Abstract
Bfl-1/A1 is generally recognized as a Bcl-2-related inhibitor of apoptosis. We show that Bfl-1 undergoes constitutive ubiquitin/proteasome-mediated turnover. Moreover, while Bfl-1 suppresses apoptosis induced by staurosporine or cytokine withdrawal, it is proapoptotic in response to tumor necrosis factor (TNF) receptor activation in FL5.12 pro-B cells. Its anti- versus proapoptotic effect is regulated by two proteolytic events: (1) its constitutive proteasome-mediated turnover and (2) its TNF/cycloheximide (CHX)-induced cleavage by μ-calpain, or a calpain-like activity, coincident with acquisition of a proapoptotic phenotype. In vitro studies suggest that calpain-mediated cleavage of Bfl-1 occurs between its Bcl-2 homology (BH)4 and BH3 domains. This would be consistent with the generation of a proapoptotic Bax-like BH1–3 molecule. Overall, our studies uncovered two new regulatory mechanisms that play a decisive role in determining Bfl-1's prosurvival versus prodeath activities. These findings might provide important clues to counteract chemoresistance in tumor cells that highly express Bfl-1.
Similar content being viewed by others
Introduction
The cellular apoptotic response to developmental or environmental cues determines whether a cell will live or die, and has obvious consequences for the fate and well-being of an organism. Disease conditions that arise from too much or too little apoptosis include autoimmune diseases, neurodegenerative disorders and cancer. Proteins in the Bcl-2 family are key regulators of the apoptotic response, owing to their ability to control mitochondrial permeability and the release of apoptogenic factors in response to death-inducing stimuli, thereby modulating the activation of caspases and amplification of the apoptotic cascade.1, 2 This family is comprised of antiapoptotic factors (Bcl-2, Bcl-xL, Bfl-1/A1, Mcl-1, Bcl-w) that possess Bcl-2 homology (BH) domains BH1, BH2, BH3 and BH4, and of multidomain proapoptotic proteins (Bax, Bak) with BH1, BH2 and BH3 domains as well as proapoptotic BH3-only members like Bid, Noxa, Puma, Nbk/Bik, Bim, Hrk, Bmf and Bad. Despite their opposite effects on the outcome of the cell, several of these factors (Bcl-xL, Bcl-2, Bcl-w, Bax, Bid) share structural homology with the pore-forming domains of bacterial toxins.3
Mechanistically, cell death results from the combined inhibition of the prosurvival activity of factors like Bcl-2 and Bcl-xL by BH3-only proteins, together with the activation of Bax or Bak.4, 5, 6 The ratio between these factors seems to determine the cell susceptibility to apoptosis.7 Indeed, synthetic peptides derived from the BH3 domains of Bid and Bim were shown to induce apoptosis by activating Bax/Bak, whereas those from Bad and Bik displaced BH3-only factors from antiapoptotic Bcl-2 proteins.5 The recent demonstration that a hydrocarbon-stapled Bid BH3 helix provoked apoptosis in human leukemia cells and inhibited tumor growth in xenografts further illustrates the important relevance of these findings.8 In addition to their mutual interaction with one another, members of the Bcl-2 family undergo post-translational modification and/or proteolytic cleavage that serve to modulate their activity. For instance, phosphorylation of the BH3-only protein Bad promotes its dissociation from Bcl-xL allowing cell survival whereas cleavage of Bcl-xL, Bcl-2, Bid or Bax by caspases, or in some cases by calpains, augments mitochondrial permeability and cell death9, 10, 11 (for reviews, see Cory and Adams,2 Wang12 and Chan and Mattson13).
The human Bfl-1 protein and its mouse homolog A1 are antiapoptotic members of the Bcl-2 family.14, 15 Akin to Bcl-2 and Bcl-xL, Bfl-1 possesses four BH domains. Although the N-terminal BH4 region involved in Bcl-2 heterodimerization with Bax is not well conserved in Bfl-1, Bfl-1 has been shown to interact with Bax in a two-hybrid assay.16, 17 Moreover, several charged residues in Bfl-1 interrupt the hydrophobic carboxyl-terminal region that is responsible for anchorage of Bcl-2 and Bcl-xL at the mitochondrial membrane. In conjunction with the fact that Bfl-1 lacks the antiproliferative property attributed to Bcl-2, these findings suggest that Bfl-1 might function somewhat differently from other members of the Bcl-2 family.18 While there are four a1 genes in the mouse genome, the unique human bfl-1 gene encodes a full-length protein that localizes at mitochondria as well as an alternatively spliced variant (Bfl-1S) lacking 12 C-terminal amino acids that localizes to the nucleus but is nevertheless protective against apoptosis.19, 20 bfl-1 transcripts are constitutively expressed in bone marrow, lymphoid organs and peripheral blood leukocytes and are induced in response to activation of NF-κB transcription factors in many cell types.14, 21, 22, 23, 24, 25, 26 These findings suggest an important role for Bfl-1 in the survival of cells in the immune system.
Bfl-1/A1 can confer protection from apoptosis induced by a variety of agents in different cells, including staurosporine (STS), FAS activation and TRAIL in Jurkat T-cells, p53 in baby rat kidney cells and antigen receptor ligation in B cells.18, 26, 27, 28, 29 However, its protective activity is not always absolute, as Bfl-1/A1 is less protective than Bcl-xL in microvascular endothelial cells treated with tumor necrosis factor (TNF)α and only delays TNFα-induced apoptosis in HT1080 fibrosarcoma cells.25, 30 Similarly, while Bfl-1 potently blocks etoposide-induced cell death in HT1080 fibrosarcoma cells, it is much less effective against this agent in Jurkat T-cells.25, 28 These results suggest that the level of protection conferred by Bfl-1 is perhaps subject to modulation in different cell contexts and in response to particular apoptotic stimuli.
Here we report that the ubiquitin/proteasome pathway regulates Bfl-1 protein stability and that Bfl-1 can act either as an antiapoptotic or as a prodeath factor, depending on the death-inducing stimulus. Our studies show that its differential effect is regulated at two levels. Experiments indicate that manifestation of its prodeath effect is dependent on Bfl-1 protein turnover, in conjunction with its cleavage by μ-calpain or a calpain-like activity in FL5.12 pro-B cells subjected to TNF receptor activation. Overall, these studies identify two new modes of Bfl-1 protein regulation, and reveal how these can influence the outcome of the apoptotic response.
Results
Bfl-1 both prevents and promotes apoptosis in response to different death-inducing stimuli in FL5.12 pro-B cells
To gain further insight into the function of Bfl-1, we compared its protective activity to that of Bcl-xL in response to different death-inducing signals by stably expressing them in the interleukin-3 (IL-3)-dependent mouse pro-B cell line FL5.12. Since none of the commercially available antibodies could recognize the human Bfl-1 protein (not shown), both Bfl-1 and Bcl-xL were expressed with an N-terminal green fluorescent protein (GFP) tag. After cell sorting, individual G418-resistant cell clones expressing GFP-Bfl-1 were isolated by limited dilution. Cell clone B3 was selected for further analysis, as it showed GFP-Bfl-1 expression closest to that of cells expressing GFP-Bcl-xL (Figure 1a). Both GFP-Bfl-1 and GFP-Bcl-xL effectively protected FL5.12 cells from apoptosis induced by the pan protein kinase inhibitor STS, as revealed by flow cytometry analysis, although the protective activity of Bfl-1 was less pronounced than that of Bcl-xL (Figure 1b). This was in contrast to control cells that showed approximately 60% of cells in the sub-G0/G1 phase. Similar to STS treatment, Bfl-1 and Bcl-xL conferred protection toward apoptosis induced by IL3 withdrawal compared to the control (Figure 1b). Whereas on average 66% of control cells died following withdrawal of IL3, approximately 34% of cells expressing GFP-Bfl-1 underwent apoptosis while only 20% of cells expressing GFP-Bcl-xL died. Overall, these results indicate that despite its sequence divergence from classical Bcl-2 family members Bcl-2 and Bcl-xL, Bfl-1 is protective in pro-B cells challenged with STS or cytokine deprivation.
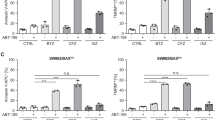
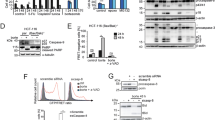
FL5.12 cells expressing GFP-Bfl-1, GFP-Bfl-1ΔC or GFP-Bcl-xL are protected from apoptosis induced by STS or IL-3 withdrawal. (a) Western blot analysis of parental FL5.12 cells (lane 1) and those expressing GFP-Bfl-1 (lane 2), GFP-Bfl-1ΔC (lane 3) or GFP-Bcl-xL (lane 4) with an anti-GFP antibody (Torrey Pines; top panel). The blot was reprobed with an anti-actin antibody as a control (bottom panel). (b) Flow cytometry analysis of FL5.12-derived cells untreated or subjected to STS treatment or IL3 withdrawal. Parental FL5.12 cells and those expressing GFP-Bfl-1, GFP-Bfl-1ΔC or GFP-Bcl-xL were treated with STS for 24 h, or cultured in medium lacking IL-3 for 48 h. After staining with propidium iodide, cells were analyzed by flow cytometry. Percent cell death represents the mean value from three independent experiments
Unlike the C-terminal transmembrane (TM) domain of Bcl-2 and Bcl-xL, several charged residues interrupt the C-terminal domain of Bfl-1. Since deletion of this region results in Bfl-1 localizing in a diffuse cytoplasmic pattern compared to the punctate mitochondrial staining observed with wild-type GFP-Bfl-1 (Simmons and Gélinas, unpublished data), we investigated the importance of mitochondrial localization for the protective function of Bfl-1 by analyzing FL5.12 cells expressing a C-terminally truncated form of Bfl-1 lacking amino acids 152–175 (GFP-Bfl-1ΔC). GFP-Bfl-1ΔC displayed increased protection toward apoptosis induced by STS or IL-3-withdrawal compared to GFP-Bfl-1, and its protective activity toward cell death induced by cytokine withdrawal was comparable to that of GFP-Bcl-xL (Figure 1b). It therefore appears that Bfl-1ΔC remains a functional apoptosis inhibitor despite its cytoplasmic relocalization compared to that of Bfl-1.
