Abstract
Putative tumor suppressor ALDH1L1, the product of natural fusion of three unrelated genes, regulates folate metabolism by catalyzing NADP+-dependent conversion of 10-formyltetrahydrofolate to tetrahydrofolate and CO2. Cryo-EM structures of tetrameric rat ALDH1L1 revealed the architecture and functional domain interactions of this complex enzyme. Highly mobile N-terminal domains, which remove formyl from 10-formyltetrahydrofolate, undergo multiple transient inter-domain interactions. The C-terminal aldehyde dehydrogenase domains, which convert formyl to CO2, form unusually large interfaces with the intermediate domains, homologs of acyl/peptidyl carrier proteins (A/PCPs), which transfer the formyl group between the catalytic domains. The 4′-phosphopantetheine arm of the intermediate domain is fully extended and reaches deep into the catalytic pocket of the C-terminal domain. Remarkably, the tetrameric state of ALDH1L1 is indispensable for catalysis because the intermediate domain transfers formyl between the catalytic domains of different protomers. These findings emphasize the versatility of A/PCPs in complex, highly dynamic enzymatic systems.
Similar content being viewed by others
Introduction
ALDH1L1 (10-formyltetrahydrofolate dehydrogenase), an enzyme of folate metabolism, regulates the availability of one-carbon groups for folate-dependent biochemical reactions1. The importance of this regulation is emphasized by high abundance of the enzyme in the liver, the main organ of folate metabolism, as well as by tight control of the ALDH1L1 expression during embryonic development and by the role of the protein as a pan-astrocyte marker1,2,3. The regulatory role of ALDH1L1 is linked to its catalytic reaction, the NADP+-dependent conversion of 10-formyltetrahydrofolate (10-fTHF) to tetrahydrofolate (THF) and CO2, which irreversibly removes one-carbon groups from the folate pool, thus diminishing the anabolic capacity1,4 (Fig. 1a). It has been proposed that this reaction interferes with rapid cellular proliferation, but at the same time supports homeostasis in non-proliferating cells by supplying THF for the reaction of conversion of serine to glycine and for the formate metabolism1,4,5. The role of the enzyme in supporting glycine production has been recently demonstrated in the Aldh1l1 knockout mouse model, with livers of ALDH1L1-deficient mice having decreased levels of THF, glycine and glycine conjugates5. Lately, the enzyme’s function has been linked to NADPH production and oxidative stress6. ALDH1L1 is also considered a putative tumor suppressor4. This role is supported by findings that the protein is strongly and ubiquitously downregulated in malignant tumors and cancer cell lines7,8, the effect associated with hypermethylation of the ALDH1L1 promoter9,10,11,12,13. Of note, expression of ALDH1L1 in cancer cell lines produces strong antiproliferative effects by activating specific apoptotic pathways7,14,15,16,17,18. In further support of the suppressive effect of ALDH1L1 on proliferation, the enzyme is strongly downregulated in S-phase of the cell cycle through proteasomal degradation but is elevated in quiescent cells19. Although the knockout of Aldh1l1 in mice did not cause the initiation of malignant lesions, it promoted the growth of larger liver tumors initiated by a chemical carcinogen20.
a ALDH1L1 converts 10-formyltetrahydrofolate (10-formylTHF) to tetrahydrofolate (THF) and CO2 in two sequential enzymatic reactions, the cleavage of the formyl group in the folate-binding N-terminal domain and the NADP+-dependent oxidation of the formyl to carbon dioxide in the aldehyde dehydrogenase C-terminal domain. The intermediate domain, which is homologous of acyl/peptidyl carrier proteins, transports the formyl group, attached to the 4′-phosphopantetheine prosthetic arm (4′-PP), from the N-terminal to the C-terminal domain. b Primary structure of ALDH1L1 with indicated domain boundaries. c Schematic of spatial organization of the ALDH1L1 tetramer based on available atomic structures of its individual domains. PDB structures 1s3i, 2cq8, and 2o2p were used for the N-terminal, intermediate, and C-terminal domains, respectively. For one protomer the domains are labeled and colored as in b. Arrows mark the positions of the hydrolase and aldehyde dehydrogenase active sites (AS) and the NADP+ binding site.
ALDH1L1 originated from a natural fusion of three unrelated genes, the phenomenon defining the structural organization of the ALDH1L1 enzyme1. The protein exists as a homotetramer, with each 902 amino acid-long protomer organized in three distinct functional domains (Fig. 1b, c). The N-terminal domain (Nt, aa 1-310) carries the folate-binding site and has sequence and structural similarity to methionyl-tRNAMet-formyltransferase (FMT), the enzyme involved in translation initiation in mitochondria21. FMT formylates the initiator Met-tRNAMet by transferring the formyl group from 10-fTHF, thus using the same substrate as ALDH1L122,23,24. The C-terminal domain (Ct, aa 405–902) belongs to the family of aldehyde dehydrogenases (ALDHs), the group of enzymes catalyzing the conversion of a large variety of aldehydes to corresponding acids using NAD+ or NADP+ as the electron acceptor25. The Ct domain shares up to 50% sequence similarity with members of this family and has a typical ALDH fold, which includes NAD(P)+-binding, catalytic, and oligomerization sub-domains26,27. The Ct domain contains all critical catalytic residues conserved in ALDHs, including Cys707, which plays the role of the catalytic center nucleophile27,28. Accordingly, the Ct domain catalyzes the conversion of short-chain aldehydes to corresponding acids in vitro, but it is not known whether ALDH1L1 participates in aldehyde oxidation in vivo1. Finally, the intermediate domain (Int, aa 314–397) linking the Nt and Ct domains is a homolog of a group of small, structurally closely related carrier proteins involved in fatty acid, polyketide, and non-ribosomal peptide biosynthesis29. A characteristic feature of these acyl/peptidyl carrier proteins (A/PCPs) is the 4′-phosphopantetheine prosthetic group (4′-PP) covalently attached to a serine residue through a phosphoester bond30,31. This prosthetic group serves as a flexible arm enabling the transfer of building blocks between subunits of multi-enzyme complexes30,31.
Functional studies of ALDH1L1, its numerous mutants and engineered constructs together with structural and functional characterization of the individual domains provided insight into the enzyme catalytic machinery21,26,27,28,29,32,33,34,35,36,37,38,39,40,41. Overall, in the ALDH1L1 catalysis, the 4′-PP arm of the Int domain transfers the formyl group cleaved from 10-fTHF in the folate-binding Nt domain to the Ct domain, where it is oxidized to carbon dioxide1 (Fig. 1a). To execute this mechanism, in addition to the flexibility of the 4′-PP moving arm, sufficient mobility of ALDH1L1 domains relative to each other is necessary. This complicates the structural analysis of the full-length ALDH1L1 protein. Indeed, while several crystal structures of Nt and Ct domains were reported21,27,38,39,40,41, and an NMR structure of a synthetic Int domain devoid of the 4′-PP arm is available (PDB 2cq8), the structure of the full-length enzyme has not been resolved so far. Here, we report the structures of ligand-free and NADP+-bound full-length ALDH1L1 at resolutions of 3.7 Å and 2.9 Å, respectively, obtained by cryo-electron microscopy (cryo-EM). This study provides insights into the ALDH1L1 structure and function by (i) demonstrating high mobility of the Nt domains, which form transient complexes with other structural units; (ii) describing the unusual mode of interaction between the Int and Ct domains, with a large contact interface atypical for A/PCPs, and (iii) revealing unique pairing of the Int and Ct domains, which requires the tetrameric organization for catalysis.
