Abstract
Productive transcriptional elongation of many cellular and viral mRNAs requires transcriptional factors to extract pTEFb from the 7SK snRNP by modulating the association between HEXIM and 7SK snRNA. In HIV-1, Tat binds to 7SK by displacing HEXIM. However, without the structure of the 7SK-HEXIM complex, the constraints that must be overcome for displacement remain unknown. Furthermore, while structure details of the TatNL4-3-7SK complex have been elucidated, it is unclear how subtypes with more HEXIM-like Tat sequences accomplish displacement. Here we report the structures of HEXIM, TatG, and TatFin arginine rich motifs in complex with the apical stemloop-1 of 7SK. While most interactions between 7SK with HEXIM and Tat are similar, critical differences exist that guide function. First, the conformational plasticity of 7SK enables the formation of three different base pair configurations at a critical remodeling site, which allows for the modulation required for HEXIM binding and its subsequent displacement by Tat. Furthermore, the specific sequence variations observed in various Tat subtypes all converge on remodeling 7SK at this region. Second, we show that HEXIM primes its own displacement by causing specific local destabilization upon binding — a feature that is then exploited by Tat to bind 7SK more efficiently.
Similar content being viewed by others
Introduction
Transcription of all class II genes is a highly regulated process within cells. Shortly after promoter clearance, RNA Polymerase II is inhibited by negative elongation factors1,2,3,4,5. Release from this stalled state requires all components to be phosphorylated by the positive elongation factor pTEFb, a heterodimeric complex consisting of Cyclin T1 and the cyclin-dependent kinase Cdk96,7,8,9,10,11,12. However, most of the pTEFb is kept catalytically inactive in the nucleus by the 7SK small nuclear ribonucleoprotein (7SK snRNP) through its interactions with the HEXIM adapter protein and the 5’ stemloop-1 of the 7SK RNA13,14,15,16,17,18,19,20,21,22,23 (7SK-SL1). Thus, productive transcriptional elongation of many genes requires transcriptional factors to extract pTEFb from the 7SK snRNP—a process that involves manipulating the interaction between HEXIM and 7SK. This association between 7SK and HEXIM tightly controls the balance between active and inactive pTEFb, and dysregulation of this interaction can have serious biological consequences, including cardiac hypertrophy and breast and pancreatic cancers24,25,26,27. Furthermore, as many viruses rely on the host transcriptional machinery to produce mRNA and genomes, they have also evolved mechanisms to capture pTEFb28,29,30. One such unique case is the human immunodeficiency virus (HIV), which utilizes the viral Tat protein to extract pTEFb by binding to the same region of 7SK as HEXIM and directly displacing it30,31,32,33,34. Structural insights into the consequence of HEXIM binding to 7SK and how positive transcriptional factors like Tat compete with it are therefore important for understanding HEXIM’s potency as a critical negative regulator.
To date, two HEXIM proteins have been identified that can carry out the same function and both bind 7SK with identical regions of their Arginine-Rich Motifs (ARMs) (residues 151–159 in HEXIM1 and 89–97 in HEXIM2)30,35,36,37,38. Although HEXIM binds 7SK as a dimer, only one ARM directly contacts 7SK by engaging the apical region of stemloop-1 (G26 to C85, 7SK-SL1apical)38,39,40,41,42,43,44. Both in vitro and in vivo studies have shown that this represents the sole interaction between the two molecules that must be modulated to release pTEFb37,38,39,41,42.
Our previous work showed that 7SK-SL1apical is enriched in arginine sandwich motifs (ASMs)45. ASMs are defined by two nucleotides that stack in a manner that allows for intercalation of arginine guanidinium moieties between the aromatic rings of the bases45,46,47,48,49,50,51. While a bulge pyrimidine forms the cap by engaging in a triple-base interaction with an n + 2 base pair in the stem, a Watson–Crick base-paired nucleotide preceding the bulge forms the base of the interaction. In the free 7SK-SL1apical, three such bulges fold into preformed arginine sandwich motifs (ASM1, ASM2, and ASM4) poised for arginine guanidinium moieties to dock into them. A fourth bulge folds into a pseudo configuration (pseudo-ASM3) where U40 can form a triple-base interaction with the A43-U66 base pair to form the cap, but the base of the sandwich is sequestered in a reverse Hoogsteen interaction, excluding it from use as a classical ASM. Our work also showed that HIV-1 Tat NL4-3 (TatNL4-3) uses its arginine-rich motif to intercalate arginines not only into the three preformed ASMs, but also to remodel the pseudo-ASM into a classical ASM45. This structural remodeling of pseudo-ASM3 is a key mechanism through which Tat displaces HEXIM.
However, without the structure of the HEXIM:7SK-SL1apical interaction, it is currently unclear what structural constraints Tat would need to overcome to access pTEFb. Furthermore, while the Tat ARM is highly conserved, sequence variations exist in different strains that allow for HEXIM displacement. For example, the ARM of Tat Finland (TatFin; KR52KHRRR) differs from HEXIM (KK151KHRRR) by only a single amino acid and would lack one of the ASM interactions from the previously described Tat NL4-3 strain (KR52RQRRR). Additionally, while Tat subtype G (TatG; KR52R53HRRR) has an equivalent number of arginines as TatNL4-3, the critical linker sequence connecting the ASM3/ASM4 and ASM1/ASM2 interactions is the same as in HEXIM. In this study, we present the structure of the 7SK-SL1apical in complex with the HEXIM, TatFin, and TatG ARMs. Despite sequence variations, the structures show deep major groove intercalations of all ARMs, albeit with differential interactions with pseudo-ASM3 and ASM4. Furthermore, we show that HEXIM causes local destabilization of ASM4, enhancing Tat’s affinity for 7SK. These studies thus uncover a feature in which HEXIM facilitates its own displacement by increasing conformational sampling, which may be a more general mechanism of pTEFb capture.
Results
Comparative binding affinities of HEXIM and Tat to 7SK
As a first step toward identifying the comparative thermodynamic properties of 7SK recognition between HEXIM and Tat, we performed binding studies using isothermal titration calorimetry (ITC). ITC traces of the ARMs into 7SK-SL1apical-AGU produce significant nonspecific heats of binding as previously observed52. In a previous study by Brillet et al., high salt conditions (0.5 M NaCl) were used to abrogate such nonspecific interactions that stem from charge-charge interactions between the positively charged peptides and the negative RNA backbone52. While such a strategy is commonly used, it is not ideal for this system as the structuring of ASMs in 7SK is highly sensitive to ionic conditions and folds only around physiological salt conditions (Supplementary Figs. 1, 2)45. Therefore, to subtract the nonspecific heats of binding, we designed a control construct that lacks all ASMs (7SK-SL1apicalΔASM). Indeed, the heats obtained from peptide titrations into this control construct completely accounted for the nonspecific heats, the subtraction of which allowed for experimental baselines to approach zero at saturation (Supplementary Fig. 2b). Titration of the N-terminal ARM residues of HEXIM (R146QLGKKKHRRR156; HEXIMN-ARM) into 7SK-SL1apical-AGU containing an AGU triloop engineered to prevent low levels of dimerization gave a Kd of 229 ± 20 nM (N = 1 ± 0.1; Fig. 1a). The redesign of the previously used GAGA tetraloop45 to an AGU triloop was done to prevent weak association between the tyrosine in the peptide and the tetraloop. Nevertheless, while the affinities obtained by AGU-triloop are 2 to 3-fold weaker, the relative difference between HEXIM and Tat are similar (see below).