In sharp contrast to the antiapoptotic activity of Bfl-1 toward STS and IL3 withdrawal, apoptosis induced by TNF receptor activation with TNFα in the presence of cycloheximide (CHX) was markedly enhanced in FL5.12 cells expressing GFP-Bfl-1 compared to SFFV-neo control cells or parental FL5.12 cells (Figure 2a, left panel, and data not shown). Indeed, cell treatment with TNF/CHX for 6 h triggered apoptosis in an average of 54% of GFP-Bfl-1-expressing cells, compared to about 23% for the SFFV-neo control. A similar increase in sensitivity to TNF/CHX-induced apoptosis was observed with another GFP-Bfl-1 cell clone as well as with a pool of GFP-Bfl-1-expressing cells, ruling out the possibility that this enhanced death phenotype resulted from clonal variation (data not shown). The incidence of apoptosis in Bfl-1-expressing cells was noticed with as little as 2 ng/ml of TNFα, indicating that a relatively low concentration of TNFα was sufficient for manifestation of this proapoptotic effect (data not shown). When assayed under the same conditions, cells expressing GFP-Bcl-xL were sensitized to TNF/CHX akin to those expressing GFP-Bfl-1 (Figure 2a, left panel). A time-course analysis revealed that enhanced apoptosis compared to the control could be detected as early as 3 h post-TNFα/CHX treatment of GFP-Bfl-1-expressing cells (Figure 2b). In agreement with our findings with GFP-Bfl-1, FL5.12 cells expressing a 2xMyc-tagged form of Bfl-1 also showed a marked increase in TNF/CHX-induced apoptosis compared to controls, thereby ruling out the possibility of nonspecific effects caused by fusion to GFP (Figure 2c).
FL5.12 cells expressing Bfl-1 are sensitized to TNFα/CHX-induced apoptosis compared to control cells, while those expressing mutant GFP-Bfl-1ΔC are protected. (a) Left panel: FL5.12 cells expressing GFP-Bfl-1 or GFP-Bcl-xL and those containing empty SFFV-neo vector were treated with TNFα/CHX for 6 h. Cell death was evaluated upon staining with propidium iodide and flow cytometry analysis. Where indicated, cells were pretreated for 1 h with the caspase inhibitor zVAD-fmk or overnight with the proteasome inhibitor MG132. Percent cell death represents the mean value from three independent experiments. Right panel: FL5.12 cells expressing GFP-Bfl-1 or GFP-Bfl-1ΔC and those containing empty SFFV-neo vector were treated with TNFα/CHX for 6 h. Cell death was evaluated upon staining with propidium iodide and flow cytometry analysis. Percent cell death represents the mean value from three independent experiments. (b) Time-course analysis of TNFα/CHX-induced apoptosis in FL5.12 cells expressing GFP-Bfl-1. Cells were treated with TNFα plus CHX as described in (a) for 1, 2, 3 or 5 h. Cells were stained with propidium iodide and analyzed by flow cytometry. (c) The TNF/CHX sensitivity of FL5.12 cells expressing 2xMyc-Bfl-1 was compared to that of FL5.12 cells expressing GFP-Bfl-1 or GFP-Bfl-1ΔC or parental FL5.12 cells treated with CHX alone or TNF/CHX for 6 h. A representative experiment is shown
As anticipated, cell pretreatment with the general caspase inhibitor carbobenzoxy-Val-Ala-Asp-fluoromethyl ketone (zVAD-fmk) significantly reduced apoptosis in GFP-Bfl-1- and GFP-Bcl-xL-expressing cells, most likely due to its ability to inhibit caspases activated downstream of TNF receptor activation (Figure 2a, left panel). Interestingly however, cell treatment with proteasome inhibitor MG132 suppressed the enhanced death phenotype observed in GFP-Bfl-1 and GFP-Bcl-xL cells treated with TNFα/CHX (Figure 2a, left panel). It is noteworthy that contrary to cells expressing full-length Bfl-1, FL5.12 cells expressing GFP-Bfl-1ΔC were not sensitized to apoptosis induced by TNF/CHX and were rather somewhat protected (Figure 2a, right panel, and Figure 2c). Western blot analyses ruled out the possibility that the different sensitivity of these cells toward TNF/CHX could be attributed to differences in endogenous c-FLIP protein levels, an important regulator of apoptosis in TNF/CHX-treated cells, as no significant changes were observed (data not shown). Taken together, the results indicate that Bfl-1 has opposite effects in response to different death-inducing stimuli in FL5.12 cells, that is, it is protective in cells challenged with STS or cytokine withdrawal, but it has a prodeath effect in cells treated with TNF/CHX. In contrast, its C-terminal mutant Bfl-1ΔC remained protective.
Bfl-1 undergoes proteolytic cleavage in FL5.12 cells treated with TNF/CHX
The prodeath phenotype that we observed in FL5.12 cells treated with TNF/CHX led us to investigate possible changes in Bfl-1 protein expression and/or stability. Western blot analysis of cells treated with CHX, TNFα or both uncovered two distinct features of the Bfl-1 protein involving its stability and processing. Full-length GFP-Bfl-1 appeared to be relatively short-lived as revealed by a very substantial decrease in the amount of GFP-Bfl-1 in cells treated with CHX alone for 6 h to block de novo protein synthesis, as was GFP-Bcl-xL (Figure 3a, left panel, compare lanes 1 and 3; 7 and 8). In contrast, GFP-Bfl-1 remained intact in cells treated with TNFα alone or with STS (lanes 2 and 4). Likewise, 2xMyc-Bfl-1 was also short-lived (Figure 3a, right panel, compare lanes 3 and 1). A time-course analysis in CHX-treated FL5.12 cells showed that the half-life of 2xMyc-Bfl-1 is less than 2 h (data now shown).
Bfl-1 undergoes constitutive proteasome-mediated turnover and it is proteolytically cleaved upon treatment of FL5.12 cells with TNFα/CHX. (a) Left panel: FL5.12 cells expressing GFP-Bfl-1 were left untreated (lane 1) or treated with STS for 24 h (lane 2), or treated for 6 h with CHX alone (lane 3), TNFα alone (lane 4) or TNFα/CHX (lane 5) and analyzed by immunoblotting. Cells expressing GFP alone (lane 6) or GFP-Bcl-xL were analyzed in parallel (lanes 7–9). Whole-cell lysates were probed with anti-GFP (Sigma; top panel) or anti-actin antibodies (bottom panel). Arrows point to the positions of full-length GFP-Bfl-1, GFP-Bcl-xL and GFP. The open arrow indicates the ∼32 kDa GFP-reactive GFP-Bfl-1 cleavage product. Asterisks mark the positions of GFP-reactive GFP-Bcl-xL cleavage products. Right panel: Parental FL5.12 cells (lane 1) or FL5.12 cells expressing 2xMyc-Bfl-1 were left untreated (lane 2), or treated with CHX alone (lane 3), TNF alone (lane 4) or TNF/CHX for 6 h (lane 5) and analyzed by immunoblotting. The blot was probed with an anti-Myc antibody (top panel) and reprobed with anti-actin (bottom panel). (b) Time-course analysis of GFP-Bfl-1 protein cleavage in response to TNF/CHX treatment. FL5.12 cells expressing GFP-Bfl-1 were left untreated (lane 1) or treated with TNF/CHX as described in (a) for 1, 2, 3, 5 or 6 h (lanes 2–6) followed by immunoblotting with an anti-GFP antibody (Sigma; top panel). The membrane was reprobed for caspase-8, -9 or -3 and actin as a control. (c) Full-length GFP-Bfl-1 is stabilized by proteasome inhibition and inhibitors of caspases or calpains block generation of its TNF/CHX-induced cleavage product. FL5.12 cells expressing GFP-Bfl-1 were left untreated or incubated with TNFα/CHX for 6 h as described in (a). Where indicated, cells were pretreated with proteasome inhibitor MG132 (10 μM for 6 h), pan-caspase inhibitor zVAD-fmk (50 μM for 30 min) or calpain inhibitor I ALLN. Lysates were analyzed by immunoblotting with an anti-GFP antibody (Torrey Pines; top and middle panels) and reprobed with anti-actin as a loading control (bottom panel). Arrows point to the position of full-length GFP-Bfl-1. Open arrows indicate the ∼32 kDa GFP-reactive GFP-Bfl-1 cleavage product that was detected upon longer exposure with this antibody (Torrey Pines; middle panel). (d) FL5.12 cells expressing GFP-Bfl-1 (lanes 2–4) or GFP-Bfl-1ΔC (lanes 5–8) were left untreated (lanes 2 and 5) or incubated with CHX alone (lanes 3 and 6) or with TNF/CHX (lanes 4 and 7). As a control, cells expressing GFP-Bfl-1ΔC were pretreated with pan-caspase inhibitor zVAD-fmk (50 μM for 30 min) prior to addition of TNF/CHX (lane 8). Cells containing the empty SFFV-neo vector were used as control (lane 1). The blot was successively probed with anti-GFP (Torrey Pines; top and middle panels) and anti-actin antibodies (bottom panel). Arrows point to the positions of full-length GFP-Bfl-1 and GFP-Bfl-1ΔC. Open arrows indicate the ∼32 kDa GFP-reactive cleavage products from GFP-Bfl-1 and GFP-Bfl-1ΔC detected upon longer exposure with this antibody (middle panel)
Our studies in TNF/CHX-treated cells also revealed a second level of regulation, as GFP-Bfl-1 appeared to undergo proteolytic cleavage in cells treated with TNF/CHX as revealed by the appearance of a GFP-reactive band of ∼32 kDa (Figure 3a, left panel, compare lane 5 to lanes 1–4). Although not obvious in this particular gel, the steady-state level of full-length GFP-Bfl-1 appeared to be further reduced in cells treated with TNFα/CHX compared to those treated with CHX alone, suggesting that TNFα/CHX might induce proteolysis of GFP-Bfl-1. In a time-course analysis, progressive disappearance of full-length GFP-Bfl-1 was detected starting at around 3 h following addition of TNF/CHX and coincided with gradual accumulation of the ∼32 kDa GFP-reactive band (Figure 3b, top panel, compare lanes 4–6 to 1–3). This coincided with activation of cellular caspase-8, -9 and -3 and the onset of ‘enhanced cell death’ in GFP-Bfl-1 cells compared to SFFV-neo control cells (Figure 3b, center panels, lanes 4–6, and Figure 2b). Importantly, no such fragment was observed in GFP-Bfl-1-expressing cells treated with STS in which GFP-Bfl-1 is protective (Figure 3a, left panel, lane 2, and Figure 1b). Together, these results suggest that GFP-Bfl-1 undergoes proteolytic cleavage in response to receptor activation by TNF/CHX. The fact that the apparent mobility of the resulting GFP-reactive band was slower than that of a GFP control in stably transfected FL5.12 cells suggested that cleavage occurred within Bfl-1 sequences (Figure 3a, left panel, compare lanes 5 and 6). When analyzed in parallel, GFP-Bcl-xL also underwent cleavage upon TNF/CHX treatment to generate GFP-reactive fragments (Figure 3a, left panel, lane 9). Taking into account the apparent mobility of the GFP tag (∼25 kDa), the size of these fragments is reminiscent of previously documented Bcl-xL cleavage products induced by m-calpain and caspase-3.11
Since caspases and calpains have been implicated in cleaving the prosurvival Bcl-2 and Bcl-xL proteins9, 10, 11, 31, 32, 33 and pretreatment with zVAD-fmk or MG132 suppressed TNFα/CHX-induced apoptosis in GFP-Bfl-1-expressing cells (Figure 2a), we investigated possible effects of proteasome inhibitor MG132, the broad-spectrum caspase inhibitor zVAD-fmk and calpain inhibitor I N-acetyl-L-leucyl-L-leucyl-norleucinal (ALLN) on the stability of the full-length Bfl-1 protein and its TNF/CHX-induced cleavage in FL5.12 cells. As shown in Figure 3c, full-length GFP-Bfl-1 was significantly stabilized by MG132 in TNFα/CHX-treated cells (top panel, compare lanes 4 and 5). Interestingly, MG132 also blocked appearance of the ∼32 kDa fragment (middle panel, compare lanes 5 and 4). In contrast, the broad-spectrum caspase inhibitor zVAD-fmk failed to prevent disappearance of full-length GFP-Bfl-1 but it inhibited appearance of the ∼32 kDa cleavage product (Figure 3c, top and middle panels, compare lane 6 to lanes 4 and 5). Thus, while inhibition of caspases promoted overall cell survival (Figure 2a), caspases are not involved in full-length Bfl-1 protein turnover. However, the results suggest that caspases somehow participate in the generation of the Bfl-1 protein fragment. Calpain inhibitor I ALLN enhanced the stability of full-length GFP-Bfl-1, most probably due to its ability to also suppress the calpain-like activity of the proteasome,34 and also inhibited formation of the ∼32 kDa cleavage fragment (Figure 3c, top and middle panels, compare lanes 7 and 5 to lane 4). Overall, these results indicate that full-length Bfl-1 undergoes turnover via the proteasome, and that caspases and calpains, or a calpain-like protease, might be implicated in its proteolytic cleavage in a direct or indirect fashion.