Results
Overall architecture of full-length ALDH1L1
To reveal the domain organization of ALDH1L1 we performed negative-stain EM (NS-EM) of the full-length ligand-free protein (Rattus norvegicus ALDH1L1 produced in insect cells using a baculovirus expression system) as well as of its individually expressed Ct domain, which forms the rigid tetrameric core of the full-length enzyme. The Ct core was clearly visible in the 2D class averages (Fig. 2a, b), enabling the identification of all four Int domains. All Int domains in full-length ALDH1L1 sat closely to the C-terminal core and were arranged in an apparently symmetrical manner. The high contrast provided by negative staining also allowed, in many cases, to resolve the Nt domains both in raw micrographs (Supplementary Fig. 1) and 2D class averages (Fig. 2b). The Nt domains assumed variable positions with respect to the rest of the protein, indicating their high mobility. Focused 2D classification revealed that the Nt domain sampled the entire range of positions between the one oriented away from the protein central core and the one in contact with the Ct domain (Fig. 2c). We did not observe any obvious coordination in positioning of the four Nt domains in the ALDH1L1 tetramer in NS-EM experiments, a finding suggesting asynchronous movement of these domains in the full-length protein. However, we noticed that a conformation in which the Nt domain is tightly packed against the Ct core was encountered repeatedly (Fig. 2b). The high mobility of the Nt domain illustrated by the NS-EM data is likely required for the multi-step catalytic mechanism of the enzyme.
a Comparison of negative-stain EM 2D class averages of the individually expressed tetrameric C-terminal domain (Ct, left panel) and the full-length protein (right panel) was used for identification of intermediate domains (Int) adjacent to the Ct domain within each protomer. Only a single N-terminal domain (Nt) is clearly visible. b Selected negative-stain 2D classes illustrate the range of positions assumed by the Nt domains within the ALDH1L1 tetramer. Arrows point to the position of the Nt domain in which it closely interacts with the Ct domain. c The range of motion of the Nt domain illustrated by a selection of negative-stain 2D classes. Below: cartoon representations of the crystal structure of Nt in the corresponding orientations. d, e Cryo-EM structures of ligand-free ALDH1L1 (d) and ALDH1L1 in complex with NADP+ (e). The maps are displayed as transparent surfaces. Atomic models are shown in cartoon representation, with Int and Ct domains colored light-blue and pink, respectively. Bound NADP+ molecules are shown in sphere representation. f Cryo-EM structure of the rigid core of ALDH1L1 corresponds well to the negative-stain EM 2D classes. Results of segmentation of the ligand-free map are shown. Scale bars in panels a–c correspond to 10 nm.
Subsequently, we used cryo-EM to characterize the structure of ligand-free ALDH1L1 at high resolution (Fig. 2d). A 3.7-Å map was obtained from a dataset of 86,276 particles when D2 symmetry was imposed. The rigid Ct core and four Int domains were clearly resolved in this structure, while the highly mobile Nt domains were not visible in the symmetrical map. To elucidate the potential effect of NADP+ on the structural organization of ALDH1L1, we also prepared cryo-EM grids after adding 1 mM NADP+ to the full-length protein. Single-particle analysis of the new dataset containing 202,398 particles produced a 2.9-Å resolution map (Fig. 2e). The arrangement of the Ct core and Int domains in this structure was identical to that in the ligand-free protein and corresponded well to the configuration of the Ct and Int modules revealed by NS-EM (Fig. 2f).
Structures of the Int and C t domains
Both the Int and Ct domains were clearly resolved in the cryo-EM maps (Fig. 3), which permitted the building of atomic models for ligand-free and NADP+-bound ALDH1L1. The entire 4′-PP prosthetic group was also well defined in the cryo-EM density (Fig. 3a). As expected based on a previous study29, the Int domain exhibits a fold typical of A/PCPs (Fig. 3a, b)42. Accordingly, its structure consists of three major α-helices (I, II, and IV) forming a loose bundle, with another short helix (III) bridging helices II and IV. Helices I and II are connected by a long 19-residue linker that forms a loop and a helical turn. Serine 354, the site of the 4′-PP group attachment, is located in the beginning of helix II.
a Angular views of the intermediate domain. The molecular model is shown as sticks, and the cryo-EM density is represented by a semi-transparent surface. 4PP: 4′-phosphopantetheine. b Superposition of the intermediate domain of ALDH1L1 (light blue) and the carrier domain of holo-non-ribosomal peptide synthetase (PDB 4zxh84, gray). The alpha-helices are labeled I to IV. The r.m.s.d. between the two structures is 1.98 Å. c Structure of the protomer of the C-terminal domain. The three sub-domains are colored individually and labeled. Bound NADP+ is shown in sphere representation. d Superposition of the Ct domains of ligand free (gray) and NADP+-bound (light blue) ALDH1L1. The binding of NADP+ induces local structural changes in one helix forming the binding cleft for the adenine moiety (dashed circle). Cryo-EM density for NADP+ is shown as semi-transparent surface.
Each Ct domain contains the catalytic, NADP+-binding and oligomerization sub-domains (Fig. 3c). The deep substrate entrance tunnel is located between the catalytic and NADP+-binding sub-domains and leads to the catalytic cysteine 707. The NADP+ binding site is situated on the side opposite to the substrate entrance tunnel. In the cryo-EM maps of ALDH1L1, the structure of the Ct domain was very similar to the crystal structures of individual Ct domains previously reported27. The root mean square deviation (r.m.s.d.) between the protomer of the Ct domain solved by cryo-EM and the protomer of the corresponding X-ray structure (PDB 2o2p or 2o2q27) was 0.46 Å and 0.75 Å, respectively, for ligand-free and NADP+-bound proteins. The r.m.s.d. between the entire tetramers was slightly higher (0.621 Å and 1.08 Å, respectively), suggesting slight differences in protomer positions in full-length ALDH1L1, which may be a consequence of the interactions between the Ct and Int domains. It is also possible that the conformation of the protein in the crystal structures was affected by crystal packing.
We found that the cryo-EM structures of ligand-free and NADP+-bound ALDH1L1 were highly similar (r.m.s.d. = 0.86 Å) (Fig. 3d). A small difference was observed in the NADP+-binding site, where the C-terminal end of the helix formed by residues 652–666 was better ordered in the presence of the coenzyme. Only the AMP portion of bound NADP+ was clearly visible in the cryo-EM map, while there was no density for the rest of the cofactor. Of note, weak density for the nicotinamide riboside of NAD+ and NADP+ is commonly observed in aldehyde dehydrogenases27,43,44.
Unique pairing of Int and C t domains
In both ligand-free and NADP+-bound ALDH1L1, each Int domain formed contacts primarily with one of the four Ct domains (Fig. 4a). The Int and Ct domains of the same protomer were separated by a 20-Å-long extended linker consisting of residues 397–404 (Fig. 4b) and did not interact with each other. Instead, the Int domain of chain A was docked at the substrate entrance tunnel of the Ct domain of chain C, whereas the Int domain of chain C was paired to the Ct domain of chain A (Fig. 4c, d). An identical arrangement was found for protomers B and D. Of note, the tetrameric Ct core of ALDH1L1 is composed of two homodimers formed by protomers A/B (dimer 1) and C/D (dimer 2). Therefore, each Int domain of ALDH1L1 is paired to a Ct domain of the opposite dimer (Fig. 4d). Out of the eight residues composing the linker in rat ALDH1L1, five are negatively charged amino acids, which makes the linker highly hydrophilic. Sequence alignments showed that this property of the linker is preserved across species as well as between ALDH1L1 and ALDH1L2 proteins (Fig. 4b), suggesting that hydrophilicity and negative charge of the linker are important for the function of the enzyme. Of note, although there was continuous cryo-EM density for the linker in the unsharpened maps (Fig. 4a), sharpening weakened this density to the extent where reliable placement of the main chain was not possible, indicating that the linker retains some flexibility.
a Cryo-EM map of NADP+-bound ALDH1L1 with the four protomers colored individually. Int of protomer A interacts with Ct of protomer C. Within each protomer, Int and Ct are connected with a long, extended linker. b Sequence alignments of the linker region for ALDH1L1 and ALDH1L2 family proteins. Invariant residues are colored green. c Pairing of Int and Ct domains illustrated using pipes-and-planks representation of the atomic model. From left to right: the entire tetrameric structure (protomers A–D), two paired protomers (A and C), and a single protomer (A). d A schematic depicting pairing of the Ct and Int domains of ALDH1L1. Each Int domain interacts with the Ct domain of a protomer from a different dimer.