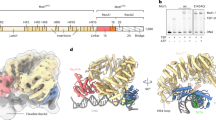
(Left) Cartoon representation of the HEXIM dimer and pTEFb heterodimer binding to the 7SK snRNP (top left). Upon introduction of Tat, HEXIM is displaced from the snRNP (bottom left). Not depicted are MEPCE and LARP7. Representative ITC data for HEXIMN-ARM binding to 7SK-SL1apical-AGU (G26-C85) (a) compared to full-length HEXIM1 bound to 7SK-SL1Full (G1-C108) with a wild-type loop (b) or an AGU triloop (c) and full-length HEXIM1 binding to 7SK-SL1apical-AGU (d) all show similar binding affinities, indicating that the loop does not play a significant role in dimeric HEXIM binding and that the HEXIMN-ARM:7SK-SL1apical-AGU complex represents the minimal binding interaction. Representative ITC traces of the TatG (e) and TatFin (f) ARMs into 7SK-SL1apical-AGU show an ~2.8 and 1.3-fold increased binding affinity compared to HEXIMN-ARM, respectively. All reported values are for n = 3 replicates.
To confirm that interactions do not extend to the loop and are represented by these minimal constructs, we performed studies with full-length HEXIM into full-length 7SK-SL1Full (G1-C108), 7SK-SL1Full-AGU, and 7SK-SL1apical-AGU, all of which give rise to similar Kds (209 ± 30 nM, Kd = 200 ± 20 nM, and Kd = 206 ± 60 nM, respectively) and bound expectedly as dimers (N = 2.1 ± 0.2, N = 2 ± 0.07, and N = 1.8 ± 0.03, respectively; Fig. 1b–d). Furthermore, NMR studies comparing full-length dimeric HEXIM1:7SK-SL1apical-AGU and HEXIMN-ARM:7SK-SL1apical-AGU complexes show that binding of either full-length HEXIM or the N-ARM gives rise to the same chemical shifts in 7SK-SL1apical-AGU, indicating that the HEXIMN-ARM:7SK-SL1apical-AGU interaction represents the biologically relevant mode of HEXIM binding to 7SK (Supplementary Fig. 2).
Our previous work showed that the TatNL4-3 (KR52RQRRR) ARM represents the interaction domain between Tat and 7SK-SL1apical and has an approximately two-fold increased affinity over the HEXIMN-ARM, which provides an explanation for HEXIM displacement45. ITC traces show that Tat Subtype G’s ARM (KR52RHRRR), which also has two N-terminal arginines, binds 7SK-SL1apical-AGU with a Kd of 81 ± 10 nM (N = 1.1 ± 0.1; Fig. 1e), which is an approximately 2.8-fold increased binding affinity over HEXIMN-ARM (Supplementary Table 1). On the other hand, Tat Finland’s ARM (KR52KHRRR), despite having an additional N-terminal arginine compared to the HEXIMN-ARM (R52 and K151, respectively), does not have a statistically significant increase in binding affinity over HEXIMN-ARM (Kd of 172 ± 10 nM, N = 1 ± 0.02; Fig. 1f). Overall, these results highlight the need for understanding the HEXIM-bound 7SK; while the increased TatG affinity would allow for HEXIM displacement, it is unclear how TatFin can achieve the same biological output.
Preformed configurations of ASM1 and ASM2 provide a common mode of interaction with C-terminal arginines
To understand how the HEXIMN-ARM and the various Tat ARMs interact with 7SK-SL1apical-AGU, we utilized a combination of small-angle X-ray scattering (SAXS) and NMR. All reconstructed ab initio SAXS envelopes showed no major overall global changes between peptide-bound and free 7SK-SL1apical-AGU (Supplementary Fig. 3). Numerous intermolecular NOEs place both HEXIM and Tat arginine-rich motifs into the major groove of the RNA and allow us to define their interactions with all ASM regions. Base pairs in the lower part of the stemloop below the G79-U32 base pair, as well as the CAGUG pentaloop do not give any intermolecular NOEs, indicating that the interactions are contained within a single turn of the helix (Fig. 2 and Table 1).
a Cartoon depicting an arginine sandwich motif. b Secondary structure of free 7SK-SL1apical-AGU with a modified AGU triloop. The base and cap residues forming ASM1, ASM2, pseudo-ASM3, and ASM4 are colored in orange, green, magenta, and blue, respectively. Dashed arcs represent triple-base interactions from the bulge to the stem, giving rise to the caps of the sandwiches. Representative NMR structures of 7SK-SL1apical-AGU bound to (c) HEXIMN-ARM, (d) TatFin, and (e) TatG show engagement with all ASMs.
In the free 7SK-SL1apical-AGU, ASM1 and ASM2 are placed in tandem orientation, and upon titration of the various ARMs, all expected NOEs for such configurations are retained. Unlike a typical ASM where the following nucleotide after the bulge is in a canonical Watson–Crick base pair, in ASM1, the residue A77 is configured into an A34-A77 base pair. A NOE from the A77 H8 proton to the H1′ of C75 positions this residue under the C75 cap (Supplementary Fig. 4). This confirms a planar orientation of C75 with the C33–G78 base pair and configures A77 in such a way that it is perfectly positioned to interact with the guanidinium moiety of R156 in HEXIMN-ARM and R57 in TatFin and TatG, which intercalate between C75 and G74 in a manner identical to canonical ASMs (Supplementary Figs. 4–7).
Similarly, in ASM2, the C71+ base also retains its protonation at the N3 position, as evidenced by a downfield shift of the N4 amino protons (Supplementary Fig. 8). The guanidinium moiety of R155 in HEXIMN-ARM and R56 of TatFin and TatG interact with G73 by intercalating between the C71+ cap and G70 base of the motif (Supplementary Figs. 5–7). Additionally, intermolecular NOEs from the aromatic protons of the C75 and C71+ caps and the G74 and G70 bases of ASM1 and ASM2 to the Hγ and the Hδ protons confirm that consecutive arginines R156 and R155 interact in a ladder-like configuration with the tandem performed motifs ASM1 and ASM2, respectively (Fig. 3a and Supplementary Fig. 6c). Such NOEs are also observed in both the TatFin and the TatG-bound complexes, confirming the similar placement of the C-terminal R57 and R56 into the tandem ASM1 and ASM2, respectively (Fig. 3a and Supplementary Fig. 7a, b, d, e). Taken together, the structures reveal a common mode of interaction between the non-varying C-terminal arginines and the tandem ASMs.
a C-terminal arginines of all ARMs dock into ASM1 (orange) and ASM2 (green) with identical tertiary structures. b The U68-A39 base pair rearranges into a cis-Hoogsteen/sugar interaction upon HEXIM binding (left) while TatFin (middle) and TatG (right) both remodel ASM3 (magenta) by rearranging the U68-A39 base pair into a Watson–Crick interaction. c K150 and K151 in HEXIM (left), R52 and K51 in TatFin (middle), and K51, R52, and R53 in TatG (right) interact with the apical ASMs. d Spacer residues between the ASM1/ASM2 and ASM3/ASM4 regions are positioned near ASM1 and ASM2. In the case of TatFin (middle), K53 also acts as a spacer residue to allow for the remodeling of ASM3 by R52.
