Abstract
We have previously suggested a central role for mitochondria in the observed sex differences in metabolic traits. However, the mechanisms by which sex differences affect adipose mitochondrial function and metabolic syndrome are unclear. Here we show that in both mice and humans, adipose mitochondrial functions are elevated in females and are strongly associated with adiposity, insulin resistance and plasma lipids. Using a panel of diverse inbred strains of mice, we identify a genetic locus on mouse chromosome 17 that controls mitochondrial mass and function in adipose tissue in a sex- and tissue-specific manner. This locus contains Ndufv2 and regulates the expression of at least 89 mitochondrial genes in females, including oxidative phosphorylation genes and those related to mitochondrial DNA content. Overexpression studies indicate that Ndufv2 mediates these effects by regulating supercomplex assembly and elevating mitochondrial reactive oxygen species production, which generates a signal that increases mitochondrial biogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA sequencing raw data can be accessed at the Gene Expression Omnibus under accession GSE64770 (HMDP expression arrays (adipose and liver)) and GSE112947 (Gonadectomized RNA-seq data). GTEx datasets can be found online at the GTEx portal website (https://gtexportal.org/home/datasets). Uncropped scans of blots are available in the source data files. HMDP data from the authors laboratories will be made available on reasonable request. Source data are provided with this paper.
References
Seldin, M., Yang, X. & Lusis, A. J. Systems genetics applications in metabolism research. Nat. Metab. 1, 1038–1050 (2019).
Bennett, B. J. et al. A high-resolution association mapping panel for the dissection of complex traits in mice. Genome Res. 20, 281–290 (2010).
Parks, B. W. et al. Genetic control of obesity and gut microbiota composition in response to high-fat, high-sucrose diet in mice. Cell Metab. 17, 141–152 (2013).
Parks, B. W. et al. Genetic architecture of insulin resistance in the mouse. Cell Metab. 21, 334–347 (2015).
Chella Krishnan, K. et al. Sex-specific metabolic functions of adipose Lipocalin-2. Mol. Metab. 30, 30–47 (2019).
Norheim, F. et al. Genetic, dietary, and sex-specific regulation of hepatic ceramides and the relationship between hepatic ceramides and IR. J. Lipid Res. 59, 1164–1174 (2018).
Norheim, F. et al. Genetic and hormonal control of hepatic steatosis in female and male mice. J. Lipid Res. 58, 178–187 (2017).
Org, E. et al. Sex differences and hormonal effects on gut microbiota composition in mice. Gut Microbes 7, 313–322 (2016).
Chella Krishnan, K. et al. Integration of multi-omics data from mouse diversity panel highlights mitochondrial dysfunction in non-alcoholic fatty liver disease. Cell Syst. 6, 103–115 e107 (2018).
Hui, S. T. et al. The genetic architecture of diet-Induced hepatic fibrosis in mice. Hepatology 68, 2182–2196 (2018).
Norheim, F. et al. Gene-by-sex interactions in mitochondrial functions and cardio-metabolic traits. Cell Metab. 29, 932–949 e934 (2019).
Seldin, M. M. et al. A strategy for discovery of endocrine interactions with application to whole-body metabolism. Cell Metab. 27, 1138–1155 e1136 (2018).
Seldin, M. M. et al. A systems genetics approach identifies Trp53inp2 as a link between cardiomyocyte glucose utilization and hypertrophic response. Am. J. Physiol. Heart Circ. Physiol. 312, H728–H741 (2017).
Yin, X. et al. Adipocyte mitochondrial function is reduced in human obesity independent of fat cell size. J. Clin. Endocrinol. Metab. 99, E209–E216 (2014).
Chouchani, E. T. et al. Complex I deficiency due to selective loss of Ndufs4 in the mouse heart results in severe hypertrophic cardiomyopathy. PLoS ONE 9, e94157 (2014).
Begriche, K., Igoudjil, A., Pessayre, D. & Fromenty, B. Mitochondrial dysfunction in NASH: causes, consequences and possible means to prevent it. Mitochondrion 6, 1–28 (2006).