Interactions between Int and C t domains
We found that the Int and Ct domains form a relatively large contact interface (Fig. 5). The base of the Int domain (including the end of helix I, the beginning of the loop connecting helices I and II, and a large part of helix II) fits into the orifice of the substrate entrance tunnel of its partner Ct and is simultaneously flanked by the tip of the oligomerization sub-domain of a second Ct domain, which contacts the end of the loop between helices I and II and, to a lesser extent, helix III (Fig. 5a, b). This secondary interaction also occurs with a Ct domain of the opposite dimer (e.g., Int of protomer A is docked into Ct of protomer C and interacts with the oligomerization sub-domain of Ct from protomer D). The total contact area between the Int domain (excluding the 4′-PP prosthetic group) and the two Ct protomers is 653 Å2. Calculation of electrostatic potentials revealed that the surface of the Int domain is mostly negatively charged, including the contact interface (Fig. 5c). This is in agreement with the known acidic nature of many A/PCPs42. In contrast, the corresponding contact area of the Ct domain that accommodates 4′-PP is charged predominantly positively. The Int and Ct domains form multiple interactions, with the closest contacts between the main chain carbonyl of Gly351 (loop connecting helices I and II) and the amide of Gln693 (2.8 Å), the guanidinium group of Arg359 (helix II) and the side-chain oxygen of Asn745 (3.2 Å), as well as a side-chain oxygen of Glu366 (helix II) and the guanidinium group of Arg742 (3.3 Å). Alignment of the cryo-EM structures of ALDH1L1 and available X-ray structures of the Ct domain revealed no significant alterations in positions of amino acid side chains in the regions that contact the Int domain. Similarly, structural superposition of the Int domain with the NMR structure of individual human Int without the 4′-PP prosthetic group (PDB 2cq8) revealed no major differences (r.m.s.d. = 0.83 Å). Therefore, complex formation between the Int and Ct domains relies on the shape and charge complementarity.
a The intermediate domain (light blue) depicted in pipes-and-planks representation contacts Ct domains of two protomers (pink and cyan). Only the oligomerization sub-domain is shown for the cyan protomer. The 4′-phosphopantetheine (4PP) arm is shown in stick representation. The alpha-helices of the Int domain are labeled I to IV. b Contacting surfaces of the Int domain and the two Ct domains. Coloring is according to panel a. Left, surface representations of the complexes. Right, the complexes are split, and the domains are shown with the interfaces towards the viewer. Contact surfaces are colored according to the color of the interacting partner. c Surfaces of the Int and Ct domains colored according to the electrostatic potentials, with red and blue corresponding to negative and positive charge, respectively. Left: the domains are shown in the same orientations as in the complex. The arrow points to the substrate entrance tunnel. Right: The same domains are rotated such that the contact interfaces face the viewer. d A comparison of negative-stain 2D classes of individual Ct domain and the Int-Ct protein expressed in bacteria. White arrows point to Int domains adjacent to the Ct core. Red arrows point to vacant sites for binding of Int domain. Scale bars represent 10 nm. e A comparison of the complex between the Int and Ct domains of ALDH1L1 with various complexes formed by acyl and peptidyl carrier proteins. The carrier proteins are aligned to the Int domain. The loop connecting helices I and II is colored orange. The PDB code is listed below each complex. 1f80: complex between Bacillus subtilis ACP and ACP synthase; 2cg585: human fatty acid synthase ACP in complex with phosphopantetheinyl transferase; 3ejb86: E. coli ACP in complex with cytochrome P450; 3ny787: E. coli ACP in complex with STAS domain of YchM anion transporter; 4zxh84: Acinetobacter baumannii PCP in complex with nonribosomal peptide synthetase; 5es888: PCP in complex with adenylation domain of nonribosomal peptide synthase from Brevibacillus parabrevis.
Interactions of the 4′-phosphopantetheine prosthetic group
The 4′-PP group covalently linked to serine 354 of the Int domain was found in the fully extended conformation, penetrating deep into the substrate entrance tunnel of the Ct domain and making multiple contacts with residues forming the tunnel (Fig. 6a). Two lysine residues of the Ct domain, Lys520 and Lys865, formed ion pairs with the phosphate of 4′-PP. The main chain carbonyl oxygen of Asn864 and the hydroxyl of Thr521 were within the hydrogen-bonding distance from the hydroxyl and carbonyl oxygens, respectively, of the pantothenic acid moiety. The amide group of asparagine 706 was positioned 3.5 Å from the carbonyl of the β-alanine moiety of 4′-PP.
a Interactions between 4PP (carbon atoms are colored light blue) and residues of the substrate entrance tunnel of the C-terminal domain (carbon atoms are colored pink). Inter-atomic distances shorter than 4 Å are labeled. b Cryo-EM density indicates the presence of a covalent bond between Cys707 and 4PP in the structure of apo-ALDH1L1. c Schematic representation of 4PP cross-linked peptides resulting from tryptic digestion of apo-ALDH1L1. d MS2 spectrum of the MS ion at m/z 1094.5 [M + 2H]2+. The fragmentation pattern reveals the presence of y- and b-ions characteristic of the amino acid sequence of the cross-linked peptides shown in panel c (breakdown along the peptide backbone). Importantly, the main ions observed in the spectrum correspond to the fragmentation along with the 4PP phosphate group, unveiling the chemical nature of the crosslink.
Curiously, we found that the sulfur atom of the catalytic nucleophile Cys707 in the ALDH active center was positioned closely to the sulfur atom of the 4′-PP, and the cryo-EM density between the two atoms appeared continuous (Fig. 6b). This suggested that a disulfide bond formed between the two atoms. To verify the presence of such a bond, we conducted trypsin digestion of ALDH1L1 followed by liquid chromatography-mass spectrometry (LC/MS). Analysis of the LC/MS data revealed readily detectable ions at m/z = 1094.5 ([M + 2H]2+) and m/z = 730.0 ([M + 3H]3+) that corresponded to a tryptic digestion product of nominal mass 2187 Da, identical to the theoretical mass of 4′-PP-crosslinked peptides 350S-R359 and 704G-R712 (Fig. 6c). The subsequent collision-induced dissociations of these ions resulted in a pattern of MS peaks, the interpretation of which allowed unequivocal identification of the chemical structure of the parent ions (Fig. 6d). The most intense peaks in the MS2 spectra resulted from the break of the labile phosphate moiety followed by the neutral loss of the phosphate group, either as phosphoric (H2PO4) or meta-phosphoric (HPO3) acid (Δ mass 98 and 80 Da, respectively)45. The presence of a series of b- and y-ions unambiguously confirmed the amino acid sequences and sites of the 4′-PP covalent modification in the crosslinked peptides. While it was not possible to quantify the prevalence of the crosslinked peptide, its presence confirmed that the disulfide bond formed between the 4′-PP prosthetic group and Cys707 in a population of ALDH1L1 molecules. The inclusion of the disulfide bond in the molecular models improved the fit of the two sulfur atoms to the cryo-EM density.
To investigate the contribution of the 4′-PP arm to the interactions between the Int and Ct domains, we designed a shorter version of ALDH1L1, containing only the Int and Ct domains (termed int-Ct), and expressed it in E. coli. It was shown previously that ALDH1L1 produced in bacteria lacks the 4′-PP prosthetic group29, and therefore only protein–protein contacts could contribute to the interactions between the Int and Ct domains in int-Ct. NS-EM of int-Ct resolved the Ct core and, in some cases, Int domains adjacent to it (Fig. 5d). However, the sites of the Ct core that had been invariably occupied by the Int domains in full-length ALDH1L1 (Fig. 2a) were predominantly vacant in the int-Ct protein lacking 4′-PP. In agreement with such an arrangement, only the tetrameric Ct core was resolved in the 2.7-Å crystal structure of int-Ct expressed in bacteria, with no electron density present for the Int domains. This indicates that the interactions of the 4′-PP arm with residues of the substrate entrance tunnel are critical for the formation of a stable complex between the Int and Ct domains of ALDH1L1.
Int domains are highly mobile when not docked into C t domains
During the initial step of the ALDH1L1 catalysis, 4′-PP arms must be accessible to interact with the Nt domains. With the arm placed outside of the Ct domain active site, the Int domain is expected to disengage the Ct domain, leaving the substrate entrance tunnel vacant. However, global 3D classification did not produce classes with vacant substrate entrance tunnels of the Ct domains. To determine the fraction of free (not occupying substrate entrance tunnels) Int domains, we performed local 3D classification within a mask encompassing each Int domain and calculated the total number of particles that contributed to empty versus occupied classes (Supplementary Fig. 2). This analysis produced estimated average occupancies of 76% and 83% for the Int domain in the ligand-free and NADP+-bound structures, respectively. This indicates that a fraction of Int domains is present in the free form and is available to shuttle the substrate between the Nt and Ct catalytic centers. However, since neither NS-EM nor cryo-EM experiments resolved Int in any other state than docked at the Ct substrate entrance tunnel, the Int domains do not seem to assume strictly defined positions while shuttling between the two catalytic domains in the ligand-free and NADP+-bound enzyme. These results also indicate that the state with the Int domains docked at the Ct substrate entrance tunnels and with the 4′-PP arms reaching into the ALDH active sites is the most favorable conformation for the resting (substrate-free) enzyme. The partial disulfide bond between the 4′-PP arm and Cys707 could serve to support this conformation.
