Abstract
SINE-VNTR-Alu (SVA) retrotransposons arose and expanded in the genome of hominoid primates concurrent with the slowing of brain maturation. We report genes with intronic SVA transposons are enriched for neurodevelopmental disease and transcribed into long non-coding SVA-lncRNAs. Human-specific SVAs in microcephaly CDK5RAP2 and epilepsy SCN8A gene introns repress their expression via transcription factor ZNF91 to delay neuronal maturation. Deleting the SVA in CDK5RAP2 initiates multi-dimensional and in SCN8A selective sodium current neuronal maturation by upregulating these genes. SVA-lncRNA AK057321 forms RNA:DNA heteroduplexes with the genomic SVAs and upregulates these genes to initiate neuronal maturation. SVA-lncRNA AK057321 also promotes species-specific cortex and cerebellum-enriched expression upregulating human genes with intronic SVAs (e.g., HTT, CHAF1B and KCNJ6) but not mouse orthologs. The diversity of neuronal genes with intronic SVAs suggest this hominoid-specific SVA transposon-based gene regulatory mechanism may act at multiple steps to specialize and achieve neoteny of the human brain.
Similar content being viewed by others
Introduction
Advances in Homo sapiens cognitive ability compared to other primates occurred in parallel with changes in brain structure (e.g., expanded cortex and cerebellum) and function. The brain’s core functional element, the glutamatergic neuron, displays neoteny or delayed maturation in human compared to other primates. Delayed neuronal maturation is considered a possible cellular basis of the extended brain growth that contributes to increased neuron numbers, size, and connectivity of the human brain1,2,3. Maturation of glutamatergic neurons from induced pluripotent stem cells (iPSCs) is delayed in human compared to its closest primate relatives, chimpanzee and bonobo4. Yet the molecular basis of this human neuronal neoteny is undefined.
SINE-VNTR-Alu (SVA) transposons are hominoid-primate-specific mobile genetic elements that transposed and became fixed in the genome during hominoid evolution5,6. As human-specific fixed transposons are considered a potential source of gene regulation that could underlie Homo Sapiens species divergence7,8,9, we postulated SVAs, particularly those unique to human, might underlie the delayed neuronal maturation found in human.
Transposable elements are also enriched in expressed transcripts of long non-coding (lnc) RNAs10, suggesting those containing SVA transposons might also have a role in hominoid-specific, transposon-based gene regulatory mechanisms.
We find that as human neurons mature, there is an increase of glutamatergic neuron-enriched SVA-lncRNA, AK057321. We increase AK057321 expression to evaluate its effects on the expression of neurodevelopmental genes with intronic SVAs. These studies were performed in a human neuronal progenitor cell to assess the effects of these interventions on neuronal maturation. We also examine effects of intervening on KRAB-domain zinc finger transcription factor, ZNF91, that binds SVA sequences and recruits protein complexes to repress genes bearing an SVA11,12. To evaluate if intronic SVAs repress neurodevelopmental genes as a potential means to delay neuronal maturation, we deleted the gene’s intronic SVAs. To complement these in vitro human neuronal progenitor cell studies, we also use humanized mice to examine whether SVA transposon-based gene regulation occurs in the brain in vivo. Finally, to begin to understand the molecular basis of this SVA transposon-based gene regulatory mechanism we evaluated for the possibility that complementary annealing of nucleic acid strands occurs between genomic and AK057321 lncRNA SVA sequences. We also examine if the genomic SVA-binding transcription factor ZNF9111,12 binds SVA-lncRNA AK057321 as a potential mechanism of displacing the transcriptional repressor.
Altogether, these studies reveal that human-specific intronic SVA transposons together with SVA-binding transcription factor ZNF91 repress expression of human brain development genes (e.g., CDK5RAP2 and SCN8A) to delay neuronal maturation. SVA sequences encoded in AK0572321 of the SVA-lncRNA gene family, possibly through RNA:DNA heteroduplex formation and ZNF91 decoy-binding, release SVA-mediated gene repression to time the initiation of neuronal maturation.
Results
SVA-lncRNA AK057321 expression increases during glutamatergic neuron maturation to upregulate expression of neurodevelopmental disease genes with intronic SVAs
SVA transposons are unique to the great ape (hominoid) primate species lineage that includes the extant human, chimpanzee, gorilla, and orangutan (Fig. 1a)5,6. SVA-lncRNA AK057321 (NONHSAT119982.2 in www.noncode.org), that is duplicated in rare cases of autism spectrum disorder (Supplementary Data 1), is found in human, chimpanzee, bonobo, and gorilla species and is encoded by three exons including a full-length SVA transposon sequence at the 3′ end (Fig. 1b, Supplementary Fig. 1, NONHSAT119982.2). SVA-lncRNA AK057321 expression is strongest in brain and testis (Supplementary Fig. 2 and Supplementary Data 2; enriched in cortex and cerebellum, see HTT gene regulatory studies below; see also the expression of NONHSAT119982.2 in www.noncode.org). We observed that SVA-lncRNA AK057321 expression increases 2-fold in NTera-2 progenitor cells as they are matured into neurons using Ngn2 transcription factor and astrocyte co-culture (Fig. 1c and Supplementary Data 3) and 10-fold as pluripotent human iPSC cells differentiate into glutamatergic neurons (Supplementary Fig. 3). Ngn2 and astrocyte co-culture promoted neuronal differentiation of NTera-2 progenitor cells as confirmed by the hyperpolarized resting membrane potential (Fig. 1d), the generation of sodium current-dependent action potential (Fig. 1d), and the visible neurite outgrowth (Fig. 1e). These results establish that SVA-lncRNA AK057321 expression increases as neuronal progenitors mature into glutamatergic neurons and validate the feasibility of using the human NTera-2 pluripotent cell line for studies of neuronal maturation.
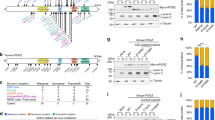
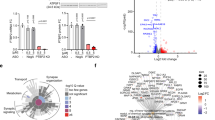
a Schematic showing SVA subtypes in primate species. b Schematic of gene AK057321, a long non-coding RNA (lncRNA) expressing a full-length SVA (UCSC genome browser, GRCh38/gh38). c–e Ngn2-driven neuronal differentiation of NTera-2 cells increases SVA-lncRNA AK057321 and CDK5RAP2 expression (c plated on astrocytes six weeks, comparing to non-treated NTera-2 cells with GAPDH as the internal reference gene, unpaired two-tailed Student’s t test, n = 2 biological replicates each measured in triplicate), generates Na+ action potentials and hyperpolarizes membrane potential (d), and increases TuJ1 protein and neurite outgrowth (e, expressed GFP and neuron-specific Class III β-tubulin immunostaining). Unpaired two-tailed Student’s t test. f Gene ontology (GO) analysis of human genes with intronic SVAs: biological process (purple bars) and disease (blue bars) term enrichment. g NTera-2 cells transfected with sh-AK057321 or sh-scramble for 24 h were sorted and analyzed for gene expression by RT-qPCR. HUNK and LSS are controls without SVAs. One-way ANOVA (F63, 298 = 11.37, p < 0.0001; n = 5 to 6 biological replicate samples were pooled and measure using n = 3 technical replicates per gene) with Bonferroni post hoc tests. Data represent the mean ± SEM. h Schematic for Crispr/Cas9 SVA_B deletion strategy with DNA agarose gel image showing expected SVA-deletion PCR product size of 269 bp present only with set 2 guides which were used for all subsequent experiments. i NTera-2 cells transfected with Cas9/sgRNAs to delete the SVA in AK057321 (vs. control, right) were sorted and analyzed for gene expression by RT-qPCR. CDK5RAP2, SCN8A, and CHAF1B expression are decreased. n = 8 biological replicates. One-way ANOVA (F7,56 = 92.36, p < 0.0001) with Bonferroni’s post hoc tests. j NTera-2 cells were transfected with sh-AK057321 or sh-scramble (left) or NTera-2 cells were transfected with Cas9/sgRNAs to delete the SVA in AK057321 (vs. control, right). After three days, cells were sorted and analyzed for gene expression by RT-qPCR with GAPDH as the internal reference gene. n = 3 biological replicates. Unpaired Student’s t test. One-way ANOVA for sh-AK057321 vs sh-scramble (F5,12 = 6.809, p = 0.0031) and one-way ANOVA for Cas9/sgRNAs (F7,16 = 10.24, p < 0.0001. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05 versus control.
Using gene ontology term enrichment analysis, we found that human genes containing intronic SVAs are enriched for those involved in neurodevelopment and neurodevelopmental and neurological disease (Fig. 1f, Supplementary Data 4–6; enrichment for other biological function and disease gene ontology terms also seen). Functionally important genes underlying microcephaly, neuron dendrite and axon growth, synapse formation, and ion channels are found to contain intronic SVAs (Supplementary Figs. 4–7), with the SVA sequences frequently being human-specific, and sometimes displaying species-specific expansions (e.g., ATRX, KDM6A), tandem insertions (CDK5RAP2, KDM6A, RAB2A, IL1RAPL1, KCNH1) or even repeated integrations at multiple sites in the same gene (KDM6A, ATP8A2, GRID1, NRXN1). These findings are consistent with a bioinformatics study suggesting SVAs are associated with genes involved in central nervous system pathways13.
Based on the SVA sequence complementarity in SVA-lncRNA AK057321 and these intronic SVAs, we postulated SVA-lncRNA AK057321 might regulate coding genes with intronic SVAs. To test this, we used a short hairpin (sh)RNA to deplete AK057321 mRNA and measured expression of several genes with intronic SVAs in the displayed categories of interest. Depleting AK057321 strongly downregulates microcephaly gene CDK5RAP2 (human-specific tandem SVA_F, Supplementary Fig. 4) and epileptic encephalopathy gene SCN8A (human-specific SVA_D, Supplementary Fig. 7), as well as several other SVA-containing genes involved in neurogenesis, neuron excitability, synapse and dendrite formation, and mitochondrial function (Fig. 1g). By contrast, genes lacking SVAs, HUNK and LSS, were unaffected. Confirming the results with the shRNA, Cas9/sgRNA-mediated deletion of SVA-lncRNA AK057321 (exon 3) reduces SVA-lncRNA AK057321 expression while downregulating CDK5RAP2, SCN8A, and CHAF1B gene expression (Fig. 1h, i and Supplementary Fig. 8). These results establish that mimicking developmental increases of SVA-lncRNA AK057321 promotes expression of neurodevelopmental genes with intronic SVAs.
Cas9 sgRNA-mediated deletion of SVA-lncRNA AK057321 (exon 3) increases pluripotency marker gene expression (Fig. 1j; POU5F1 and NANOG three days post transfection). These results suggest SVA-lncRNA AK057321 promotes the switch from progenitor to neuron.