Sanyal, A. J. et al. Nonalcoholic steatohepatitis: association of insulin resistance and mitochondrial abnormalities. Gastroenterology 120, 1183–1192 (2001).
Kim, J. A., Wei, Y. & Sowers, J. R. Role of mitochondrial dysfunction in insulin resistance. Circ. Res. 102, 401–414 (2008).
Mittelstrass, K. et al. Discovery of sexual dimorphisms in metabolic and genetic biomarkers. PLoS Genet. 7, e1002215 (2011).
White, U. A. & Tchoukalova, Y. D. Sex dimorphism and depot differences in adipose tissue function. Biochim. Biophys. Acta 1842, 377–392 (2014).
Yang, X. et al. Tissue-specific expression and regulation of sexually dimorphic genes in mice. Genome Res. 16, 995–1004 (2006).
Karp, N. A. et al. Prevalence of sexual dimorphism in mammalian phenotypic traits. Nat. Commun. 8, 15475 (2017).
Wallace, D. C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: a dawn for evolutionary medicine. Annu Rev. Genet. 39, 359–407 (2005).
Franzen, O. et al. Cardiometabolic risk loci share downstream cis- and trans-gene regulation across tissues and diseases. Science 353, 827–830 (2016).
Consortium, G. T. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Robin, E. D. & Wong, R. Mitochondrial DNA molecules and virtual number of mitochondria per cell in mammalian cells. J. Cell. Physiol. 136, 507–513 (1988).
Laakso, M. et al. The Metabolic Syndrome in Men study: a resource for studies of metabolic and cardiovascular diseases. J. Lipid Res. 58, 481–493 (2017).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016).
Perry, R. J. et al. Hepatic acetyl CoA links adipose tissue inflammation to hepatic insulin resistance and type 2 diabetes. Cell 160, 745–758 (2015).
Lapuente-Brun, E. et al. Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 340, 1567–1570 (2013).
Garcia-Poyatos, C. et al. Scaf1 promotes respiratory supercomplexes and metabolic efficiency in zebrafish. Embo Rep. 21, e50287 (2020).
Cheng, C.-F., Ku, H.-C. & Lin, H. PGC-1α as a pivotal factor in lipid and metabolic regulation. Int. J. Mol. Sci. 19, 3447 (2018).
Lee, J. H. et al. The role of adipose tissue mitochondria: regulation of mitochondrial function for the treatment of metabolic diseases. Int. J. Mol. Sci. 20, 4924 (2019).
Heinonen, S., Jokinen, R., Rissanen, A. & Pietilainen, K. H. White adipose tissue mitochondrial metabolism in health and in obesity. Obes. Rev. 21, e12958 (2020).
Benador, I. Y. et al. Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metab. 27, 869–885.e866 (2018).
Acin-Perez, R. et al. A novel approach to measure mitochondrial respiration in frozen biological samples. EMBO J. 39, e104073 (2020).
Wang, T. et al. Respiration in adipocytes is inhibited by reactive oxygen species. Obesity 18, 1493–1502 (2010).
St-Pierre, J. et al. Suppression of reactive oxygen species and neurodegeneration by the PGC-1 transcriptional coactivators. Cell 127, 397–408 (2006).
Dogan, S. A. et al. Perturbed redox signaling exacerbates a mitochondrial myopathy. Cell Metab. 28, 764–775 e765 (2018).
Moreno-Loshuertos, R. et al. Differences in reactive oxygen species production explain the phenotypes associated with common mouse mitochondrial DNA variants. Nat. Genet. 38, 1261–1268 (2006).
Swierczek, M. et al. An electronic bus bar lies in the core of cytochrome bc1. Science 329, 451–454 (2010).
Tormos, K. V. et al. Mitochondrial complex III ROS regulate adipocyte differentiation. Cell Metab. 14, 537–544 (2011).
Acin-Perez, R., Fernandez-Silva, P., Peleato, M. L., Perez-Martos, A. & Enriquez, J. A. Respiratory active mitochondrial supercomplexes. Mol. Cell 32, 529–539 (2008).
Acı́n-Pérez, R. et al. Respiratory complex III is required to maintain complex I in mammalian mitochondria. Mol. Cell 13, 805–815 (2004).