Transient interactions of the N-terminal domain
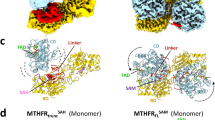
In an attempt to resolve the Nt domain of ALDH1L1, we performed refinement of the cryo-EM dataset of ligand-free ALDH1L1 without enforcing symmetry. The resulting 4.4-Å-resolution map was very similar to its symmetrical counterpart, except that weak density for a single Nt domain became visible (Fig. 7a). This Nt domain appeared to interact with the NADP+-binding sub-domain of one of the Ct domains. Low r.m.s.d. values between the chains of this structure (0.67–0.74 Å) indicated that this interaction did not induce large structural rearrangements in the protein core. A subsequent local 3D classification of the same dataset isolated a conformation with two clearly visible Nt domains in the same orientation, corresponding to 5517 particles. 3D refinement of this smaller dataset resulted in a 6.8-Å map with strong density for the two Nt domains. (Fig. 7b). The middle section of each Nt domain was positioned within 8 Å from several secondary structure elements of the Ct domain, and the amino-terminal portion of the Nt domain sat directly above the α-helix (residues 653–664) of the Ct domain that forms one side of the cleft accommodating the adenine moiety of NADP+ 27. Notably, although most α-helices of the Ct core were well resolved in the map, there was no cryo-EM density for this key helix.
a Relaxation of the D2 symmetry led to the appearance of weak density for one Nt domain in the cryo-EM map of ligand-free ALDH1L1. b Conformation with two symmetrically located Nt domains. The Nt domains interact with the regions of the Ct core responsible for the binding of NADP+. c Conformation of ALDH1L1 with one Nt domain interacting with the linker between the intermediate and the Ct domains (Int-Ct linker) of a different protomer. The linker is represented with the black spring. The 4′-PP arms are in sphere representation. In the two images to the right, the surface of the Nt domain is colored according to the electrostatic potential. The positively charged patch interacting with the Int-Ct linker is enclosed in dashed ovals. In the remaining images, the Nt, Int, and Ct domains are colored gold, blue, and salmon, respectively.
In another 3D class, containing 17,499 particles and refined to 7.0 Å resolution, a single Nt domain was found to straddle the Ct core between two Int domains, with both ends of Nt in contact with the ALDH1L1 core (Fig. 7c). As expected, the carboxyl-terminal region of the Nt domain was positioned close to the amino-terminal end of the Int domain of the same protomer. Interestingly, the amino-terminal portion of the Nt domain interacted with the linker connecting the Int and Ct domains of a different protomer. Although the resolution of the map is insufficient for interpreting this interaction at atomic details, it is clear that residues 55–61 of the Nt domain, composing a loop and a short beta-strand, were supported by the Int-Ct linker. Of note, while the linker is negatively charged (Fig. 3B), the complementary interface of Nt contains a positively charged patch along the contact interface (Fig. 7c).
To confirm the formation of transient complexes between Nt domains and the ALDH1L1 core, we performed chemical cross-linking of the full-length protein with 0.1% glutaraldehyde followed by NS-EM and 2D classification (Supplementary Fig. 3a). This treatment resulted in gradual disappearance of 2D classes displaying Nt moieties not in contact with the protein core (Supplementary Fig. 3b), indicating that glutaraldehyde cross-linking stabilized the transient complexes formed by the N-terminal domains. Of note, Nt domains attached to the core were reliably resolved in the 2D class averages even after prolonged glutaraldehyde treatment, suggesting that cross-linking occurred at specific positions.
Discussion
ALDH1L1 has two catalytic centers located in separate domains and utilizes a carrier protein (evolutionarily incorporated as a domain) to transfer the reaction intermediate between these centers. This carrier protein domain is highly similar to A/PCPs employed in the biosynthesis of fatty acids, non-ribosomal peptides and polyketides, reactions performed by large and complex multi-enzymatic molecular machines46,47,48. This type of modular organization implies extensive domain movements accompanying the transport of substrate between the active sites. Likewise, we found that the tetrameric aldehyde dehydrogenase module of ALDH1L1, located at the C terminus, forms the rigid core of the enzyme, whereas the N-terminal hydrolase domains assume a continuum of positions apparently constrained mainly by the length of the inter-domain linkers. In our cryo-EM structures of ALDH1L1 in the resting state (i.e., in the absence of substrate), the Int (carrier) domains were resolved docked at the substrate entrance tunnels of the Ct core, but the incomplete occupancy of these anchored carriers indicates that they operate as highly mobile units. The complex between the Nt and Int domains was not detected by cryo-EM or NS-EM, which suggests it is transient in nature, a common phenomenon for interactions involving A/PCPs49. Overall, our findings provide strong structural support for the catalytic mechanism wherein (i) the Nt domain transfers the formyl group from 10-fTHF to the 4′-PP arm of the Int domain and (ii) the Int domain delivers the formyl to the Ct domain where (iii) the formyl is oxidized to CO229 (Supplementary Fig. 4). Of note, in vitro both the Nt and Ct domains, either expressed individually or within the full-length enzyme, are capable of independent catalysis, 10-formylTHF hydrolysis or small chain aldehyde oxidation, respectively26,34. It is not clear whether such independent activities take place in the cell since the hydrolase catalysis in vitro requires high concentrations of non-physiological sulfhydryls while putative substrates for the ALDH reaction are unknown. Thus, the complex mechanism enabled by the merging of the three domains is likely the only catalytic function of ALDH1L1.
Most aldehyde dehydrogenases are known to exist as either homodimers, typified by the members of the ALDH3 family, or homotetramers, represented mainly by ALDH1/2 families, which also include ALDH1L150. Such homotetramer is organized as a dimer of homodimers formed by protomers A/B and C/D as schematically presented in Fig. 4d. Although the enzymatic mechanism of ALDH1L1 does not dictate a specific quaternary organization of the enzyme, the 3-dimensional structure revealed that the tetrameric state of the C-terminal ALDH module is indispensable for the enzyme function. In our cryo-EM structures, the carrier domains of protomers A and B were paired with the ALDH domains of protomers C and D, respectively, while the carrier domains of protomers C and D interacted with the Ct domains of protomers A and B. This pairing scheme can only be realized in a tetrameric enzyme. Furthermore, we found that the length and composition of the linker connecting the Int and Ct domains are preserved in cytosolic (ALDH1L1) and mitochondrial (ALDH1L2) enzymes as well as across multiple species, suggesting that this intricate domain pairing is a universal characteristic of 10-formyltetrahydrofolate dehydrogenases. Of note, although multiple studies analyzed the oligomeric state of ALDH proteins (recently reviewed in Shortall et al.51), the physiological significance of oligomerization is unclear for most of these enzymes. One possible exception is tetrameric ALDH from Thermus thermophilus, which has a ~30 amino-acid-long C-terminal extension that interacts with the N-terminal region of a protomer from a different homodimer52. Other ALDHs, including fatty aldehyde dehydrogenase (FALDH) and ALDH7A1, also feature short C-terminal extensions, but they interact with the protomer within the same homodimer53,54. In contrast, ALDH1L1 is the example of an ALDH with two additional domains spanning 400 amino acids at the N-terminus of the enzyme, with the tetrameric state being a prerequisite for its complex function.
The acyl and peptidyl carrier proteins evolved to shuttle catalytic intermediates between reaction centers, which requires interaction with multiple partner proteins. This functional versatility necessitates that the nature of such interactions is transient, which is crucial for the uninterrupted action of molecular machines that employ A/PCPs49. Accordingly, the contact area between the A/PCP and the partner protein is usually small, with most interacting residues confined to helix II of the carrier protein and, to a lesser extent, helix III and the part of the linker between helices I and II that is close to helix II42,55,56,57. The small size of the contact interface often requires cross-linking to enable structural investigation57,58,59. In contrast, the relatively large contact interface between the Int and Ct domains of ALDH1L1 was resolved in its native, non-cross-linked form (Fig. 5a–c). In addition to helix II, this interface also involves the base of helix I. Moreover, distinct to other A/PCPs, the beginning of the loop connecting helices I and II protrudes towards the Ct domain to interact with two helices forming the orifice of the substrate entrance tunnel (Fig. 5e). Thus, the structure of ALDH1L1 illustrates, to our knowledge, a new mode of interaction between an A/PCP-like carrier protein and its catalytic partner. Importantly, while this interaction favors the state with Int docked at the ALDH substrate entrance tunnel, the incomplete occupancy of this anchored Int domain indicates that this interaction is reversible and does not preclude the shuttling of free (undocked) Int between the catalytic domains during catalysis.
