Abstract
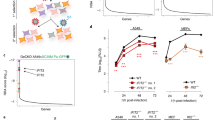
Activation of the transcription factor interferon regulatory factor-3 (IRF-3) is an essential event in the innate immune response to viral infection. To understand the contribution of IRF-3 to host defense, we used a systems biology approach to analyze global gene expression dependent on IRF-3. Comparison of expression profiles in cells from IRF-3 knockout animals or wild-type siblings following viral infection revealed three sets of induced genes, those that are strictly dependent on IRF-3, augmented with IRF-3, or not responsive to IRF-3. Products of identified IRF-3 target genes are involved in innate or acquired immunity, or in the regulation of cell cycle, apoptosis and proliferation. These results reveal the global effects of one transcription factor in the immune response and provide information to evaluate the integrated response to viral infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Honda K, Taniguchi T . IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat Rev 2006; 6: 644–658.
Kawai T, Akira S . Innate immune recognition of viral infection. Nat Immunol 2006; 7: 131–137.
Hiscott J . Triggering the innate antiviral response through IRF-3 activation. J Biol Chem 2007; 282: 15325–15329.
Weaver BK, Kumar KP, Reich NC . Interferon regulatory factor 3 and CREB-binding protein/p300 are subunits of double-stranded RNA-activated transcription factor DRAF1. Mol Cell Biol 1998; 18: 1359–1368.
Yoneyama M, Suhara W, Fukuhara Y, Fukuda M, Nishida E, Fujita T . Direct triggering of the type I interferon system by virus infection: activation of a transcription factor complex containing IRF-3 and CBP/p300. EMBO J 1998; 17: 1087–1095.
Wathelet MG, Lin CH, Parekh BS, Ronco LV, Howley PM, Maniatis T . Virus infection induces the assembly of coordinately activated transcription factors on the IFN-beta enhancer in vivo. Mol Cell 1998; 1: 507–518.
Sato M, Tanaka N, Hata N, Oda E, Taniguchi T . Involvement of the IRF family transcription factor IRF-3 in virus-induced activation of the IFN-beta gene. FEBS Lett 1998; 425: 112–116.
Lin R, Heylbroeck C, Pitha PM, Hiscott J . Virus-dependent phosphorylation of the IRF-3 transcription factor regulates nuclear translocation, transactivation potential, and proteasome-mediated degradation. Mol Cell Biol 1998; 18: 2986–2996.
Sato M, Suemori H, Hata N, Asagiri M, Ogasawara K, Nakao K et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-alpha/beta gene induction. Immunity 2000; 13: 539–548.
Honda K, Yanai H, Negishi H, Asagiri M, Sato M, Mizutani T et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 2005; 434: 772–777.
Honda K, Takaoka A, Taniguchi T . Type I interferon [corrected] gene induction by the interferon regulatory factor family of transcription factors. Immunity 2006; 25: 349–360.
Fitzgerald KA, McWhirter SM, Faia KL, Rowe DC, Latz E, Golenbock DT et al. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat Immunol 2003; 4: 491–496.
Sharma S, tenOever BR, Grandvaux N, Zhou GP, Lin R, Hiscott J . Triggering the interferon antiviral response through an IKK-related pathway. Science 2003; 300: 1148–1151.
Kumar KP, McBride KM, Weaver BK, Dingwall C, Reich NC . Regulated nuclear-cytoplasmic localization of interferon regulatory factor 3, a subunit of double-stranded RNA-activated factor 1. Mol Cell Biol 2000; 20: 4159–4168.
Daly C, Reich NC . Double-stranded RNA activates novel factors that bind to the interferon-stimulated response element. Mol Cell Biol 1993; 13: 3756–3764.
Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD . How cells respond to interferons? Annu Rev Biochem 1998; 67: 227–264.
Levy DE, Darnell Jr JE . Stats: transcriptional control and biological impact. Nat Rev Mol Cell Biol 2002; 3: 651–662.
Sen GC . Viruses and interferons. Annu Rev Microbiol 2001; 55: 255–281.
Stetson DB, Medzhitov R . Type I interferons in host defense. Immunity 2006; 25: 373–381.