Maranzana, E., Barbero, G., Falasca, A. I., Lenaz, G. & Genova, M. L. Mitochondrial respiratory supercomplex association limits production of reactive oxygen species from complex I. Antioxid. Redox Signal 19, 1469–1480 (2013).
Nookaew, I. et al. Adipose tissue resting energy expenditure and expression of genes involved in mitochondrial function are higher in women than in men. J. Clin. Endocrinol. Metab. 98, E370–E378 (2012).
Hirst, J. Mitochondrial complex I. Annu. Rev. Biochem. 82, 551–575 (2013).
Fiedorczuk, K. et al. Atomic structure of the entire mammalian mitochondrial complex I. Nature 538, 406–410 (2016).
Zhu, J., Vinothkumar, K. R. & Hirst, J. Structure of mammalian respiratory complex I. Nature 536, 354–358 (2016).
Karamanlidis, G. et al. Mitochondrial complex I deficiency increases protein acetylation and accelerates heart failure. Cell Metab. 18, 239–250 (2013).
Urra, F. A., Munoz, F., Lovy, A. & Cardenas, C. The mitochondrial complex(i)ty of cancer. Front Oncol. 7, 118 (2017).
Papa, S. & De Rasmo, D. Complex I deficiencies in neurological disorders. Trends Mol. Med. 19, 61–69 (2013).
Dröse, S. & Brandt, U. Advances in experimental medicine and biology. Adv. Exp. Med Biol. 748, 145–169 (2012).
Lazarou, M., McKenzie, M., Ohtake, A., Thorburn, D. R. & Ryan, M. T. Analysis of the assembly profiles for mitochondrial- and nuclear-DNA-encoded subunits into complex I. Mol. Cell. Biol. 27, 4228–4237 (2007).
Szczepanowska, K. et al. A salvage pathway maintains highly functional respiratory complex I. Nat. Commun. 11, 1643 (2020).
Latorre-Pellicer, A. et al. Mitochondrial and nuclear DNA matching shapes metabolism and healthy ageing. Nature 535, 561–565 (2016).
Boudina, S. & Graham, T. E. Mitochondrial function/dysfunction in white adipose tissue. Exp. Physiol. 99, 1168–1178 (2014).
Krawczyk, S. A., Haller, J. F., Ferrante, T., Zoeller, R. A. & Corkey, B. E. Reactive oxygen species facilitate translocation of hormone sensitive lipase to the lipid droplet during lipolysis in human differentiated adipocytes. PLoS ONE 7, e34904 (2012).
Masschelin, P. M., Cox, A. R., Chernis, N. & Hartig, S. M. The impact of oxidative stress on adipose tissue energy balance. Front Physiol. 10, 1638 (2019).
Acín-Pérez, R. et al. Fgr kinase is required for proinflammatory macrophage activation during diet-induced obesity. Nat. Metab. 2, 974–988 (2020).
Reaven, G. M. Banting Lecture 1988. Role of insulin resistance in human disease. Diabetes 37, 1595–1607 (1988).
Lusis, A. J., Attie, A. D. & Reue, K. Metabolic syndrome: from epidemiology to systems biology. Nat. Rev. Genet. 9, 819–830 (2008).
Gagnon, D. & Kenny, G. P. Does sex have an independent effect on thermoeffector responses during exercise in the heat? J. Physiol. 590, 5963–5973 (2012).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The SVA package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nat. Methods 8, 833–835 (2011).
Vergnes, L., Chin, R., Young, S. G. & Reue, K. Heart-type fatty acid-binding protein is essential for efficient brown adipose tissue fatty acid oxidation and cold tolerance. J. Biol. Chem. 286, 380–390 (2011).
Rooney, J. P. et al. PCR based determination of mitochondrial DNA copy number in multiple species. Methods Mol. Biol. 1241, 23–38 (2015).
Venegas, V. & Halberg, M. C. Measurement of mitochondrial DNA copy number. Methods Mol. Biol. 837, 327–335 (2012).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Link, V. M., Romanoski, C. E., Metzler, D. & Glass, C. K. MMARGE: Motif Mutation Analysis for Regulatory Genomic Elements. Nucleic Acids Res. 46, 7006–7021 (2018).