Importantly, the 4′-PP arm of the carrier domain was fully resolved in our cryo-EM maps. It spanned the entire 12-Å-deep substrate entrance tunnel of the ALDH domain, extending towards the catalytic cysteine. We found that the contacts formed by the 4′-PP prosthetic group are critical for the formation of a stable complex between the Int and ALDH domains of ALDH1L1. During catalysis, the extended 4′-PP conformation would place the formyl group transported from the Nt domain precisely in the ALDH active site, allowing the nucleophilic attack by Cys707. Curiously, in the absence of the substrate, a partial covalent bond formed between the sulfur atoms of 4′-PP and Cys707. While formation of this disulfide link is likely prevented by the formyl group attached to the 4′-PP arm during catalysis, it could be hypothesized that in the resting state of the enzyme such a bond prevents irreversible oxidation of both the catalytic cysteine and 4′-PP sulfur atoms. This disulfide could be reduced by cellular glutathione accessing the active center through the NADP+ binding site. Of note, in the individually expressed Ct domain, Cys707 was shown to form a transient covalent adduct with the C4 atom of the nicotinamide ring of NADP+ 27, which suggests that this cysteine is highly reactive beyond the immediate catalytic step. It could also be hypothesized that maintaining the 4′-PP arm within the substrate entrance tunnel prevents the entrance of small aldehydes into the ALDH catalytic center in vivo, thus preserving the enzyme for the 10-formylTHF dehydrogenase catalysis. Alternatively, we cannot exclude the possibility that the observed disulfide bond is the result of oxidation in our experimental setting.
The high mobility of the Nt domains suggests that ALDH1L1 catalysis is driven primarily by stochastic domain movements. However, the cryo-EM maps of the states with Nt domains resolved in fixed positions provide evidence of non-random interactions of these functional modules, which could play a role in the enzymatic mechanism. In one such cryo-EM map, two Nt domains were shown to interact with the NADP+ binding regions of the ALDH domains (Fig. 7b). Since a key helix forming the NADP+ binding site was disordered in each involved ALDH domain, it is possible that in this conformation the Nt domain interferes with the binding of NADP+. In support of such a possibility, this conformation was not detected in the ALDH1L1-NADP+ dataset. Based on these results, we hypothesize that the hydrolase domains of ALDH1L1 may be involved in regulating the enzymatic reaction performed by the ALDH domains. It has to be noted, however, that full-length ALDH1L1 and the individually expressed ALDH domain displayed similar affinities for NADP+ (Kd of 0.3 µM versus 0.2 µM, respectively)26, suggesting that the proposed effect is likely small. In the second cryo-EM map, the N-terminal moiety of the hydrolase domain rested on the linker connecting the carrier and ALDH domains of a different protomer (Fig. 7c), with a remarkable charge complementarity between the linker, carrying a strong negative charge (Fig. 4b), and the positively charged region of the Nt domain contacting it (Fig. 7c). We surmise that in this ALDH1L1 conformation the Nt domain may be involved in the extraction of the Int domain from the substrate entrance tunnel of the Ct core, with the positively charged patch acting as a hook. Alternatively, this domain arrangement may create a scaffold for the formation of the complex between the Nt and Int domains. In the latter scenario, a large-scale rotation and shift of the Int domain would be necessary to bring together the sulfhydryl of the 4′-PP arm and the Nt active site residues, which are ~50 Å apart. While the above interpretations are speculative, the existence of scarcely populated states with firmly positioned hydrolase domains alludes to an intricate mechanism of catalysis that may involve various auxiliary inter-domain interactions guiding the overall random domain movements during catalysis.
In summary, in this study cryo-EM revealed the unusual architecture of the multi-domain enzyme ALDH1L1, which enables the complex catalytic mechanism. Protein oligomerization and multidomain organization are common phenomena in eukaryotes60,61,62,63. While a modular organization can expand the enzyme functionality61, oligomerization provides benefits such as efficiency, regulation and stability63. In some cases, oligomerization is required because catalytic centers are formed by residues from different protomers or because oligomers enable additional non-catalytic regulatory sites. Metabolic enzymes can also form structures of higher degree of order like filaments, which might not directly affect the catalysis within a single unit64. Of note, all such examples were reported in folate metabolism where ALDH1L1 belongs65,66,67,68. Here we uncovered another mechanism in which tetrameric organization allows modular catalysis bypassing spatial restrictions within a single protomer. Thus, the tetrameric state of ALDH1L1 is indispensable for the enzyme functionality, which also involves transient domain interactions and large-scale domain movements. Finally, the complex between the intermediate and aldehyde dehydrogenase domains of ALDH1L1 demonstrates, to our knowledge, a new mode of interaction between an A/PCP-like carrier protein and a catalytic domain, emphasizing the versatility of A/PCPs.
Methods
Protein expression and purification
Full-length rat ALDH1L1 was expressed following a previously developed protocol69. Specifically, High Five insect cells (Invitrogen) grown as monolayer (Grace’s insect medium supplemented with 10% fetal bovine serum/175-cm2 cell culture flasks) at 27 °C were infected with a high titer recombinant baculovirus stock produced as previously described69. Five days after infection, the culture medium was collected, and detached cells were removed by centrifugation (10,000 × g, 10 min). To purify ALDH1L1, the cell culture medium was applied to a column containing 5-formyl-THF-Sepharose affinity resin equilibrated with 10 mM Tris-HCl buffer, pH 7.4, containing 10 mM 2-ME and 1 mM NaN3 (buffer A). The column was washed with buffer A and then with the same buffer containing 1.0 M KCl; the enzyme was eluted with buffer A containing 1.0 M KCl and 20 mM folic acid. The eluate was concentrated and excess KCl removed using a spin concentrator. Additional purification was then carried out using FPLC/Mono-Q column (GE) chromatography with a linear KCl gradient (0–0.5 M in buffer A) and Sephacryl S-300 (GE) size-exclusion chromatography in buffer A with 0.2 M NaCl. The individual Ct domain and Int-Ct protein were expressed as 6xHis tagged constructs in E. coli (Invitrogen) from pRSET vectors. Protein expression was carried out at 22 °C, and the soluble cell fraction was separated by sonication and centrifugation. The proteins were purified using Ni-NTA or Co-NTA agarose (Qiagen) using a 5–20 mM imidazole gradient to remove impurities followed by elution with 100 mM imidazole in buffer A supplemented with 100 mM KCl. Additional purification was done by size-exclusion chromatography on Sephacryl S-300. The purity of all proteins was confirmed by SDS-PAGE with Coomassie staining. Purified full-length ALDH1L1 was tested for the 10-formylTHF dehydrogenase activity as we previously described33. Ct domain and Int-Ct protein were tested for the aldehyde dehydrogenase activity using propanal as the substrate and NADP+ as the cofactor essentially as we described26. After purification, all protein preparations used in the present study had specific activities close to previously reported values26,33 and were stored at −80 °C in the presence of 10 mM 2-ME and 20% glycerol.
Liquid chromatography/mass spectrometry
In total, 30 µg of ALDH1L1 (50 µL of protein solution) was combined with 25 µL of 9 M urea and 10 µL of acetonitrile and incubated for 10 min at 42 °C. This mixture was diluted with 250 µL of 100 mM ammonium bicarbonate prior to the addition of 5 µg of sequencing grade trypsin (Promega). The proteolytic digestion was carried out for 4 h at 37 °C. The resulting peptides were loaded onto a reverse-phase C4 (2.1 mm × 50 mm) column (Thermo Scientific). Peptides were resolved and eluted with a gradient of acetonitrile in water (from 98% H2O with 0.1% (v/v) formic acid (A) and 2% acetonitrile with 0.1% (v/v) formic acid (B) to 100% B) developed over 20 min. Separation was achieved at a flow rate of 0.3 mL/min using an Agilent Technology 1100 Series HPLC system. The eluent was directed into an LTQ Velos linear trap quadropole mass spectrometer (Thermo Scientific) equipped with an electrospray ionization source operated in positive ion mode. Parameter settings of the mass spectrometer for peptide detection were as follows: activation type, collision-induced dissociation; normalized collision energy, 35 kV; capillary temperature, 370 °C; source voltage, 5 kV; capillary voltage, 43 V; tube lens, 105 V. MS spectra were collected over a 200–2000 m/z range. The raw MS data were analyzed using Qual Browser for Thermo Xcalibur version 2.1.