Rearrangement of pseudo-ASM3 allows for HEXIM N-terminal interactions
In the free 7SK-SL1apical-AGU, pseudo-ASM3 and ASM4 adopt a pseudo-symmetrical architecture where the two motifs are spatially opposed. Upon HEXIMN-ARM binding, the pseudo-ASM3 maintains its U40:A43-U66 triple-base interaction although the base of the sandwich, A39, rearranges from a reverse Hoogsteen interaction with U68 into a cis-Hoogsteen/sugar interaction, giving rise to an alternate pseudo configuration. (Fig. 3b). This is evidenced both by NOEs from the U68 imino proton to the A39 amino protons and NOEs from the U68 H2′ and H3′ protons to the A39 H8 proton (Supplementary Fig. 8b). This frees up the U68 imino proton to engage the backbone carbonyl of K152 while simultaneously bringing the N1 proton acceptor of A39 into the major groove to hydrogen-bond with the side chain Hε protons of K151 (Fig. 3c). Thus, both K151 and 152 can enter deep into the major groove by remodeling the pseudo-ASM3.
The amino side chain of K151 is within hydrogen-bonding distance of the A39 N1 nitrogen as evidenced by NOEs from the K151 Hγ and Hβ protons to the C37 H6 and H5 protons, respectively, and from the K151 Hε protons to the C38 H6 and H5 protons (Fig. 3c and Supplementary Fig. 6f). Additionally, NOEs between the K152 Hβ protons with the U68 H5 proton, the K152 Hδ protons with the C67 and U66 H5 protons, and the K152 Hε protons with the C67 H5 and H6 protons position the amino side chain of K152 within hydrogen-bonding distance of the C67 backbone (Fig. 3c and Supplementary Fig. 6e, f). This gives rise to a forked configuration of the two lysines, orienting the side chain amino groups towards the phosphate backbones on opposite ends of the groove.
Unlike the other three ASMs, where the NOEs clearly define a single predominant structural configuration as described above, multiple dynamic states exist for ASM4 (see below). In the most abundant form, the preformed nature found in the free state is retained as evidenced by a direct imino-to-imino connectivity between U44 and U63 along with maintenance of the G46–C62 Watson–Crick base pair (Supplementary Fig. 8c). In fact, this interaction is stabilized by K150, which displays NOEs between the Hε protons with the U63 and the U40 H5 protons, positioning the amino side chain within hydrogen-bonding distance of the O4 atoms of both U63 and U40 (Supplementary Fig. 6d). Additional intermolecular interactions between the U40 H5 proton and the U63 H5 and H1′ protons with the K150 Hδ, Hγ, and Hβ protons places K150 directly under the U63 cap of ASM4 (Supplementary Fig. 6d, e). Taken together, these data show that despite the lack of arginines, the lysine-rich N-terminus of HEXIMN-ARM can be accommodated by 7SK: the Watson–Crick face of A39 turns from the minor into the major groove to interact with K151 and 152, which then positions K150 to interact with the oxygen-rich environment of the U63 and U40 caps.
Conformational plasticity of the ASM3/ASM4 region provides differential mode of interactions with N-terminal and spacer residues
Our previous study showed that TatNL4-3 displaces HEXIM by remodeling the pseudo-ASM3 into a canonical ASM3 to allow for arginine intercalation45. Furthermore, an additional arginine docks into the preformed ASM4. While the mechanism of remodeling pseudo-ASM3 is conserved upon binding of both TatFin and TatG ARMs (Fig. 3b, f and Supplementary Fig. 6), both the drivers of the conformational switch and the engagement of the ASM4 vary depending on differences in amino acid sequences.
While TatFin has two major differences from TatNL4-3 (K53 to R53 and spacer H54 to Q54, respectively), it only differs by a single amino acid from HEXIM (R52 and K151, respectively). Like TatNL4-3, R52 is responsible for remodeling pseudo-ASM3 (Fig. 3c and Supplementary Fig. 7a). However, while R53 in TatNL4-3 flips over R52 and engages ASM4, the equivalent K53 stays in the spacer region between the ASM1/ASM2 and ASM3/ASM4 regions in a manner similar to HEXIM as evidenced by NOEs between the K53 Hβ protons with the U68 H5 proton, the K53 Hδ protons with the C67 and U66 H5 protons, and the K53 Hε protons with the C67 H5 and H6 protons, which position the amino side chain of K53 within hydrogen-bonding distance of the C67 backbone (Fig. 3c and Supplementary Fig. 7a).
As for HEXIM, ASM4 remains unoccupied upon binding TatFin and the structure shows that the K51 amino side chain is positioned to hydrogen-bond with the U63 ribose ring in a stabilizing interaction (Fig. 3c). This is evidenced by NOEs of the K51 Hδ protons with the U63 H5 and H1′ protons and the K51 Hε protons with the U63 2′ hydroxyl proton (Supplementary Fig. 7c). Furthermore, the N-terminal K50 exits near the apical loop, with NOEs observed of the K50 Hδ and Hε protons with the C38 and C37 H5, and H1′ protons position the amino side chain of K50 to the C38 phosphate backbone (Fig. 3c and Supplementary Fig. 7a).
Finally, in evaluating the structural consequences of the spacer substitution, we see that H54 and R55 in TatFin remain near ASM1 and ASM2, similar to what is found in HEXIM. This is evidenced by NOEs of the H54 (H153 in HEXIM) Hβ protons with the C35, C36, and C37 H5 protons, placing H54 near ASM2, whereas the R55 (R154 in HEXIM) Hδ protons display NOEs with the A34 H1′ proton and the C33 H1’, H5, and H6 protons, positioning this spacer residue near ASM1 (Fig. 3d and Supplementary Figs. 6, 7). This is in contrast with the binding mode of TatNL4-3 in which the intercalation of R53 into ASM4 drags both the Q54 and R55 spacer residues towards the apical ASMs.
The importance of the histidine H54 spacer is even more evident in the TatG strain where it represents the only difference from TatNL4-3. This single difference changes the identity of the arginine that remodels pseudo-ASM3. In this ARM, the positioning of H54 near ASM2 precludes R53 from reaching ASM4 to accomplish the inverse intercalation seen in NL4-3 (Fig. 3d and Supplementary Fig. 7d, e). The interactions with the apical ASMs thus occur in a ladder-like manner where R53 intercalates into the remodeled ASM3 whereas R52 intercalates into ASM4 (Fig. 3c, d and Supplementary Fig. 7a, d, e). K51 makes the final stabilizing interaction with NOEs seen between the Hε protons and the U63 2′ hydroxyl proton, indicating a hydrogen-bonding interaction between the K51 amino side chain and the U63 ribose ring (Fig. 3d and Supplementary Fig. 7f). Taken together, these studies show that arginine sandwich motifs provide mini domains that arginine-rich motifs of proteins can differentially interact with to achieve deep major groove binding into the stem of 7SK-SL1apical-AGU.
HEXIM allows for increased conformational sampling of apical ASMs
While titration of all four arginine-rich motifs stabilizes the majority of 7SK-SL1apical-AGU into one predominant configuration, the HEXIM ARM is an outlier wherein binding causes ASM1 and ASM4 to become destabilized and exhibit multiple conformations (Fig. 4a, b and Supplementary Fig. 4). In such conformations, the NOEs between the imino protons of U63 and U44 disappear, indicating the disruption of the U63:U44-A65 triple and loss of ASM4 (Supplementary Fig. 8c). The destabilization of this region is also indicated by the line-broadening of K150, which interacts with U63 in the folded configuration (Supplementary Fig. 6e).