Agalioti T, Lomvardas S, Parekh B, Yie J, Maniatis T, Thanos D . Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 2000; 103: 667–678.
Grandvaux N, Servant MJ, tenOever B, Sen GC, Balachandran S, Barber GN et al. Transcriptional profiling of interferon regulatory factor 3 target genes: direct involvement in the regulation of interferon-stimulated genes. J Virol 2002; 76: 5532–5539.
Elco CP, Guenther JM, Williams BR, Sen GC . Analysis of genes induced by Sendai virus infection of mutant cell lines reveals essential roles of interferon regulatory factor 3, NF-kappaB, and interferon but not toll-like receptor 3. J Virol 2005; 79: 3920–3929.
Daly C, Reich NC . Characterization of specific DNA-binding factors activated by double-stranded RNA as positive regulators of interferon alpha/beta-stimulated genes. J Biol Chem 1995; 270: 23739–23746.
Terenzi F, deVeer MJ, Ying H, Restifo NP, Williams BR, Silverman RH . The antiviral enzymes PKR and RNase L suppress gene expression from viral and non-viral based vectors. Nucleic Acids Res 1999; 27: 4369–4375.
Marie I, Durbin JE, Levy DE . Differential viral induction of distinct interferon-alpha genes by positive feedback through interferon regulatory factor-7. EMBO J 1998; 17: 6660–6669.
Moser B, Wolf M, Walz A, Loetscher P . Chemokines: multiple levels of leukocyte migration control. Trends Immunol 2004; 25: 75–84.
Arenzana-Seisdedos F, Virelizier JL, Rousset D, Clark-Lewis I, Loetscher P, Moser B et al. HIV blocked by chemokine antagonist. Nature 1996; 383: 400.
Starling GC, Whitney GS, Siadak AW, Llewellyn MB, Bowen MA, Farr AG et al. Characterization of mouse CD6 with novel monoclonal antibodies, which enhance the allogeneic mixed leukocyte reaction. Eur J Immunol 1996; 26: 738–746.
Chien YH, Jores R, Crowley MP . Recognition by gamma/delta T cells. Annu Rev Immunol 1996; 14: 511–532.
Kambayashi T, Kraft-Leavy JR, Dauner JG, Sullivan BA, Laur O, Jensen PE . The nonclassical MHC class I molecule Qa-1 forms unstable peptide complexes. J Immunol 2004; 172: 1661–1669.
Wang X, Johansen LM, Tae HJ, Taparowsky EJ . IFP 35 forms complexes with B-ATF, a member of the AP1 family of transcription factors. Biochem Biophys Res Commun 1996; 229: 316–322.
Weichenhan D, Kunze B, Zacker S, Traut W, Winking H . Structure and expression of the murine Sp100 nuclear dot gene. Genomics 1997; 43: 298–306.
Espert L, Degols G, Lin YL, Vincent T, Benkirane M, Mechti N . Interferon-induced exonuclease ISG20 exhibits an antiviral activity against human immunodeficiency virus type 1. J Gen Virol 2005; 86 (Part 8): 2221–2229.
Terenzi F, Pal S, Sen GC . Induction and mode of action of the viral stress-inducible murine proteins, P56 and P54. Virology 2005; 340: 116–124.
Chin KC, Cresswell P . Viperin (cig5), an IFN-inducible antiviral protein directly induced by human cytomegalovirus. Proc Natl Acad Sci USA 2001; 98: 15125–15130.
MacMicking JD, Taylor GA, McKinney JD . Immune control of tuberculosis by IFN-gamma-inducible LRG-47. Science 2003; 302: 654–659.
Zhu H, Cong JP, Shenk T . Use of differential display analysis to assess the effect of human cytomegalovirus infection on the accumulation of cellular RNAs: induction of interferon-responsive RNAs. Proc Natl Acad Sci USA 1997; 94: 13985–13990.
Spengler D, Villalba M, Hoffmann A, Pantaloni C, Houssami S, Bockaert J et al. Regulation of apoptosis and cell cycle arrest by Zac1, a novel zinc finger protein expressed in the pituitary gland and the brain. EMBO J 1997; 16: 2814–2825.
Unoki M, Nakamura Y . EGR2 induces apoptosis in various cancer cell lines by direct transactivation of BNIP3L and BAK. Oncogene 2003; 22: 2172–2185.