Negative-stain electron microscopy
Protein samples were diluted with buffer containing 10 mM HEPES, pH 7, and 150 mM NaCl to ~0.02 mg/ml. A 4.7-µl drop of the diluted sample was placed on a freshly glow-discharged carbon-coated copper grid and left for 15 s. Excess liquid was removed using filter paper, and the grid was washed three times with 4.7-µl drops of the same buffer. After the final wash, the buffer drop was removed in the same manner, and the protein was negatively stained by applying a 4.7-µl drop of 0.75% uranyl acetate for 30 s. Excess negative stain was removed using filter paper, and the grid was allowed to dry. Data were collected using SerialEM70 on a Tecnai T20 electron microscope (FEI, the Netherlands) equipped with a LaB6 filament operated at 200 kV and a 2k × 2k FEI Eagle CCD camera. The nominal magnification was 100,000x, which corresponded to a pixel size of 2.2 Å. EMAN271 was used to semi-automatically select 249,416 particles from 3920 micrographs. The selected particles were extracted into 128 × 128-pixel boxes and subjected to reference-free 2D classification into 256 classes using Relion 2.172. For separate visualization of the Nt domains, 30,186 peripheral domains (arms) of negatively stained FDH molecules were selected manually using EMAN2 from 392 micrographs of the same dataset. The selected particles were extracted into 64×64-pixel boxes and classified using Relion 2.1 into 256 classes.
Chemical cross-linking and comparative quantification of mobile N-terminal domains
Full-length ALDH1L1 was diluted to 0.01 mg/ml with buffer containing 10 mM HEPES, pH 7, and 150 mM NaCl, followed by the addition of 0.1% glutaraldehyde. Aliquots were taken before the addition of glutaraldehyde and after 1 min, 5 min, 10 min, 30 min, and 60 min of incubation at 4 °C, and negative staining and NS-EM data collection were performed as described above. All datasets were subjected to 2D classification in Relion. After discarding 2D classes that did not represent intact ALDH1L1 molecules, the final NS-EM datasets contained 57,888 (control), 36,538 (1 min), 32,722 (5 min), 39,150 (10 min), 47,877 (30 min), and 30,449 (60 min) particles. 2D class averages displaying ALDH1L1 molecules with at least one arm that was not in contact with the protein core were identified by visual inspection, and their fractions were calculated based on the total number of particles that contributed to these classes. These fractions were used solely for the purpose of comparing the cross-linking time points because not all mobile Nt domains could be captured by 2D classification due to their dynamic nature.
Cryo-electron microscopy specimen preparation and data collection
ALDH1L1 was vitrified at a concentration of 0.4 mg/ml in 20 mM HEPES, pH 7.6 (apo-ALDH1L1) or 40 mM HEPES, pH 7, 1 mM NADP+ (ALDH1L1-NADP+ complex). Cryo-EM specimens were prepared by plunge-freezing in liquid ethane using Vitrobot Mark IV (FEI) at room temperature and 90% humidity. The grids (Quantifoil R2/2 with gold support) were glow-discharged for 30 s at a pressure of 37 mBar and with the current set to 30 mA. The drop volume was 3 µl. Data were collected at the National Cryo-Electron Microscopy Facility (NCEF) at National Cancer Institute on a Titan Krios electron microscope (FEI) operated at 300 kV and equipped with a K2-Summit direct electron detector (Gatan). The detector was used in the super-resolution mode. For apo-ALDH1L1, 2202 movies were collected with a nominal dose of 40 e−/Å2 equally distributed between 40 frames of a 12-s movie, and the pixel size (super-resolution mode) was 0.66 Å (magnification: 105,000x). The defocus range was −1 to −3 µm. For ALDH1L1-NADP+ complex, 2381 movies were collected with a nominal dose of 40 e−/Å2 equally distributed between 40 frames of a 14-s movie, and the pixel size (super-resolution mode) was 0.532 Å (magnification: 130,000x). The defocus range was −1 to −2.5 µm (Table 1).
Single-particle analysis of cryo-electron microscopy data
Motion correction and dose weighting were performed using MotionCor273. For local motion correction, frames were divided into 25 tiles. Images were binned 2x (apo-ALDH1L1) and 1.5x (ALDH1L1-NADP+) during motion correction, resulting in pixel sizes of 1.32 Å and 0.76 Å, respectively. Contrast transfer function parameters were estimated using ctffind 4.174. All other image processing steps were performed in Relion 3.072 unless stated otherwise. Particles were picked automatically using projections of an X-ray structure of the tetrameric C-terminal domain of ALDH1L1 (PDB 2o2p27) low-pass filtered to 40 Å, resulting in datasets containing 1,082,600 (apo-ALDH1L1) and 1,050,740 (ALDH1L1/NADP+) particles. The particles were extracted, with 2x binning, into 80 × 80 (apo- ALDH1L1) or 140 × 140 (ALDH1L1/NADP+) pixel boxes and subjected to reference-free 2D classification into 128 classes with selection of high-resolution classes corresponding to a complete, undistorted tetramer of the Ct domain that appeared symmetrical. This selection reduced the size of the datasets to 424,239 and 640,945 particles, respectively. The corresponding particles were re-extracted, without binning, into 160 × 160 (apo-ALDH1L1) or 280 × 280 (ALDH1L1/NADP+) pixel boxes, and reference-free 2D classification into 128 classes was repeated. Selection of best-looking classes resulted in datasets of 147,837 (apo-FDH) and 594,883 (FDH/NADP+) particles. 3D classification into 10 classes was performed next with the above-mentioned X-ray structure of the tetramer of the Ct domain low-pass filtered to 40 Å serving as the initial model. No symmetry was imposed at this stage. The presence of additional density at the substrate entrance tunnel of the Ct domain was obvious in the resulting 3D classes, and in all 3D classes with sufficiently high resolution this density consisted of four α-helices and an arm protruding deep into the substrate entrance tunnel. Additional density consistent with the size and shape of the Nt domain of ALDH1L1 (PDB 1s3i21) was observed in several 3D classes. High-resolution 3D classes, as well as 3D classes with density for one or more Nt domains, were subjected to 3D auto-refinement with D2 or C2 symmetry imposed as well as without imposing symmetry. Post-processing included automatic B-factor sharpening and detector modulation transfer function correction, and the gold-standard resolution was determined within a soft mask using a 0.143 FSC threshold. Local resolution was estimated using ResMap75. Representative micrographs and 2D class averages, FSC curves, and local resolution data are presented in Supplementary Figs. 5 and 6. Supplementary Fig. 7 illustrates the cryo-EM density for 4′-PP and NADP+.
Model building
The crystal structure of the tetrameric C-terminal domain of ALDH1L1 (residues 405-902) in the apo form (PDB 2o2p27) or in complex with NADP+ (PDB 2o2q27) and four instances of a homology model of the Int domain of rat FDH (residues 306–402) obtained using the SWISS-MODEL server76 were fit into the corresponding cryo-EM density using USCF Chimera77. This was followed by one round of real-space refinement in PHENIX78 and alternating rounds of model building in Coot79 and restrained model refinement in Refmac80. Molprobity81 was used to assess the quality of the atomic models. Map-model correlations were evaluated using phenix.mtriage82.
Estimation of occupancy of Int domains
A soft mask was prepared for each of the four Int domains by segmenting the symmetrical ALDH1L1 map in UCSF Chimera (Supplementary Figure 2). Before 3D classification, both apo-ALDH1L1 and ALDH1L1-NADP+ maps were refined without symmetry imposed. 3D classification into 8–10 classes without particle alignment was then performed in Relion 3.0 for each Int domain separately using the final map low-pass filtered to 40 Å as the reference. The resulting 3D classes were examined visually, and total particle counts for classes with occupied and vacant Int domain binding sites were determined. Int domain occupancy was calculated as the fraction of the particles contributing to the classes representing occupied sites, averaged across the four sites within the tetramer.
Other methods
Protein structure similarity search was performed with the mTM-align server (35). Figures were prepared in UCSF Chimera, UCSF ChimeraX83, and Coot.
Statistics and reproducibility
LC/MS experiments were repeated four times. The cross-linked peptides were detected in all these experiments.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Cryo-EM maps of ligand-free ALDH1L1 and ALDH1L1 in complex with NADP+ have been deposited to the EMDB with accession codes EMDB-24540 and EMDB-24547, respectively. Fitted coordinates have been deposited to the PDB with accession codes 7RLT and 7RLU, respectively. All other data are available from the corresponding authors upon request.
References
Krupenko, S. A. FDH: an aldehyde dehydrogenase fusion enzyme in folate metabolism. Chem. Biol. Interact. 178, 84–93 (2009).