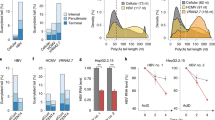
a Comparison of enthalpic and entropic contributions between TatG, TatFin, and HEXIMN-ARM in complex with 7SK-SL1apical-AGU, and full-length HEXIM in complex with 7SK snRNA. Entropy values were calculated using a T value of 298 K. The reversal in the entropic and enthalpic contribution for Tat ARM compared to HEXIM is evident with HEXIM having an entropically-driven binding profile. NMR competition titration analysis showing binding of 7SK by TatG (b) and TatFin (c) concomitant with the total displacement of HEXIMN-ARM. Data are shown for the A39 (left) and A34/77 (right) h2-c2 correlations. The increase in Tat engagement of ASM1 for the HEXIMN-ARM-bound complex is evident by the lack of free-RNA populations for the A34 resonance in the competition experiment compared to binding to free 7SK. Furthermore, the destabilization of A34 in ASM1 by HEXIM is indicated by multiple bound states. Also shown for comparison is the complete engagement of A39 by all ARMs. d Representative ITC data for full-length HEXIM bound to 7SK snRNP demonstrating expected stoichiometry and specific binding. All reported values are for n = 3 replicates.
The destabilization of 7SK-SL1apical-AGU only by HEXIM is further evident when comparing the thermodynamic profiles between Tat and HEXIM. The binding of TatFin and TatG strains is enthalpically driven (ΔH = −7.5 ± 0.2 and −8.9 ± 2.2 kcal mol−1, respectively; Fig. 4c) with a modest entropic contribution (−TΔS = −1.7 ± 0.3 and −2 ± 0.8 kcal mol−1, respectively; Fig. 4c). On the other hand, HEXIM binding is entropically enhanced by ~2.5-fold over both Tat strains (−TΔS = −4.6 ± 0.8 kcal mol−1, ΔH = −4.4 ± 0.7 kcal mol−1; Fig. 4c). The Brillet et al. study performed in 0.5 M salt saw an unfavorable entropic contribution for Tat and a negligible entropic for HEXIM binding, underscoring the importance of maintaining native ASM folding for a mechanistic understanding of this biological process52. Nevertheless, the overall observation that HEXIM binding is comparatively more entropic than Tat agrees with our results52.
To evaluate the implication of HEXIM’s ability to locally destabilize ASM1 and ASM4 in the context of its displacement required for transcriptional regulation, we compared TatFin and TatG ARM binding to 7SK-SL1apical both free and in the presence of HEXIMN-ARM. Due to the modest differences in binding energetics between the different ARMs, competition experiments using ITC were not tractable. A 1:1 titration of both TatG and TatFin into 7SK in the NMR shows the ability to completely engage ASM2 and ASM3, while a significant fraction of ASM1 and ASM4 shows the presence of free configurations, indicating reduced access for the termini. However, upon titration of both Tats into the HEXIM-bound 7SK complex, we observe not only complete engagement of all ASMs but also a total displacement of HEXIM (Fig. 4a, b). This is especially striking given that the binding affinities of TatFin and HEXIM for free 7SK are equivalent. Taken together, these data indicate that Tat can better engage 7SK that is destabilized by HEXIM at the outer ASMs. Finally, ITC data of full-length HEXIM bound to full-length 7SK snRNA (N = 1.9 ± 0.1; Fig. 4d) show that an entropy-driven interaction is maintained and, in fact, is even more pronounced (−TΔS = −6.4 ± 1.3 kcal mol−1, ΔH = −2.6 ± 1 kcal mol-1; Fig. 4c), suggesting that HEXIM binding may globally increase the conformational space sampled by the 7SK snRNP complex. These studies suggest that destabilization by HEXIM may play an important role in how transcription factors access 7SK for pTEFb capture.
Discussion
The 7SK snRNP represents a central biomolecule that a wide range of transcriptional factors needs to interact with to access pTEFb to control transcriptional elongation. In particular, pTEFb extraction by HIV Tat from this complex requires manipulating the interaction between the 7SK snRNA and the HEXIM adapter protein. In this study, we solved the structures of the RNA binding domains of HEXIM and Tat bound to 7SK and gained several insights into their functional significance, including the malleability of 7SK, the local destabilization by HEXIM, and the specific sequence variations of Tat.
The structures show that both HEXIM and Tat directly bind the stem of 7SK-SL1apical through intercalation of arginine-rich motifs into an entire helical turn of the major groove. This is unusual as RNA major grooves are deep and narrow, making them generally inaccessible for protein binding. The architecture of the four sandwich motifs in 7SK allows for transcriptional regulators to differentially utilize their ARMs. On the one hand, the tandem preformed ASMs, ASM1 and ASM2, remain unchanged from their free configuration upon encountering the C-terminal arginines of Tat and HEXIM. On the other hand, the apical pseudo-symmetrical ASMs, pseudo-ASM3, and ASM4, reconfigure depending on their binding partners. The structures show that the ASM3 region can adopt at least three different base pair interactions: a reverse Hoogsteen in the free state, a cis-Hoogsteen/sugar interaction upon HEXIM binding, and a Watson–Crick base pair upon Tat binding. The cis-Hoogsteen/sugar interaction is especially significant because it allows HEXIM to enter the major groove despite the lack of arginines in the N-terminus. Similarly, while ASM4 retains its preformed configuration found in the free state upon Tat binding, it can be destabilized in the presence of HEXIM and adopt multiple flexible states. Taken together, these studies show that 7SK is adaptable in its ASM architecture, which can be modulated upon encountering different transcription factors.
Comparative analyses of HEXIM and Tat provide insights into how both positive and negative regulators can manipulate 7SK to carry out their transcription roles. Our studies implicate HEXIM as potentially having a dual structural role. On the one hand, it can bind with high affinity to the apical portion of 7SK-stemloop-1, and on the other hand, it simultaneously causes local destabilization of this region, enhancing the binding of a positive regulator such as Tat. In comparison to Tat, the thermodynamic profile and solution-state characteristics of HEXIM binding show an entropy-driven mode of interaction that is particularly attributed to the destabilization of ASM1 and ASM4 regions, indicating a mechanism in line with conformational selection. Indeed, mutational studies have shown that deletion of U63 significantly reduces HEXIM binding37,43. This expansion in the dynamic state of 7SK surrounding the ASM1 and ASM4 region is also supported both by in vivo SHAPE analysis where U63 and C75 become ultra-reactive upon HEXIM:pTEFb binding53. Such increased conformational sampling was also demonstrated by structural and molecular dynamics modeling45,52,53,54,55,56,57. Furthermore, we show that Tat capitalizes on this increased dynamic state, binding to more motifs with greater affinity to the HEXIM-bound complex than to free 7SK. While the use of a HEXIM-displacement mechanism for pTEFb capture by binding to 7SK-SL1 has yet to be discovered for cellular factors, the destabilization-driven preparation of 7SK snRNP may potentially be a general feature exploited by specialized transcriptional factors.
Comparative analysis of HEXIM and Tat also sheds light on the sequence requirements of ARMs for 7SK binding. While N-terminal lysines of HEXIM allow for destabilization of ASM4, the anchoring required to enter the major groove can only be provided by the stacking of C-terminal arginines within ASM1 and ASM2. Indeed, the importance of these C-terminal arginines for HEXIM binding is supported by their nearly complete conservation across metazoan species58. Conversely, the equivalent arginines in HIV-1 Tat occur as a consecutive pair only in ~50% of reported strains, albeit with the strong requirement of at least one arginine. The structures show that these variations may be possible due to the anchoring provided by the arginines that intercalate into the apical ASMs. Nevertheless, when two arginines are present in Tat, the interactions with the tandem ASMs mirror HEXIM.