Neuronal maturation is initiated by increasing SVA-lncRNA AK057321 or deleting transcription factor ZNF91 that binds SVAs to repress gene expression
Based on our observations that SVA-lncRNA AK057321 increases during neuronal maturation (Fig. 1c, Supplementary Fig. 3) and increases expression of neuronal genes with intronic SVAs such as the microcephaly gene CDK5RAP2 (Fig. 1c, g–i), we reasoned SVA-lncRNA AK057321 might function to initiate neuronal maturation by relieving the repression of genes by their intronic SVAs. For a second independent test of the potential role of SVAs in human neuronal maturation, we also investigated the effects of manipulating ZNF91, a primate-specific Krüppel-associated box (KRAB) domain containing zinc finger protein (KZNF) that binds to intronic SVA sequences to repress gene expression11,14,15. Importantly, ZNF91 is expressed in human neuronal progenitor NTera-2 cells16. Deleting ZNF91 gene in human embryonic stem cells increases the expression of genes located near SVA sequences12. Prior luciferase reporter studies have shown ZNF91 binds SVAs to repress gene expression11,16. We predicted deletion of ZNF91 would increase expression of genes containing intronic SVA sequences in human NTera-2 progenitor cells.
We compared the effects of deleting ZNF91 to those of increasing SVA-lncRNA AK057321, each intervention hypothesized to increase expression of genes repressed by ZNF91 repressor complex binding to intronic SVA sequences (Fig. 2a). Over-expressing SVA-lncRNA AK057321 or deleting ZNF91, each independently, increases CDK5RAP2 gene expression (Fig. 2b, c), a microcephaly gene with an intronic human-specific SVA_F that increases during neuronal maturation (Fig. 1c).
a schematic illustrates ZNF91 transcription factor binds to the Alu-VNTR and VNTR-SINE borders within SVAs12 and inhibits nearby gene expression11,12,16. When SVA-lncRNA AK057321 expression is increased, or ZNF91 is deleted, SVA-containing genes are upregulated, and stem/progenitor cells mature into neurons (Created with Biorender.com). b AK057321 and CDK5RAP2 expression in NTera-2 cells transfected with either SVA-lncRNA AK057321 cDNA (OE) or control plasmid for 24 h (GAPDH, reference; n = 6 biological replicates). Unpaired Student’s t test. c, Gene regulation by Cas9/sgRNA deletion of ZNF91 in NTera-2 cells with GAPDH as the internal reference gene. n = 5 biological replicates. Unpaired Student’s t test. d Representative inward current traces in ZNF91 sgRNA and AK057321 OE treatment groups in response to step depolarization (−90 mV to 0 mV). Inward currents were recorded in voltage clamp mode. e Tetrodotoxin (TTX) sensitive inward currents (control vs. TTX) were significantly larger in AK057321 OE and ZNF91 sgRNA and treatment groups than that in the control group. f, g Resting membrane potentials (RMP) is hyperpolarized in AK057321 OE (n = 7) and ZNF91 sgRNA (n = 7) compared to control. h, i Cell capacitance is increased in AK057321 OE (n = 7) and ZNF91 sgRNA (n = 7) compared to control. j, k Representative images of NTera-2 cell morphology with or without AK057321 overexpression 15 days after plating. Scale bar represents 20 μm. l Sholl analysis on the processes of Ntera-2 cells with or without AK057321 OE. m Quantification of cell area from control and AK057321 OE in NTera-2 cells. n Quantification of average mean length of processes. o Quantification of circularity based on cell morphology. p Quantification of average maximum length of processes. All graphs are presented as Mean ± SEM. Control, N = 31, AK overexpression, N = 25. In (l), ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05 two-way ANOVA with Sidak’s post hoc test. b, c, e–i, m–p Unpaired two-tailed Student’s t test. Data represent the mean ± SEM.****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05 versus control.
To quantify neuronal maturation, we measure neurophysiologic (sodium and potassium currents, resting membrane potential, membrane capacitance, and action potential firing) and morphologic (cell size, shape and dendrite formation) features of the maturing neuron17,18,19. We found human NTera-2 progenitor cells undergo neuronal maturation in response to transcription factor Ngn2 when co-cultured with astrocytes (Fig. 1c–e). We observed that increasing SVA-lncRNA AK057321 expression or deleting ZNF91 gene in human NTera-2 progenitor cells co-cultured with astrocytes (but without added Ngn2) initiates early stages of neuronal maturation. Both interventions increase inward voltage-gated sodium current (Fig. 2d) and outward potassium current (Supplementary Fig. 9). The sodium current is inhibited by voltage-gated sodium channel blocker tetrodotoxin (Fig. 2e). These interventions also hyperpolarize resting membrane potential (RMP; Fig. 2f, g) and increase membrane capacitance (Fig. 2h, i). Consistent with the increased membrane capacitance reflecting increased membrane area, we observed neurite outgrowth, a morphologic property of maturing neurons, by confocal microscopy in cells treated with SVA-lncRNA AK057321 (Fig. 2j, k). Quantification of neuronal morphology (Sholl analysis) confirmed SVA-lncRNA AK057321 overexpression increases process length (Fig. 2l), cell area (Fig. 2m), and average mean and maximum process length (Fig. 2n, p) while decreasing circularity (Fig. 2o). SVA-lncRNA AK057321 also increases average mean length and minimum length of processes, and average process and tip number, while increasing microtubule-associated protein 2 (MAP2), a marker of neuronal differentiation (Supplementary Fig. 10). The results indicate increasing SVA-lncRNA AK057321 or depleting ZNF91 increases CDK5RAP2 expression while initiating functional and structural neuron maturation.
SVA-lncRNA AK057321 initiates neuronal maturation by increasing CDK5RAP2 expression otherwise repressed by an intronic tandem SVA_F
We reasoned that removing specific genomic SVAs might identify the pathway whereby SVA-lncRNA AK057321 drives neuronal differentiation. We first focused on CDK5RAP2 because of its major role in human microcephaly and requirement during neurogenesis20,21,22,23,24,25,26, its human-specific tandem SVA_F (Fig. 3a), and its strong downregulation with AK057321 loss (Fig. 1g–i). CDK5RAP2 expression was also upregulated by multiple interventions that promote neuronal maturation [increased expression of Ngn2 (Fig. 1c), increased SVA-lncRNA AK057321 expression (Fig. 2b) or decreased ZNF91 expression (Fig. 2c)].
a Tandem human-specific SVA_F in microcephaly gene CDK5RAP2 gene intron (UCSC genome browser, GRCh38/gh38). b Schematic for Crispr/Cas9 tandem SVA_F, F deletion strategy with DNA agarose gel image showing expected SVA-deletion PCR product size of 947 bp present only in set 2 guides which were used for all subsequent experiments. c, NTera-2 cells transfected with set 2 guides were sorted and analyzed for gene expression by RT-qPCR. n = 4 biological replicates. d RT-qPCR of NTera-2 cells transfected with CDK5RAP2 shRNA show decreased CDK5RAP2 mRNA expression compared to control cells. n = 7 biological replicates. e RT-qPCR from NTera-2 cells transfected with CDK5RAP2 cDNA and induced with doxycycline have increased CDK5RAP2 gene expression. n = 8 biological replicates. c–e Unpaired two-tailed Student’s t test for each with GAPDH as the internal reference gene. f TTX-sensitive currents were recorded in voltage clamp mode. Representative sodium current traces in response to step depolarization were shown. g TTX-sensitive inward currents were larger in AK057321 OE (n = 7), CDK5RAP2-OE (n = 7) and CDK5RAP2-SVA del (n = 6) del groups than those in the control group. AK057321 OE + CDK5RAP2 shRNA has no significant difference compare to control. One-way ANOVA (F5,36 = 29.44, p < 0.0001) with Bonferroni post hoc tests. h Resting membrane potential (RMP) is hyperpolarized in AK057321 OE (n = 7), CDK5RAP2-OE (n = 7) and CDK5RAP2-SVA del (n = 6) del groups than those in the control group. AK057321 OE + CDK5RAP2 shRNA has no significant difference compared to control. One-way ANOVA (F5,36 = 29.44, p < 0.0001) with Bonferroni post hoc tests. i Capacitance was larger in AK057321 OE (n = 7), CDK5RAP2-OE (n = 7) and CDK5RAP2-SVA del (n = 6) del groups than those in the control group. AK057321 OE + CDK5RAP2 shRNA has no significant difference compared to control. One-way ANOVA (F5,36 = 29.44, p < 0.0001) with Bonferroni post hoc tests. Data represent the mean ± SEM. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05.
Using the CRISPR/Cas9-sgRNA approach, we deleted the tandem SVA_F in CDK5RAP2 in NTera-2 cells (Fig. 3a, b and Supplementary Fig. 11). Loss of the intronic SVA sequences increases CDK5RAP2 expression (Fig. 3c), underscoring the inhibitory effect of this tandem SVA_F on CDK5RAP2 expression. The intervention also decreases stem/progenitor cell gene expression (NANOG and POU5F1; Fig. 3c), suggesting deletion of this 2.4 kb intronic sequence might suffice to initiate neuronal maturation.
As an additional test of the role of CDK5RAP2 expression on neuronal maturation, we used an shRNA to decrease, and a cDNA to increase CDK5RAP2 expression (RT-qPCR, Fig. 3d, e). Because CDK5RAP2 overexpression is toxic26, CDK5RAP2 cDNA was expressed from a Tet-On controlled promoter (RT-qPCR, Fig. 3e).
As observed previously (Fig. 2d, e), and repeated experimentally here, increasing SVA-lncRNA AK057321 increases inward voltage-gated sodium currents (Fig. 3f, g). Given that SVA-lncRNA AK057321 increases CDK5RAP2 expression (Fig. 2b), we blocked this increase with the CDK5RAP2 shRNA (Fig. 3d, f–i). CDK5RAP2 shRNA blocked AK057321-induced increases of sodium currents (Fig. 3f, g). Moreover, over-expressing CDK5RAP2 itself increases sodium currents (Fig. 3f, g). Most importantly, deleting just the tandem SVA_F sequences in CDK5RAP2 increases sodium currents (Fig. 3f, g). As observed in response to increases of SVA-lncRNA AK057321 or decreases of ZNF91 expression, the interventions that increase CDK5RAP2 expression (overexpression and SVA deletion) also hyperpolarize resting membrane potential (Fig. 3h) and increase membrane capacitance (Fig. 3i). CDK5RAP2 shRNA blocks the SVA-lncRNA AK057321-induced hyperpolarization (Fig. 3h) and increase of membrane capacitance (Fig. 3i). These results show the human-specific intronic tandem SVA_F in CKD5RAP2 represses its expression to block neuronal maturation and SVA-lncRNA AK057321 releases this block by derepressing CDK5RAP2.
