Abstract
Lipoprotein lipase (LPL) is a rate-limiting enzyme for the hydrolysis of triglycerides (TG). Hundreds of genetic variants including single nucleotide polymorphisms have been identified across the 30Kb gene locus on chromosome 8q22. Several of these variants have been demonstrated to have genetic association with lipid level variation but many remain unresolved. Controversial reports on the genetic association of variants among different populations pose a challenge to which variants are informative. This study aimed to investigate “common” LPL variants (rs1121923, rs258, rs328, rs13702) and their possible role in plasma lipid level. Genotyping was performed using Realtime PCR. Based on the observed genotypes, the minor allele frequencies were A: 0.065 for rs1121923; C: 0.379 for rs258; G: 0.087 for rs328 and C: 0.337 for rs13702. Using linear regression, a lowering effect of rs1121923 (p = 0.024) on TG levels (−0.14 B coefficient: CI: −0.27–−0.019) and rs258 (p = 0.013) on VLDL levels (B: −0.046; CI: −0.082–−0.009) was observed indicating a “protective” role for the two variants. Moreover, the findings indicate the potential for including rs1121923 and rs258 in diagnostic panels for use as an estimator of “risk” scores for dyslipidemia.
Similar content being viewed by others
Introduction
Plasma lipid levels play an important role in maintaining homeostasis and are useful risk markers for cardio-metabolic disorders such as the metabolic syndrome and type 2 diabetes mellitus (T2DM). Persistent elevation of plasma lipid levels often results in dyslipidemia that may lead to further complications such as coronary heart disease (CHD). It has been reported that hypertriglyceridemia (HTG) is a risk factor for CHD through mediating decreased levels of high density lipoprotein-cholesterol (HDL-C) and increased levels of low density lipoprotein-C (LDL-C) that may facilitate thrombogenicity leading to atherosclerosis1. Although numerous studies have demonstrated the complex etiology of dyslipidemia implicating a variety of environmental factors such as nutrition, conflicting results persist with regards to the role of genetic factors. More recently, studies have indicated that the heritably of plasma lipid levels is not only influenced by numerous genetic variants but also by ethnicity2,3,4,5,6. In fact, Deo et al. reported that local ancestry contributed significantly (p < 0.05) to variation in lipid levels3. Furthermore, Johansen et al. (2010) indicated that both common and rare genetic variants could explain 41.6% of total variation in HTG with common genetic variants explaining 20.8% (specifically for 7 gene loci) and the rare genetic variants explaining only 1.1% (at 4 gene loci) while the other associated factors explained 19.7% of the cases7. Among all these studies, the gene for lipoprotein lipase (LPL) has always been implicated with a significant influence on one or multiple variations in lipid parameters2,3,4,6,7,8,9.
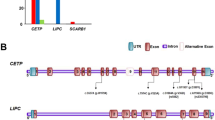
The LPL gene codes for the 475-amino acid enzyme responsible for the hydrolysis of triglycerides to free fatty acid and is an important catalyst in lipid metabolism and transport pathways. The gene has been fully mapped to chromosome 8q22 and has been fully sequenced in different ethnic groups2,5,6,10,11,12. The gene spans 30Kb including 10 exons with the first exon encoding the 5′ untranslated region (UTR) and signal peptide and the last exon encoding the full 3′ UTR. Most of the gene comprises noncoding sequences mainly localized to intron 16,10. Some studies have reported associations of numerous variants in both the coding and the non-coding regions, especially in introns 2, 3, 5, 6 and 8 with variation in lipid levels2,5,6,10,11,12,13,14,15.