O’Neill, S. M. et al. Targeting adipose tissue via systemic gene therapy. Gene Ther. 21, 653–661 (2014).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT–PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034 (2002).
Rogers, G. W. et al. High throughput microplate respiratory measurements using minimal quantities of isolated mitochondria. PLoS ONE 6, e21746 (2011).
Brand, M. D. & Nicholls, D. G. Assessing mitochondrial dysfunction in cells. Biochem. J. 435, 297–312 (2011).
Affourtit, C., Jastroch, M. & Brand, M. D. Uncoupling protein-2 attenuates glucose-stimulated insulin secretion in INS-1E insulinoma cells by lowering mitochondrial reactive oxygen species. Free Radic. Biol. Med. 50, 609–616 (2011).
Osto, C. et al. Measuring mitochondrial respiration in previously frozen biological samples. Curr. Protoc. Cell Biol. 89, e116 (2020).
Garaude, J. et al. Mitochondrial respiratory-chain adaptations in macrophages contribute to antibacterial host defense. Nat. Immunol. 17, 1037–1045 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Acknowledgements
We thank Z. Zhou, Y. Meng, S. Charugundla, D. W. Jayasekera, S. Nand and J. Ure for assistance in experiments. This work was supported by NIH grants NIH-P01HL028481, NIH-R01DK117850 and NIH-R01HL144651 (A.J.L.), NIH-R01HL125863 (J.L.M.B.), NIH-R01HL147187 (C.E.R.), NIH-1R01AA026914-01A1 (M. Liesa), NIH-R00DK120875 (K.C.K) and NIH-K99DK120875 (K.C.K); UCLA/UCSD/CTSI grants P30DK41301 (M. Liesa), UL1TR001881 (M. Liesa), P30DK063491 (M. Liesa); the American Heart Association grants A14SFRN20840000 (J.L.M.B.) and 18POST33990256 (K.C.K.); the Academy of Finland 321428 (M. Laakso); the Swedish Research Council 2018-02529 (J.L.M.B); Heart Lung Foundation 20170265 (J.L.M.B.); Astra-Zeneca through ICMC, Karolinska Institutet, Sweden (J.L.M.B.) and the Foundation Leducq 12CVD04 (L.V. and K.R.) and 18CVD02 (J.L.M.B.). The funders had no role in the study design, data collection and interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
K.C.K. and A.J.L. conceived the study. K.C.K., L.V., R.A.P., L.S., M.S., L.M., E.M., C.P., T.M.M., M.P., C.E.R., K.R., J.L.M.B., M. Laakso, M.Liesa and A.J.L. performed experiments or analysed the data. K.C.K. and A.J.L. drafted the manuscript. All authors read or revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary handling editors: George Caputa and Isabella Samuelson. Nature Metabolism thanks Lawrence Kazak and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chr17 locus is not associated with liver mitochondria.
Related to Fig. 3. High-resolution association mapping of 911 (1312 probes) mitochondrial gene abundance levels from (a) male and (b) female liver tissues isolated from ~100 HMDP strains to identify eQTL networks. The X- and Y- axis represent SNP and gene position on the mouse genome, respectively. Each dot represents a significant association. (c) Frequency distributions of P values of association between the chr17 trans-eQTL lead eSNP (rs48062344) and mitochondria-related transcripts in the male and female liver tissues. P values of association were calculated using (A – C) FaST-LMM that uses Likelihood-Ratio test.
Extended Data Fig. 2 Chr17 trans-eQTL do not affect the expression profiles of OXPHOS genes in male adipose or both sexes of liver tissues.
Related to Fig. 4. Volcano plots showing genetic (lead SNP of chr17 locus) differences in the expression profiles of OXPHOS genes in the (a) male adipose (n = 98 male strains; CC: 56 & TT: 42); (b) female and (c) male liver (n = 97 sex-matched strains; CC: 55 & TT: 42) isolated from HMDP. Genes corresponding to individual OXPHOS complexes are color-coded. Horizontal dotted lines represent 5% FDR-corrected significance threshold. Data are presented as log2 fold change between genotype. P values were calculated using (A – C) DESeq2 Bioconductor package that uses Wald test.