Deleting the human-specific SVA_D in SCN8A selectively increases voltage-gated sodium current
We reasoned that intronic SVAs in other genes regulated by SVA-lncRNA AK057321 (Fig. 1g) might delay specific aspects of neuronal maturation and chose to examine SCN8A because it encodes the pore forming α subunit of a tetrodotoxin-sensitive, voltage-gated sodium channel and is needed to generate action potentials during early neuronal maturation27. SCN8A’s role in early neuronal excitability is evidenced by the ability of gain-of-function mutations to cause early infantile epileptic encephalopathy and loss-of-function mutations to cause intellectual disability28,29,30. Increasing and decreasing SVA-lncRNA AK057321 was found to upregulate and down-regulate, respectively, SCN8A expression (Supplementary Fig. 12a, b, Fig. 1g–i). Moreover, CRISPR/Cas9-sgRNA-mediated deletion of ZNF91 upregulates SCN8A expression (Supplementary Fig. 12c). To determine if the human-specific intronic SVA in SCN8A impacts any aspects of neuronal maturation as observed for CDK5RAP2, we used CRISPR/Cas9-sgRNA to delete this SVA in NTera-2 cells co-cultured with astrocytes (Fig. 4a and Supplementary Figs. 12–13). We analyzed TTX-sensitive sodium currents in voltage clamp mode and representative traces are shown (Fig. 4b). Deleting the SVA in SCN8A increases sodium currents (Fig. 4c) without altering resting membrane potential (Fig. 4d) or capacitance (Fig. 4e). The results suggest the human-specific intronic SVA_D in SCN8A selectively delays maturation of voltage-gated sodium channel currents.
a Schematic showing SVA_D located within the SCN8A gene is a human-specific insertion (UCSC genome browser, GRCh38/gh38) and strategy for Crispr/Cas9 SVA_D deletion strategy with DNA agarose gel image showing expected SVA-deletion PCR product size of 515 bp present only with transfection of SCN8A guides which were used for all subsequent experiments. b Representative sodium current traces in SCN8A-SVA del group in response to step depolarization were shown. c TTX-sensitive current in SCN8A-SVA del group (n = 7) is higher than in control (n = 7). Unpaired Student’s t test. ***p < 0.001. d Resting membrane potential (RMP) in SCN8A-SVA del (n = 7) and control (n = 7) group has no significant difference. Unpaired Student’s t test. e, Capacitance in SCN8A-SVA del (n = 7) and control (n = 7) group has no significant difference. Unpaired Student’s t test. Whole-cell patch-clamp recording of NTera-2 cell up to day 15 in culture in different treatment groups: f Current injection (300 pA) at day 15. g Change in voltage (Delta V). h TTX-sensitive inward currents. i Resting membrane potential. j Cell membrane capacitance. Two-way ANOVA with Bonferroni post hoc tests. Data represent the mean ± SEM. ****p < 0.0001, ***p < 0.001, ns p > 0.05.
We compared the time course of neuronal maturation across different interventions to understand their kinetics and to evaluate if these parameters change spontaneously in control cells. SVA-lncRNA AK057321 overexpression, ZNF91 depletion, SCN8A SVA deletion, CDK5RAP2 overexpression, or CDK5RAP2 SVA deletion (but not control or CHAF1B SVA deletion, see also below) increase sodium spike amplitude over the 15-day time period (Fig. 4f, g and Supplementary Fig. 12d, e). Increases of inward voltage-sensitive sodium currents displayed a similar time course across all interventions but the increase is absent in control and with CHAF1B SVA deletion (Fig. 4h). Importantly, unlike the other interventions, SCN8A SVA deletion did not hyperpolarize resting membrane potential or increase membrane capacitance at any time point, consistent with its specialized role in regulating voltage-gated sodium currents (Fig. 4i, j).
Deleting the SVA_B in CHAF1B increases its expression and promotes proliferation and stem cell gene expression
Some intronic SVAs may control functions beyond neuronal maturation. We examined functions of the SVA in chromatin assembly factor 1B (CHAF1B) because of its intronic SVA_B (Supplementary Fig. 14a) and strong downregulation when SVA-lncRNA AK057321 is depleted (Fig. 1g–i). CHAF1B is a component of a complex required for DNA replication and chromatin formation31,32,33 that promotes proliferation33 and maintains cell identity34. Deleting SVA-repressive ZNF91, or increasing SVA-lncRNA AK057321 upregulates CHAF1B expression (Supplementary Fig. 14b). Deleting the SVA_B in CHAF1B (CRISPR/Cas9-gRNA technique, Supplementary Fig. 14c and Supplementary Fig. 15) also increases CHAF1B expression and increases expression of pluripotency-promoting genes (POU5F1, NANOG, and SOX2) while reducing expression of neuron-differentiation promoting transcription factor FOXG1 (Supplementary Fig. 14d). Deleting the CHAF1B SVA also fails to increase sodium spikes (Fig. 4f, 15 days) or currents (Supplementary Fig. 14e, f), to hyperpolarize resting membrane potential (Supplementary Fig. 14g), or to increase membrane capacitance (Supplementary Fig. 14h). Interestingly, deleting the SVA in CHAF1B (but not in CDK5RAP2 or SCN8A) also increases the number of cells at 3 and 5 days following the intervention (Supplementary Fig. 14i–n). These results suggest the intronic SVA in CHAF1B may repress its expression to inhibit proliferation of a stem cell pool generated by human NTera-2 progenitor cells.
Human-specific cortex and cerebellum-enriched expression of Huntingtin’s disease gene HTT with an intronic SVA is driven by SVA-lncRNA AK057321
HTT mRNA is expressed in the developing and adult brain in both mice and humans35,36 and contains an intronic SVA_B shared with chimpanzee and gorilla (Fig. 5a, length expanded in human and chimpanzee). Dominantly inherited mutations of huntingtin (HTT) that expand a poly-glutamine tract in exon 1 of the HTT protein underlie the neurological disorder Huntingtin’s disease (HD)37 with degeneration of medium spiny neurons of the striatum, motor deficits, and impaired cognitive executive functioning37. To investigate if SVA-lncRNA AK057321 not only acts during neuronal maturation but also in the fully mature brain to regulate expression of genes with an intronic SVA, we generated transgenic mice with a full-length human SVA-lncRNA AK057321 gene [derived from a bacterial artificial chromosome (BAC) vector human gene library, corresponding to the region duplicated in autism38 (Fig. 1b, Supplementary Data 1, and Supplementary Fig. 2)]. We then crossed these SVA-lncRNA AK057321 transgenic mice to mice containing the human HTT transgene (carried in yeast artificial chromosome vector and bearing an HD model mutation, YAC128 mice39).
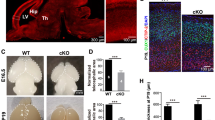
a Illustration of SVA_B retrotransposon located within the HTT gene, UCSC genome browser (GRCh38/gh38). b Native SVA-lncRNA AK057321 transcript levels differ across human brain regions compared to levels in Bs. One-way ANOVA (F9,20 = 43.19, versus control Bs). c HTT transcript analysis with quantitative RT-qPCR on human brain tissue shows HTT transcript expression varies across human brain regions, compared to levels in Bs. One-way ANOVA (F9,20 = 14.20, p < 0.0001) with Bonferroni tests. d HTT is increased more in the cortex and cerebellum compared to other brain regions in 2x AK057321/HTT mice. One-way ANOVA (F9,20 = 15.77, p < 0.0001) with Bonferroni tests. No difference is seen in 1x AK057321/HTT and HTT control mice. One-way ANOVA (F9,20 = 1.934, p = 0.1052) with Bonferroni tests. e Mouse Htt transcript across brain regions. One-way ANOVA with Bonferroni tests for 2x AK057321/HTT (F9,20 = 1.840, p = 0.1229); for 1x AK057321/HTT, (F9,20 = 0.9679, p = 0.4935); for 2x AK057321 (F9,20 = 0.6754, p = 0.7218); and for 1x AK057321 (F9,20 = 1.107, p = 0.4014). c–e RPL13A was used as the internal reference gene. For all experiments above, 6 mice per genotype were used. qRT-PCR performed on n = 3 RNA samples (each pooled from two mice). Data represent the mean ± SEM. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05. AK: AK057321, F: frontal cortex, T: temporal cortex, P: parietal cortex, O: occipital cortex, Cx: cortex, V: vermis, H/Hip: hippocampus, Cereb/Ce: cerebellum, C: cerebellar cortex, V: cerebellar vermis, Sc: subcortex, Ca: caudate, Th: thalamus, Bs: brainstem.
In human brain tissue, we found SVA-lncRNA AK057321 and HTT mRNA each display a 1.5 to 2-fold enriched expression in human cortex and cerebellum (Cereb) relative to brainstem (Bs) (Fig. 5b, c). This cortex and cerebellum-enriched expression of SVA-lncRNA AK057321 in human brain is reconstituted in AK057321 transgenic mice (Supplementary Fig. 16). Mouse brains expressing both SVA-lncRNA AK057321 and human HTT transgenes also reconstitute the human-specific cortex and cerebellum-enriched expression pattern of HTT mRNA (Fig. 5d). Whereas in mice expressing human HTT alone, the HTT mRNA transcript lacks cortex and cerebellum-enriched expression being equal across brain regions (Fig. 5d), as observed for native mouse Htt mRNA (Fig. 5e). Thus, SVA-lncRNA AK057321 drives a cortex and cerebellum-enriched expression pattern of human HTT (Fig. 5d) without altering expression of mouse Htt that lacks the intronic SVA (Fig. 5e). The results suggest SVA-lncRNA AK057321 may confer cortex and cerebellum-enriched of multiple genes contain an intronic SVA (Fig. 1f, g and Supplementary Figs. 4–7).
SVA-lncRNA AK057321 also upregulates expression of human Chromosome 21 genes with intronic SVAs in the brain
Effects of SVA-lncRNA AK057321 on genes with intronic SVAs on human chromosome 21 were also examined in the mouse brain. We crossed SVA-lncRNA AK057321 transgenic mice to mice with one copy of human chromosome 21, a transchromosomic model of the human intellectual disability disorder Down Syndrome40. SVA transposons on human chromosome 21 (Chr21) were identified using Repeat Masker in the UCSC genome browser (GRCh38/hg38). SVA-lncRNA AK057321 upregulates the expression of 3 of 7 genes with an intronic SVA in mouse cortex including POFUT2, CHAF1B, and KCNJ6 (Fig. 6a, b). AK057321 did not regulate mouse orthologs (Fig. 6c) or genes on human Chr21 nearby but lacking SVAs (Supplementary Fig. 17). Other human genes on Chr21 lacking intronic SVAs were also unaffected (Fig. 6d). AK057321 also increases CHAF1B expression in cortex of mice with a full-length human CHAF1B transgene (Fig. 6e). These data provide further examples where SVA-lncRNA AK057321 upregulates expression of human genes carrying but not mouse orthologs lacking an intronic SVA in the brain.
a RT-qPCR of AK057321 transcript levels shown in mouse cortex from human Chr21 transchromosomic/1x AK057321 mice compared to human Chr21 transchromosomic mice. n = 9 biological replicates. Unpaired Student’s t test. b RT-qPCR expression levels of human genes with intronic SVAs from mouse cortex of human Chr21 transchromosomic/1x AK057321 mice compared to human Chr21 transchromosomic mice. n = 11–14 biological replicates. One-way ANOVA (F13,165 = 10.38, p < 0.0001) with Bonferroni post hoc tests. c RT-qPCR of ortholog mouse gene expression from mouse cortex in AK057321 SVA-lncRNA transgenic mice compared to wild-type mice. n = 4 biological replicates. One-way ANOVA (F13,42 = 0.5728, p = 0.8616) with Bonferroni post hoc tests. d Expression levels of two human Chr21 genes without intronic SVAs shown from mouse cortex of human Chr21 transchromosomic/1x AK057321 mice compared to human Chr21 transchromosomic mice. Measured by RT-qPCR. n = 6 biological replicates. Unpaired Student’s t test. e Expression level of CHAF1B from mouse cortex of human CHAF1B transgenic mice compared to human CHAF1B transgenic/AK057321 transgenic mice. Measured by RT-qPCR. n = 7 biological replicates. Unpaired Student’s t test. a–e B2M used as the internal reference gene. Data represent the mean ± SEM. ****p < 0.0001, ***p < 0.001, **p < 0.01, ns p > 0.05.