Furthermore, differences in N-terminal and spacer ARM residues orchestrate the structural modulations of the apical ASMs. To accommodate the continuation of the HEXIM ARM chain from the ASM1/ASM2 to the ASM3/ASM4 region required for U63 destabilization, K152 induces the reconfiguration of pseudo-ASM3 from a reverse Hoogsteen to a cis-Hoogsteen/sugar interaction. In all variations of N-terminal Tat sequences studied, binding is concomitant with the rearrangement of pseudo-ASM3 into a canonical ASM3 through the intercalation of an arginine.
The structures also provide insights into specific sequence variations that occur in the highly conserved Tat ARM to displace HEXIM. When two arginines are available in the N-terminal residues, both are involved in arginine sandwich interactions, providing a twofold increase in affinity; however, either R52 (TatNL4-3) or R53 (TatG) can act as the remodeler. This can be explained by the presence of either glutamine or histidine spacer, respectively, which is the only amino acid difference between the two strains. As glutamine (75%) and histidine (15%) make up most of the sequence variation in this spacer, the structures show that these two spacer residues drive the differential positioning of the arginine remodeler. In the TatFin strain, which has a histidine spacer, it is the R52 that acts as a remodeler. In this case, the R53K substitution provides the stabilizing interactions to reposition the single R52 arginine near pseudo-ASM3.
It is also interesting to compare the mode of binding of TatFin to HEXIM. First, the single residue difference (R52 vs K151) provides TatFin with the additional ASM intercalation required for displacement. Thus, Tat has evolved specific sequence variations that allow for the reconfiguration of pseudo-ASM3, even in cases where there is only a single variation from HEXIM. Second, despite both ARMs having lysines positioned near ASM4, only HEXIM leads to local destabilization. Our studies, therefore, provide HEXIM as an example of a negative regulator that primes its own displacement by locally destabilizing 7SK. Overall, these studies have broader implications for 7SK snRNP-mediated regulation. Given that the destabilization-driven displacement is a robust mechanism, it is possible that other yet-to-be-identified cellular and viral transcriptional regulators recruit pTEFb through direct intercalation of ARMs into 7SK-SL1apical. Furthermore, as a destabilized state of 7SK snRNP is what is presented to all transcriptional regulators, the mechanisms necessary to extract pTEFb may converge on capitalizing on this conformational heterogeneity.
Methods
RNA sample preparation
RNA samples used for biophysical experiments were synthesized by in vitro transcription using T7 RNA polymerase with either plasmid DNA or with synthetic DNA templates containing 2′-O-methylated (Integrated DNA Technologies) containing the T7 promoter and the desired sequences. Plasmid DNA for 7SK-SL1Full-WT and 7SK-SL1Full-AGU containing the T7 promoter, insert, and SmaI recognition sequence were cloned by Genscript in between the EcoRI and BamHI restriction sites of a puc19 vector. Plasmid DNA was prepared for in vitro transcription from a 5 mL overnight culture of NEB 5α Competent E.coli (C29871) transformed with the plasmid using Qiaprep Spin Miniprep Kit (Qiagen 27104). 10 μL of purified DNA were combined with 25 μL of 2′-O-methylated reverse primer at 100 μM (5′-mGmGAGCGGTGAGG GAGGAAG-3′ where m indicates 2′ O-methylated nucleotides), 25 μL of forward primer at 100 μM (5′-GACAAGCCCGTCAGGG-3′), 2.44 mL of water, and two tubes of EconoTaq PLUS 2X Master Mix (Lucigen 30035-2). The 5 mL mixture was then aliquoted into 50 μL increments in a 96-well PCR plate and the templates for in vitro transcription reactions were amplified using the following PCR protocol: 95 °C for 5 min, 34 cycles of (95 °C for 30 s, 50 °C for 1 min, and 68 °C for 90 s), and 68 °C for 5 min. After PCR amplification, reactions were pooled into 5 mL volume in a 50 mL Falcon tube and 0.5 mL of 3 M sodium acetate, pH 5 and 32 mL of 100% ethanol were added to the mixture and chilled at −80 °C for at least 30 min before spinning down at 9000×g for 10 min at 4 °C. The ethanol was decanted, and the pellet was left to dry overnight before in vitro transcription use. Template preparation for 7SK-SL1apical-AGU using 2′-O-methylated reverse primers in order to suppress the heterogeneity at the 3′ end of the transcripts involved combining 15 mL of both forward (5′-TAATACGACTCACTA TAGGGATCTGTCACCCCATTGATCGCCAGTGGCTGATCTGGCTGGCTAGGCGGGTCCC-3′) and reverse (5′-mGmGGACCCGCCTAGCCAGCCAGATCAGCCACTGGC GATCAATGGGGTGACAGATCCCTATAGTGAGTCGTATTA-3′ where m indicates 2′ O-methylated nucleotide) primers at 1 mM stock solution with 470 mL of water59. The mixture was heated at 95 °C for 5 min and cooled at room temperature for 30 min before assembling the in vitro transcription reaction. Samples were either unlabeled or residue-specifically labeled with 13C/15N- or 2H (Cambridge Isotope Laboratories, Inc.). After transcription, RNA samples were heat denatured and purified by using urea-denaturing polyacrylamide gels. The same in vitro transcription reaction protocol was done for 7SK-SL1apicalΔASM using a forward (5′-TAATACG ACTCACTATAGGGATCTGTCACCCCAGATCGCCAGTGGCGATCTGGGGAGGCGGGTCCC-3′) and reverse (5′-mGmGGACCCGCCTCCCCAGATCGCCACTGGCGATCT GGGGTGACAGATCCCTATAGTGAGTCGTATTA-3′ where m indicates 2′ O-methylated nucleotide).
HEXIM ARM and Tat ARM peptide preparation
Unlabeled HEXIMN-ARM (GISYGRQLGKKKHRRRAHQ), TatFin ARM (GISYGRKKRKHRRRAHQ), and TatG ARM (GISYGRKKRRHRRRAHQ) peptides were purchased from Tufts University Core Facility at a 0.1 mmol scale. Tat adapters were placed around the HEXIMN-ARM sequence to prevent non-physiological aggregation in solution-state NMR studies. HEXIMN-ARM peptides containing selective 13C/15N-labeled residues, underlined, (GISYGRQLGKKKHRRRAHQ and GISYGRQLGKKKHRRRAHQ) were purchased from New England Peptide.
Full-length HEXIM1 preparation
Synthetic DNA encoding HEXIM1 (2-359) was cloned into a bacterial pMCSG7 expression vector59 encoding an N-terminal tobacco etch virus (TEV) protease-cleavable His6 tag and was expressed in E. coli BL21 AI cells in an overnight culture at 20 °C. Cells were lysed by sonication in buffer containing 50 mM Tris pH 8.0, 500 mM NaCl, 0.1% β-mercaptoethanol, 50 mM (NH4)2SO4 and protease inhibitor aprotinin and leupeptin. His6-HEXIM1 was purified from the cleared cell lysate using Ni-NTA resin (Qiagen) and the His6 tag was cleaved with TEV protease. The HEXIM was run over a second Ni-NTA column, followed by anion exchange on a 5 mL HiTrap Q HP column (Cytiva) and gel filtration on a Superdex 200 16/60 column (Cytiva) in a final buffer containing 25 mM HEPES pH 7.5, 200 mM NaCl, 5% glycerol, 1 mM TCEP. HEXIM was flash frozen in liquid nitrogen and stored at −80 °C.