Despite consistency in the implication of LPL and variation in lipid levels with associated clinical manifestations2,16,17,18,19,20,21,22, the effect of many of the significant variants identified were not observed in different populations and ethnic groups2,3,5,6,8,9,12,13. It has been suggested that the role of LPL and its variants be further investigated in different populations with reference to ethnic backgrounds2. However, challenges are encountered when deciding which variants should be selected for genetic association with lipid levels3,6,7,9. A common approach would be to select a representative SNP that is in strong Linkage disequilibrium with other SNPs and that the selected tagged SNPs would represent the different haplogroups across the LPL gene. Such an approach may prove to be time and cost ineffective. Johansen et al. (2010) indicated that 20.8% of variation in TG levels could be explained by common variants. In addition, Deo et al. identified 12 risk variants at the LPL gene locus associated with TG levels, four of which are included in this study. Moreover, Evans et al. (2013) suggested the use of the common disease common variant (CDCV) model is applicable to studying LPL variants. The model implies that genetic variants which occur at high frequencies among the general population would increase the susceptibility to the disease but with a small “effect size”7.
With this approach in mind, the present study aimed to investigate common LPL variants with a global minor frequency above 5% among the general Kuwaiti population to assess their role in contributing to fluctuations in plasma lipid levels, specifically triglycerides (TG) and high-density lipoprotein-Cholesterol (HDL-C). The four variants selected have been previously reported for their effect size in different ethnic backgrounds3,5,12. Two of the variants selected (rs1121923 and rs258) have not been extensively studied in different populations21,22 while the other two (rs328 and rs13702) have been extensively investigated with conflicting results2,3,4,5,8,22. Deo et al. (2009) identified rs328 as a strong indicator (p = 2.7 × 10−6) of increased TG levels with a higher impact in populations of African ancestry when compared to Europeans. The other commonly studied variant rs325 was not included as it showed strong LD with rs328. However, conflicting results have been reported with regards to rs3283,4,5,12,22. Several studies have suggested that rs13702 is a strong candidate for TG and HDL-C and TG levels4,8,23.
The present study investigated LPL common variants (rs1121923; rs258; rs328 and rs13702) (Table 1) in a Kuwaiti cohort that have been implicated in disorders associated with plasma lipid levels and that yielded either conflicting or inconclusive findings. The SNPs are in regions reported to be involved in gene expression or splice regulation. Common variants, based on the minor allele frequency reported in the GenBank database and based on Genome Reference Consortium Human Build 38 patch release 12 (Grch38.p12), are more likely to indicate a genetic association with variation in lipid levels, if present, among a small sample size which is representative of a heterogenous population. The cohort in this study is a representative of an Arab admixed population24 in which common variants are more likely to yield informative and conclusive results than rare variants among a population known for demonstrating high prevalence of dyslipidemia and the metabolic syndrome25,26,27.
Results
Genotyping, Hardy-Weinberg Equilibrium and Linkage Disequilibrium
The most frequent genotype observed was that for the homozygous wildtype allele except for variant rs258 in which the heterozygous form had the highest frequencies (Table 2). All the frequencies were found to be in HWE (p > 0.05). The MAF was found to be lowest for rs1121923 A allele (A: 0.065) while the highest was for rs258 (C: 0.379).
Analysis of LD revealed that the four selected variants were not correlated and didn’t form a common haploblock (Fig. 1). The four SNPs at the LPL locus (rs1121923, rs258, rs328 and rs13702) were analyzed in a total of 12 pair-wise combinations and the resulting r2 values showed no significant LD between any SNP pairs (r2 < 0.8).
Haploblock scheme between LPL rs1121923, rs258, rs328 and rs13702 based on linkage disequilibrium and recombination frequencies. The confidence Bounds Color Scheme shows evidence of LD (dark grey, black), uninformative (shades of light grey) and strong evidence of recombination (white) based on the r2 values, all of which were found insignificant. The image was generated by Haploview (version 4.2.).