SVA-containing gene transcripts are increased in human compared to mouse cerebellum
Based on the finding that SVA-lncRNA AK057321 drives a cerebellum-enriched expression of SVA-containing human HTT but not mouse Htt lacking an SVA, we compared the expression of several genes with intronic SVAs in human and mouse cerebellum samples. We designed RT-qPCR primers to conserved sequences in human and mouse for genes containing intronic SVAs and identified five genes CHAF1B, IARS, PI4K2B, OSBP, and FNIP2 where RT-qPCR primers amplified a product in cerebellum of both species. Transcripts for each gene were increased in human compared to mouse in cerebellum tissues (Supplementary Fig. 18a). Expression of control genes for gray matter (PSD95) and white matter (APOD) were equivalent suggesting similar tissue composition and RNA sampling. Moreover, SVA-lncRNA AK057321 increases expression of each gene increased in human compared to mouse cerebellum: CHAF1B, IARS, PI4K2B, OSBP, and FNIP2 (full-length genes in bacterial artificial chromosomes stably transformed in mouse 3T3 fibroblast cells, Supplementary Fig. 18b). These results provide additional evidence suggesting SVA-lncRNA AK057321 upregulates genes containing an intronic SVAs to confer species-specific expression patterns in brain.
SVA-lncRNA AK057321 opposes the transcription repressing effects of SVA sequences and ZNF91 in a luciferase reporter
SVA-lncRNA AK057321 upregulates genes with an intronic SVA in NTera-2 cells where ZNF91 is also expressed (Fig. 1g–i). ZNF91 is specific to primates11,14 and deleting ZNF91 gene increases expression of SVA-containing transcripts12. ZNF91 binds and represses transcription from genes containing SVA sequences11,12. We have provided many examples showing SVA-lncRNA AK057321 and ZNF91 have opposing effects on the expression of genes with intragenic SVAs (see diagram, Fig. 2a): CDK5RAP2 (Fig. 2b, c), SCN8A (Supplementary Fig. 12a–c) and CHAF1B (Supplementary Fig. 14b). To examine these effects of the SVA sequences in a simplified system, we examined the effects of SVA-lncRNA AK057321 and ZNF91 on a luciferase reporter with SVAs from either CHAF1B or HTT. Both SVAs repress luciferase reporter transcription in human embryonic kidney HEK293T cells (Supplementary Fig. 19a, b). Deleting the VNTR sequence in the SVA eliminates the transcriptional repressive effects of the CHAF1B SVA (Supplementary Fig. 19c). SVA-lncRNA AK057321 partially reverses the repressive effects of the SVAs (CHAF1B and HTT, Supplementary Fig. 19d, e) and counteracts the further repression produced by ZNF91 (Supplementary Fig. 19e). ZNF91 represses full-length but not VNTR deleted CHAF1B SVA (Supplementary Fig. 19f). Intronic SVAs of JAM2, AGPAT3, POFUT2 and CDK5RAP2 also repress transcription of the luciferase reporter compared to an empty vector (Supplementary Fig. 19g). These results indicate SVA sequences repress gene expression, likely by binding to repressor complexes associated with transcription factors like ZNF91. This effect is further magnified by increases of ZNF91. SVA-lncRNA AK057321 antagonizes the transcriptional repressive effects of SVA sequences that act at least in part by binding transcription factors like ZNF91 that recruits chromatin modifiers to repress gene expression.
SVA-lncRNA AK057321 forms RNA:DNA heteroduplexes with genomic SVA sequences in the CDK5RAP2, SCN8A, and CHAF1B genes it regulates
We postulated that SVA ribonucleic acid sequences in the SVA-lncRNA AK057321 might bind complementary SVA deoxyribonucleic acid sequences in the genome. We used two independent approaches (method flow chart in Fig. 7a): (1) an antibody (S9.6) that binds to RNA:DNA heteroduplexes41 and to a lesser extent AU-rich RNA:RNA duplexes42; and (2) an RNA-encoded aptamer tag with affinity to streptavidin added to the sequence of AK05732143 (Supplementary Figs. 20–21).
a Schematic showing two approaches to isolate genomic DNA associated with SVA-lncRNA AK057321. b RT-qPCR of human CHAF1B mRNA levels in 3T3 clones stably transfected with CHAF1B BAC DNA or CHAF1B BAC del-SVA DNA and infected SVA-lncRNA AK057321 lentivirus or control virus. n = 3 biological replicates for clone 1, n = 6 biological replicates clones 2–3. Unpaired Student’s t test. c qPCR for CHAF1B genomic DNA from eluted from RNA:DNA heteroduplex antibody or control antibody pulldowns in CHAF1B BAC or CHAF1B SVA-del BAC 3T3 clones transfected with AK057321 cDNA or control. n = 4 biological replicates. d qPCR for CHAF1B genomic DNA in eluted RNA:DNA complexes isolated by streptavidin pulldowns from CHAF1B BAC or CHAF1B SVA-del BAC 3T3 clones transfected with 5’-apatamer-AK057321, untagged AK057321 or control. n = 4 biological replicates. e qPCR for CHAF1B genomic DNA in eluted RNA:DNA complexes from streptavidin pulldowns performed in NTera-2 cells transfected with 5’-apatamer-AK057321 or untagged AK057321 n = 8 biological replicates. c–e Results are shown as folds enrichment compared to control. f, g qPCR for CDK5RAP2 or SCN8A genomic DNA from eluted RNA:DNA heteroduplex antibody and control antibody pulldowns in NTera-2 cells co-transfected with AK057321 cDNA with or without Crispr/Cas9 sgRNAs to delete the SVA within CDK5RAP2 (f) or SCN8A(g). n = 6 biological replicates each. h Approach for co-precipitation of myc-tagged ZNF91 with SVA-lncRNA AK057321 (Created with Biorender.com). i RT-qPCR for AK057321 RNA levels in eluted anti-myc vs. control antibody pulldowns performed in 293T cells co-transfected with myc-tagged ZNF91 and either AK057321 cDNA or control vector. Results are graphed as folds enrichment of AK057321 compared to control antibody pulldown. n = 3 biological replicates. Data represent the mean ± SEM. ****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05.
We examine mouse 3T3 fibroblast clones stably expressing a human full-length CHAF1B gene with or without the intronic SVA deleted. SVA-lncRNA AK057321 upregulates expression of intact but not SVA-deleted CHAF1B (Fig. 7b). qPCR primers target CHAF1B genomic sequences adjacent to the SVA present in both wild-type and SVA-deleted CHAF1B (Fig. 7a). S9.6 RNA:DNA heteroduplex antibody were found to pull down CHAF1B genomic sequences when the SVA is present but not when deleted (Fig. 7c). To confirm SVA-lncRNA AK057321 requires CHAF1B SVA sequences for binding, the 5′-aptamer tagged AK057321 (5′-apt-AK057321) was tested in CHAF1B clones with or without the SVA. Genomic DNA isolated from each condition was digested with restriction enzymes and precipitated with magnetic streptavidin beads (Fig. 7a). 5′-apt-AK057321 SVA-lncRNA precipitates CHAF1B genomic DNA from clones with the SVA present but not those with the SVA deleted (Fig. 7d). In addition, 5′-aptamer tagged AK057321 co-precipitates the CHAF1B genomic DNA in NTera-2 cells (Fig. 7e). Altogether these data show SVA-lncRNA AK057321 affinity purifies intronic SVA genomic sequences in CHAF1B forming a structure recognized by an RNA:DNA heteroduplex antibody.
To provide evidence that SVA-lncRNA AK057321 binds intronic SVAs in the CDK5RAP2 and SCN8A genes, CRISPR/Cas9-sgRNAs was used to delete the native SVA sequences in NTera-2 cells expressing SVA-lncRNA AK057321. qPCR was performed with primers to genomic sequences outside of the SVAs in CDK5RAP2 and SCN8A. S9.6 RNA:DNA heteroduplex antibody co-precipitated CDK5RAP2 (Fig. 7f) and SCN8A (Fig. 7g) genomic sequences. Notably, Cas9-gRNA deletion of the SVA sequences decreases the recovery of CDK5RAP2 and SCN8A genomic sequence (Fig. 7f, g). The results suggest SVA-lncRNA AK057321 may form RNA:DNA heteroduplexes with intronic SVAs in the CDK5RAP2 and SCN8A genes (see diagram in Fig. 8).
a Schematic showing SVA-lncRNA AK05732 increases as neural progenitor cells mature into glutamatergic neurons. b Schematic where SVA-lncRNA AK05732 forms an RNA:DNA heteroduplex with intronic SVAs in CDK5RAP2 and SCN8A and binds to the ZNF91 transcription factor that otherwise represses expression of genes containing an intronic SVA (Created with Biorender.com).
The presence of an entire SVA sequence in SVA-lncRNA AK057321 (Fig. 1b; exon 3) suggested the possibility that ZNF91 protein, that binds SVA deoxyribonucleic acid sequences in the genome, might also bind SVA ribonucleic acid sequences in SVA-lncRNA AK057321. Myc-tagged ZNF91 protein was co-expressed with SVA-lncRNA AK057321 or control plasmid in HEK293T cells, immunoprecipitated with anti-myc or control antibodies, and analyzed for SVA-lncRNA AK057321 by RT-qPCR (methods in Fig. 7h). The results reveal that myc-ZNF91 pulldown enriches for SVA-lncRNA AK057321 (Fig. 7i). The findings suggest SVA-lncRNA may anneal to the genomic SVA sequences and also bind ZNF91 to disrupt its repressive effects on SVA-bearing genes like CDK5RAP2 and SCN8A thereby promoting neuronal differentiation (biological pathway diagrammed in Fig. 8).
Discussion
To identify genes repressed by their intronic SVAs to delay neuronal maturation and derepressed by SVA-lncRNA AK057321 to drive neuronal maturation, we focused on genes with the following properties: (1) those with human-specific intronic SVAs; (2) that underlie human neurodevelopmental disease; and (3) that are strongly regulated by SVA-lncRNA AK057321 and ZNF91. This led us to CDK5RAP2, a gene with a human-specific tandem SVA-F important to neurogenesis20,21,22,23,24,25,26. Homozygous inactivating mutations of CDK5RAP2 underlie human congenital primary microcephaly with reduced cerebral cortex growth44,45,46. We found SVA-lncRNA AK057321 depletion (shRNA or Cas9/sgRNA) downregulates, while its overexpression upregulates CDK5RAP2 expression. Bioinformatic genomics studies previously suggested microcephaly genes might underlie the enlarged neocortex of higher primates specifically suggesting a possible role for CDK5RAP247,48,49. Consistent with such ideas, we find the human-specific, tandem SVA_F in the CDK5RAP2 intron represses its expression to slow neuronal maturation. We further find that SVA-lncRNA AK057321 induces neuronal maturation by derepressing CDK5RAP2.