Isothermal titration calorimetry
Binding constants for the interactions of 7SK-SL1apical-AGU with the HEXIMN-ARM and TatFin and TatG ARMs and full-length HEXIM1 with 7SK-SL1apical-AGU, 7SK-SL1Full-WT, and 7SK-SL1Full-AGU were measured using an ITC-200 microcalorimeter (MicroCal). 68 μM HEXIMN-ARM peptide was titrated into 5 μM solutions of 7SK-SL1apical-AGU or 7SK-SL1apicalΔASM in 10 mM sodium phosphate, 70 mM NaCl, 0.1 mM EDTA, pH 5.2 at 25 °C. Titrations of Tat ARMs into 7SK-SL1apical-AGU or 7SK-SL1apicalΔASM were also performed in the same buffer conditions as the HEXIMN-ARM titration, although the Tat ARM concentration was at 2.5 μM and the 7SK-SL1apical-AGU concentration was at 45 μM. Titrations with full-length HEXIM1 were done at 100 μM of HEXIM1 into 3 μM of either 7SK-SL1apical-AGU, 7SK-SL1Full-WT, and 7SK-SL1Full-AGU in a buffer of 25 mM HEPES pH 7.5, 200 mM NaCl, 5% glycerol, and 1 mM TCEP. Titration curves were analyzed using ORIGIN (OriginLab) and all thermodynamic parameters are reported with n = 3 experiments.
Small-angle X-ray scattering
SAXS data for the 7SK-SL1apical-AGU:HEXIMN-ARM, 7SK-SL1apical-AGU: TatFin ARM, and 7SK-SL1apical-AGU:TatG ARM complexes were obtained at SIBYLS beamline of Advanced Light Source at Lawrence Berkeley National Laboratory. Measurements were performed in a buffer containing 10 mM sodium phosphate, 70 mM NaCl, 0.1 mM EDTA, pH 5.2, and the background scattering was subtracted from the sample scattering to obtain the scattering intensity from the solute molecules. Data from three different concentrations (50, 75, and 100 μM) were compared with scattering intensities at q = 0 Å−1 [I(0)], as determined by Guinier analysis, to detect possible interparticle interactions. Data were analyzed by using ScÅtter software, and the presented DAMAVER envelope structures were reconstructed by using DAMMIF/DAMMIN software from 23 independent DAMMIF runs. Chi-squared values of SAXS profiles were analyzed on FoXS60,61.
NMR data acquisition, resonance assignment, and structural calculations
For NMR experiments, the Tat ARM/HEXIMN-ARM:7SK-SL1apical-AGU complexes were dissolved in a buffer containing 10 mM potassium phosphate, 70 mM NaCl, and 0.1 mM EDTA, pH 5.2, whereas the full-length HEXIM1:7SK-SL1apical complex was in a buffer with 25 mM HEPES pH 7.5, 200 mM NaCl, 5% 2H-glycerol, and 1 mM TCEP. All NMR experiments were acquired by using Bruker 700 or 800 MHz instruments equipped with cryogenic probes. Spectra for observing non-exchangeable protons were collected at 298 K in 99.96% D2O, whereas those for exchangeable protons were at 283 K and 298 K in 10% D2O. For NOESY experiments, mixing times were set to 200 ms. To help unambiguously assign the intermolecular NOEs of the HEXIMN-ARM with 7SK-SL1apical-AGU, we used both specifically protonated GA, AC, and GU samples of 7SK-SL1apical-AGU and two HEXIMN-ARM peptides synthesized by with different combinations of 13C/15N-labeled amino acids. Samples of the 7SK-SL1apical-AGU:HEXIMN-ARM, the 7SK-SL1apical-AGU:TatFin ARM, and 7SK-SL1apical-AGU:TatG ARM were prepared at 1:0.9 equivalents, whereas the full-length HEXIM1:7SK-SL1apical-AGU complex was prepared at 1:0.3 equivalents to avoid any nonspecific binding or aggregation of the protein to the RNA. Assignments for non-exchangeable 1H, 13C, 15N signals of 7SK-SL1apical-AGU in complex with HEXIMN-ARM and Tat ARMs were obtained by analyzing two-dimensional 1H-1H NOESY recorded with non-labeled samples and two-dimensional 13C-HMQC and 15N-HSQC and three-dimensional 13C-edited HMQC-NOESY spectra for labeled samples.
Initial structural models were generated using manually assigned restraints in CYANA, where upper-limit distance restraints of 2.7, 3.3, and 5.0 A were employed for direct NOE cross-peaks of a strong, medium, and weak intensities, respectively62. However, for cross-peaks pairs associated with the intra-residue H8/6 to H2′ and H3′, upper distance limits of 4.2 and 3.2 Å were employed for NOEs of medium and strong intensity, respectively. To prevent the generation of structures with collapsed major grooves, cross-helix P–P distance restraints (with 20% weighting coefficient) were employed for A-form helical segments. Standard torsion angle restraints were used for regions of A-helical geometry, allowing for ±50° deviations from ideality (α = −62°, β = 180°, γ = 48°, δ = 83°,ɛ = −152°,ζ = −73°)63. Standard hydrogen-bonding restraints with approximately linear NH–N and NH–O bond distances of 1.85 ± 0.05 Å and N–N and N–O bond distances of 3.00 ± 0.05 Å, and two lower-limit restraints per base pair (G–C base pairs: G-C4 to C-C6 ≥ 8.3 Å and G-N9 to C-H6 ≥ 10.75 Å; A–U base pairs: A-C4 to U-C6 ≥ 8.3 Å and A-N9 to U-H6 ≥ 10.75 Å) were employed in order to weakly enforce base-pair planarity (20% weighting coefficient).
The CYANA structure with the lowest target function was used as the initial model for structure calculations Xplor-NIH to incorporate electrostatic constraints. First, structures were calculated using annealing from 2000 °C to 25 °C in steps of 12.5 °C. Standard energy potential terms for bonds, angles, torsion angles, van der Waals interactions, and interatomic repulsions were included. The statistical backbone H-bond potential was utilized for protein residues. Energy potentials for NOEs, hydrogen bonds, and planarity were incorporated with restraints derived from NMR data. All restraints used in CYANA were included except for phosphate-phosphate distances. The structures were sorted by energy using bond, angle, dihedral, and NOE energy potential terms, and the ten percent of the structures with the lowest sort energy were further minimized with SAXS terms to incorporate orientation restraints. For this step, minimization started at 1500 °C to 25 °C in steps of 12.5 °C. The lowest ten percent of these were deposited in the RCSB databank.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Atomic coordinates have been deposited in the Protein Data Bank under accession codes PDB 7T1N (7SK-SL1apical-AGU:HEXIMN-ARM), PDB 7T1P (7SK-SL1apical-AGU:TatFin), and PDB 7T1O (7SK-SL1apical-AGU:TatG). Chemical shifts have been deposited in the Biological Magnetic Resonance Data Bank under accession codes 30971 (7SK-SL1apical-AGU:HEXIMN-ARM), 30973 (7SK-SL1apical-AGU:TatFin), and 30972 (7SK-SL1apical-AGU:TatG). SAXS data were submitted to and validated by SASBDB (https://www.sasbdb.org/)64 under accession codes SASDME9 (7SK-SL1apical-AGU:HEXIMN-ARM), SASDMF9 (7SK-SL1apical-AGU:TatFin), and SASDMD9 (7SK-SL1apical-AGU:TatG). Source data are available through Dryad (https://doi.org/10.5061/dryad.12jm63z17) and upon reasonable request from the corresponding author.