Genetic Association and Regression Analysis
Linear regression analysis based on the additive genetic model of the four variants against variation in plasma lipid levels detected significant associations (p < 0.05) for only rs1121923 (Table 3) and rs258 with TG and VLDL levels respectively (Table 4). Significant (p = 0.02), however the significance level was comprised after Bonferroni’s correction (p = 0.0125), lower levels of TG (0.94 mmol/L ± 0.74 for GA and 1.09 mmol/L ± 0.84 for GG) and border line significance of VLDL (0.38 mmol/L ± 0.30 for GA and 0.45 mmol/L ± 0.35 for G) were observed for the heterozygous individuals. In addition, significantly lower VLDL levels were observed in both heterozygous (0.41 mmol/L ± 0.31) and homozygous individuals for the minor allele C (0.43 mmol/L ± 0.27) when compared to the homozygous wildtype allele G (0.48 mmol/L ± 0.41) for rs258. A borderline significant association was also observed for rs328 (Table 5) with LDL-C levels in which individuals with the homozygous wildtype had significantly lower levels (3.09 mmol/L ± 0.81) than carriers (3.24 mmol/L ± 0.86 for GC and 3.2 mmol/L ± 0.52 for GG) of the minor G allele. No significant association was observed for rs13702 (Table 6).
A multivariate analysis using linear regression was conducted on the significant SNPs to indicate the predictor variables associated with lipid levels. BMI, age and gender along with rs1121923 genotype were all found to be predictors of TG levels (Table 7). The AA genotype of rs258 was associated with a lowering effect indicated by a B coefficient of −0.14 (95% CI: −0.27 – −0.019; p = 0.24). In addition, BMI, age and gender along with rs1121923 were significantly associated with VLDL levels (Table 8). The genotype CC of rs258 was associated with a lowering effect as indicated by a B coefficient of −0.046 (95% CI: −0.082 – −0.009; p = 0.013).
Discussion
The present study is the first to report a significant association of an LPL intronic variant rs258 with a lowering effect on VLDL levels as well as a lowering effect of the synonymous variant rs1121923 on plasma TG levels amongst an apparently healthy individual. Both variants have been previously implicated for their potential effects on LPL activity and role in affecting plasma lipid levels3,6,22. However, the novel findings from the present study clearly demonstrated the significant “protective” effect of the minor alleles of the two variants in lowering TG and VLDL levels.
The positive association of these non-structural variants may be the outcome of their interaction with other loci that modulate LPL expression levels. It has been suggested that although the synonymous variant rs1121923 located in exon 3 doesn’t alter the enzyme structure, it may however affect LPL levels22,28 and/or activity. In a recent study, a strong positive association of these variants was reported with higher HDL-C levels for carriers of the minor allele21. The present study also supports a “protective” effect of rs1121923 variants on plasma lipid levels despite the effect of Bonferroni’s correction on the initial significance observed (p = 0.02). It worth noting that there are concerns when applying Bonforroni’s correction especially since it relies on the assumption that the same variants are all simultaneously significant which is not the case in this type of study in addition to its contribution in increasing Type II errors29. The haplotype analysis clearly demonstrated that the four variants are segregating independently. In addition, the authors opted to perform haplotype analysis that takes into consideration several significant variants that may occur simultaneously but through linkage disequilibrium.
The mechanism by which the minor allele of this variant lower’s TG is more likely to be the outcome of interaction with other variants either at the same gene locus or other gene loci involved in lipid metabolism and transport. It has been suggested that simple variants within regions encompassing consensus and important sequences may exert a pathogenic effect through the inactivation of splice and/or activation of cryptic splice sites leading to undesirable alternative splicing2. This is a likely scenario for rs1121923 as exon three codes for a fraction of the 20 amino acids that form the B loop of the LPL enzyme, which is important for its catalytic activity30.