Contrary to wild-type GFP-Bfl-1 that sensitized cells to apoptosis induced by TNF/CHX, FL5.12 cells expressing the GFP-Bfl-1ΔC mutant were protected (Figure 2a). To gain further insight into this observation, we examined the status of the GFP-Bfl-1ΔC protein in TNFα/CHX-treated cells. In Western blots, full-length GFP-Bfl-1ΔC protein was noticeably more stable (∼2.8-fold) than wild-type GFP-Bfl-1 in cells treated with CHX (Figure 3d, top panel, compare lanes 5 and 6 to 2 and 3). Interestingly, TNFα/CHX treatment of GFP-Bfl-1ΔC-expressing cells also gave rise to a GFP-reactive cleavage fragment of ∼32 kDa similar to that observed in cells expressing wild-type GFP-Bfl-1 (Figure 3d, top and middle panels, lanes 7 and 4). As with GFP-Bfl-1, caspase inhibition with zVAD-fmk suppressed appearance of the GFP-Bfl-1ΔC cleavage product, suggesting that caspase activation is implicated either directly or indirectly in the proteolytic cleavage of Bfl-1 (Figure 3d, compare lanes 8 and 7). Moreover, the results suggest that deletion of the C-terminal domain abrogates Bfl-1 protein turnover but not its TNF/CHX-induced proteolytic cleavage. Taken together, the data indicate that full-length Bfl-1 undergoes proteasome-mediated turnover and that cell treatment with TNF/CHX promotes cleavage of the protein, coincident with a marked enhancement in cell death compared to control cells. Furthermore, stabilization of GFP-Bfl-1 by proteasome inhibition or deletion of the C-terminal domain counteracted the proapoptotic effects associated with TNFα/CHX treatment of GFP-Bfl-1-expressing cells.
Bfl-1 is a substrate for ubiquitination and its modification involves lysines and serine/threonine residues in the C-terminal domain
The stabilizing effects of proteasome inhibitor MG132 on the GFP-Bfl-1 protein, together with the increased stability of the GFP-Bfl-1ΔC mutant compared to wild type, led us to explore if Bfl-1 was a substrate for ubiquitination. Human 293T cells were cotransfected with GFP-Bfl-1 or GFP-Bfl-1ΔC together with Myc-tagged ubiquitin, followed by immunoprecipitation with an anti-GFP antibody and Western blotting with anti-Myc. A significant amount of polyubiquitinated GFP-Bfl-1 was observed in cells cotransfected with Myc-Ub in the presence or absence of MG132 (Figure 4a, top panel, lanes 2 and 3). In contrast, very low-level ubiquitination was seen for GFP-Bfl-1ΔC, even in the presence of proteasome inhibitor (compare lanes 4 and 5 to 2 and 3). Furthermore, when FL5.12 cells stably expressing wild-type GFP-Bfl-1 or mutant GFP-Bfl-1ΔC were analyzed for endogenous ubiquitination in the absence or presence of MG132, immunoprecipitation with anti-GFP followed by Western blotting with anti-ubiquitin revealed significant polyubiquitination of GFP-Bfl-1 in the presence of MG132 (Figure 4b, top panel, lane 2). In contrast, GFP-Bfl-1ΔC showed dramatically lower levels of endogenous ubiquitin conjugation in MG132-treated cells (lane 4). These data indicate that Bfl-1 is a substrate for polyubiquitination and suggest that deletion of its C-terminal domain interferes with its efficient modification.
GFP-Bfl-1 and the S152A.T156A mutant are efficiently ubiquitinated in contrast to C-terminal mutants GFP-Bfl-1ΔC, S152D.T156D or K151,163,172R. (a) Immunoblot analysis of Bfl-1 ubiquitination in transfected 293T cells. Human 293T cells were transiently transfected with constructs expressing GFP-Bfl-1 (lanes 1–3), GFP-Bfl-1ΔC (ΔC; lanes 4 and 5) or GFP-Bfl-1 point mutants S152A.T156A (ST/AA; lanes 6 and 7), S152D.T156D (ST/DD; lanes 8 and 9) or K151,163,172R (KKK/RRR; lanes 10 and 11), alone (lane 1) or together with Myc-tagged ubiquitin (lanes 2–11). Where indicated, cells were pretreated with proteasome inhibitor MG132 (20 μM) for 4 h (lanes 1, 3, 5, 7, 9 and 11). Extracts were immunoprecipitated with an anti-GFP antibody (Clontech) and immunoblotted with anti-Myc. Total lysates (20% of input) were immunoblotted with anti-GFP as a control (Torrey Pines; bottom panel). (b) Effects of C-terminal mutations on the endogenous ubiquitination of Bfl-1 in FL5.12 pro-B cells. FL5.12 cells stably expressing wild-type GFP-Bfl-1 (lanes 1 and 2), GFP-Bfl-1ΔC (lanes 3 and 4) or GFP-Bfl-1 point mutant K151,163,172R (KKK/RRR; lanes 5 and 6) were left untreated (lanes 1, 3 and 5) or were incubated with MG132 for 4 h (lanes 2, 4 and 6). Extracts were immunoprecipitated with an anti-GFP antibody (Clontech) followed by immunoblotting with an antibody to ubiquitin (Santa-Cruz). Total lysates (20% of input) were immunoblotted with an anti-GFP antibody (lower panel). (c) Alignment of human Bfl-1 and mouse A1 protein sequences. The BH1–3 domains and putative BH4 sequences are indicated.17, 18 The C-terminal domain is underlined. Amino acids marked in bold were substituted by site-directed mutagenesis with alanine (A), aspartic acid (D) or arginine (R) residues. (d) Effects of C-terminal lysine mutations on the stability of Bfl-1 in FL5.12 pro-B cells. FL5.12 cells stably expressing wild-type GFP-Bfl-1 (lanes 1–3) or GFP-Bfl-1 point mutant K151,163,172R (KKK/RRR; lanes 4–6) were left untreated (lanes 1 and 4) or treated for 6 h with CHX alone (lanes 2 and 5) or with MG132 for 4 h (lanes 3 and 6). Extracts were immunoprecipitated with an anti-GFP antibody (Clontech) followed by immunoblotting with an anti-GFP antibody (Torrey Pines; top panel). The blot was reprobed for actin as a loading control (bottom panel)
Site-directed mutagenesis was then used to identify residues in the C-terminal domain of Bfl-1 that might be involved in regulating Bfl-1 ubiquitination and degradation. The C-terminal domain of Bfl-1 contains three lysines that could serve as potential substrates for covalent modification by ubiquitin, two of which are conserved in its mouse homolog A1 (Figure 4c). Substitution of arginine for lysines 151, 163 and 172 in the C-terminal domain of Bfl-1 slightly reduced its ubiquitination in 293T cells cotransfected with Myc-ubiquitin (Figure 4a, compare lanes 10 and 11 to 2 and 3). The effect of this mutation was more pronounced when ubiquitination of mutant K151,163,172R was analyzed endogenously in FL5.12 pro-B cells. Akin to the GFP-Bfl-1ΔC mutant, the C-terminal lysine mutant protein K151,163,172R showed dramatically lower levels of endogenous ubiquitin conjugation in the presence of MG132 compared to wild-type GFP-Bfl-1 when stably expressed in FL5.12 cells (Figure 4b, top panel, compare lanes 5, 6 to 1, 2 and 3, 4). Overall, these results are consistent with the identification of lysine residues involved in ubiquitination and suggest that the residual ubiquitination of mutant K151,163,172R that we observed in 293T cells transiently transfected with Myc-ubiquitin might reflect ubiquitination at alternate sites.