References
Wada, T. et al. DSIF, a novel transcription elongation factor that regulates RNA polymerase II processivity, is composed of human Spt4 and Spt5 homologs. Genes Dev. 12, 343–356 (1998).
Yamaguchi, Y. et al. NELF, a multisubunit complex containing RD, cooperates with DSIF to repress RNA polymerase II elongation. Cell 97, 41–51 (1999).
Missra, A. & Gilmour, D. S. Interactions between DSIF (DRB sensitivity inducing factor), NELF (negative elongation factor), and the Drosophila RNA polymerase II transcription elongation complex. Proc. Natl Acad. Sci. USA 107, 11301–11306 (2010).
Renner, D. B., Yamaguchi, Y., Wada, T., Handa, H. & Price, D. H. A highly purified RNA polymerase II elongation control system. J. Biol. Chem. 276, 42601–42609 (2001).
Ehara, H. et al. Structure of the complete elongation complex of RNA polymerase II with basal factors. Science 357, 921–924 (2017).
Marshall, N. F. & Price, D. H. Purification of P-TEFb, a transcription factor required for the transition into productive elongation. J. Biol. Chem. 270, 12335–12338 (1995).
Peng, J., Zhu, Y., Milton, J. T. & Price, D. H. Identification of multiple cyclin subunits of human P-TEFb. Genes Dev. 12, 755–762 (1998).
Kim, J. B. & Sharp, P. A. Positive transcription elongation factor B phosphorylates hSPT5 and RNA polymerase II carboxyl-terminal domain independently of cyclin-dependent kinase-activating kinase. J. Biol. Chem. 276, 12317–12323 (2001).
Fujinaga, K. et al. Dynamics of human immunodeficiency virus transcription: P-TEFb phosphorylates RD and dissociates negative effectors from the transactivation response element. Mol. Cell Biol. 24, 787–795 (2004).
Vos, S. M. et al. Structure of activated transcription complex Pol II-DSIF-PAF-SPT6. Nature 560, 607–612 (2018).
Yamada, T. et al. P-TEFb-mediated phosphorylation of hSpt5 C-terminal repeats is critical for processive transcription elongation. Mol. Cell 21, 227–237 (2006).
Krueger, B. J. et al. LARP7 is a stable component of the 7SK snRNP while P-TEFb, HEXIM1 and hnRNP A1 are reversibly associated. Nucleic Acids Res. 36, 2219–2229 (2008).
Yang, Z., Zhu, Q., Luo, K. & Zhou, Q. The 7SK small nuclear RNA inhibits the CDK9/cyclin T1 kinase to control transcription. Nature 414, 317–322 (2001).
Nguyen, V. T., Kiss, T., Michels, A. A. & Bensaude, O. 7SK small nuclear RNA binds to and inhibits the activity of CDK9/cyclin T complexes. Nature 414, 322–325 (2001).
Michels, A. A. et al. MAQ1 and 7SK RNA interact with CDK9/cyclin T complexes in a transcription-dependent manner. Mol. Cell Biol. 23, 4859–4869 (2003).
Yik, J. H. et al. Inhibition of P-TEFb (CDK9/Cyclin T) kinase and RNA polymerase II transcription by the coordinated actions of HEXIM1 and 7SK snRNA. Mol. Cell 12, 971–982 (2003).
Chen, R., Yang, Z. & Zhou, Q. Phosphorylated positive transcription elongation factor b (P-TEFb) is tagged for inhibition through association with 7SK snRNA. J. Biol. Chem. 279, 4153–4160 (2004).
Michels, A. A. et al. Binding of the 7SK snRNA turns the HEXIM1 protein into a P-TEFb (CDK9/cyclin T) inhibitor. EMBO J. 23, 2608–2619 (2004).
Kobbi, L. et al. An evolutionary conserved Hexim1 peptide binds to the Cdk9 catalytic site to inhibit P-TEFb. Proc. Natl Acad. Sci. USA 113, 12721–12726 (2016).
Fujinaga, K., Luo, Z. & Peterlin, B. M. Genetic analysis of the structure and function of 7SK small nuclear ribonucleoprotein (snRNP) in cells. J. Biol. Chem. 289, 21181–21190 (2014).
Prasanth, K. V. et al. Nuclear organization and dynamics of 7SK RNA in regulating gene expression. Mol. Biol. Cell 21, 4184–4196 (2010).
Kohoutek, J., Blazek, D. & Peterlin, B. M. Hexim1 sequesters positive transcription elongation factor b from the class II transactivator on MHC class II promoters. Proc. Natl Acad. Sci. USA 103, 17349–17354 (2006).
Barboric, M. et al. Interplay between 7SK snRNA and oppositely charged regions in HEXIM1 direct the inhibition of P-TEFb. EMBO J. 24, 4291–4303 (2005).
Sano, M. et al. Activation and function of cyclin T-Cdk9 (positive transcription elongation factor-b) in cardiac muscle-cell hypertrophy. Nat. Med. 8, 1310–1317 (2002).
Kretz, A. L. et al. CDK9 is a prognostic marker and therapeutic target in pancreatic cancer. Tumour Biol. 39, 1010428317694304 (2017).
Schlafstein, A. J. et al. CDK9 expression shows role as a potential prognostic biomarker in breast cancer patients who fail to achieve pathologic complete response after neoadjuvant chemotherapy. Int. J. Breast Cancer 2018, 6945129 (2018).
Shao, H. et al. HEXIM1 controls P-TEFb processing and regulates drug sensitivity in triple-negative breast cancer. Mol. Biol. Cell 31, 1867–1878 (2020).
Cho, W. K., Jang, M. K., Huang, K., Pise-Masison, C. A. & Brady, J. N. Human T-lymphotropic virus type 1 Tax protein complexes with P-TEFb and competes for Brd4 and 7SK snRNP/HEXIM1 binding. J. Virol. 84, 12801–12809 (2010).
Zhou, M. et al. Tax interacts with P-TEFb in a novel manner to stimulate human T-lymphotropic virus type 1 transcription. J. Virol. 80, 4781–4791 (2006).
Muniz, L., Egloff, S., Ughy, B., Jady, B. E. & Kiss, T. Controlling cellular P-TEFb activity by the HIV-1 transcriptional transactivator Tat. PLoS Pathog. 6, e1001152 (2010).
Sobhian, B. et al. HIV-1 Tat assembles a multifunctional transcription elongation complex and stably associates with the 7SK snRNP. Mol. Cell 38, 439–451 (2010).
Krueger, B. J., Varzavand, K., Cooper, J. J. & Price, D. H. The mechanism of release of P-TEFb and HEXIM1 from the 7SK snRNP by viral and cellular activators includes a conformational change in 7SK. PLoS ONE 5, e12335 (2010).
Barboric, M. et al. Tat competes with HEXIM1 to increase the active pool of P-TEFb for HIV-1 transcription. Nucleic Acids Res. 35, 2003–2012 (2007).
Yik, J. H., Chen, R., Pezda, A. C., Samford, C. S. & Zhou, Q. A human immunodeficiency virus type 1 Tat-like arginine-rich RNA-binding domain is essential for HEXIM1 to inhibit RNA polymerase II transcription through 7SK snRNA-mediated inactivation of P-TEFb. Mol. Cell Biol. 24, 5094–5105 (2004).