It is worth pointing out that the novel findings in the present study is in establishing a significant association of LPL, and specifically variant rs258, with variation in VLDL levels. Other studies have reported an association of other variants of LPL with VLDL14,31. Salinelli et al. (1998) demonstrated the role the LPL enzyme plays in the uptake of VLDL followed by its hydrolysis through the binding of specific domains in the enzyme to a lipoprotein receptor14. The association of rs258 with its lowering effect on VLDL level may also be through affecting protein binding and/or regulation of expression levels of LPL similar to other reported intronic variants17,32. A few studies have reported novel and rare variants in intronic regions to be associated with VLDL levels6,33. Both rs261 and rs263 in intron 5 have been reported to be associated with changes in both TG and HDL levels3 thus supporting the effect variants might have on plasma lipid levels. The mechanism behind this effect may be due to intron 5 harboring sites for regulatory elements as has been observed with intron 8 and the effect of rs32515. Another important consideration is that intron 5 flanks the coding sequences for the binding sites of APOC2 and that can affect the catalytic activity of the enzyme11. Intron 5 has the least number of reported variants as compared to intron 1, 6 and 9 indicating that it has highly conserved regions due to its important regulatory role and/or due to the relative size of this intron. It has been suggested that regulatory elements, believed to span across the LPL gene locus at the 3′ and 5′ UTR as well as within intronic regions, could be sensitive to trans acting regulatory factors which may be either intrinsic or extrinsic15. This may affect the modulation of binding of transcription factors needed during gene expression, and in turn contribute to the risk of dyslipidemia and the subsequent clinical manifestation of the metabolic syndrome and CHD6,19,20.
Although several studies have reported positive genetic association of rs328 and rs137023,5,8,17,21,22 with plasma lipid levels, the present study did not identify any such association. This is very likely a consequence of choosing apparently healthy subjects in the studied cohort. Both rs328 and rs13702 variants have been documented to be associated with a pathogenic effect increasing the risk to clinical dyslipidemia and the metabolic syndrome. Another interesting finding is the detected association between the two LPL variants (rs1121923 and rs258) is likely to be the outcome of the number of heterozygous individuals since very low numbers of homozygous for the minor allele were identified in the cohort. This suggests that the alleles at these two loci are co-dominant which is supported by the additive genetic modeling (Tables 7 and 8). Accordingly, it is likely that the pathogenic potential of these two variants may be observed in clinical cases of dyslipidemia, the metabolic syndrome and CHD where there may be a higher frequency of homozygosity for the minor allele than individuals devoid of such diseases.
Studies that supported a genetic association with lipid levels included numerous LPL variants and tagged SNPs, based on the haplogroups, that included rs13702, rs320, rs325 and rs328. These studies had excluded rs1121923 and rs258 from the association studies which in this study were identified to be worthy of analysis along with rs13702 and rs328 (Table 9). In one study that reported a significant association of rs320 with TG levels in Hispanics28 was often in linkage disequilibrium with rs3283,12 suggested that some variants maybe independent of other variable sites in the LPL gene locus. This supports the haplotype analysis in this study which revealed that the four selected variants (Fig. 1) are not in linkage disequilibrium thus suggesting that rs1121923 and rs258 a high direct and an independent association with TG and VLDL levels.
Conclusion
The findings in the present study confirm the role of genetic variants in the noncoding regions with variation in plasma lipid levels. These variants may play a direct role in exerting their effect either on the expression of the gene directly or through the interaction with other variants. In addition, the identification of a significant association of two SNPs with variation in plasma lipid levels may be specific to the Arab ethnic group represented by the Kuwaiti population. Previous studies have emphasized the role of local ancestry in generating significant association of genetic variants with plasma lipid levels that may be ethnic specific3,6. This in turn highlights the importance of identifying ethnic specific genetic variants that can affect lipid metabolism. Once identified with a confirmed effect, either “risk” or “protective”, such variants could be used to form molecular diagnostic panels for screening for dyslipidemia and associated clinical diseases. The findings in this study suggest that rs1121923 and rs258 should be considered for such panels and for the estimation of “risk” to dyslipidemia in admixed populations. Although no significant findings were observed for rs328 and rs13702, despite numerous reports on their association, the results from this study on those two variants are inconclusive. This is probably due to the fact that the cohort in the present study did not include patients with confirmed clinical diagnosis of dyslipidemia, T2DM and/or CHD. It is important to point out that one of the strengths of the study was to use “common” variants to assess the effect of LPL on plasma lipid levels in a reasonably sized cohort whilst allowing the identification of homozygosity for the minor alleles and the analysis of its frequency distribution. The other strength was the selection of four variants across the LPL gene locus that were not in linkage disequilibrium to demonstrate the effect of the various regions may play in altering LPL activity and subsequent regulation of plasma lipid levels. It is strongly recommended that the functional role of rs258 in protein binding of transcription factors be investigated.