Extended Data Fig. 3 Association mapping of adipose mtDNA levels in HMDP.
Related to Fig. 4. Association mapping of mtDNA levels from (a) female and (b) male adipose in the HMDP cohort (n = 216 female and 260 male mice) Red line represents significance threshold (HMDP: P = 4.1E-06). P values of association were calculated using (A – B) FaST-LMM that uses Likelihood-Ratio test.
Extended Data Fig. 4 Liver Ndufv2 is unaffected by the chr17 trans-eQTL locus.
Related to Fig. 5. Association mapping of (a) male and (b) female liver Ndufv2 expression from HMDP. Red line represents significance threshold (P = 4.1E-06). (c – e) Sex and genetic (lead SNP of chr17 locus) differences in the liver Ndufv2 expression from HMDP (n = 97 sex-matched strains; CC: 55 & TT: 42). Data are presented as median and interquartile range (boxplots). P values were calculated using (A – B) FaST-LMM that uses Likelihood-Ratio test; (C – E) Unpaired two-tailed Student’s t test.
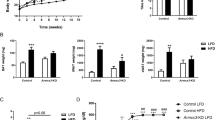
Extended Data Fig. 5 Changes in body weight and insulin sensitivity mediated by adipose Ndufv2 overexpression.
Related to Fig. 6. Eight-week old females or males of (a – b) C57BL/6 J or (c – j) A/J mice were injected with AAV vectors expressing either GFP or NDUFV2 in an adipose-specific manner and fed a HF/HS diet for eight additional weeks. Changes in (A - D) fat mass and total body weight measured every two-weeks (females n = 7 per group; males n = 8 per group), (E and H) ITT, (F and I) end-point HOMA-IR and (G and J) fasting insulin levels (F – A/J n = 8 per group; M – A/J GFP n = 7, NDUFV2 n = 8) are shown. Data are presented as mean ± SEM. P values were calculated using (A – E and H) Repeated measures 2-factor ANOVA corrected by post-hoc ‘Holm-Sidak’s’ multiple comparisons test; (F, G, I and J) Unpaired two-tailed Student’s t test.
Extended Data Fig. 6 Adipose Ndufv2 overexpression regulated adiposity in a sex-by-strain manner.
Related to Fig. 6. Eight-week old males of (a – e) C57BL/6 J or (f – j) A/J mice were injected with AAV vectors expressing either GFP or NDUFV2 in an adipose-specific manner and fed a HF/HS diet for eight additional weeks. Metabolic traits such as (A and F) total mass and food intake; (B and G) fat and lean mass were monitored over eight weeks. Comparisons of (C and H) adipose Ndufv2 expression; (D and I) plasma free glycerol; (E and J) tissue weights from C57BL/6 J and A/J males, respectively. Data are presented as mean ± SEM (n = 8 per group). P values were calculated using (A and F) Repeated measures 2-factor or (B and G) 3-factor or (E and J) 2-factor ANOVA corrected by post-hoc ‘Holm-Sidak’s’ multiple comparisons test; (C, D, H and I) Unpaired two-tailed Student’s t test.
Extended Data Fig. 7 Adipose Ndufv2 overexpression regulated adipocyte size in a sex-by-strain manner.
Related to Fig. 7. Comparisons of adipocyte size distribution in the gonadal adipose tissues from females and males of (a – b) C57BL/6 J or (c – d) A/J mice overexpressing GFP or NDUFV2, respectively. Representative histological sections are shown for each group (scale: 100 µm). Red and blue shades (or bars) represent female and male NDUFV2 datapoints, while brown represents respective GFP datapoints, respectively. Data are presented as frequency distribution of adipocyte sizes or mean ± SEM (F – C57BL/6 J GFP n = 5064 cells from 7 mice, NDUFV2 n = 5249 cells from 7 mice; M – C57BL/6 J GFP n = 11721 cells from 8 mice, NDUFV2 n = 7290 cells from 7 mice; F – A/J GFP n = 5423 cells from 7 mice, NDUFV2 n = 4687 cells from 7 mice; M – A/J GFP n = 4084 cells from 8 mice, NDUFV2 n = 4863 cells from 8 mice). P values were calculated using Unpaired two-tailed Student’s t test.