Byers, S. A., Price, J. P., Cooper, J. J., Li, Q. & Price, D. H. HEXIM2, a HEXIM1-related protein, regulates positive transcription elongation factor b through association with 7SK. J. Biol. Chem. 280, 16360–16367 (2005).
Yik, J. H., Chen, R., Pezda, A. C. & Zhou, Q. Compensatory contributions of HEXIM1 and HEXIM2 in maintaining the balance of active and inactive positive transcription elongation factor b complexes for control of transcription. J. Biol. Chem. 280, 16368–16376 (2005).
Egloff, S., Van Herreweghe, E. & Kiss, T. Regulation of polymerase II transcription by 7SK snRNA: two distinct RNA elements direct P-TEFb and HEXIM1 binding. Mol. Cell Biol. 26, 630–642 (2006).
Dulac, C. et al. Transcription-dependent association of multiple positive transcription elongation factor units to a HEXIM multimer. J. Biol. Chem. 280, 30619–30629 (2005).
Li, Q. et al. Analysis of the large inactive P-TEFb complex indicates that it contains one 7SK molecule, a dimer of HEXIM1 or HEXIM2, and two P-TEFb molecules containing Cdk9 phosphorylated at threonine 186. J. Biol. Chem. 280, 28819–28826 (2005).
Schonichen, A. et al. A flexible bipartite coiled coil structure is required for the interaction of Hexim1 with the P-TEFB subunit cyclin T1. Biochemistry 49, 3083–3091 (2010).
Blazek, D., Barboric, M., Kohoutek, J., Oven, I. & Peterlin, B. M. Oligomerization of HEXIM1 via 7SK snRNA and coiled-coil region directs the inhibition of P-TEFb. Nucleic Acids Res. 33, 7000–7010 (2005).
Martinez-Zapien, D. et al. Intermolecular recognition of the non-coding RNA 7SK and HEXIM protein in perspective. Biochimie 117, 63–71 (2015).
Lebars, I. et al. HEXIM1 targets a repeated GAUC motif in the riboregulator of transcription 7SK and promotes base pair rearrangements. Nucleic Acids Res. 38, 7749–7763 (2010).
Czudnochowski, N., Vollmuth, F., Baumann, S., Vogel-Bachmayr, K. & Geyer, M. Specificity of Hexim1 and Hexim2 complex formation with cyclin T1/T2, importin alpha and 7SK snRNA. J. Mol. Biol. 395, 28–41 (2010).
Pham, V. V. et al. HIV-1 Tat interactions with cellular 7SK and viral TAR RNAs identifies dual structural mimicry. Nat. Commun. 9, 4266 (2018).
Chen, L. & Frankel, A. D. An RNA-binding peptide from bovine immunodeficiency virus Tat protein recognizes an unusual RNA structure. Biochemistry 33, 2708–2715 (1994).
Puglisi, J. D., Tan, R., Calnan, B. J., Frankel, A. D. & Williamson, J. R. Conformation of the TAR RNA-arginine complex by NMR spectroscopy. Science 257, 76–80 (1992).
Puglisi, J. D., Chen, L., Blanchard, S. & Frankel, A. D. Solution structure of a bovine immunodeficiency virus Tat-TAR peptide-RNA complex. Science 270, 1200–1203 (1995).
Brown, J. A. Unraveling the structure and biological functions of RNA triple helices. Wiley Interdiscip. Rev. RNA 11, e1598 (2020).
Chavali, S. S., Cavender, C. E., Mathews, D. H. & Wedekind, J. E. Arginine forks are a widespread motif to recognize phosphate backbones and guanine nucleobases in the RNA major groove. J. Am. Chem. Soc. 142, 19835–19839 (2020).
Chavali, S. S. et al. Cyclic peptides with a distinct arginine-fork motif recognize the HIV trans-activation response RNA in vitro and in cells. J. Biol. Chem. 297, 101390 (2021).
Brillet, K. et al. Different views of the dynamic landscape covered by the 5’-hairpin of the 7SK small nuclear RNA. RNA 26, 1184–1197 (2020).
Brogie, J. E. & Price, D. H. Reconstitution of a functional 7SK snRNP. Nucleic Acids Res. 45, 6864–6880 (2017).
Martinez-Zapien, D. et al. The crystal structure of the 5 functional domain of the transcription riboregulator 7SK. Nucleic Acids Res. 45, 3568–3579 (2017).
Bourbigot, S. et al. Solution structure of the 5’-terminal hairpin of the 7SK small nuclear RNA. RNA 22, 1844–1858 (2016).
Roder, K., Stirnemann, G., Dock-Bregeon, A. C., Wales, D. J. & Pasquali, S. Structural transitions in the RNA 7SK 5’ hairpin and their effect on HEXIM binding. Nucleic Acids Res. 48, 373–389 (2020).
Roder, K. & Pasquali, S. RNA modeling with the computational energy landscape framework. Methods Mol. Biol. 2323, 49–66 (2021).
Marz, M. et al. Evolution of 7SK RNA and its protein partners in metazoa. Mol. Biol. Evol. 26, 2821–2830 (2009).
Stols, L. et al. A new vector for high-throughput, ligation-independent cloning encoding a tobacco etch virus protease cleavage site. Protein Expr. Purif. 25, 8–15 (2002).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. Accurate SAXS profile computation and its assessment by contrast variation experiments. Biophys. J. 105, 962–974 (2013).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. FoXS, FoXSDock and MultiFoXS: single-state and multi-state structural modeling of proteins and their complexes based on SAXS profiles. Nucleic Acids Res. 44, W424–W429 (2016).
Guntert, P., Mumenthaler, C. & Wuthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Tolbert, B. S. et al. Major groove width variations in RNA structures determined by NMR and impact of 13C residual chemical shift anisotropy and 1H-13C residual dipolar coupling on refinement. J. Biomol. NMR 47, 205–219 (2010).
Kikhney, A. G., Borges, C. R., Molodenskiy, D. S., Jeffries, C. M. & Svergun, D. I. SASBDB: towards an automatically curated and validated repository for biological scattering data. Protein Sci. 29, 66–75 (2020).
Acknowledgements
This work was supported by HHMI grant 55108516 (to V.M.D'S.) and NIH grant U54 AI50470 (formerly U54 GM103297) (to V.M.D'S. and J.L.S.). SAXS data were collected at the Advanced Light Source (ALS), SIBYLS beamline on behalf of US DOE-BER through the Integrated Diffraction Analysis Technologies (IDAT) program. Additional support comes from the NIGMS project ALS-ENABLE (P30 GM124169) and a High-End Instrumentation Grant S10OD018483.
Author information
Authors and Affiliations
Contributions
V.V.P, M.G., and V.M.D'S. conceived and designed the experiments. V.V.P., M.G., and J.L.M. purified the samples and V.V.P performed the NMR, ITC, and SAXS experiments. V.V.P., M.G., and V.M.D’S. performed the structural analyses, interpreted the data, and wrote the manuscript. V.V.P, M.G., J.L.M., J.L.S., and V.M.D'S. analyzed data and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Shintaro Aibara and the other, anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Anam Akhtar and Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pham, V.V., Gao, M., Meagher, J.L. et al. A structure-based mechanism for displacement of the HEXIM adapter from 7SK small nuclear RNA. Commun Biol 5, 819 (2022). https://doi.org/10.1038/s42003-022-03734-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-03734-w
This article is cited by
-
The cell biology of HIV-1 latency and rebound
Retrovirology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.