Addition of polyubiquitin chains is often regulated by phosphorylation or, in some instances, by dephosphorylation of serine(s) or threonine(s) in the vicinity of lysines that are substrate for ubiquitination (reviewed by Fuchs et al.35). Alignment of human Bfl-1 with its mouse homolog A1 identified serine 152 and threonines 156 and 161 to be conserved in both species (Figure 4c). The combined mutation of serine 152 and threonine 156 to alanine or aspartic acid was used to mimic unphosphorylated or constitutively phosphorylated Bfl-1 at these positions. As shown in Figure 4a, GFP-Bfl-1 mutant S152A.T156A (ST/AA) was highly ubiquitinated in the presence or absence of MG132 when cotransfected in 293T cells with Myc-tagged ubiquitin (top panel, compare lanes 6 and 7 to 2 and 3). In contrast, its phosphomimetic counterpart S152D.T156D (ST/DD) showed a marked decrease in ubiquitination compared to wild type, akin to the GFP-Bfl-1ΔC deletion mutant (Figure 4a, top panel, compare lanes 8, 9 and 4, 5 to 2, 3). Overall, these assays identified some critical substrate lysines and putative serine or threonine phosphorylation sites in the C-terminus of Bfl-1 that appear to be important for regulating its ubiquitination.
Mutation of C-terminal lysines attenuates the proteasome-mediated turnover of Bfl-1 in FL5.12 cells and suppresses its prodeath phenotype in cells treated with TNFα/CHX
We next went on to assess the effects of C-terminal lysine mutations on the endogenous stability of GFP-Bfl-1 in FL5.12 pro-B cells. Protein levels in cells treated with protein synthesis inhibitor CHX alone or with proteasome inhibitor MG132 were assessed by immunoblotting with anti-GFP. As anticipated, wild-type GFP-Bfl-1 was unstable in cells treated with CHX for 6 h, whereas cell treatment with MG132 stabilized the protein (Figure 4d, lanes 1–3). In contrast, C-terminal mutant K151,163,172R was significantly more stable than wild type in the presence of CHX, consistent with its reduced susceptibility to undergo ubiquitination (KKK/RRR; Figure 4d, lanes 3–6, and Figure 4b, lanes 5 and 6). Overall, these data indicate that mutation of C-terminal lysines that interfere with Bfl-1's ubiquitination leads to increased stability in FL5.12 pro-B cells.
The activity of Bfl-1 ubiquitination mutants toward TNF-induced apoptosis was explored in stably transfected FL5.12 cells expressing wild-type or mutant proteins at equivalent levels. As seen in Figure 5, the prodeath phenotype of Bfl-1 in FL5.12 cells subjected to TNFα/CHX treatment was reduced in cells expressing its degradation-resistant mutant K151,163,172R. Similarly, the S152D.T156D mutant also conferred increased survival compared to GFP-Bfl-1. Together with the increased stability of the GFP-Bfl-1ΔC mutant and its protective activity, these results indicate that mutations that diminish proteasome-dependent degradation of Bfl-1 help to counteract its proapoptotic effect in FL5.12 cells treated with TNF/CHX.
Mutations that interfere with efficient ubiquitination of GFP-Bfl-1 suppress its prodeath phenotype in FL5.12 cells treated with TNFα/CHX. Flow cytometry analysis of FL5.12 cells expressing mutant Bfl-1 proteins following TNFα/CHX treatment. FL5.12 cells stably expressing GFP-Bfl-1 or mutants K151,163,172R (KKK/RRR) or S152D.T156D (ST/DD) were left untreated (left) or treated with CHX alone (middle) or with TNFα/CHX (right) for 6 h, followed by staining with propidium iodide and flow cytometry analysis. Cells containing the empty SFFV-neo vector served as a control. A representative experiment is shown
Bfl-1 is a substrate for μ-calpain and its endogenous cleavage in FL5.12 cells treated with TNF/CHX coincides with acquisition of a prodeath phenotype
Our observation that GFP-Bfl-1 protein cleavage during a time-course analysis (Figure 3b) coincided with the onset of enhanced cell death in FL5.12 cells treated with TNF/CHX (Figure 2b) raised the possibility that its proteolytic cleavage might be correlated with its prodeath phenotype in this context. Studies in Figure 3c suggested that caspases and calpains, or a calpain-like protease, might be implicated in a direct or indirect fashion in its cleavage, as zVAD-fmk, MG132 or ALLN inhibited appearance of the ∼32 kDa GFP-Bfl-1 cleavage product in TNF/CHX-treated FL5.12 cells (middle panel, lanes 5–7). Moreover, considering that both caspases and calpains have been implicated in cleaving the prosurvival Bcl-2 and Bcl-xL proteins.9, 10, 11, 31, 32, 33 we first assayed whether GFP-Bfl-1 was a substrate for cleavage by recombinant caspase-3, -8 or -9. Although caspases were functional for cleaving a control Myc-tagged Bid protein in vitro, none of them could induce cleavage of radiolabeled GFP-Bfl-1 (data not shown). It therefore appears that caspases might play an indirect role in Bfl-1 protein cleavage, perhaps by acting at an upstream step leading to processing of Bfl-1 in the TNFR cascade.
Next, we investigated the possible implication of calpains. Calpains are calcium-dependent cysteine proteases whose μ- and m-isoforms regulate several important intracellular processes. Calpains have been shown to cleave several proteins involved in apoptosis including members of the Bcl-2 family like proapoptotic members Bid and Bax, and cleavage of the prosurvival factor Bcl-xL leads to the emergence of a proapoptotic fragment.11, 36, 37, 38 Pretreatment with the cell-permeable calpain inhibitor I ALLN significantly decreased the ‘enhanced death’ phenotype of GFP-Bfl-1 in TNFα/CHX-treated FL5.12 cells (Figure 6a), similar to proteasome inhibitor MG132 that incidentally can also suppress the activity of calpains39 (Figure 2a; R Mellgren, personal communication). However, since ALLN is a competitive inhibitor of calpain proteases40 (Ki 190 nM) and can also suppress the activity of the proteasome34 (Ki 670 nM), we do not rule out the possibility that its protective activity in this context might result from suppression of Bfl-1 cleavage, stabilization of full-length GFP-Bfl-1 or both.
Bfl-1 is a substrate for μ-calpain and is cleaved by a calpain-like activity in FL5.12 cells treated with TNF/CHX, coincident with acquisition of a prodeath phenotype. (a) Cell pretreatment with calpain inhibitor I ALLN suppresses the prodeath phenotype of GFP-Bfl-1 in FL5.12 cells treated with TNFα/CHX. Cells were left untreated or treated with TNFα/CHX for 6 h followed by staining with propidium iodide and flow cytometry analysis. Where indicated, cells were pretreated with calpain inhibitor I ALLN. The results are representative of three independent experiments. (b) In vivo processing of GFP-Bfl-1 in FL5.12 cells during a time course of TNF/CHX treatment coincides with disappearance of the 80 kDa proform of endogenous μ-calpain. FL5.12 cells stably expressing GFP-Bfl-1 were treated with TNFα/CHX for 1, 2, 3, 5 or 6 h in the absence (lanes 1–5) or pretreated for 1 h with ALLN before incubation with TNF/CHX (lane 6). Cell extracts were analyzed by immunoblotting with an anti-GFP antibody (Sigma) and reprobed with anti-μ-calpain to monitor cleavage-induced activation of endogenous pro-μ-calpain. The blot was reprobed with anti-actin as a loading control. An asterisk marks a nonspecific band that is occasionally seen above GFP-Bfl-1. (c) GFP-Bfl-1 is a substrate for μ-calpain cleavage in vitro. 35S-labeled GFP-Bfl-1 produced by in vitro translation was incubated with μ-calpain (lanes 2–4) or m-calpain (lanes 5 and 6) in the absence (lanes 2, 3 and 5) or presence of calpain inhibitor I ALLN (lane 4) or calpain inhibitor II ALLM (lane 6). Buffer lacking calcium and containing the chelating agent EGTA was used as a control (lane 3). In vitro-translated 35S-labeled Flag-Bax (lanes 7 and 8) and GFP-Bcl-xL (lanes 9 and 10) were used as positive controls to verify the activity of μ-calpain (lane 8) or m-calpain (lane 10). An open arrowhead marks the GFP-tagged calpain-induced cleavage product of GFP-Bfl-1. (d) Contrary to GFP-Bfl-1 and GFP-Bfl-1ΔC, GFP-Bcl-xL and a GFP-Bfl-1/xL hybrid containing amino acids 7–22 of Bfl-1 are not efficiently cleaved by μ-calpain in vitro. 35S-labeled GFP-Bfl-1 (lanes 1 and 2), GFP-Bfl-1ΔC (lanes 3 and 4), GFP-Bcl-xL (lanes 5 and 6) or GFP-Bfl-1/xL (lanes 7 and 8) was incubated with purified μ-calpain as described in (c), followed by PAGE and autoradiography. Arrows point to the positions of the full-length proteins. (e) Like GFP-tagged Bfl-1, untagged Bfl-1 is a substrate for μ-calpain in vitro. 35S-labeled untagged Bfl-1 was incubated with μ-calpain (lanes 2–4) in the absence (lanes 2 and 3) or presence of calpain inhibitor I ALLN (lane 4). Buffer lacking calcium and containing the chelating agent EGTA was used as a control (lane 3). An open arrowhead marks the ∼16 kDa 35S-labeled C-terminal cleavage product of Bfl-1 generated by μ-calpain
Interestingly, the onset of endogenous GFP-Bfl-1 processing, as indicated by appearance of the ∼32 kDa GFP-reactive fragment, during a time course of cell treatment with TNF/CHX coincided with disappearance of the 80 kDa proform of the large subunit of μ-calpain, suggestive of cleavage-mediated proteolytic activation of μ-calpain (Figure 6b, top and middle panels, lanes 3–5). Pretreatment with ALLN partially suppressed cleavage of pro-μ-calpain in vivo, to levels seen at ∼3 h, and concomitantly reduced TNF/CHX-induced production of the GFP-Bfl-1 cleavage fragment (compare lanes 6 and 5). These findings raised the possibility that calpain, or a calpain-like activity, was involved in Bfl-1 processing. This suggestion would agree with the fact that calpains can be activated by caspases.
We further explored this hypothesis by testing the ability of either μ-calpain or m-calpain to cleave radiolabeled GFP-Bfl-1 produced by in vitro translation. μ-Calpain completely cleaved full-length GFP-Bfl-1 to produce a fragment ∼32 kDa, reminiscent of the fragment observed in TNF/CHX-treated cells (Figure 6c, compare lanes 1 and 2). As expected for a calpain protease, this activity was dependent on the presence of calcium, as omission of calcium in the reaction blocked Bfl-1 processing (lane 3). Likewise, calpain inhibitor I ALLN abolished in vitro cleavage of GFP-Bfl-1 by μ-calpain (lane 4). Consistent with previous reports,37, 38 μ-calpain also cleaved a control Bax protein (compare lanes 7 and 8). m-Calpain was much less efficient in inducing cleavage of GFP-Bfl-1, whereas its control target Bcl-xL was cleaved under the same conditions (Figure 6c, lanes 5 and 10).
