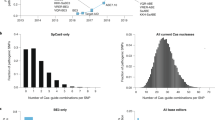
Abstract
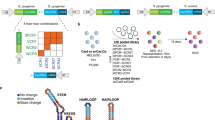
The continued improvement of combinatorial CRISPR screening platforms necessitates the development of new computational pipelines for scoring combinatorial screening data. Unlike for single-guide RNA (sgRNA) pooled screening platforms, combinatorial scoring for multiplexed systems is confounded by guide design parameters such as the number of gRNAs per construct, the position of gRNAs along constructs, and additional features that may impact gRNA expression, processing or capture. In this protocol we describe Orthrus, an R package for processing, scoring and analyzing combinatorial CRISPR screening data that addresses these challenges. This protocol walks through the application of Orthrus to previously published combinatorial screening data from the CHyMErA experimental system, a platform we recently developed that pairs Cas9 with Cas12a gRNAs and enables programmed targeting of multiple genomic sites. We demonstrate Orthrus’ features for screen quality assessment and two distinct scoring modes for dual guide RNAs (dgRNAs) that target the same gene twice or dgRNAs that target two different genes. Running Orthrus requires basic R programming experience, ~5–10 min of computational time and 15–60 min total.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Data availability
The example dataset is downloadable with the Orthrus package at https://github.com/csbio/Orthrus. The expected output from all procedures is provided under a CC-BY 4.0 license at https://zenodo.org/record/4527616.
Code availability
The Orthrus package is available at https://github.com/csbio/Orthrus, and the version of the code run in the protocol is available at https://zenodo.org/record/4827171 (ref. 20). All code presented from all procedures is also available in separate scripts along with their expected output, and are provided under a CC-BY 4.0 license in the Zenodo repository at https://zenodo.org/record/4527616. Code contained in this repository generated Figs. 4–10. The code in this protocol has been peer reviewed.
References
VanderSluis, B. et al. Integrating genetic and protein–protein interaction networks maps a functional wiring diagram of a cell. Curr. Opin. Microbiol. 45, 170–179 (2018).
Costanzo, M. et al. A global genetic interaction network maps a wiring diagram of cellular function. Science 353, aaf1420–aaf1420 (2016).
Simpkins, S. W. et al. Predicting bioprocess targets of chemical compounds through integration of chemical-genetic and genetic interactions. PLoS Comput. Biol. 14, e1006532–e1006532 (2018).
Piotrowski, J. S. et al. Functional annotation of chemical libraries across diverse biological processes. Nat. Chem. Biol. 13, 982–993 (2017).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Ruiz, S. et al. A genome-wide CRISPR screen identifies CDC25A as a determinant of sensitivity to ATR inhibitors. Mol. Cell 62, 307–313 (2016).
Wong, A. S. L. et al. Multiplexed barcoded CRISPR-Cas9 screening enabled by CombiGEM. Proc. Natl Acad. Sci. 113, 2544–2549 (2016).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
Aregger, M. et al. Systematic mapping of genetic interactions for de novo fatty acid synthesis identifies C12orf49 as a regulator of lipid metabolism. Nat. Metab 2, 499–513 (2020).
Gonatopoulos-Pournatzis, T. et al. Genetic interaction mapping and exon-resolution functional genomics with a hybrid Cas9–Cas12a platform. Nat. Biotechnol. 38, 638–648 (2020).
Najm, F. J. et al. Orthologous CRISPR-Cas9 enzymes for combinatorial genetic screens. Nat. Biotechnol. 36, 179–189 (2018).
Shen, J. P. et al. Combinatorial CRISPR-Cas9 screens for de novo mapping of genetic interactions. Nat. Methods 14, 573–576 (2017).
Han, K. et al. Synergistic drug combinations for cancer identified in a CRISPR screen for pairwise genetic interactions. Nat. Biotechnol. 35, 463–474 (2017).
Gier, R. A. et al. High-performance CRISPR-Cas12a genome editing for combinatorial genetic screening. Nat. Commun. 11, 3455 (2020).
Horlbeck, M. A. et al. Mapping the genetic landscape of human cells. Cell 174, 953–967.e22 (2018).