SVA-lncRNA AK057321 upregulates a wide variety of neurodevelopmental and neurological disease genes containing intronic SVAs beyond CDK5RAP2 including SCN8A, an epilepsy and intellectual disability sodium channel gene with a human-specific intronic SVA_D. Heterozygous gain-of-function mutations in SCN8A underlie early infantile epileptic encephalopathy and heterozygous loss-of-function mutations underlie intellectual disability, suggesting an early dose-dependent role of this gene in neuronal function28,29. We find that deleting the intronic SVA_D in SCN8A drives selective maturational increases of voltage-gated sodium currents and sodium spikes (immature action potentials) but not other neuronal properties (i.e., hyperpolarized resting membrane potential or increased membrane capacitance). Based on the wide variety of neurodevelopmental disease genes containing intronic SVAs that are regulated by SVA-lncRNA AK057321, we suggest intragenic SVAs may delay maturation of multiple specialized neuronal functions (e.g., synaptogenesis involving NRXN1, NRXN2, and SHANK2 and the developmental change of chloride reversal potential that converts GABAergic transmission from excitatory to inhibitory involving WNK3).
We show the neuronal maturation-promoting effects of SVA-lncRNA AK057321 are mimicked by decreasing expression of the SVA-repressive transcription factor ZNF91. These data suggest a biological pathway where SVA-lncRNA AK057321 opposes the transcriptional repressive effects of ZNF91 repressor complex bound to the intronic SVA sequences. KZNFs such as ZNF91 transcriptionally repress genomic regions containing retrotransposons by binding TRIM2850 leading to repressive epigenetic modifications51,52 with some components mediated by the human silencing hub (HUSH) complex16.
We provide evidence that SVA ribonucleic acid sequences in SVA-lncRNA AK057321 may form an RNA:DNA heteroduplex with complementary genomic SVA deoxyribonucleic acid sequences in the introns of CDK5RAP2, SCN8A and CHAF1B genes to increased their expression. SVA-lncRNA AK057321 also binds ZNF91 suggesting a possible decoy effect that might displace ZNF91 transcriptional repressor complex from genomic SVA sequences. Taken together, these data reveal the existence of an SVA transposon-based gene regulatory system involving intronic SVAs bound by either ZNF91 repressor complex to repress, or by SVA-lncRNA AK057321 to enhance gene expression, thereby regulating the timing of neuronal maturation in the hominoid species lineage.
Human Huntingtin disease gene HTT39 contains an intronic SVA. We find HTT gene expression is enriched in cortex and cerebellum relative to brainstem in human (not seen for native mouse Htt gene or human HTT transgene in mouse brain) and that this cortex and cerebellum-enriched expression pattern is driven by SVA-lncRNA AK057321. Using mice transgenic for human SVA-lncRNA AK057321 and human chromosome 21, we show that SVA-lncRNA AK057321 upregulates expression of genes that harbor intronic SVAs on human chromosome Chr21 in mice40, increasing POFUT2, CHAF1B and KCNJ6 genes but not mouse orthologs lacking SVAs. These results provide evidence that SVA-lncRNA AK057321 may increase the expression of multiple neurodevelopmental genes with intronic SVAs in the cortex and cerebellum relative to subcortical brain regions. We speculate that increasing expression of neurodevelopmental genes in cortex and cerebellum glutamatergic neurons relative to the other brain regions might enable these neurons to become the drivers of subcortical neuronal circuit activities (e.g., in thalamus, hypothalamus, and brainstem).
Polymorphic SVA insertions found in the human population in protein-coding or regulatory regions have resulted in insertional mutagenesis or aberrant RNA splicing to cause disease. Examples include intragenic SVA insertions in Fukuyama muscular dystrophy53, X-linked dystonia Parkinsonism54, Neurofibromatosis type 155, and autosomal recessive hypercholesterolemia56. De novo SVA insertions have also been reported in non-coding regions of the genome including introns57. Here we provide insights into the functional significance of such intronic SVAs acting to repress gene expression during neuronal maturation and in subcortical brain regions in adulthood. We speculate that newly acquired intronic SVAs might promote previously unrecognized genetic forms of neurodevelopmental, neurological, or psychiatric disease.
We find SVA-lncRNA AK057321 is expressed in glutamatergic but not cholinergic or dopaminergic neurons. We speculate that other SVA-lncRNA family members might be restricted to other neuronal and glial cell types enabling independent regulation of the timing of gene expression in those cells.
SVA-lncRNA AK0574321 is also expressed in testis and multiple genes regulating spermatogenesis and fertility contain intronic SVAs (e.g., SPATA558,59 and CDK5RAP260). This observation suggests the SVA transposon-based gene regulatory system might influence species-specific aspects of male fertility and new intronic SVAs acquired during speciation may confer fertility advantages to those species.
Given the numerous genes with intronic SVAs in hominoids (many specific to human), future studies should examine the functional role of the other intronic SVAs in regulating glutamatergic neuron progenitor growth, maturation, structure and function. Such studies could provide further insights into how this SVA transposon-based gene regulatory system may have helped shape structures and functions unique to the human brain.
Methods
Statistics and reproducibility
Statistical analysis of datasets showing unpaired two-tailed Student’s t test, one-way ANOVA with Bonferroni post hoc tests, and two-way ANOVA with Bonferroni post hoc tests were calculated on datasets in this study using Graphpad Prism 9.3.1 software. Graphed data represent the mean ± SEM.****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05, ns p > 0.05 versus control unless otherwise indicated. Blinding was performed during data collection and analysis. All cell, animal, and human samples were randomized. Group sizes were chosen based on adequate statistical power to detect differences. Details of biological replicates described in figure legends. No data was excluded. Experiments were independently replicated at least twice (with the exception of Fig. 1g (a subset of genes), Fig. 7i and Supplementary Fig. 3). For mice, a biological n indicates tissue from an independent animal with the same genotype. For cells, a biological n indicates cells from an independent set of transfection/FACS sort with the same vectors.
GO term enrichment analysis
GO term enrichment analysis was performed at the following websites: pantherdb.org and g:Profiler – a web server for functional enrichment analysis and conversions of gene lists (ut.ee).
NTera-2 transfection and cell sorting
NTera-2, CRL-1973, was purchased from ATCC. NTera-2 cells were plated at a density of 5000 cells/cm3 12–16 h prior to transfection with the indicated plasmids using Lipofectamine 3000 (ThermoFisher) for the indicated times. Cells were harvested and FACS sorting of NTera-2 transfected fluorescent cells was performed at the BIDMC Flow cytometry Core on a MOFLO Astrios EQ and on a BD FACSAria II. Each experiment utilized a fluorescent empty vector control, and un-transfected NTera2 cells were used as the negative control to set the gate for positive cell collection in each experiment.
NTera-2 neuronal maturation with mouse astrocyte co-plating
For Ngn2-induced differentiation, two lentiviruses were each produced from expression plasmids pLV-TetO-hNGN2-eGFP-Puro (Addgene 79823) and FUGW-UbCp-rtTA3 (Addgene 105171) using the method of co transfection HEK293T cells with psPAX2 (Addgene 12260) and pMD2.g (Addgene 12259) (Addgene: Lentivirus Production Protocol). For all co-culture studies with transfected NTera-2 cells, mouse glial cells were isolated from the forebrains of 1-3 day-old CD1 pups as described61. Isolated forebrains were put into calcium and magnesium-free HANKS (ThermoFisher 14175015) and dissociated with trypsin-EDTA (ThermoFisher 25200056). After 20 min, trypsin was inhibited by suspension of the cells into MEM (ThermoFisher 51200038) supplemented with 10% Fetal Bovine Serum (ThermoFisher). Cells were triturated through a pipette and passed through a 0.40 μM cell strainer, pelleted and washed twice, re-suspended and then cultured in T75 flasks. Media was changed the next day and cells were passaged one time to avoid neuronal contamination before their use in co-culture experiments. For co-culture experiments of Ngn2-induced NTera-2 cells with mouse astrocytes, NTera-2 cells were transduced similarly as described62 with the following daily steps. On day −5, glass coverslips were coated with Matrigel (BD Biosciences 354230) within wells of a 24-well plate and on day −4, isolated glial cells were trypsinized and plated onto coverslips with matrigel and set aside until they reached 70–80% confluency on day 2. On day −2, Ntera-2 cells were plated at 30-50% confluency in T75 flasks. On day 1—NTera-2 cells were co-infected with Ngn2 and rtTA containing lentiviral supernatants as prepared above, in DMEM supplemented with polybrene (8 μg/mL, MilliporeSigma). On day 0, doxycycline was added at a concentration of 2 μg/mL to induce Ngn2 and puromycin resistance gene expression. On Day 1 puromycin (ThermoFisher A1113803) was added at a concentration of 1 μg/mL to select cells expressing Ngn2. On day 2, Ngn2-positive NTera-2 cells were removed from T75 flasks with TrypLE (ThermoFisher 12604013) and were co-plated at a density of 1.5 × 104 cells per well in Neurobasal medium (ThermoFisher 21103049) supplemented with Glutamax (ThermoFisher 35050079), B27 (ThermoFisher 17504044) and BDNF (R&D Systems 248-BD) along with 2 μg/mL doxycycline. Ara-C (2 g/l, Sigma) was added to inhibit astrocyte proliferation once astrocytes reached confluency. 50% of the media (0.3 ml) was changed every 3–4 days until experiments were performed. In experiments lacking Ngn2, FACS-sorted NTera-2 cells were plated at a low density (5000 cells per cm2) along with mouse astrocytes onto matrigel-coated glass coverslips in Neurobasal medium supplemented with B27, Glutamax and BDNF. Ara-C (2 g/l, MilliporeSigma) was added to inhibit astrocyte proliferation once astrocytes reach confluency. 50% of the media (0.3 ml) per well was changed every 2-3 days.
Immunostaining
Ngn-2 induced neurons at 90 days after co-plating were fixed in 4% PFA in PBS for 1 h at room temperature. After PBS washing, anti-beta 3 Tubulin (TuJ1) Antibody (2G10, Saint Cruz, 1 to 2000 dilution) was used for staining TuJ1 in blocking solution at 4 °C temperature overnight. Sections were washed and incubated with Alexa-conjugated secondary antibodies (1:500, Invitrogen) for 3 h at room temperature. Cells then were mounted in Vectashield with DAPI (Vector). Fluorescent images were taken using a LSM510 confocal microscope (Zeiss) and were processed with Image J. For immunostaining of neuronal maturation due to AK057321 overexpression, NTera-2 cells were FACS-sorted NTera-2 cells, AK057321 overexpression vs. control cells, were individually co-plated with mouse astrocytes for 13 days as described above. Cells were then fixed in 4% PFA in PBS, followed by permeabilization with 0.3% Triton X-100, blocking in 5% goat serum/PBS for 1 h. MAP2 antibody (1:100, MilliporeSigma AB5622) was diluted in 2.5% goat serum in PBS and incubated with cells for 1 h at room temperature, washed three times in PBS, followed by incubation with secondary antibody (1:500, ThermoFisher A-11011) for 1 h at room temperature. After washing in PBS, the cells were mounted with anti-fade medium (Vectorshield plus H-1900-10) and stored at 4 degrees overnight prior to microscopy with LSM510 confocal microscope (Zeiss). Images were processed with Image J.