Our study highlights the complex interaction between coding and non-coding regions, and the summative or subtractive effects different variants may have on an outcome such as plasma lipid levels. This may explain the ambiguous and sometimes conflicting results obtained by different studies when dealing with different populations.
Methods
Sample Description and Biochemical Parameters
This study was approved (Reference number: VDR/JC/256) by the Joint Committee for the Protection of Human Subjects in Research (Health Sciences Center, Kuwait University and Kuwait Institute for Medical Specializations) and conducted in accordance with the procedures set in the Helsinki guidelines. Each study subject was required to give voluntary informed consent to participate in the study and provide a blood sample. A total of 702 blood samples were collected in EDTA tubes by a certified nurse in the biochemistry laboratory at several medical clinics/hospitals in Kuwait during the period from 2012 to 2016. The subjects were randomly recruited with the following inclusion criteria: age above 18 years, willingness to provide blood for fasting lipid profile and Kuwaiti nationality (Table 10). The exclusion criteria were confirmed clinical diagnosis of T2DM, CHD, taking any medication that may alter plasma lipid levels and refusal to give informed consent. The diagnosis for T2DM was based on the HbA1c criteria of the American Diabetes Association (American Diabetes Association. Diagnosis and classification of diabetes mellitus34. Using the criteria, HbA1c ≥6.5% (48 mmol/L) was indicative of T2DM. Presence or absence of CHD was determined as previously described35. Briefly, subjects with history of myocardial infarction or angina were evaluated with the Rose questionnaire36. Subjects without history of CHD were evaluated with resting electrocardiographic (ECG) coded using the Minnesota codes37 and CHD was defined as probable ischemia (code 1.1–1.2) or possible ischemia (code 1.3, 4.1–4.4, 5.1–5.3, or 7.1).
Biochemical Parameters and Lipid Level Determination
Standard plasma lipid parameters (plasma total cholesterol (TC); Triglycerides (TG), HDL-C, low-density lipoproteins (LDL-C) and very low-density lipoprotein (VLDL) were determined on an automated chemistry analyzer (Beckman Unicel DxC 800, Beckman Corporation, Brea, CA, USA) using commercially available reagents. TC was measured using a multi-step enzymatic end point method that breaks down cholesterol into Quinonimine and water. TG was measured using a timed-endpoint method in a sequence of multi-enzyme reactions using glycerol kinase, glycerophosphate oxidase and horseradish peroxidase. In the final step of the reactions, formation of a red quinonimine dye measurement at 520 nm. HDL-C in the sample was released by a detergent which solubilizes only the HDL-C lipoprotein particles. Released HDL-C was reacted with cholesterol esterase and cholesterol oxidase to produce a color product that measured at 560 nm. LDL-C and (VLDL-C were calculated using the Friedewald formula: LDL cholesterol = Total cholesterol – HDL cholesterol – (Total triglyceride ÷ 2.2). Total triglyceride ÷ 2.2 provides a good estimate of VLDL38. The reference values used in the present study are those set by the Kuwait Ministry of Health where: TC = 3.0–5.17 mmol/L, TG = 0.40–1.7 mmol/L, HDL-C = 0.91–2.5 mmol/L and LDL-C = 1.8–3.2 mmol/L.
DNA Extraction and Genotyping
Total genomic DNA was isolated from 5 ml of whole blood using a salt extraction method39. All DNA samples were analyzed qualitatively and quantitively for suitability of use in the genotyping assay by Realtime PCR27. All suitable DNA samples were standardized to give a final concentration of 10 ng/ul. The genotyping assay for the four selected LPL variants (rs1121923; rs258; rs328 and rs13702) was achieved by the Taqman Allele Discrimination Assay with Realtime PCR (ABI 7900HT FAST REAL TIME PCR) on a 10 ng/µl DNA with commercially available primer and probe Kits (Table 11) as described by the manufacturer’s protocol (Thermo fisher Scientific, Applied Biosystems).