Unlike caspases, μ-calpain does not recognize a well-defined cleavage motif although it cleaves the Bax protein after an aspartic acid. However, mutant proteins with alanine substitutions for each of the seven aspartic acids in Bfl-1 were cleaved by μ-calpain in vitro as efficiently as wild-type GFP-Bfl-1 (data not shown). It therefore appeared that μ-calpain cleavage of Bfl-1 did not occur at a caspase-like motif. Since GFP alone was not processed by μ-calpain (data not shown), we investigated if the cleavage occurred within the BH4 domain of Bfl-1. As shown in Figure 6d, full-length GFP-Bfl-1 and the GFP-Bfl-1ΔC mutant were cleaved with similar efficiency by μ-calpain (compare lanes 2 and 4). In contrast, a hybrid protein containing amino acids 7–22 derived from the BH4 domain of Bfl-1 substituted for amino acids 2–18 in Bcl-xL (GFP-Bfl-1/xL) did not undergo significant proteolysis, similar to the control GFP-Bcl-xL protein under the same conditions (compare lanes 6 and 8). Combined with fact that in vitro cleavage of GFP-Bfl-1 by μ-calpain generated an N-terminal GFP-reactive product that was ∼32 kDa (Figure 6c), these results suggested that cleavage by μ-calpain occurs downstream of amino acid 22 in Bfl-1.
In vitro cleavage of untagged Bfl-1 confirmed that Bfl-1 is a bona fide substrate of μ-calpain and that its cleavage by this protease was not an artifact of its fusion to GFP (Figure 6e, lane 2). As anticipated, cleavage was dependent on calcium and was suppressed by ALLN (lanes 3 and 4). In conjunction with the ∼32 kDa N-terminal GFP-reactive fragment detected upon processing of GFP-Bfl-1 in vitro and in vivo (Figures 3 and 6c), the ∼16 kDa 35S-labeled cleavage product derived from untagged Bfl-1 suggested that μ-calpain cleaves in the N-terminus of Bfl-1 somewhere in the vicinity of amino acids ∼25–35. Altogether, these results suggest that cleavage by μ-calpain occurs between the BH4 and BH3 domains of Bfl-1. This would be consistent with the generation of a Bax-like multi-domain proapoptotic Bfl-1 fragment. However, since none of the commercially available antibodies could recognize the human Bfl-1 protein, we do not rule out the possibility that μ-calpain or a calpain-like activity might cleave Bfl-1 at multiple sites in vivo. Future studies will help to determine the cleavage site(s) in Bfl-1.
Overall, our findings implicate μ-calpain or a calpain-like activity in the post-translational processing of Bfl-1 in FL5.12 cells subjected to TNFα/CHX treatment and its conversion into a multidomain BH1–3 proapoptotic factor in this context. Since impairment of proteasome-mediated turnover of full-length Bfl-1 antagonized its enhanced death phenotype in TNFα/CHX-treated cells, it appears that these two distinct proteolytic activities might act in concert to produce the proapoptotic phenotype of Bfl-1 in this context.
Discussion
In this study, we demonstrate that while Bfl-1 generally suppresses apoptosis triggered by several death-inducing stimuli including STS or cytokine withdrawal, it can behave as a prodeath factor in certain cells in response to TNF receptor activation in the presence of CHX. Our experiments demonstrate that manifestation of Bfl-1's proapoptotic effect is regulated by the combined action of two different proteolytic events: (1) ubiquitin-dependent proteasome-mediated turnover of full-length Bfl-1 via modification of residues in its C-terminal domain, and (2) proteolytic cleavage by μ-calpain or a calpain-like activity in vivo. The observation that mutations that interfered with ubiquitin-mediated turnover of Bfl-1 suppressed its prodeath phenotype, despite cleavage by μ-calpain, underscores the fact that these two different proteolytic events are both necessary to convert Bfl-1 from an antiapoptotic protein into a prodeath factor. Consistent with our findings with GFP-tagged Bfl-1 in the FL5.12 cell model, cells expressing a Myc-tagged form of Bfl-1 were also sensitized to TNF receptor-mediated apoptosis, but not STS, compared to control FL5.12 cells. Overall, this study uncovered two new regulatory mechanisms for Bfl-1 that play a decisive role in determining the extent of its antiapoptotic activity.
The protective activity of Bfl-1 in FL5.12 cells against apoptosis induced by STS or cytokine deprivation is in sharp contrast to its capacity to enhance their demise following treatment with TNFα/CHX. This suggests that Bfl-1 can exert differential effects depending on the nature of the death-inducing stimulus. While Bfl-1 was previously reported to potently inhibit cell death caused by the chemotherapeutic agent etoposide, it only partially suppressed apoptosis induced by TNFα in the HT1080 fibrosarcoma cell line.25 Bfl-1 also delays endothelial cell apoptosis in response to TNFα but does not abrogate it.41 These observations with Bfl-1 agree with the capacity of other Bcl-2 family proteins like Bcl-2 and Bcl-xL to either suppress cell death (fully or partially) or promote apoptosis depending on the stimulus and/or the cell type.9, 10, 31 Contrary to a recent report indicating that GFP-Bfl-1 is proapoptotic in transiently transfected 293T cells in the absence of a death-inducing signal,42 we found no evidence that GFP-Bfl-1 is by itself proapoptotic, as FL5.12 cells stably expressing GFP-Bfl-1 were viable and proliferated normally. Moreover, GFP-Bfl-1 showed no toxicity when transiently expressed in other cell lines such as MCF-7 or HeLa cells (data not shown). Importantly, we obtained similar results with a Myc-tagged form of Bfl-1. Thus, the phenotype that was reported in 293T cells is perhaps cell specific.
Proteasome-mediated degradation regulates the stability of certain pro- and antiapoptotic Bcl-2 family members, and is often triggered in response to a death-inducing stimulus. For example, the prosurvival Mcl-1 protein is rapidly degraded early in apoptosis and is necessary for cell death to ensue.43, 44 In our study, we found that Bfl-1 undergoes constitutive proteasome-mediated turnover. However, we do not rule out the possibility that Bfl-1 might also undergo signal-induced proteasome-mediated turnover to accelerate its elimination in response to certain death-inducing stimuli and/or in certain cells.
Dephosphorylated Bcl-2 undergoes proteasomal degradation in endothelial cells treated with TNFα.45, 46 Alanine substitution of S152.T156 in Bfl-1 led to a remarkable increase in GFP-Bfl-1 ubiquitination. This suggests that dephosphorylation of Bfl-1 might trigger its ubiquitination and proteasome-dependent decay, and T156 is a potential phosphorylation site for casein kinase I or II. Interestingly, although the S152D/T156D and K151,163,172R mutations decreased ubiquitination of Bfl-1 and reduced its proapoptotic phenotype, they did not confer full resistance to TNF/CHX-induced apoptosis in FL5.12 cells, unlike the Bfl-1ΔC mutant that was protective. This raises the possibility that additional modifications elsewhere in the protein might also be required to stabilize fully Bfl-1 in FL5.12 cells. Incidentally, mutation of potential phosphorylation sites in Bcl-2, distinct from those previously described to be targeted by MAP kinase, also affected Bcl-2 protein stability.47 Based on these results, we do not rule out the possibility that additional sites within Bfl-1 might also be involved in regulating its turnover. Future studies will help to address this issue and to identify the E3 ligase and kinase(s) responsible for its modification.
There is increasing evidence supporting a role for calpains in apoptosis, although their function in this context appears to be cell- and stimulus-dependent and has been the subject of conflicting observations (for review, see Wang12 and Goll et al.48). For instance, calpains have been described by some to act upstream of caspase processing, but others implicated them downstream of caspase activation.49, 50, 51, 52, 53, 54 Likewise, some found apoptosis induced by FAS ligand to be independent of calpains, but others implicated them in FAS receptor-mediated apoptosis of neutrophils.55, 56 Our observation that Bfl-1 sensitizes cells to TNF/CHX-induced death, coincident with its proteolytic processing by μ-calpain or a calpain-like activity in vivo, is consistent with reports indicating that caspases and calpains can cleave prosurvival Bcl-xL or Bcl-2 proteins leading to the emergence of a prodeath fragment.9, 10, 11, 31, 32, 33 However, it should be noted that these proteins are often partially cleaved in vivo and that coexistence of death-promoting fragments along with the full-length proteins can yield an overall antiapoptotic outcome, akin to the behavior of the GFP-Bfl-1ΔC mutant. It is therefore possible that cleaved Bfl-1 readily manifests its proapoptotic phenotype because its half-life is fairly short (<2 h for 2xMyc-Bfl-1) compared to that of Bcl-2 (∼10 h).57, 58
Although the cleavage site(s) in Bfl-1 remain(s) to be mapped precisely, our studies raise the possibility that Bfl-1 might be converted into a proapoptotic Bax-like multi-BH1–3 protein. For example, cleavage of Bcl-2 and Bcl-xL by caspases or calpain generates a Bax-like fragment, and the proapoptotic activity of Bcl-2-derived fragment was shown to depend on the integrity of the BH3 and TM domains.10, 11, 59 The fact that our Bfl-1ΔC mutant was not converted into a proapoptotic factor although it was still processed by μ-calpain suggests that Bfl-1 might be processed into a Bax-like factor whose death-inducing activity requires anchoring at the mitochondria (Figure 7). In addition, we do not rule out the possibility that the greater stability of full-length GFP-Bfl-1ΔC compared to GFP-Bfl-1 might also be responsible for the positive outcome of the cells, since Bfl-1 protein turnover is required for manifestation of its prodeath phenotype and residues that regulate Bfl-1 ubiquitination map to its putative C-terminal TM domain (Figure 7).