Sholl analysis
SVA-lncRNA AK057321 (or control) transfected NTera-2 cells, matured for 13 days with mouse astrocytes on matrigel-coated glass coverslips, were fixed in 4% PFA in PBS for 10 min at room temperature, followed by 3 times wash with PBS, and mounted with anti-fade medium (Vectorshield plus; Cat. #: H-1900-10) and placed at 4 degrees overnight before subjected to immunofluorescence microscopy. Images were acquired with Leica DM6 confocal microscopy by Leica Application Suite X (LAS X) software. Exposure time was fixed throughout the imaging process. Images were converted to 8-bit format and processes were traced using NeuroJ plugin. The center of the cell body was used as a reference point and 94 concentric circles were generated on the tracings: the starting radium was 30 pixels (9.2 μm) and the ending radium was 380 pixels (121.6 μm) at the step of 5 pixels (1.6 μm). The number of intersections at each concentric circle was registered and plotted. The total length of processes, mean length of processes, the average maximum/ minimum length of processes, number of processes were quantified from the tracing files. Cell morphology was quantified with ImageJ plugins for circularity. Cell area was measure with ImageJ with fixed threshold.
Electrophysiology
Recordings were made on the indicated days after NTera-2 and mouse astrocytes were co-plated on matrigel and poly-lysine-coated glass coverslips immediately after sorting. Membrane potential and currents were recorded using the whole-cell configuration of the patch-clamp technique with an EPC-10 HEKA amplifier and the HEKA PatchMaster 9 data acquisition software. The extracellular solution consisted of (in mM): 140 NaCl, 5.4 KCl, 0.5 MgCl2, 2.5 CaCl2, 5.5 HEPES, 11 glucose, and 10 sucrose, pH 7.4 with NaOH for current clamp mode. Resistance of the patch pipette was 3–4 MΩ when filled with (in mM) 105 K-gluconate, 20 KCl, 1 CaCl2, 5 MgATP, 10 HEPES, 10 EGTA, and 25 glucose, pH 7.4 with KOH. Action potential was elicited by a 100 ms square current injection from 0- 600 pA with 100 pA increment. Cells were held at −90 mV by hyperpolarizing currents to induce sodium-based action potential. \(\varDelta\)V was calculated by measuring the difference between peak amplitude and 100 ms after current injection. To record Na+ currents, the extracellular solution was changed to a solution consisting of (in mM): 130 NaCl, 20 TEA-Cl, 1 CaCl2, 5 MgCl2, 10 HEPES, and 5.56 glucose, pH 7.4 with NaOH. Internal solution was changed to (in mM) 10 NaCl, 130 CsCl, 5 MgCl2, 5 EGTA and 10 HEPES. Na+ currents were evoked from a holding potential of −90 mV by stepping to voltages −30 mV for 20 ms. Series resistance of 5–10 MΩ was electronically compensated 80-90%. Current traces were sampled at 10 kHz and filtered at 5 kHz. Comparisons were made at day 15 unless mentioned otherwise.
SVA-lncRNA AK057321 expression and knockdown
SVA-lncRNA AK057321, Homo sapiens cDNA FLJ32759 fis, clone TESTI2001793, was subcloned into BamHI and EcoRI sites of pLVX-mcherry, and also into XbaI and BamHI sites of BioSettia pLV-09. For knockdown of SVA-lncRNA AK057321, target sequences listed in Supplementary Table 1 were cloned into a modified version of pLVX-shRNA2-mcherry in which the CMV promoter was excised with BamHI and AgeI digestion and replaced with the UbC promoter, P2A, the zeomycin resistance gene and T2A in frame with mcherry using HiFi DNA Assembly (primers listed in Supplementary Table 1). UbC, P2A, and T2A sequences were PCR-amplified from pLV-TetO-hNGN2-eGFP-Puro (Addgene #79823) and the zeocin resistance gene sequence was amplified by PCR from FRT-Em7-Zeo-FRT plasmid (Thermo Fisher).
myc-tagged human ZNF91
ORF clone expressing myc-tagged human ZNF91, accession number NM_003430.3 was purchased from GeneCopoeia.
CRISPR/Cas9-mediated deletions
For CRISPR-Cas9 deletion of ZNF91, published gRNAs12 were cloned into a modified version of lentiCRISPRv2 hygro (Addgene #98291) in which the hygromycin resistance gene was excised by digestion with PshAI and MluI and replaced with PCR-amplified mcherry from pLVX-shRNA2-mcherry using NEB HiFi DNA assembly (primers listed in Supplementary Table 2). All PCR amplifications for cloning were performed with Phusion (NEB; with cycling conditions: 1, 98 °C for 30 s, 25–35, 45–72 °C for 10–30 s, 72 °C for 15–30 s per Kb, last cycle, 72 °C for 5–10 min). ZNF91 sgRNAs were first cloned individually into the BsmBI site of lentiCRISPRv2 hygro. To ensure delivery of both sgRNAs into every transfected cell, both sgRNAs were cloned in tandem into lentiCRISPRv2-mcherry, each with their own U6 promoter, using long primers to amplify one sgRNA and short primers for the other. lentiCRISPRv2-mcherry with no sgRNAs was digested with KpnI and NheI and each guide was PCR-amplified from the corresponding Addgene #98291 vector (lentiCRISPRv2 hygro) it was originally cloned into. The digested lentiCRISPRv2-mcherry vector (KpnI and NheI), along with PCR-amplified U6-sgRNA (up; long_guide) and PCR-amplified U6-sgRNA (down; short_guide) were assembled using NEB HiFi DNA assembly (primers listed in Supplementary Table 2). For CRISPR-Cas9 deletion of the tandem SVA located within the AK057321 gene, candidate sgRNAs (located 5′ and 3′ to the SVA sequence) were each individually cloned into lentiCRISPRv2 hygro with the recommended protocol: lentiCRISPRv2 puro was a gift from Brett Stringer (Addgene plasmid # 98290; http://n2t.net/addgene:98290; RRID:Addgene_98290). Cloned sgRNAs were tested in pairs (5′ sgRNA plus 3′ sgRNA) to determine an optimal combination to achieve SVA deletion. These sgRNAs (5′ sgRNA plus 3′ sgRNA) were then cloned in tandem using the same HiFi DNA assembly strategy as was done for lentiCRISPRv2-mcherry, but into a modified version of lentiCRISPRv2 hygro in which T2A and blue fluorescent protein (BFP) were inserted in frame after the MluI site in lentiCRISPRv2 hygro, (primers listed in Supplementary Table 2). sgRNAs used to delete each SVA located within introns of CDK5RAP2, SCN8A and CHAF1B were similarly cloned and tested, with optimized sgRNA pairs cloned in tandem into lentiCRISPRv2-BFP cut with KpnI and NheI. Supplementary Table 3 lists the sgRNA sequences used in this study (PAM sequences are underlined). “Left” vs. “right” refers to sgRNA sequences used to cut on each side of the SVA; one sgRNA on the left (5′ to the SVA) and one sgRNA on the right (3′ from the SVA). Supplementary Table 4 lists the genomic PCR primers used to detect the CRISPR-Cas9 deletion of SVAs located within the AK057321 lncRNA sequence, and within CDK5RAP2, SCN8A and CHAF1B introns. Genomic PCRs were performed using Azura 2X Taq Red Mix, using cycling conditions: 1; 95 °C, 1 min, 25–40; 95 °C, 15 s, 60 °C, 15 s, 72 °C, 30 s per Kb).
RNA aptamer tag added to SVA-lncRNA AK057321
A sephadex and a streptavidin dual aptamer tag was constructed by PCR with overlapping oligos and added to SVA-lncRNA-AK057321 at the 5′ end with the spacer sequence “GGAAGAGGAAGA” after the tag and before the start of the AK057321 sequence, internally between exons 2 and 3 with the same spacer sequence flanking the 5′ and 3′ end of the dual tag sequence, and at the 3′ end with the spacer sequence after the AK057321 sequence and before the start of the dual aptamer tag sequence in Supplementary Table 5.
CDK5RAP2 knockdown and overexpression
CDK5RAP2-set of piLenti (Human) siRNAs: piLenti-siRNA-GFP that targets four CDK5RAP2 mRNA sequences were purchased from abm, catalog number i004292. Inducible Expression of CDK5RAP2: Overexpression of CDK5RAP2 has reported to be toxic, so an inducible construct to express lower levels of CDK5RAP2 was generated by us. For inducible expression of CDK5RAP2, the CMV promoter within pRcCMV Cep215 (Addgene #41152) was excised with PciI and AfeI digestion. The TetO sequence was amplified using TetO FWD and TetO REV primers listed in Supplementary Table 6 from pLV-TetO-hNGN2-eGFP-Puro (Addgene #79823) vector and was inserted into digested pRcCMV Cep215 using NEB HiFi Assembly with HiFi PCR primers listed in Supplementary Table 6. For expression studies, TetO-CDK5RAP2 was co-transfected with FUW-rtTA (Addgene #20342) Ntera-2 cells and CDK5RAP2 expression was induced with 1 μg/mL doxycycline for 24 h.
Luciferase assays
All luciferase assays were performed in HEK293T cells. After 48 h cells were assayed for luciferase activity using ONE-Glo EX Luciferase Assay System (Promega) on a BioTek Synergy luminometer using Gen5 software. The CHAF1B SVA was amplified from bacterial artificial chromosome (BAC) clone RP11-108J14 purchased from BACPAC https://bacpacresources.org/order_clones.php using Phusion polymerase (NEB) the CHAF1B FWD and CHAF1B REV primers listed in Supplementary Table 7. All PCR amplifications for SVA cloning were performed with Phusion (NEB; with the GC buffer and DMSO). Cycling conditions: 1, 98 °C for 30 s, 25–35, 45–72 °C for 10–30 s, 72 °C for 15–30 s per Kb, last cycle, 72 °C for 5–10 min). The SVA was then cloned into the KpnI and NheI sites within the pGL3-luciferase promoter vector (Promega) using PCR fragment amplified with the FWD and REV HiFi primers (Phusion) with HiFi NEBuilder Assembly (NEB). Positive clones were selected and screened for the presence of the SVA by PCR with Phusion (NEB) using the CHAF1B SVA FWD and REV primer sets. The CHAF1BminusVNTR version of the CHAF1B SVA was assembled by PCR from two separate pieces (Part A and Part B) that lacked the VNTR region, each PCR-amplified from bacterial artificial chromosome (BAC) clone RP11-108J14 using Phusion polymerase (NEB). The Part A and Part B SVA pieces were then cloned into the KpnI and NheI sites within the pGL3-luciferase promoter vector (Promega) using PCR fragments amplified with the FWD and REV HiFi primers (Supplementary Table 8) with HiFi NEBuilder Assembly (NEB). Positive clones were selected and screened for the presence of the SVA by PCR with Phusion (NEB) using the CHAF1B SVA FWD and REV primer sets (Supplementary Table 7). The HTT SVA was amplified by gradient PCR using Phusion (NEB) from the bacterial artificial chromosome (BAC) clone RP11-866L6 purchased from BACPAC https://bacpacresources.org/order_clones.php., using the HTT SVA FWD and REV primers listed in Supplementary Table 9. The PCR fragment containing the HTT SVA was then TA-cloned into TOPO pcR2.1 (Invitrogen) and positive clones were selected with screening primers listed in Supplementary Table 9. The HTT SVA-containing fragment was then excised from TOPO pcR2.1 with KpnI and XhoI and cloned directly into these sites of KpnI/XhoI digested pGL3-luciferase promoter vector (Promega). Positive clones were screened for insertion with the screening primers listed in Supplementary Table 9. JAM2, AGPAT3, POFUT2, and CDK5RAP2 SVA clonings: SVAs were amplified from individual bacterial artificial chromosome (BAC) clones obtained from BACPAC https://bacpacresources.org/order_clones.php using Phusion polymerase (NEB) the primers listed in Supplementary Table 10. The SVAs were then similarly cloned into the KpnI and NheI sites within the pGL3-luciferase promoter vector (Promega) using PCR fragment amplified with the FWD and REV HiFi primers (Phusion) with HiFi NEBuilder Assembly (NEB) using primers listed in Supplementary Table 10. Positive clones were selected and screened for the presence of the SVA by PCR with Phusion (NEB) using the CHAF1B SVA FWD and REV primer sets.