Genotyping, Hardy-Weinberg Equilibrium and Linkage Disequilibrium
The genotypes for each sample (Supplementary Table S1) was relatively simple to determine based on the SDS plots generated following the completion of Realtime PCR. Each sample was genotyped as homozygote for the wildtype allele (WW), heterozygous (WM) or homozygous for the minor allele (MM). Based on the generated genotypes, genotype and allele frequencies were estimated using a simple gene counting method in which the minor allele frequency (MAF) was determined for each variant in the cohort. Deviation from Hardy-Weinberg Equilibrium was estimated using the web-based calculator available at http://www.tufts.edu/.
Haploview (version 4.2.) based on the method described by Gabriel et al. (2002) was used to determine linkage disequilibrium (LD) patterns and haplotype blocks. LD was measured as r-squared value (r2) by an estimation of each pair-wise combination of SNPs. R2 values greater than 0.8 indicate a significant LD between two loci whereas a value of 0 indicates that the two loci are in linkage equilibrium. All pairs of markers (SNPs) following one of those conditions were said to be informative markers, whereas other markers falling outside that value were said to be non-informative. A haplotype block was then created if 95% of informative comparisons were in strong LD.
Genetic Association and Regression Analysis
Kruskal-Wallis ANOVA was used to analyze differences in mean between genotypes and lipid levels. The values were reported as mean ± standard error. This was followed by linear regression to assess the association of the four variants with lipid levels after controlling for age, gender and BMI. The values of the regression analysis were represented as beta (B) coefficient and 95% confidence intervals (CI). Multivariate analysis using linear regression to assess predictor factors was followed for significant lipid variables. Normality was assessed using Kolmogorov-Smirnov test. All lipid parameters were log-transformed for their association with LPL variants to ensure an approximate normal distribution. After Bonferroni correction for multiple testing of the four SNPs, the modified significance level = 0.5/4 = 0.0125. All statistical analyses reported in this study were performed using both the Statistical Package for the Social Sciences software (version 23; SPSS Inc., Chicago, IL, USA) and “SNPassoc” package from R software (R Stats Package, Version 3.3.0) where appropriate.
Ethical Approval
This study was approved (Reference number: VDR/JC/256) by the Joint Committee for the Protection of Human Subjects in Research (Health Sciences Center, Kuwait University and Kuwait Institute for Medical Specializations) in accordance to the procedures set and that are based on the Helsinki guidelines. The sample and medical data collection protocol and informed consents used were in accordance with the revised version (2000) of the 1975 Helsinki guidelines. Informed consent in this study was obtained from each participant.
Data Availability
The raw genotypic data is provided in Supplementary Table S1. Any additional may be available upon request.
References
Oberman, A. Hypertriglyceridemia and Coronary Heart Disease. J Clin Endocrinol Metab. 85, 2098–2105 (2000).
Yang, Y. et al. Genetic screening of the lipoprotein lipase gene for mutations in Chinese subjects with or without Hypertriglyceridemia. J Genet Genomics. 34, 381–91 (2007).
Deo, R. C. et al. Genetic differences between the determinants of lipid profile phenotypes in African and European Americans: the Jackson Heart Study. PloS Genet. 5, e1000342 (2009).
Shetty, P. B. et al. Variants for HDL-C, LDL-C, and Triglycerides Identified from Admixture Mapping and Fine-Mapping Analysis in African American Families. Circ Cardiovasc Genet. 8, 106–113 (2015).
Pirim, D. et al. Resequencing of LPL in African Blacks and associations with lipoprotein-lipid levels. Eur J Hum Genet. 23, 1244–53 (2015).
Al-Bustan, S. A. et al. A novel LPL intronic variant: g. 18704C > A identified by re-sequencing Kuwaiti Arab samples is associated with high-density lipoprotein, very low-density lipoprotein and triglyceride lipid levels. PloS One. 13, e0192617 (2018).