Extended Data Fig. 8 Adipose Ndufv2 overexpression regulated mitochondrial respiration in a sex-by-strain manner.
Related to Fig. 7. Comparisons of mitochondrial RCR and coupling efficiency in the gonadal adipose tissues from males of (a – b) C57BL/6 J or (c – d) A/J mice overexpressing GFP or NDUFV2, respectively. Coupling assays of isolated gonadal adipose mitochondria from females and males of (e – f) C57BL/6 J or (g – h) A/J mice overexpressing GFP or NDUFV2, respectively. Data are presented as mean ± SEM (n = 4 per group). P values were calculated using 2-factor ANOVA.
Extended Data Fig. 9 Adipose Ndufv2-mediated mitochondrial regulation is not a consequence of body weight.
Related to Fig. 7. Eight-week old female C57BL/6 J mice were fed a HF/HS diet for the first six weeks without any intervention, after which were injected with AAV vectors expressing either GFP or NDUFV2 in an adipose-specific manner and diet continued for six additional weeks. Metabolic traits such as (a) total mass; (b) fat and lean mass were monitored over 12 weeks. Comparisons of (c) tissue weights; (d) adipose Ndufv2 expression; (e) mitochondrial RCR; (f) coupling efficiency and (g) coupling respiration rates between GFP and NDUFV2 animals, respectively. Data are presented as mean ± SEM (n = 6 per group). P values were calculated using (A) Repeated measures 2-factor or (B) 3-factor or (C and G) 2-factor ANOVA corrected by post-hoc ‘Holm-Sidak’s’ multiple comparisons test; (D – F) Unpaired two-tailed Student’s t test.
Extended Data Fig. 10 Ndufv2 overexpression regulated mitochondrial function in both AML12 (liver) and differentiated 3T3-L1 (adipose) cells.
Related to Fig. 8. Comparisons between GFP and NDUFV2 overexpressing AML12 (liver) cells in (a) relative normalized expression values of Ndufv2 (n = 9 per group); Coupling assays and RCR with all three complex I substrates (Pyruvate, Palmitoyl carnitine and Glutamate) either added (b – e) together (n = 5 per group) or (f - m) separately (n = 4 per group except Glutamate, n = 8 per group); and (N) relative normalized expression values of Ndufs4 (complex I), Sdhc (complex II), Atp5a1 (complex V) and Cpt1a (FAO) (n = 9 per group). Similarly, comparisons between control and NDUFV2 overexpressing differentiated 3T3-L1 cells in (O – R) coupling assays and RCR with different substrates (Pyruvate, Palmitoyl carnitine and Succinate) added separately (n = 6 per group). Data are presented as mean ± SEM. P values were calculated using (A and J – N) Unpaired two-tailed Student’s t test; (B – I and O – R) 2-factor ANOVA corrected by post-hoc ‘Holm-Sidak’s’ multiple comparisons test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Tables 4 and 5
Supplementary Table 1
Supplementary Tables 1–3
Source data
Source Data Fig. 7
Unprocessed western blots
Source Data Fig. 8
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Chella Krishnan, K., Vergnes, L., Acín-Pérez, R. et al. Sex-specific genetic regulation of adipose mitochondria and metabolic syndrome by Ndufv2. Nat Metab 3, 1552–1568 (2021). https://doi.org/10.1038/s42255-021-00481-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00481-w
This article is cited by
-
Sarcopenic obesity is part of obesity paradox in dementia development: evidence from a population-based cohort study
BMC Medicine (2024)
-
Sex differences in energy metabolism: natural selection, mechanisms and consequences
Nature Reviews Nephrology (2024)
-
Mitochondrial heterogeneity in diseases
Signal Transduction and Targeted Therapy (2023)
-
Maternal high-fat diet programs white and brown adipose tissue lipidome and transcriptome in offspring in a sex- and tissue-dependent manner in mice
International Journal of Obesity (2022)
-
Sex differences in the intergenerational inheritance of metabolic traits
Nature Metabolism (2022)