Dede, M., McLaughlin, M., Kim, E. & Hart, T. Multiplex enCas12a screens detect functional buffering among paralogs otherwise masked in monogenic Cas9 knockout screens. Genome Biol 21, 262 (2020).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Ward., H. csbio/orthrus: Protocol Release (v0.4.3.3). Zenodo. https://doi.org/10.5281/zenodo.4827171 (2021).
Zamanighomi, M. et al. GEMINI: a variational Bayesian approach to identify genetic interactions from combinatorial CRISPR screens. Genome Biol. 20, 137 (2019).
Aregger, M. et al. Application of CHyMErA Cas9-Cas12a combinatorial genome-editing platform for genetic interaction mapping and gene fragment deletion screening. Nat. Protoc. https://doi.org/10.1038/s41596-021-00595-1 (2021).
Imkeller, K., Ambrosi, G., Boutros, M. & Huber, W. gscreend: modelling asymmetric count ratios in CRISPR screens to decrease experiment size and improve phenotype detection. Genome Biol 21, 53 (2020).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733 (2014).
Boucher, B. & Jenna, S. Genetic interaction networks: better understand to better predict. Front. Genet. 4, 290 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Yang, Y. H. et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 30, e15 (2002).
Acknowledgements
We are grateful to K. Lin for his help testing the Orthrus package. H.N.W, M.B. and C.L.M. were partially supported by grants from the National Science Foundation (1818293) and the National Institutes of Health (R01HG005084, R01HG005853). T.G.-P. was supported by the NIH Earl Stadtman Investigator Program and the NIH Distinguished Scholars Program. M.A. was supported by a Swiss National Science Foundation fellowship (P300PA_164667). M.B. was supported by the German Research Foundation DFG (Bi 2086/1-1). B.J.B was supported by a Canadian Institutes for Health Research Foundation grant (FDN-148434) and by an Ontario Institute of Regenerative Medicine grant. J.M. was supported by grants from the Canadian Institutes for Health Research (MOP-142375) and by Genome Canada (OGI-157).
Author information
Authors and Affiliations
Contributions
H.N.W. wrote the software and performed all analyses. H.N.W. drafted the protocol, and C.L.M., M.A., T.G.-P., M.B., K.R.B. and T.K.O. provided revisions. H.N.W. and M.B. developed the scoring procedure implemented in Orthrus based on conceptual contributions from M.B. T.K.O. provided feedback to improve the software. M.A., T.G.-P., M.B., K.R.B., J.M., B.J.B. and C.L.M. developed the CHyMErA experimental platform. M.A., T.G.-P. and K.R.B. performed experiments to generate the data analyzed in Procedure 2. K.R.B. contributed data, code and text to Procedure 1. J.M., B.J.B. and C.L.M. acquired funding to support this work and provided supervision throughout the project.
Corresponding author
Ethics declarations
Competing interests
A patent application (no. GB 1907733.8) describing the development and applications of CHyMErA, to the University of Toronto and T.G.-P., M.A., K.R.B., J.M. and B.J.B., is pending. J.M. previously performed sponsored research for Repare Therapeutics.
Additional information
Peer review information Nature Protocols thanks Max Horlbeck and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Gonatopoulos–Pournatzis, T. et al. Nat. Biotechnol. 38, 638–648 (2020): https://doi.org/10.1038/s41587-020-0437-z
Aregger, M. et al. Nat. Protoc. (2021): https://doi.org/10.1038/s41596-021-00595-1
Dede, M. et al. Genome Biol. 21, 262 (2020): https://doi.org/10.1186/s13059-020-02173-2
Supplementary information
Supplementary Video 1
Tutorial on how to set up input files to run the Orthrus scoring pipeline
Rights and permissions
About this article
Cite this article
Ward, H.N., Aregger, M., Gonatopoulos-Pournatzis, T. et al. Analysis of combinatorial CRISPR screens with the Orthrus scoring pipeline. Nat Protoc 16, 4766–4798 (2021). https://doi.org/10.1038/s41596-021-00596-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00596-0
This article is cited by
-
A scalable platform for efficient CRISPR-Cas9 chemical-genetic screens of DNA damage-inducing compounds
Scientific Reports (2024)
-
High-content CRISPR screening
Nature Reviews Methods Primers (2022)
-
Application of CHyMErA Cas9-Cas12a combinatorial genome-editing platform for genetic interaction mapping and gene fragment deletion screening
Nature Protocols (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.