BAC recombineering
BAC recombineering was used to insert neomycin resistance gene into various BAC clones and generation of stable NIH3T3 clones. Genomic DNA bacterial artificial chromosome (BAC) clones were purchased from was purchased from BACPAC (RP11) and Invitrogen (CTD), https://bacpacresources.org/order_clones.php and https://clones.thermofisher.com as listed in Supplementary Table 11. These BACs are on the pBACe3.6 backbone (RP11) or on the pBeloBAC11 backbone (CTD). A a neomycin resistance gene cassette was added to each BAC in Supplementary Table 12. The neomycin resistance gene was amplified by PCR with Phusion (NEB) using NeomycinR FWD and REV primers from the PGK-Em7-Neo plasmid. All primer sequences used in recombineering are shown in Supplementary Table 12. For the RP11 and CTD clones, PCR-amplified arms were generated and selection primers as shown in Supplementary Table 12. The neomycin cassette was combined with these arms by PCR creating an amplified fragment containing the upstream arm – neomycin resistance gene – and downstream arm. This PCR fragment was gel-purified and inserted into the clones by homologous recombination using electrocompetent SW105 cells. Cells were then plated on chloramphenicol/kamamycin plates for selection. Individual clones were screened for insertion with the following primers that amplify on the BAC backbone around the loxP site listed in Supplementary Table 13. BAC recombineering to delete the SVA in a CHAF1B BAC clone: The BAC clone RP11-108J14, chr21:36384336-36558420 (174,085 bp; Hg18 UCSC Genome Browser) contains the entire human CHAF1B genomic sequence, including an intragenic SVA was first modified by the addition of a neomycin drug resistance cassette. To replace the SVA sequence within the CHAF1B on this BAC clone, PCR (Phusion polymerase; NEB) was used to generate three pieces (Supplementary Table 14); a left arm with sequence identity to the BAC upstream of the SVA, a right arm with identity to the BAC downstream of the SVA and then a middle piece that contains the zeocin drug resistance cassette and Frt sites, from FRT-Em7-Zeo-FRT plasmid. BAC recombinations were performed using electrocompetent SW105 with 200-300 of gel-purified assembled DNA, followed by plating on chlro/kan/zeo plates. Clones were expanded and tested for insertion by PCR from the bacteria cultures using a zeo oligo and a CHAF1B oligo (Supplementary Tables 8 and 15). The zeocin resistance gene was amplified by PCR from FRT-Em7-Zeo-FRT clone GF-21 with the primers listed in Supplementary Table 15. In summary, the overall strategy generated three pieces, a left arm with sequence identity to the BAC upstream of the SVA, a right arm with identity to the BAC downstream of the SVA and then a middle piece that contains the zeo drug resistance cassette using the primers listed in Supplementary Tables 14–15. These three overlapping pieces were then assembled in the PCR machine using Phusion polymerase (NEB).
Gene expression analysis
Total RNA was isolated using Trizol (Fisher Scientific). All RNA pellets were resuspended and stored in RNASecure (Fisher Scientific) at −80 °C. Reverse transcription of RNA was performed after DNAse treatment (NEB) with M-MuLV Reverse Transcriptase (NEB). Quantification of mRNA levels was performed using PowerUp SYBR (Applied Biosystems; 50 °C 2 min, 95 °C 2 min, 45 cycles of 95 °C for 3 s and then 60 °C for 20 s) or Taqman Gene Expression master mix (Applied Biosystems; 50 °C 2 min, 95 °C 10 min, 45 cycles of 95 °C for 15 s and then 60 °C for 1 min) on a BIO-RAD CFX384 Real-Time System. Primers and probes were ordered from Integrated DNA technologies (IDT, Supplementary Table 16). Other primers used for expression analysis were ordered from Eurofins (Supplementary Tables 17 and 18). Primers that recognize both human and mouse transcripts (data shown in Supplementary Fig. 18) are bolded in Supplementary Table 17. Supplementary Table 18 lists the primers used for relative-quantitative PCR (rqPCR) of AK057321 expression (Supplementary Fig. 16). Supplementary Table 19 lists the thermocycler conditions for reverse transcriptase and polymerase chain reactions. Multiple bands were amplified in the PCR from transgenic mice and human cDNA samples with different primer sets. The band at the expected size of AK057321 amplicons and two additional bands were isolated from agarose gel with QIAquick Gel Extraction Kit (Qiagen, 28704) and sequenced at the Beth Israel Deaconess Medical Center DNA Sequencing Core. The band that was identified as AK057321 was used in relative-quantitative PCR experiments to determine expression levels in transgenic mouse and human brain regions. Classic and Universal QuantumRNA 18 S Internal Standard kits (Ambion, AM1716 and AM1718) were used according to manufacturer’s instructions. PCRs were performed with Phire Hot Start DNA polymerase (Finnzymes F-120) on cDNA prepared from brain tissue of AK057321 transgenic mice (1 or 2 copies of the transgene, 3 mice for each genotype) or human brain tissue. Optimization reactions were performed to determine the linear range and cycle number of PCR amplification, and 18 S Primer:Competimer ratio. The reactions were run on a DNA agarose gel, and images were acquired with a BioRad ChemiDoc XRS + system. The band intensities were quantified with ImageJ software. RNA from iPSCs and each differentiated neuron type was obtained from Elixirgen Scientific, https://www.elixirgensci.com.
Transgenic mice
For generation of AK057321 transgenic mice, genomic DNA bacterial artificial chromosome (BAC) clone RP11-1043N4 32,604,995-32,769,907 (164,912 bp; Hg18 UCSC Genome Browser) was purchased from BACPAC (https://bacpacresources.org/order_clones.php). BAC DNA was prepared by double sodium acetate precipitation and CsCl2 gradient method and confirmed by sequencing analysis. BAC DNA was linearized by restriction enzyme PI-SceI (NEB) and then microinjected in the BIDMC transgenic core facility. The studies use human mutant HTT transgenic mice bearing an HD model mutation and carried in a yeast artificial chromosome vector, YAC128 mice (Jackson Laboratory, mouse strain #004938). For generation of CHAF1B transgenic mice, genomic DNA bacterial artificial chromosome (BAC) clone RP11-108J14, chr21:36384336-36558420 (174,085 bp; Hg18 UCSC Genome Browser) contains the entire human CHAF1B genomic sequence, including an intronic SVA (BACPAC, https://bacpacresources.org/order_clones.php). BAC DNA was prepared by purification with Nucleobond BAC 100 kit (Machery-Nagel). BAC DNA was linearized by restriction enzyme PI-SceI (NEB) and then microinjected in the BIDMC transgenic core facility. All protocols were approved by the BIDMC IACUC and Harvard Medical Area Standing Committee on Animals.
Sample tissue preparation
Adult mice (between 2-6 months in age) were deeply anesthetized with isoflurane and rapidly decapitated. For studies involving human mutant HTT transgenic and matched controls, males were chosen. For studies involving human Chr21 transchromosomic mice and human CHAF1B transgenic mice, animals were chosen without regard to sex. The brain was removed and cortex, cerebellum, hippocampus, subcortical, and brain stem were microdissected with fine scissors and forceps. Samples were flash frozen on dry ice and stored at −80 °C until RNA was prepared. Human brain tissue was collected randomly from sequential postmortem cases with next-of-kin consent in the Beth Israel Deaconess Medical Center Pathology Department with IRB approval. Samples from males (50-65 years of age) cortex (frontal, temporal, parietal, occipital), cerebellum (cortex and vermis), hippocampus, caudate, thalamus and pons were collected, flash frozen on dry ice, and stored at −80 °C until RNA was prepared.
SVA-lncRNA AK057321 binding to genomic SVA studies
For SVA-lncRNA AK057321 binding to the intronic SVA in CHAF1B, NIH3T3 clones stably expressing with the human CHAF1B gene from genomic BAC RP11-108J14, or from a version of genomic BAC RP11-108J14 in which the SVA was deleted were first used. For SVA-lncRNA AK057321 binding to the intronic SVA in endogenous CHAF1B, CDK5RAP2 and SCN8A genes, NTera-2 cells were used. After each transfection, cell pellets were harvest and flash frozen at −80 °C. Genomic DNA was extracted from each cell pellet using phenol/chloroform/isoamyl alcohol (FisherSci) and genomic DNA was washed with 70% ethanol, dried and resuspended in RNASecure (InVitrogen). 20 μg of genomic DNA from each condition were digested with an enzyme cocktail containing BamHI, HindIII, XbaI, PstI and purified RNAse inhibitor. The RNAse inhibitor (Addgene #153314) was grown in bacteria and was purified using NEB’s pMAL Protein Fusion and Purification System, E8200. To assess RNA:DNA heteroduplex formation, each sample was resuspended in TE buffer with 0.1% NP-40 equivalent detergent (FisherSci) and immune-precipitated with S9.6 or control antibody with Protein G magnetic beads (FisherSci) overnight, supplemented with RNAse inhibitor. Beads were then washed 5 times in buffer supplemented RNAse inhibitor. Complexes were eluted by heating the washed beads at 80 °C for 5 min. The presence of the genomic DNA adjacent to the respective intronic SVAs (CHAF1B, CDK5RAP2, SCN8A) are assessed by qPCR using the genomic primer and probes listed in Supplementary Table 20. For pulldown of 5′-aptamer tagged SVA-lncRNA AK057321, the same protocol was followed except pulldowns were performed with magnetic streptavidin beads (FisherSci) for only 1 h.
SVA-lncRNA AK057321 binding to ZNF91 protein
myc-tagged ZNF91 and SVA-lncRNA AK057321 were co-expressed in 293T cells for 48 h, harvested and then cell pellets were flash frozen at −80 °C. Cells were lysed and immune-precipitated with anti-myc or control antibodies for 2 h in the presence of RNAse inhibitor, followed by 5 washes with buffer and then elution of complexes by heating at −80C for 5 min. RT-qPCR was performed to detect the presence of SVA-lncRNA AK057321 in each condition using the probes listed in Supplementary Table 16.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Rakic, P. Evolution of the neocortex: a perspective from developmental biology. Nat. Rev. Neurosci. 10, 724–735 (2009).
Somel, M. et al. Transcriptional neoteny in the human brain. Proc. Natl Acad. Sci. USA 106, 5743–5748 (2009).
Bakken, T. E. et al. A comprehensive transcriptional map of primate brain development. Nature 535, 367–375 (2016).
Schornig, M. et al. Comparison of induced neurons reveals slower structural and functional maturation in humans than in apes. Elife 10, https://doi.org/10.7554/eLife.59323 (2021).