Johansen, C. T. et al. Excess of rare variants in Non-Genome-Wide association study of candidate genes in patients with hypertriglyceridemia. Nat Genet. 42, 684–7 (2010).
Evans, D., Beil, F. U. & Aberle, J. Resequencing the untranslated regions of the lipoprotein lipase (LPL) gene reveals that variants in microRNA target sequences are associated with triglyceride levels. J Clin Lipidol. 7, 610–4 (2013).
Bentley, A. R. et al. Gene-based sequencing identifies lipid – influencing variants with ethnicity-specific effects in African Americans. PloS Genet. 10, e1004190 (2014).
Deeb, S. S. & Peng, R. L. Structure of the human lipoprotein lipase gene. Biochemistry. 28, 4131–5 (1989).
Kirchgessner, T. G. et al. Organization of the human lipoprotein lipase gene and evolution of the lipase gene family. Proc Natl Acad Sci USA 86, 9647–51 (1989).
Pirim., D. et al. Lipoprotein lipase gene sequencing and plasma lipid profile. J Lipid Res. 55, 85–93 (2014).
Nickerson, D. A. et al. DNA sequence diversity in a 9. 7-kb region of the human lipoprotein lipase gene. Nat Genet. 19, 233–40 (1998).
Salinelli, S. et al. Structure-function relationship of lipoprotein lipase-mediated enhancement of very low density lipoprotein binding and catabolism by the low density lipoprotein receptor. Functional importance of a properly folded surface loop covering the catalytic center. J Biol. Chem. 271, 21906–13 (1996).
Chen, Q., Razzaghi, H., Demirci, F. Y. & Kamboh, M. I. Functional significance of lipoprotein lipase HindIII polymorphism associated with the risk of coronary artery disease. Atherosclerosis 200, 102–8 (2008).
Wang, J. et al. Resequencing genomic DNA of patients with severe hypertriglyceridemia (MIM 144650). Arterioscler Thromb Vasc Biol. 27, 2450–5 (2007).
Smith, A. J. et al. Application of statistical and functional methodologies for the investigation of genetic determinants of coronary heart disease biomarkers: lipoprotein lipase genotype and plasma triglycerides as an exemplar. Hum Mol Genet. 19, 3936–47 (2010).
Wang, K. et al. A genome-wide association study on obesity and obesity-related traits. PloS One. 6, e18939 (2011).
Richardson, K. et al. Gain-of-Function Lipoprotein Lipase Variant rs13702 Modulates Lipid Traits through Disruption of a MicroRNA-410 Seed Site. Am J Hum Genet. 92, 5–14 (2013).
Rabacchi, C. et al. Spectrum of mutations of the LPL gene identified in Italy in patients with severe Hypertriglyceridemia. Atherosclerosis. 241, 79–86 (2015).
Rodrigues, R. et al. Pathogenic classification of LPL gene variants reported to be associated with LPL deficiency. J Clin Lipidol. 10, 394–409 (2016).
Ayyappa, K. A. et al. High fat diet modifies the association of lipoprotein lipase gene polymorphism with high density lipoprotein cholesterol in an Asian Indian population. Nutr Metab(Lond). 14, 8 (2017).
Merkel, M., Eckel, R. H. & Goldberg, I. J. Lipoprotein lipase: genetics, lipid uptake, and regulation. J Lipid Res. 43, 1997–2006 (2002).
Al-Bustan, S. A. et al. Genetic association of APOB polymorphisms with variation in serum lipid profile among the Kuwait population. Lipids Health Dis. 13, 157 (2014).
Olusi, S. O., Al-Awadi, A. M. & Abraham, M. Baseline population survey data on the prevalence of risk factors for coronary artery disease among Kuwaitis aged 15 years and older. Ann Saudi Med. 23, 162–6 (2003).
Al Rashdan, I. & Al Nesef, Y. Prevalence of overweight, obesity, and metabolic syndrome among adult Kuwaitis: results from community-based national survey. Angiology 61, 42–8 (2010).
