
This work was supported by the National Natural Science Funds for Distinguished Young Scholar (81425025), National Key Programme for Basic Research (2012CB910800), National Natural Science Foundation of China (81171925, 81201537, U1202224 and 81460441) and grants from the State Key Laboratory of Medical Genomics. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
The project was conceived by G.B.Z. The experiments were designed by G.B.Z. The experiments were conducted by H.L.P., X.C., G.Z.W. and M.M.W. Patient samples were provided by Z.S.W., Y.C.H., Y.C.Z., Z.Y.W. and Y.Q.G. Data were analysed by G.B.Z. and Y.C. The manuscript was written by G.B.Z.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Pan, HL., Wen, ZS., Huang, YC. et al. Down-regulation of microRNA-144 in air pollution-related lung cancer. Sci Rep 5, 14331 (2015). https://doi.org/10.1038/srep14331
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14331
This article is cited by
-
MicroRNA-1915-3p inhibits cell migration and invasion by targeting SET in non-small-cell lung cancer
BMC Cancer (2021)
-
Differential expression of circulating serum miR-1249-3p, miR-3195, and miR-3692-3p in non-small cell lung cancer
Human Cell (2020)
-
Bituminous coal combustion and Xuan Wei Lung cancer: a review of the epidemiology, intervention, carcinogens, and carcinogenesis
Archives of Toxicology (2019)
-
MiR-144-3p promotes the tumor growth and metastasis of papillary thyroid carcinoma by targeting paired box gene 8
Cancer Cell International (2018)
-
Profiling of 179 miRNA Expression in Blood Plasma of Lung Cancer Patients and Cancer-Free Individuals
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.