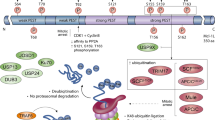
Model illustrating the interplay between constitutive proteasome-mediated turnover of Bfl-1 and its TNF/CHX-induced cleavage in Fl5.12 cells. Left panel: (a) Wild-type Bfl-1 localized at mitochondria is antiapoptotic and undergoes constitutive proteasome-mediated turnover as a result of ubiquitination in its C-terminal domain. (b) TNF receptor activation in FL5.12 cells by TNF/CHX treatment leads to activation of μ-calpain, or a calpain-like activity, that cleaves Bfl-1 to generate an ∼3 kDa N-terminal fragment and a multidomain Bax-like C-terminal BH1–3 cleavage product that localizes to mitochondria and is proapoptotic. Right panel: (a) Mutant Bfl-1ΔC does not localize to mitochondria but is nevertheless antiapoptotic. Because it lacks the C-terminal domain of Bfl-1, Bfl-1ΔC is less susceptible to ubiquitin/proteasome-mediated turnover than wild-type Bfl-1. (b) TNF/CHX-mediated activation of μ-calpain, or a calpain-like activity, cleaves Bfl-1ΔC to generate an ∼3 kDa N-terminal fragment and a multidomain C-terminal BH1–3 cleavage product that fails to localize to mitochondria. We propose that the increased stability of full-length Bfl-1ΔC together with the failure of its BH1–3 cleavage product to localize to mitochondria may be responsible for the fact that Bfl-1ΔC is not proapoptotic in FL5.12 cells treated with TNF/CHX, contrary to wild-type Bfl-1. The apical steps proposed in this model are suggested by prior studies56
Our experiments in FL5.12 cells expressing GFP-Bfl-1 suggest that TNF receptor activation in the presence of CHX triggers a pathway leading to cleavage of Bfl-1 by μ-calpain or a calpain-like activity in vivo. Although it remains to be determined how calpain proteases might be activated in this context, FAS-induced activation of caspase-8 in neutrophils was reported to result in inactivation of the calpain inhibitor calpastatin, thereby promoting cleavage of Bax by μ-calpain.56 Moreover, previous work showed that calpastatin is sensitive to proteolysis by recombinant caspase-1, -3 and -7 and that FAS-induced cell death in Jurkat T-cells or in human neuroblastoma cells converges on calpastatin degradation.54 Thus, by converging on caspase activation, death receptor pathways might activate calpain via degradation of calpastatin. A model positioning caspase-8 upstream of calpastatin and μ-calpain activation is supported by the inability of recombinant caspase-8 to cleave directly GFP-Bfl-1 in our in vitro assays (Figure 7 and data not shown). Alternatively, we do not rule out the possibility that phospholipid mediators, including second messengers implicated in TNFα signaling, might activate the catalytic activity of calpain by binding to its 30 kDa regulatory subunit to lower its calcium requirement to a range normally found intracellularly.60, 61, 62 Future studies will undoubtedly help to define the pathway(s) involved.
Overexpression of antiapoptotic genes including those encoding prosurvival Bcl-2 family proteins is a characteristic of many chemoresistant tumor cells, including diffuse large B-cell lymphoma and Hogdkin's Reed–Sternberg cells in which endogenous bfl-1 transcripts are highly expressed.63, 64, 65 Our results raise the possibility that, in some instances, high endogenous expression of Bfl-1 could perhaps be exploited to facilitate cell death in otherwise chemoresistant cells. It will therefore be interesting to see whether these findings could be extended to human tumor cells that display high endogenous levels of bfl-1 expression. In conclusion, our studies uncovered two proteolytic mechanisms involving the proteasome and calpains or a calpain-like activity that are interdependently necessary to convert Bfl-1 from an antiapoptotic protein into a prodeath factor.
Materials and Methods
Plasmids and mutagenesis
Plasmid pSFFV-neo66 was used to stably express Bfl-1 or Bcl-xL under the control of the spleen focus-forming virus LTR promoter in FL5.12 cells. The cDNA for Bcl-xL was a gift from C Labrie (CHUL). GFP-tagged forms of wild-type or mutant Bfl-1 or Bcl-xL were generated by cloning into the pEGFP-C3 vector (Clontech), followed by PCR-mediated subcloning into the EcoRI site of pSFFV-neo. This plasmid was also used in in vitro translation reactions in rabbit reticulocyte lysates from the T3 promoter. GFP-Bfl-1 mutants were generated in the pEGFP vector using the QuickChange Site-Directed Mutagenesis Kit (Stratagene). 2xMyc-Bfl-1 was generated by fusing two consecutive 10-amino-acid Myc epitopes to the N-terminus of Bfl-1 by PCR, followed by cloning into the EcoRI site of pSFFV-neo. Bid-Myc was expressed from the CMV promoter of pcDNA3.1/HisB (Invitrogen) and was a gift from E White (CABM & Rutgers Univ.). Flag-Bax was generated by subcloning Bax from a His-Myc-Bax construct along with a Flag tag into pcDNA3.1/HisC (Invitrogen). They were used for in vitro translation using the T7 promoter. Plasmid PCW7, used for transient transfection of 293T cells, expressed 6-His-Myc-tagged ubiquitin from the CMV promoter of pRBG4 and was a gift from J-Y Springael (Université Libre de Bruxelles).
Cell culture and transfection
Mouse WEHI-3B cells and the IL3-dependent mouse FL5.12 pro-B cells were maintained in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS), 20 mM HEPES, 2 mM L-glutamine, penicillin (100 U/ml) and streptomycin (100 μg/ml). The culture medium for FL5.12 cells was supplemented with 10% supernatant from confluent WEHI-3B cell cultures as a source of IL-3. FL5.12 cells transfected with SFFV-neo-derived vectors were selected and maintained in the presence of G418 (1 mg/ml). The human adenovirus-transformed embryonic kidney cell line 293T, expressing the SV40 large T antigen, was cultured in DMEM medium supplemented with 10% FBS, penicillin (100 U/ml) and streptomycin (100 μg/ml). Cell culture reagents were purchased from Invitrogen, unless indicated otherwise, and cells were maintained at 37°C in an atmosphere of 5% CO2.
For transfection, FL5.12 cells (1 × 107) in 0.4 ml of complete culture medium were incubated with 15 μg of pSFFV-neo-derived vector DNA for 10 min on ice and electroporated at 200V and a capacitance of 960 μF with a Gene Pulser (Biorad). Cells were incubated on ice for 10 min, transferred to 10 ml of complete culture medium and incubated for 36–48 h at 37°C before selection for neomycin resistance with G418 (1 mg/ml). Drug-resistant GFP-expressing FL5.12 cells were isolated on a Beckman Coulter EPICS Elite cell sorter. Pools of GFP-positive cells were generally used in this study. Where indicated, GFP-Bfl-1-expressing cell clone B3 was used, as it matched most closely the expression levels of GFP-Bcl-xL in FL5.12 cells. Its behavior was identical to that of a pool of sorted GFP-Bfl-1-expressing cells, ruling out the possibility of clonal variation. Human 293T cells used for analysis of Bfl-1 ubiquitination were transfected with a total of 10 μg DNA using Lipofectamine 2000 (Invitrogen).
Cell death assays and flow cytometry analysis
FL5.12-derived cell lines were treated with STS (20 nM; Sigma) for 24 h or with mouse recombinant TNFα (10 ng/ml; Sigma) plus CHX (10 μg/ml; Sigma) for 6 h unless specified otherwise. To assay cell death in response to cytokine deprivation, FL5.12-derived cell lines were washed three times with RPMI-1640 medium and resuspended in complete FL5.12 culture medium lacking IL-3 at a density of 5 × 105 cells/ml. Where indicated, cell death assays were performed following cell pretreatment overnight with proteasome inhibitor MG132 (20 μM; Calbiochem). Cell pretreatment with the pan-caspase inhibitor zVAD-fmk (Enzyme Systems Products) was at 50 μM for 1 h. Pretreatment with calpain inhibitor I ALLN (Calbiochem) or calpain inhibitor II N-acetyl-L-leucyl-L-leucyl-methioninal (ALLM; Calbiochem) was for 1 h at 20 μM. Cell death was monitored by flow cytometry analysis of cells (1–2 × 106) fixed in 70% ethanol for 30 min at 4°C, washed with PBS and resuspended in 0.5 ml PBS in the presence of DNase-free RNase (50 μg; Sigma) and propidium iodide (5 μg; Sigma) for 30 min at room temperature before analysis of DNA content on a Beckman Coulter Cytomics FC500 flow cytometer.
Immunoblotting
Cell extracts were prepared in EBC lysis buffer (50 mM Tris pH 8.0, 120 mM NaCl, 0.5% NP-40, 10% glycerol) in the presence of 1 × protease inhibitor cocktail (Roche). Total protein concentration was determined by the method of Bradford.67 Proteins were resolved by sodium dodecyl sulfate (SDS)–polyacrylamide gel electrophoresis (PAGE), transferred to nitrocellulose membranes (Schleicher & Schuell) and incubated with primary monoclonal anti-GFP (Sigma G-6539; 1 : 2000), polyclonal anti-GFP (Torrey Pines #TP401; 1 : 5000), monoclonal anti-Myc (Invitrogen 46-0603; 1 : 5000), polyclonal anti-caspase-3 (BD Pharmingen 552785; 1 : 1000), polyclonal anti-caspase-8 (BD Pharmingen 559932; 1 : 1000), monoclonal anti-caspase-9 (Stressgen AAM-139; 1 : 1000), polyclonal anti-μ-calpain (Calbiochem 208751; 1 : 1000), anti-ubiquitin (Santa-Cruz SC-8017; 1 : 1000) or polyclonal anti-actin (Sigma A-2066; 1 : 500). Secondary antibody detection was performed by enhanced chemiluminescence (ECL; Amersham Life Sciences).
Ubiquitination assays
Human 293T cells at 90% confluence in a 10 cm dish were transiently transfected with PCW7 expressing Myc-tagged ubiquitin (5 μg) along with GFP-tagged constructs expressing wild-type or mutant Bfl-1 (5 μg) using Lipofectamine 2000 (45 μl). Cells were lysed in RIPA buffer (50 mM Tris pH 8.0, 120 mM NaCl, 1% NP-40, 0.5% deoxycholic acid, 0.1% SDS) in the presence of 1 × protease inhibitor cocktail, 1 mM phenylmethanesulfonyl fluoride (PMSF) and 0.005 M iodoacetamide (Calbiochem). After preclearing with protein A-sepharose beads (Amersham), lysates (500 μl) were immunoprecipitated with a polyclonal anti-GFP antibody (Clontech 8372-2; 3 μl) and protein A-sepharose for 4 h each, washed four times in RIPA buffer and boiled in SDS loading buffer. Immunoprecipitates were resolved by 7.5% SDS–PAGE and transferred to nitrocellulose membranes followed by immunoblotting with an anti-Myc antibody. For each experiment, straight lysates (20% of input) were analyzed by immunoblotting with anti-GFP (Torrey Pines; 1 : 5000) as a control. Endogenous ubiquitination of Bfl-1 was monitored in stably transfected FL5.12 cells (1 × 107) expressing wild-type or mutant GFP-Bfl-1. Cell extracts were immunoprecipitated with an anti-GFP antibody (Clontech) as described above and immunoblotted with an anti-ubiquitin antibody (Santa-Cruz).