Anthony, T. E. & Heintz, N. The folate metabolic enzyme ALDH1L1 is restricted to the midline of the early CNS, suggesting a role in human neural tube defects. J. Comp. Neurol. 500, 368–383 (2007).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Krupenko, S. A. & Krupenko, N. I. Loss of ALDH1L1 folate enzyme confers a selective metabolic advantage for tumor progression. Chem. Biol. Interact. 302, 149–155 (2019).
Krupenko, N. I. et al. Cytosolic 10-formyltetrahydrofolate dehydrogenase regulates glycine metabolism in mouse liver. Sci. Rep. 9, 14937 (2019).
Damal Villivalam, S. et al. A necessary role of DNMT3A in endurance exercise by suppressing ALDH1L1-mediated oxidative stress. EMBO J. 40, e106491 (2021).
Krupenko, S. A. & Oleinik, N. V. 10-formyltetrahydrofolate dehydrogenase, one of the major folate enzymes, is down-regulated in tumor tissues and possesses suppressor effects on cancer cells. Cell Growth Differ. 13, 227–236 (2002).
Tackels-Horne, D. et al. Identification of differentially expressed genes in hepatocellular carcinoma and metastatic liver tumors by oligonucleotide expression profiling. Cancer 92, 395–405 (2001).
Oleinik, N. V., Krupenko, N. I. & Krupenko, S. A. Epigenetic silencing of ALDH1L1, a metabolic regulator of cellular proliferation, in cancers. Genes Cancer 2, 130–139 (2011).
Beniaminov, A. D. et al. Deep sequencing revealed a CpG methylation pattern associated With ALDH1L1 suppression in breast cancer. Front. Genet. 9, 169 (2018).
Dmitriev, A. A. et al. Genetic and epigenetic analysis of non-small cell lung cancer with NotI-microarrays. Epigenetics 7, 502–513 (2012).
Dmitriev, A. A. et al. Epigenetic alterations of chromosome 3 revealed by NotI-microarrays in clear cell renal cell carcinoma. Biomed. Res. Int. 2014, 735292 (2014).
Senchenko, V. N. et al. Novel tumor suppressor candidates on chromosome 3 revealed by NotI-microarrays in cervical cancer. Epigenetics 8, 409–420 (2013).
Oleinik, N. V. & Krupenko, S. A. Ectopic expression of 10-formyltetrahydrofolate dehydrogenase in a549 cells induces g(1) cell cycle arrest and apoptosis. Mol. Cancer Res. 1, 577–588 (2003).
Oleinik, N. V., Krupenko, N. I., Priest, D. G. & Krupenko, S. A. Cancer cells activate p53 in response to 10-formyltetrahydrofolate dehydrogenase expression. Biochem. J. 391, 503–511 (2005).
Oleinik, N. V., Krupenko, N. I. & Krupenko, S. A. Cooperation between JNK1 and JNK2 in activation of p53 apoptotic pathway. Oncogene 26, 7222–7230 (2007).
Hoeferlin, L. A., Oleinik, N. V., Krupenko, N. I. & Krupenko, S. A. Activation of p21-dependent G1/G2 arrest in the absence of DNA damage as an antiapoptotic response to metabolic stress. Genes Cancer 2, 889–899 (2011).
Hoeferlin, L. A., Fekry, B., Ogretmen, B., Krupenko, S. A. & Krupenko, N. I. Folate stress induces apoptosis via p53-dependent de novo ceramide synthesis and up-regulation of ceramide synthase 6. J. Biol. Chem. 288, 12880–12890 (2013).
Khan, Q. A. et al. CHIP E3 ligase mediates proteasomal degradation of the proliferation regulatory protein ALDH1L1 during the transition of NIH3T3 fibroblasts from G0/G1 to S-phase. PLoS ONE 13, e0199699 (2018).
Krupenko N. I., et al. Knockout of putative tumor suppressor Aldh1l1 in mice reprograms metabolism to accelerate growth of tumors in a diethylnitrosamine (DEN) model of liver carcinogenesis. Cancers 13, 3219 (2021).
Chumanevich, A. A., Krupenko, S. A. & Davies, C. The crystal structure of the hydrolase domain of 10-formyltetrahydrofolate dehydrogenase: mechanism of hydrolysis and its interplay with the dehydrogenase domain. J. Biol. Chem. 279, 14355–14364 (2004).
Schmitt, E., Blanquet, S. & Mechulam, Y. Structure of crystalline Escherichia coli methionyl-tRNA(f)Met formyltransferase: comparison with glycinamide ribonucleotide formyltransferase. EMBO J. 15, 4749–4758 (1996).
Schmitt, E., Panvert, M., Blanquet, S. & Mechulam, Y. Crystal structure of methionyl-tRNAfMet transformylase complexed with the initiator formyl-methionyl-tRNAfMet. EMBO J. 17, 6819–6826 (1998).
Tucker, E. J. et al. Mutations in MTFMT underlie a human disorder of formylation causing impaired mitochondrial translation. Cell Metab. 14, 428–434 (2011).
Koppaka, V. et al. Aldehyde dehydrogenase inhibitors: a comprehensive review of the pharmacology, mechanism of action, substrate specificity, and clinical application. Pharmacol. Rev. 64, 520–539 (2012).
Krupenko, S. A., Wagner, C. & Cook, R. J. Expression, purification, and properties of the aldehyde dehydrogenase homologous carboxyl-terminal domain of rat 10-formyltetrahydrofolate dehydrogenase. J. Biol. Chem. 272, 10266–10272 (1997).
Tsybovsky, Y., Donato, H., Krupenko, N. I., Davies, C. & Krupenko, S. A. Crystal structures of the carboxyl terminal domain of rat 10-formyltetrahydrofolate dehydrogenase: implications for the catalytic mechanism of aldehyde dehydrogenases. Biochemistry 46, 2917–2929 (2007).
Krupenko, S. A., Wagner, C. & Cook, R. J. Cysteine 707 is involved in the dehydrogenase activity site of rat 10-formyltetrahydrofolate dehydrogenase. J. Biol. Chem. 270, 519–522 (1995).
Donato, H., Krupenko, N. I., Tsybovsky, Y. & Krupenko, S. A. 10-formyltetrahydrofolate dehydrogenase requires a 4′-phosphopantetheine prosthetic group for catalysis. J. Biol. Chem. 282, 34159–34166 (2007).
Lai, J. R., Koglin, A. & Walsh, C. T. Carrier protein structure and recognition in polyketide and nonribosomal peptide biosynthesis. Biochemistry 45, 14869–14879 (2006).
Finzel, K., Lee, D. J. & Burkart, M. D. Using modern tools to probe the structure-function relationship of fatty acid synthases. Chembiochem 16, 528–547 (2015).
Krupenko, S. A. & Wagner, C. Aspartate 142 is involved in both hydrolase and dehydrogenase catalytic centers of 10-formyltetrahydrofolate dehydrogenase. J. Biol. Chem. 274, 35777–35784 (1999).
Krupenko, S. A., Wagner, C. & Cook, R. J. Recombinant 10-formyltetrahydrofolate dehydrogenase catalyses both dehydrogenase and hydrolase reactions utilizing the synthetic substrate 10-formyl-5,8-dideazafolate. Biochem. J. 306, 651–655 (1995). Pt 3.
Krupenko, S. A., Wagner, C. & Cook, R. J. Domain structure of rat 10-formyltetrahydrofolate dehydrogenase. Resolution of the amino-terminal domain as 10-formyltetrahydrofolate hydrolase. J. Biol. Chem. 272, 10273–10278 (1997).
Krupenko, S. A., Vlasov, A. P. & Wagner, C. On the role of conserved histidine 106 in 10-formyltetrahydrofolate dehydrogenase catalysis: connection between hydrolase and dehydrogenase mechanisms. J. Biol. Chem. 276, 24030–24037 (2001).
Reuland, S. N., Vlasov, A. P. & Krupenko, S. A. Disruption of a calmodulin central helix-like region of 10-formyltetrahydrofolate dehydrogenase impairs its dehydrogenase activity by uncoupling the functional domains. J. Biol. Chem. 278, 22894–22900 (2003).
Reuland, S. N., Vlasov, A. P. & Krupenko, S. A. Modular organization of FDH: exploring the basis of hydrolase catalysis. Protein Sci. 15, 1076–1084 (2006).
Tsybovsky, Y. & Krupenko, S. A. Conserved catalytic residues of the ALDH1L1 aldehyde dehydrogenase domain control binding and discharging of the coenzyme. J. Biol. Chem. 286, 23357–23367 (2011).