Ostertag, E. M., Goodier, J. L., Zhang, Y. & Kazazian, H. H. Jr. SVA elements are nonautonomous retrotransposons that cause disease in humans. Am. J. Hum. Genet. 73, 1444–1451 (2003).
Wang, H. et al. SVA elements: a hominid-specific retroposon family. J. Mol. Biol. 354, 994–1007 (2005).
Glinsky, G. V. Mechanistically distinct pathways of divergent regulatory DNA creation contribute to evolution of human-specific genomic regulatory networks driving phenotypic divergence of Homo sapiens. Genome Biol. Evol. 8, 2774–2788 (2016).
Trizzino, M. et al. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 27, 1623–1633 (2017).
Trizzino, M., Kapusta, A. & Brown, C. D. Transposable elements generate regulatory novelty in a tissue-specific fashion. BMC Genomics 19, 468 (2018).
Kapusta, A. et al. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet. 9, e1003470 (2013).
Jacobs, F. M. et al. An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons. Nature 516, 242–245 (2014).
Haring, N. L. et al. ZNF91 deletion in human embryonic stem cells leads to ectopic activation of SVA retrotransposons and up-regulation of KRAB zinc finger gene clusters. Genome Res. 31, 551–563 (2021).
Vasieva, O. et al. Potential Impact of primate-specific SVA retrotransposons during the evolution of human cognitive function. Trends Evol. Biol. 6514, https://doi.org/10.4081/ni.2017.6514 (2017).
Hamilton, A. T. et al. Evolutionary expansion and divergence in the ZNF91 subfamily of primate-specific zinc finger genes. Genome Res. 16, 584–594 (2006).
Imbeault, M., Helleboid, P. Y. & Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 543, 550–554 (2017).
Robbez-Masson, L. et al. The HUSH complex cooperates with TRIM28 to repress young retrotransposons and new genes. Genome Res. 28, 836–845 (2018).
Kasten, M. R., Rudy, B. & Anderson, M. P. Differential regulation of action potential firing in adult murine thalamocortical neurons by Kv3.2, Kv1, and SK potassium and N-type calcium channels. J. Physiol. 584, 565–582 (2007).
Zhou, Y. D. et al. Arrested maturation of excitatory synapses in autosomal dominant lateral temporal lobe epilepsy. Nat. Med. 15, 1208–1214 (2009).
Czupryn, A. et al. Transplanted hypothalamic neurons restore leptin signaling and ameliorate obesity in db/db mice. Science 334, 1133–1137 (2011).
Bond, J. & Woods, C. G. Cytoskeletal genes regulating brain size. Curr. Opin. Cell Biol. 18, 95–101 (2006).
Cox, J., Jackson, A. P., Bond, J. & Woods, C. G. What primary microcephaly can tell us about brain growth. Trends Mol. Med. 12, 358–366 (2006).
Thornton, G. K. & Woods, C. G. Primary microcephaly: do all roads lead to Rome? Trends Genet. 25, 501–510 (2009).
Lizarraga, S. B. et al. Cdk5rap2 regulates centrosome function and chromosome segregation in neuronal progenitors. Development 137, 1907–1917 (2010).
Buchman, J. J. et al. Cdk5rap2 interacts with pericentrin to maintain the neural progenitor pool in the developing neocortex. Neuron 66, 386–402 (2010).
Barrera, J. A. et al. CDK5RAP2 regulates centriole engagement and cohesion in mice. Dev. Cell 18, 913–926 (2010).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Meisler, M. H., Hill, S. F. & Yu, W. Sodium channelopathies in neurodevelopmental disorders. Nat. Rev. Neurosci. 22, 152–166 (2021).
Veeramah, K. R. et al. De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP. Am. J. Hum. Genet. 90, 502–510 (2012).
Blanchard, M. G. et al. De novo gain-of-function and loss-of-function mutations of SCN8A in patients with intellectual disabilities and epilepsy. J. Med. Genet. 52, 330–337 (2015).
Lopez-Santiago, L. F. et al. Neuronal hyperexcitability in a mouse model of SCN8A epileptic encephalopathy. Proc Natl Acad Sci USA 114, 2383–2388 (2017).
Smith, S. & Stillman, B. Purification and characterization of CAF-I, a human cell factor required for chromatin assembly during DNA replication in vitro. Cell 58, 15–25 (1989).
Kaufman, P. D., Kobayashi, R., Kessler, N. & Stillman, B. The p150 and p60 subunits of chromatin assembly factor I: a molecular link between newly synthesized histones and DNA replication. Cell 81, 1105–1114 (1995).
Volk, A. & Crispino, J. D. The role of the chromatin assembly complex (CAF-1) and its p60 subunit (CHAF1b) in homeostasis and disease. Biochim. Biophys. Acta 1849, 979–986 (2015).
Cheloufi, S. et al. The histone chaperone CAF-1 safeguards somatic cell identity. Nature 528, 218–224 (2015).
Bhide, P. G. et al. Expression of normal and mutant huntingtin in the developing brain. J. Neurosci. 16, 5523–5535 (1996).
Reiner, A., Dragatsis, I., Zeitlin, S. & Goldowitz, D. Wild-type huntingtin plays a role in brain development and neuronal survival. Mol. Neurobiol. 28, 259–276 (2003).
Walker, F. O. Huntington’s disease. Semin. Neurol. 27, 143–150 (2007).
Glessner, J. T. et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 459, 569–573 (2009).
Slow, E. J. et al. Selective striatal neuronal loss in a YAC128 mouse model of Huntington disease. Hum. Mol. Genet. 12, 1555–1567 (2003).
O’Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science 309, 2033–2037 (2005).
Boguslawski, S. J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986).
Phillips, D. D. et al. The sub-nanomolar binding of DNA-RNA hybrids by the single-chain Fv fragment of antibody S9.6. J. Mol. Recognit. 26, 376–381 (2013).
Walker, S. C., Good, P. D., Gipson, T. A. & Engelke, D. R. The dual use of RNA aptamer sequences for affinity purification and localization studies of RNAs and RNA-protein complexes. Methods Mol. Biol. 714, 423–444 (2011).
Bond, J. et al. A centrosomal mechanism involving CDK5RAP2 and CENPJ controls brain size. Nat. Genet. 37, 353–355 (2005).
Moynihan, L. et al. A third novel locus for primary autosomal recessive microcephaly maps to chromosome 9q34. Am. J. Hum. Genet. 66, 724–727 (2000).
Abdullah, U. et al. A novel mutation in CDK5RAP2 gene causes primary microcephaly with speech impairment and sparse eyebrows in a consanguineous Pakistani family. Eur. J. Med. Genet. 60, 627–630 (2017).
Montgomery, S. H., Capellini, I., Venditti, C., Barton, R. A. & Mundy, N. I. Adaptive evolution of four microcephaly genes and the evolution of brain size in anthropoid primates. Mol. Biol. Evol. 28, 625–638 (2011).
Montgomery, S. H. & Mundy, N. I. Microcephaly genes evolved adaptively throughout the evolution of eutherian mammals. BMC Evol. Biol. 14, 120 (2014).
Evans, P. D., Vallender, E. J. & Lahn, B. T. Molecular evolution of the brain size regulator genes CDK5RAP2 and CENPJ. Gene 375, 75–79 (2006).
Rowe, H. M. et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463, 237–240 (2010).
Schultz, D. C., Ayyanathan, K., Negorev, D., Maul, G. G. & Rauscher, F. J. 3rd SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 16, 919–932 (2002).
Turelli, P. et al. Interplay of TRIM28 and DNA methylation in controlling human endogenous retroelements. Genome Res. 24, 1260–1270 (2014).
Kobayashi, K. et al. An ancient retrotransposal insertion causes Fukuyama-type congenital muscular dystrophy. Nature 394, 388–392 (1998).
Makino, S. et al. Reduced neuron-specific expression of the TAF1 gene is associated with X-linked dystonia-parkinsonism. Am. J. Hum. Genet. 80, 393–406 (2007).
Vogt, J. et al. SVA retrotransposon insertion-associated deletion represents a novel mutational mechanism underlying large genomic copy number changes with non-recurrent breakpoints. Genome Biol. 15, R80 (2014).
Wilund, K. R. et al. Molecular mechanisms of autosomal recessive hypercholesterolemia. Hum. Mol. Genet. 11, 3019–3030 (2002).
Borges-Monroy, R. et al. Whole-genome analysis reveals the contribution of non-coding de novo transposon insertions to autism spectrum disorder. Mob. DNA 12, 28 (2021).
Ge, L. Q. et al. Suppressing male spermatogenesis-associated protein 5-like gene expression reduces vitellogenin gene expression and fecundity in Nilaparvata lugens Stal. Sci. Rep. 6, 28111 (2016).
Liu, Y., Black, J., Kisiel, N. & Kulesz-Martin, M. F. SPAF, a new AAA-protein specific to early spermatogenesis and malignant conversion. Oncogene 19, 1579–1588 (2000).
Zaqout, S., Bessa, P., Kramer, N., Stoltenburg-Didinger, G. & Kaindl, A. M. CDK5RAP2 is required to maintain the germ cell pool during embryonic development. Stem Cell Rep. 8, 198–204 (2017).
Franke, B., Figiel, M. & Engele, J. CNS glia are targets for GDNF and neurturin. Histochem. Cell Biol. 110, 595–601 (1998).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Acknowledgements
The authors thank Anderson Lab members Oriana DiStefano, Greg Salimando, Rebecca Broadhurst and Scott Rochard for mouse colony upkeep and genotyping. We thank David Stoppel for mouse brain dissections and Annmarie McKeon for SVA genome annotation. We thank the BIDMC Flow Cytometry Facility, the Harvard Center for Biological Imaging at Harvard University, the Neurobiology Imaging Facility at Neurobiology Department of Harvard Medical School for consultation and instrument availability that supported this work (in part through a NINDS P30 Core Center grant #NS07203) and Boston Children’s Hospital IDDRC (1U54HD090255, P30HD18655). This work was supported by funding to M.P.A. from The National Institute of Mental Health (R01MH112714, R01MH114858, and 1R21MH100868), The National Institute of Neurological Disorders and Stroke (1R01NS08916), The Eunice Kennedy Shriver National Institute of Child Health and Human Development (1R21HD079249), The Nancy Lurie Marks Family Foundation, Landreth Foundation, Autism Speaks/National Alliance for Autism Research, and the Simons Foundation.
Author information
Authors and Affiliations
Contributions
M.N., W.C., E.O., Y.H., Y.N., M.J., M.B., A.M., and M.P.A. designed the study. M.N and M.P.A wrote the manuscript. M.N., W.C., E.O., Y.N., Y.H., M.J., M.B., A.M. performed all the experiments and analyses except for the following: W.C. and Y.N. performed the electrophysiology.
Corresponding author
Ethics declarations
Competing interests
Y.H., Y.N., and M.P.A. became employees of Regeneron Pharmaceutical. The other authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Xiaodong Liu, Gemma Carvill and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nadler, M.J.S., Chang, W., Ozkaynak, E. et al. Hominoid SVA-lncRNA AK057321 targets human-specific SVA retrotransposons in SCN8A and CDK5RAP2 to initiate neuronal maturation. Commun Biol 6, 347 (2023). https://doi.org/10.1038/s42003-023-04683-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04683-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.