Al-Bustan, S. A. et al. Re-sequencing of the APOAI promoter region and the genetic association of the -75GA Apo polymorphism with increased cholesterol and low density lipoprotein levels among a sample of the Kuwaiti population. BMC Med Genet. 14, 90 (2013).
Razzaghi, H., Aston, C. E., Hamman, R. F. & Kamboh, M. I. Genetic screening of the lipoprotein lipase gene for mutations associated with high triglyceride/low HDL-cholesterol levels. Hum Genet. 107, 257–67 (2000).
Perneger, T. V. What’s wrong with Bonferroni adjustments. BMJ (Clinical research ed.) 316, 1236–8 (1998).
Mead, J. R., Irvine, S. A. & Ramji, D. P. Lipoprotein lipase: structure, function, regulation, and role in disease. J Mol Med (Berl). 80, 753–69 (2002).
Wood, A. C. et al. Lipoprotein Lipase S447X variant associated with VLDL, LDL and HDL diameter clustering in the MetS. Lipids Health Dis. 10, 143 (2011).
Cho, Y. S. et al. Association of lipoprotein lipase (LPL) single nucleotide polymorphisms with type 2 diabetes mellitus. Exp Mol Med. 40, 523–32 (2008).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science. 296, 2225–9 (2002).
American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care. 37, S81–90 (2014).
Mojiminiyi, O. A., Al Mulla, F. & Abdella, N. A. Which obesity index best explains the link between adipokines, coronary heart disease risk and metabolic abnormalities in type 2 diabetes mellitus? Med Princ Pract. 18, 123–9 (2009).
Rose, G. A. The diagnosis of ischaemic heart pain and intermittent claudication in field surveys. Bull World Health Organ. 27, 645–58 (1962).
Blackburn, H. Electrocardiographic classification for population comparisons. The Minnesota code. J Electrocardiol. 2, 5–9 (1969).
Friedewald, W. T., Levy, R. I. & Fredrickson, D. S. Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin Chem. 18, 499–502 (1972).
Miller, S. A., Dykes, D. D. & Polesky, H. F. A simple salting-out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Acknowledgements
The authors would like to extend their appreciation and gratitude to Kuwait University Research Sector and to the Kuwait Ministry of Health for facilitating the ethical approval and sample collection. This work was achieved with the kind assistance of the technical staff in the medical clinics by collecting the samples and analyzing the lipid profiles and at the Biotechnology Center (Dr. Betsy Sheena Cherian, Mrs Sheela Thankakkon and Mr. Philip Koshy) for the use of the general Realtime PCR and sequencing facility (Project GS 01/02). The authors would also like to acknowledge the assistance of Ms. Arwa Ayman and Mrs Sahar A. Barhoush, graduate students in the Department of Biological Sciences, for their assistance with the linkage disequilibrium analysis and haplogroups scoring. Finally, the authors extend their gratitude to all the participants in this study for providing consent and relevant information. This research was funded by Kuwait University Research Sector, Projects SL04/11.
Author information
Authors and Affiliations
Contributions
S.A. prepared the project proposal and study design, supervised the molecular genetic studies and phenotypic documentation, performed most of the data analysis and drafted the manuscript. A.A. performed the statistical analysis on the cohort and allele frequencies and participated in writing the manuscript. M.A. assisted with final data analysis and contributed in drafting, editing and finalizing the manuscript. B.A. performed all the sequence and real-rime experiments as well as assisted in preparation of some figures and tables. O.M. Provided advise and consults on the study subjects, sample collection and revised the manuscript. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41598_2019_42021_MOESM1_ESM.pdf
Genetic association of LPL rs1121923 and rs258 with plasma TG and VLDL levels Al-Bustan S; Al-Serri A, Alnaqeeb M; Annice B; Mojiminiyi O - Supplementary Table S1: LPL variants genotypes in the Kuwai
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Al-Bustan, S.A., Al-Serri, A., Alnaqeeb, M.A. et al. Genetic association of LPL rs1121923 and rs258 with plasma TG and VLDL levels. Sci Rep 9, 5572 (2019). https://doi.org/10.1038/s41598-019-42021-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42021-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.