In vitro cleavage assays of in vitro-translated proteins by calpains and caspases
GFP-Bfl-1, GFP-Bfl-1ΔC, GFP-Bfl-1/-xL, GFP-Bcl-xL, GFP or untagged Bfl-1 was produced by in vitro transcription/translation from the T3 promoter of SFFV-neo using the TNT-coupled reticulocyte lysate system (Promega) in the presence of [35S]methionine (Amersham). Control Flag-Bax was expressed from the T7 promoter of pcDNA3.1/HisC. Radiolabeled GFP-Bfl-1, GFP-Bfl-1ΔC, GFP-Bfl-1/-xL, GFP-Bcl-xL, GFP or untagged Bfl-1 proteins (5 μl) or Flag-Bax control (1 μl) were incubated with purified porcine μ-calpain (Calbiochem; 10 μl) in 30 μl of μ-calpain reaction buffer (30 mM Tris-HCl pH 7.5, 750 μM CaCl2, 1.5 mM DTT),68 or with recombinant rat m-calpain (10 μl; Calbiochem) in 30 μl of m-calpain reaction buffer (20 mM HEPES pH 7.5, 50 mM KCl, 2 mM MgCl2, 5 mM CaCl2, 1 mM DTT).36 Control reactions included buffer lacking CaCl2 and supplemented with calcium chelator EGTA (1.5 mM), or containing calpain inhibitor I ALLN (Calbiochem) or calpain inhibitor II ALLM (Calbiochem) at 10 μM each. Reactions were incubated at 30°C for 2 h for μ-calpain or at 37°C for 2 h for m-calpain, and terminated on ice by addition of 2 × SDS loading buffer. The susceptibility of radiolabeled GFP-Bfl-1 to serve as substrate for caspase cleavage was assayed upon incubation with 1 U of recombinant caspase-3, caspase-9 (Apotech) or caspase-8 (Biovision) in 2 × caspase reaction buffer (Biovision) and incubated at 37°C for 2 h. Bid-Myc expressed from the T7 promoter of pcDNA3.1/HisB was used as a positive control. The reaction was terminated by addition of SDS loading buffer. Proteins were resolved by 15% SDS–PAGE and the gels were vacuum dried and exposed to XOMAT film.
Abbreviations
- STS:
-
staurosporine
- TNF:
-
tumor necrosis factor
- BH:
-
Bcl-2 homology
- TM:
-
transmembrane
- CHX:
-
cycloheximide
- zVAD-fmk:
-
carbobenzoxy-Val-Ala-Asp-fluoromethyl ketone
- GFP:
-
green fluorescent protein
- ALLN:
-
N-acetyl-L-leucyl-L-leucyl-norleucinal
- ALLM:
-
N-acetyl-L-leucyl-L-leucyl-methioninal
- IL-3:
-
interleukin-3
References
Adams JM and Cory S (1998) The Bcl-2 protein family: arbiters of cell survival. Science 281: 1322–1326
Cory S and Adams JM (2002) The Bcl2 family: regulators of the cellular life-or-death switch. Nat. Rev. Cancer 2: 647–656
Petros AM, Olejniczak ET and Fesik SW (2004) Structural biology of the Bcl-2 family of proteins. Biochim. Biophys. Acta 1644: 83–94
Cheng EH, Wei M, Weiler S, Flavell RA, Mak TW, Lindsten T and Korsmeyer SJ (2001) BCL-2, BCL-XL sequester BH3 domain-only molecules preventing BAX- and BAK-mediated mitochondrial apoptosis. Mol. Cell 8: 705–711
Letai A, Bassik MC, Walensky LD, Sorcinelli MD, Weiler S and Korsmeyer SJ (2002) Distinct BH3 domains either sensitize or activate mitochondrial apoptosis, serving as prototype cancer therapeutics. Cancer Cell 2: 183–192
Chen L, Willis SN, Wei A, Smith BJ, Fletcher JI, Hinds MG, Colman PM, Day CL, Adams JM and Huang DCS (2005) Differential targeting of prosurvival Bcl-2 proteins by their BH3-only ligands allows complementary apoptotic function. Mol. Cell 17: 393–403
Chittenden T (2002) BH3 domains: intracellular death-ligands critical for initiating apoptosis. Cancer Cell 2: 165–166
Walensky LD, Kung AL, Escher I, Malia TJ, Barbuto S, Wright RD, Wagner G, Verdine GL and Korsmeyer SJ (2004) Activation of apoptosis in vivo by a hydrocarbon-stapled BH3 helix. Science 305: 1466–1470
Clem RJ, Cheng EH, Karp CL, Kirsch DG, Ueno K, Takahashi A, Kastan MB, Griffin DE, Earnshaw WC, Veliuona MA and Hardwick JM (1998) Modulation of cell death by Bcl-xL through caspase interaction. Proc. Natl. Acad. Sci. USA 95: 554–559
Cheng EH, Kirsch DG, Clem RJ, Ravi R, Kastan MB, Bedi A, Ueno K and Hardwick JM (1997) Conversion of Bcl-2 to a Bax-like death effector by caspases. Science 278: 1966–1968
Nakagawa T and Yuan J (2000) Cross-talk between two cysteine protease families. Activation of caspase-12 by calpain in apoptosis. J. Cell Biol. 150: 887–894
Wang K (2000) Calpain or caspase: can you tell the difference? Trends Neurosci. 23: 20–26
Chan SL and Mattson MP (1999) Caspase and calpain substrates: roles in synaptic plasticity and cell death. J. Neurosci. Res. 58: 167–190
Choi SS, Park I-C, Yun JW, Sung YC, Hong S-I and Shin H-S (1995) A novel Bcl-2 related gene, Bfl-1, is overexpressed in stomach cancer and preferentially expressed in bone marrow. Oncogene 11: 1693–1698
Lin EY, Orlofsky A, Berger MS and Prystowsky MB (1993) Characterization of A1, a novel hemopoietic-specific early-response gene with sequence similarity to bcl-2. J. Immunol. 151: 1979–1988
Hirotani M, Zhang Y, Fujita N, Naito M and Tsuruo T (1999) NH2-terminal BH4 domain of Bcl-2 is functional for heterodimerization with Bax and inhibition of apoptosis. J. Biol. Chem. 274: 20415–20420
Zhang H, Cowan-Jacob SW, Simonen M, Greenhalf W, Heim J and Meyhack B (2000) Structural basis of Bfl-1 for its interaction with Bax and anti-apoptotic action in mammalian and yeast cells. J. Biol. Chem. 275: 11092–11099 (a) Sedlak TW, Oltvai ZN, Yang E, Wang K, Boise LH, Thompson CB and Korsmeyer SJ (1995) Multiple Bcl-2 family members demonstrate selective dimerizations with Bax. Proc. Natl. Acad. Sci. USA 92: 7834–7838
D’Sa-Eipper C and Chinnadurai G (1998) Functional dissection of Bfl-1, a Bcl-2 homolog: anti-apoptosis, oncogene-cooperation and cell proliferation activities. Oncogene 16: 3105–3114
Hatakeyama S, Hamasaki A, Negishi I, Loh DY, Sendo F and Nakayama K (1998) Multiple gene duplication and expression of mouse bcl-2-related genes, A1. Int. Immunol. 10: 631–637
Ko JK, Lee MJ, Cho SH, Cho JA, Lee BY, Koh JS, Lee SS, Shim YH and Kim CW (2003) Bfl-1S, a novel alternative splice variant of Bfl-1, localizes in the nucleus via its C-terminus and prevents cell death. Oncogene 22: 2457–2465
Kenny JJ, Knobloch TJ, Augustus M, Carter KC, Rosen CA and Lang JC (1997) GRS, a novel member of the Bcl-2 gene family, is highly expressed in multiple cancer cell lines and in normal leukocytes. Oncogene 14: 997–1001
Zong WX, Edelstein LC, Chen C, Bash J and Gélinas C (1999) The prosurvival Bcl-2 homolog Bfl-1/A1 is a direct transcriptional target of NF-κB that blocks TNFα-induced apoptosis. Genes Dev. 13: 382–387
Hu X, Yee E, Harlan JM, Wong F and Karsan A (1998) Lipopolysaccharide induces the antiapoptotic molecules, A1 and A20, in microvascular endothelial cells. Blood 92: 2759–2765
Lee HH, Dadgostar H, Cheng Q, Shu J and Cheng G (1999) NF-kappaB-mediated up-regulation of Bcl-x and Bfl-1/A1 is required for CD40 survival signaling in B lymphocytes. Proc. Natl. Acad. Sci. USA 96: 9136–9141
Wang CY, Guttridge DC, Mayo MW and Baldwin Jr AS (1999) NF-kappaB induces expression of the Bcl-2 homologue A1/Bfl-1 to preferentially suppress chemotherapy-induced apoptosis. Mol. Cell. Biol. 19: 5923–5929
Grumont RJ, Rourke IJ and Gerondakis S (1999) Rel-dependent induction of A1 transcription is required to protect B cells from antigen receptor ligation-induced apoptosis. Genes Dev. 13: 400–411 (a) Edelstein LC, Lagos L, Simmons M, Tirumalai H and Gelinas C (2003) NF-κB-dependent assembly of an enhanceosome-like complex on the promoter region of apoptosis inhibitor Bf1-1/A1, Mol. Cell. Biol. 23: 2749–2761
Somogyi RD, Wu Y, Orlofsky A and Prystowsky MB (2001) Transient expression of the Bcl-2 family member, A1-a, results in nuclear localization and resistance to staurosporine-induced apoptosis. Cell Death Diff. 8: 785–793
Werner AB, de Vries E, Tait SWG, Bontjer I and Borst J (2002) Bcl-2 family member Bfl-1/A1 sequesters truncated Bid to inhibit its collaboration with pro-apoptotic Bak or Bax. J. Biol. Chem. 277: 22781–22788
D’Sa-Eipper C, Subramanian T and Chinnadurai G (1996) Bfl-1, a Bcl-2 homologue, suppresses p53-induced apoptosis and exhibits potent cooperative transforming activity. Cancer Res. 56: 3879–3882
Karsan A, Yee E and Harlan JM (1996) Endothelial cell death induced by tumor necrosis factor-alpha is inhibited by the Bcl-2 family member, A1. J. Biol. Chem. 271: 27201–27204
Liang Y, Nylander KD, Yan C and Schor NF (2002) Role of caspase-3-dependent Bcl-2 cleavage in potentiation of apoptosis by Bcl-2. Mol. Pharmacol. 61: 142–149
Kirsch DG, Doseff A, Chau BN, Lim DS, de Souza-Pinto NC, Hansford R, Kastan MB, Lazebnik YA and Hardwick JM (1999) Caspase-3-dependent cleavage of Bcl-2 promotes release of cytochrome c. J. Biol. Chem. 274: 21155–21161
Johnson BW and Boise LH (1999) Bcl-2 and caspase inhibition cooperate to inhibit tumor necrosis factor-alpha-induced cell death in a Bcl-2 cleavage-independent fashion. J. Biol. Chem. 274: 18552–18558
Rock KL, Gramm C, Rothstein L, Clark K, Stein R, Dick L, Hwang D and Goldberg AL (1994) Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell 78: 761–771