Tsybovsky, Y., Malakhau, Y., Strickland, K. C. & Krupenko, S. A. The mechanism of discrimination between oxidized and reduced coenzyme in the aldehyde dehydrogenase domain of Aldh1l1. Chem. Biol. Interact. 202, 62–69 (2013).
Lin, C. C. et al. Structures of the hydrolase domain of zebrafish 10-formyltetrahydrofolate dehydrogenase and its complexes reveal a complete set of key residues for hydrolysis and product inhibition. Acta Crystallogr. D Biol. Crystallogr. 71, 1006–1021 (2015).
Kursula, P. et al. Structures of the hydrolase domain of human 10-formyltetrahydrofolate dehydrogenase and its complex with a substrate analogue. Acta Crystallogr. D. Biol. Crystallogr. 62, 1294–1299 (2006).
Byers, D. M. & Gong, H. Acyl carrier protein: structure-function relationships in a conserved multifunctional protein family. Biochem. Cell Biol. 85, 649–662 (2007).
Perez-Miller, S. J. & Hurley, T. D. Coenzyme isomerization is integral to catalysis in aldehyde dehydrogenase. Biochemistry 42, 7100–7109 (2003).
Wyatt, J. W. et al. Inhibition, crystal structures, and in-solution oligomeric structure of aldehyde dehydrogenase 9A1. Arch. Biochem. Biophys. 691, 108477 (2020).
Potel, C. M., Lemeer, S. & Heck, A. J. R. Phosphopeptide fragmentation and site localization by mass spectrometry: an update. Anal. Chem. 91, 126–141 (2019).
Leibundgut, M., Jenni, S., Frick, C. & Ban, N. Structural basis for substrate delivery by acyl carrier protein in the yeast fatty acid synthase. Science 316, 288–290 (2007).
Maier, T., Leibundgut, M. & Ban, N. The crystal structure of a mammalian fatty acid synthase. Science 321, 1315–1322 (2008).
Tanovic, A., Samel, S. A., Essen, L. O. & Marahiel, M. A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008).
Beld, J., Lee, D. J. & Burkart, M. D. Fatty acid biosynthesis revisited: structure elucidation and metabolic engineering. Mol. Biosyst. 11, 38–59 (2015).
Vasiliou, V., Thompson, D. C., Smith, C., Fujita, M. & Chen, Y. Aldehyde dehydrogenases: from eye crystallins to metabolic disease and cancer stem cells. Chem. Biol. Interact. 202, 2–10 (2013).
Shortall, K., Djeghader, A., Magner, E. & Soulimane, T. Insights into aldehyde dehydrogenase enzymes: a structural perspective. Front. Mol. Biosci. 8, 659550 (2021).
Hayes, K. et al. The quaternary structure of Thermus thermophilus aldehyde dehydrogenase is stabilized by an evolutionary distinct C-terminal arm extension. Sci. Rep. 8, 13327 (2018).
Keller, M. A. et al. A gatekeeper helix determines the substrate specificity of Sjogren-Larsson Syndrome enzyme fatty aldehyde dehydrogenase. Nat. Commun. 5, 4439 (2014).
Luo, M. & Tanner, J. J. Structural basis of substrate recognition by aldehyde dehydrogenase 7A1. Biochemistry 54, 5513–5522 (2015).
Crosby, J. & Crump, M. P. The structural role of the carrier protein−active controller or passive carrier. Nat. Prod. Rep. 29, 1111–1137 (2012).
Hur, G. H., Vickery, C. R. & Burkart, M. D. Explorations of catalytic domains in non-ribosomal peptide synthetase enzymology. Nat. Prod. Rep. 29, 1074–1098 (2012).
Gulick, A. M. & Aldrich, C. C. Trapping interactions between catalytic domains and carrier proteins of modular biosynthetic enzymes with chemical probes. Nat. Prod. Rep. 35, 1156–1184 (2018).
Nguyen, C. et al. Trapping the dynamic acyl carrier protein in fatty acid biosynthesis. Nature 505, 427–431 (2014).
Miyanaga, A., Iwasawa, S., Shinohara, Y., Kudo, F. & Eguchi, T. Structure-based analysis of the molecular interactions between acyltransferase and acyl carrier protein in vicenistatin biosynthesis. Proc. Natl Acad. Sci. USA 113, 1802–1807 (2016).
Ali, M. H. & Imperiali, B. Protein oligomerization: how and why. Bioorg. Med. Chem. 13, 5013–5020 (2005).
Han, J. H., Batey, S., Nickson, A. A., Teichmann, S. A. & Clarke, J. The folding and evolution of multidomain proteins. Nat. Rev. Mol. Cell Biol. 8, 319–330 (2007).
Bordin, N., Sillitoe, I., Lees, J. G. & Orengo, C. Tracing evolution through protein structures: nature captured in a few thousand folds. Front. Mol. Biosci. 8, 668184 (2021).
Goodsell, D. S. & Olson, A. J. Structural symmetry and protein function. Annu. Rev. Biophys. Biomol. Struct. 29, 105–153 (2000).
Lynch, E. M., Kollman, J. M. & Webb, B. A. Filament formation by metabolic enzymes-A new twist on regulation. Curr. Opin. Cell Biol. 66, 28–33 (2020).
Renwick, S. B., Snell, K. & Baumann, U. The crystal structure of human cytosolic serine hydroxymethyltransferase: a target for cancer chemotherapy. Structure 6, 1105–1116 (1998).
Luka, Z. et al. 5-methyltetrahydrofolate is bound in intersubunit areas of rat liver folate-binding protein glycine N-methyltransferase. J. Biol. Chem. 282, 4069–4075 (2007).
Mao, Y. et al. Structure of the bifunctional and Golgi-associated formiminotransferase cyclodeaminase octamer. EMBO J. 23, 2963–2971 (2004).
Pedley, A. M. & Benkovic, S. J. A new view into the regulation of purine metabolism: the purinosome. Trends Biochem. Sci. 42, 141–154 (2017).
Krupenko, S. A., Horstman, D. A., Wagner, C. & Cook, R. J. Baculovirus expression and purification of rat 10-formyltetrahydrofolate dehydrogenase. Protein Expr. Purif. 6, 457–464 (1995).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Pettersen, E. F. et al. UCSF Chimera−a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D Biol. Crystallogr. 71, 136–153 (2015).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr 66, 12–21 (2010).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D Struct. Biol. 74, 814–840 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Drake, E. J. et al. Structures of two distinct conformations of holo-non-ribosomal peptide synthetases. Nature 529, 235–238 (2016).
Bunkoczi, G. et al. Mechanism and substrate recognition of human holo ACP synthase. Chem. Biol. 14, 1243–1253 (2007).
Cryle, M. J. & Schlichting, I. Structural insights from a P450 Carrier Protein complex reveal how specificity is achieved in the P450(BioI) ACP complex. Proc. Natl Acad. Sci. USA 105, 15696–15701 (2008).
Babu, M. et al. Structure of a SLC26 anion transporter STAS domain in complex with acyl carrier protein: implications for E. coli YchM in fatty acid metabolism. Structure 18, 1450–1462 (2010).
Reimer, J. M., Aloise, M. N., Harrison, P. M. & Schmeing, T. M. Synthetic cycle of the initiation module of a formylating nonribosomal peptide synthetase. Nature 529, 239–242 (2016).
Acknowledgements
This work was supported in part by federal funds from the Frederick National Laboratory for Cancer Research, NIH, under Contract HHSN261200800001 (YT) and the NIH R01 DK54388 grant (SAK). M.G. was supported by the NIH R01 EY023948 grant. Cryo-EM datasets were collected at the National CryoEM Facility (NCEF) of the National Cancer Institute. We would like to thank Dr. Ulrich Baxa for collecting cryo-EM data. This research was, in part, supported by the National Cancer Institute’s National Cryo-EM Facility at the Frederick National Laboratory for Cancer Research under contract HSSN261200800001E.
Funding
Open Access funding provided by the National Institutes of Health (NIH).
Author information
Authors and Affiliations
Contributions
Y.T. and S.A.K. conceived and planned the research. V.S. and N.I.K. produced the proteins. Y.T. and V.S. performed negative-stain EM and prepared cryo-EM specimens. Y.T. performed single-particle analysis of negative-stain and cryo-EM datasets. M.G. performed LC/MS experiments. Y.T. and S.A.K. wrote the manuscript, with all authors providing revisions and comments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Janesh Kumar and Anam Akhtar. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsybovsky, Y., Sereda, V., Golczak, M. et al. Structure of putative tumor suppressor ALDH1L1. Commun Biol 5, 3 (2022). https://doi.org/10.1038/s42003-021-02963-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02963-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.