Calfon M, Zeng H, Urano F, Till JH, Hubbard SR, Harding HP et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 2002; 415: 92–96.
Trubia M, Sessa L, Taramelli R . Mammalian Rh/T2/S-glycoprotein ribonuclease family genes: cloning of a human member located in a region of chromosome 6 (6q27) frequently deleted in human malignancies. Genomics 1997; 42: 342–344.
Shiloh Y . ATM and related protein kinases: safeguarding genome integrity. Nat Rev Cancer 2003; 3: 155–168.
Monaco JJ, Nandi D . The genetics of proteasomes and antigen processing. Annu Rev Genet 1995; 29: 729–754.
Genin P, Algarte M, Roof P, Lin R, Hiscott J . Regulation of RANTES chemokine gene expression requires cooperativity between NF-kappa B and IFN-regulatory factor transcription factors. J Immunol 2000; 164: 5352–5361.
Bluyssen HA, Vlietstra RJ, Faber PW, Smit EM, Hagemeijer A, Trapman J . Structure, chromosome localization, and regulation of expression of the interferon-regulated mouse Ifi54/Ifi56 gene family. Genomics 1994; 24: 137–148.
Honda K, Yanai H, Takaoka A, Taniguchi T . Regulation of the type I IFN induction: a current view. Int Immunol 2005; 17: 1367–1378.
Lee AH, Hong JH, Seo YS . Tumour necrosis factor-alpha and interferon-gamma synergistically activate the RANTES promoter through nuclear factor kappaB and interferon regulatory factor 1 (IRF-1) transcription factors. Biochem J 2000; 350 (Part 1): 131–138.
Weaver BK, Ando O, Kumar KP, Reich NC . Apoptosis is promoted by the dsRNA-activated factor (DRAF1) during viral infection independent of the action of interferon or p53. FASEB J 2001; 15: 501–515.
Taniguchi T, Takaoka A . The interferon-alpha/beta system in antiviral responses: a multimodal machinery of gene regulation by the IRF family of transcription factors. Curr Opin Immunol 2002; 14: 111–116.
Lomvardas S, Thanos D . Modifying gene expression programs by altering core promoter chromatin architecture. Cell 2002; 110: 261–271.
Maniatis T, Falvo JV, Kim TH, Kim TK, Lin CH, Parekh BS et al. Structure and function of the interferon-beta enhanceosome. Cold Spring Harb Symp Quant Biol 1998; 63: 609–620.
Acknowledgements
We thank all the members of the laboratory for support and discussions. We gratefully acknowledge Dr Tadasugu Taniguchi (University of Tokyo) for providing us with cells from WT and IRF-3 KO animals. We also thank John Schwedes at the Affymetrix core facility for his role in processing and analyzing the microarrays, and Michelle Massoth for help with Genesifter Software (www.genesifter.net). We are grateful to Dr Herb Lewis and Robert Andersen for writing a Visual Basic code used for secondary comparisons. Human IFN-αA was a kind gift of Hoffman-LaRoche, NJ, USA). These studies were supported by grants from NIH (R21AI067885 and PO1AI0555621).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Genes and Immunity website (http://www.nature.com/gene)
Supplementary information
Rights and permissions
About this article
Cite this article
Andersen, J., VanScoy, S., Cheng, TF. et al. IRF-3-dependent and augmented target genes during viral infection. Genes Immun 9, 168–175 (2008). https://doi.org/10.1038/sj.gene.6364449
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364449
Keywords
This article is cited by
-
Antiviral responses are shaped by heterogeneity in viral replication dynamics
Nature Microbiology (2023)
-
Cross-species analysis of viral nucleic acid interacting proteins identifies TAOKs as innate immune regulators
Nature Communications (2021)
-
Structural and mechanistic basis of mammalian Nudt12 RNA deNADding
Nature Chemical Biology (2019)
-
IRF3 and type I interferons fuel a fatal response to myocardial infarction
Nature Medicine (2017)
-
Mycobacterium tuberculosis-triggered Hippo pathway orchestrates CXCL1/2 expression to modulate host immune responses
Scientific Reports (2016)