Fuchs SY, Fried VA and Ronai Z (1998) Stress-activated kinases regulate protein stability. Oncogene 17: 1483–1490
Mandic A, Viktorsson K, Strandberg L, Heiden T, Hansson J, Linder S and Shoshan MC (2002) Calpain-mediated Bid cleavage and calpain-independent Bak modulation: two separate pathways in cisplatin-induced apoptosis. Mol. Cell. Biol. 22: 3003–3013
Wood DE, Thomas A, Devi LA, Berman Y, Beavis RC, Reed JC and Newcomb EW (1998) Bax cleavage is mediated by calpain during drug-induced apoptosis. Oncogene 17: 1069–1078
Choi WS, Lee EH, Chung CW, Jung YK, Jin BK, Kim SU, Oh TH, Saido TC and Oh YJ (2001) Cleavage of Bax is mediated by caspase-dependent or -independent calpain activation in dopaminergic neuronal cells: protective role of Bcl-2. J. Neurochem. 77: 1531–1541
Mellgren RL (1997) Specificities of cell permeant peptidyl inhibitors for the proteinase activities of μ-calpain and the 20S proteasome. J. Biol. Chem. 272: 29899–29903
Griscavage JM, Wilk S and Ignarro LJ (1995) Serine and cysteine proteinase inhibitors prevent nitric oxide production by activated macrophages by interfering with transcription of the inducible NO synthase gene. Biochem. Biophys. Res. Commun. 215: 721–729
Duriez PJ, Wong F, Dorovini-Zis K, Shahidi R and Karsan A (2000) A1 functions at the mitochondria to delay endothelial apoptosis in response to tumor necrosis factor. J. Biol. Chem. 275: 18099–18107
Ko JK, Choi KH, Kim HJ, Choi HY, Yeo DJ, Park SO, Yang WS, Kim YN and Kim CW (2003) Conversion of Bfl-1, an anti-apoptotic Bcl-2 family protein, to a potent pro-apoptotic protein by fusion with green fluorescent protein (GFP). FEBS Lett. 551: 29–36
Cuconati A, Mukherjee C, Perez D and White E (2003) DNA damage response and MCL-1 destruction initiate apoptosis in adenovirus-infected cells. Genes Dev. 17: 2922–2932
Nijhawan D, Fang M, Traer E, Zhong Q, Gao W, Du F and Wang X (2003) Elimination of Mcl-1 is required for the initiation of apoptosis following ultraviolet irradiation. Genes Dev. 17: 1475–1486
Dimmeler S, Breitschopf K, Haendeler J and Zeiher AM (1999) Dephosphorylation targets Bcl-2 for ubiquitin-dependent degradation: a link between the apoptosome and the proteasome pathway. J. Exp. Med. 189: 1815–1822
Breitschopf K, Haendeler J, Malchow P, Zeiher AM and Dimmeler S (2000) Posttranslational modification of Bcl-2 facilitates its proteasome-dependent degradation: molecular characterization of the involved signaling pathway. Mol. Cell. Biol. 20: 1886–1896
Deng X, Gao F, Flagg T and May WS (2004) Mono- and multisite phosphorylation enhances Bcl-2’s antiapoptotic function and inhibition of cell cycle entry functions. Proc. Natl. Acad. Sci. USA 101: 153–158
Goll DE, Thompson VF, Li H, Wei W and Cong J (2003) The calpain system. Physiol. Rev. 83: 731–801
Ruiz-Vela A, Gonzalez de Buitrago G and Martinez AC (1999) Implication of calpain in caspase activation during B cell clonal deletion. EMBO J. 18: 4988–4998
Wolf BB, Goldstein JC, Beere H, Amarante-Mentes GP, Salvesen GS and Green DR (1999) Calpain functions in a caspase-independent manner to promote apoptosis-like events during platelet activation. Blood 94: 1683–1692
Chua BT, Guo K and Li P (2000) Direct cleavage by the calcium-activated protease calpain can lead to inactivation of caspases. J. Biol. Chem. 275: 5131–5135
McGinnis KM, Gnegy ME, Park YH, Mukerjee N and Wang KK (1999) Procaspase-3 and poly(ADP)ribose polymerase (PARP) are calpain substrates. Biochem. Biophys. Res. Commun. 263: 94–99
Porn-Ares MI, Samali A and Orrenius S (1998) Cleavage of the calpain inhibitor, calpastatin, during apoptosis. Cell Death Differ. 5: 1028–1033
Wang KK, Posmantur R, Nadimpalli R, Nath R, Mohan P, Nixon RA, Talanian RV, Keegan M, Herzog L and Allen H (1998) Caspase-mediated fragmentation of calpain inhibitor protein calpastatin during apoptosis. Arch. Biochem. Biophys. 356: 187–196
Saido TC, Sorimachi H and Suzuki K (1994) Calpain: new perspectives in molecular diversity and physiological-pathological involvement. FASEB J. 8: 814–822
Altznauer F, Conus S, Cavalli A, Folkers G and Simon HU (2004) Calpain-1 regulates Bax and subsequent Smac-dependent caspase-3 activation in neutrophil apoptosis. J. Biol. Chem. 279: 5947–5957
Merino R, Ding L, Veis DJ, Korsmeyer SJ and Nunez G (1994) Developmental regulation of the Bcl-2 protein and susceptibility to cell death in B lymphocytes. EMBO J. 13: 683–691
Reed JC (1996) A day in the life of the Bcl-2 protein: does the turnover rate of Bcl-2 serve as a biological clock for cellular lifespan regulation? Leuk. Res. 20: 109–111
Fujita N, Nagahashi A, Nagashima K, Rokudai S and Tsuruo T (1998) Acceleration of apoptotic cell death after cleavage of Bcl-xL protein by caspase-3-like proteases. Oncogene 17: 1295–1304
Chakrabarti AK, Dasgupta S, Gadsden RHS, Hogan EL and Banik NL (1996) Regulation of brain m-calpain Ca2+ sensitivity by mixtures of mambrane lipids: activation at intracellular Ca2+ level. J. Neurosci. Res. 44: 374–380
Melloni E, Pontremoli S, Michetti M, Sacco O, Sparatore B and Horecker BL (1986) The involvement of calpain in the activation of protein kinase C in neutrophils stimulated by phorbol myrstic acid. J. Biol. Chem. 261: 4101–4105
Eto A, Akita Y, Saido TC, Suzuki K and Kawashima S (1995) The role of the calpain–calpastatin system in thyrotropin-releasing hormone-induced selective down-regulation of a protein kinase C isozyme, nPKC epsilon, in rat pituitary GH4C1 cells. J. Biol. Chem. 270: 25115–25120
Alizadeh A, Eisen M, Davis R, Ma C, Lossos I, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson Jr J, Lu L, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R, Levy R, Wilson WH, Greyer MR, Byrd JC, Botstein D, Brown PO and Staudt LM (2000) Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403: 503–511
Debatin KM, Poncet D and Kroemer G (2002) Chemotherapy: targeting the mitochondrial cell death pathway. Oncogene 21: 8786–8803
Hinz M, Loser P, Mathas S, Krappmann D, Dorken B and Scheidereit C (2001) Constitutive NF-kappaB maintains high expression of a characteristic gene network, including CD40, CD86, and a set of antiapoptotic genes in Hodgkin/Reed–Sternberg cells. Blood 97: 2798–2807
Boise L, Gonzalez-Garcia M, Postema C, Ding L, Lindsten T, Turka L, Mao X, Nunez G and Thompson C (1993) bcl-x, a bcl-2-related gene that functions as a dominant regulator of apoptotic cell death. Cell 74: 597–608
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal. Biochem. 72: 248–254
Shumway SD, Maki M and Miyamoto S (1999) The PEST domain of IkBa is necessary and sufficient for in vitro degradation by μ-calpain. J. Biol. Chem. 274: 30874–30881
Acknowledgements
We thank C Labrie, J-Y Springael, E White and R Sundararajan for the gifts of cells and reagents. We are very grateful to E White, R Sundararajan and members of the Gélinas laboratory for discussion and suggestions during the course of this work, to C Gauthier-Rouvière for allowing some experiments to be carried out in her laboratory, and to Petra Pham and Hyejeong Choi for help with FACS analysis. We also thank E White, Y Fan, J Dutta, N Gupta and G Fan for helpful comments on the manuscript. This work was supported by Public Health Service Grant CA83937 from the National Cancer Institute to CG. We acknowledge partial support from the Flow Cytometry Core Facility of the Cancer Institute of New Jersey. JFK was a postdoctoral fellow of the New Jersey Commission on Cancer Research and The Foundation of the UMDNJ. MJS was partially supported by NIH predoctoral training grant in Biochemistry and Molecular Biology GM08360.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by ME Peter
Rights and permissions
About this article
Cite this article
Kucharczak, J., Simmons, M., Duckett, C. et al. Constitutive proteasome-mediated turnover of Bfl-1/A1 and its processing in response to TNF receptor activation in FL5.12 pro-B cells convert it into a prodeath factor. Cell Death Differ 12, 1225–1239 (2005). https://doi.org/10.1038/sj.cdd.4401684
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401684
Keywords
This article is cited by
-
BCL-2 protein family: attractive targets for cancer therapy
Apoptosis (2023)
-
What can we learn from mice lacking pro-survival BCL-2 proteins to advance BH3 mimetic drugs for cancer therapy?
Cell Death & Differentiation (2022)
-
Characterization of a novel human BFL-1-specific monoclonal antibody
Cell Death & Differentiation (2020)
-
TRIM17 and TRIM28 antagonistically regulate the ubiquitination and anti-apoptotic activity of BCL2A1
Cell Death & Differentiation (2019)
-
BFL1 modulates apoptosis at the membrane level through a bifunctional and multimodal mechanism showing key differences with BCLXL
Cell Death & Differentiation (2019)