Abstract
Recently, there has been an explosion of scientific literature describing the use of colorimetry for monitoring the progression or the endpoint result of colorimetric reactions. The availability of inexpensive imaging technology (e.g., scanners, Raspberry Pi, smartphones and other sub-$50 digital cameras) has lowered the barrier to accessing cost-efficient, objective detection methodologies. However, to exploit these imaging devices as low-cost colorimetric detectors, it is paramount that they interface with flexible software that is capable of image segmentation and probing a variety of color spaces (RGB, HSB, Y’UV, L*a*b*, etc.). Development of tailor-made software (e.g., smartphone applications) for advanced image analysis requires complex, custom-written processing algorithms, advanced computer programming knowledge and/or expertise in physics, mathematics, pattern recognition and computer vision and learning. Freeware programs, such as ImageJ, offer an alternative, affordable path to robust image analysis. Here we describe a protocol that uses the ImageJ program to process images of colorimetric experiments. In practice, this protocol consists of three distinct workflow options. This protocol is accessible to uninitiated users with little experience in image processing or color science and does not require fluorescence signals, expensive imaging equipment or custom-written algorithms. We anticipate that total analysis time per region of interest is ~6 min for new users and <3 min for experienced users, although initial color threshold determination might take longer.
Similar content being viewed by others
Introduction
As described by the Young–Helmholtz theory, human vision is a tristimulus system in which three different types of cone cells function as spectrally sensitive receptors (~6 million per eye). Photopigments within the three classes of cone cell are stimulated by different wavelength ranges that correspond to one of three colors: red, green or blue (RGB)1,2. In humans with normal trichromatic vision, the brain combines the three independent wavelength inputs to reconstruct the observed color, which falls within a vast kaleidoscope of roughly 2 million distinguishable colors3. Many on-site detection methods exploit this visual dynamic range by relying on the human eye as a portable sensor, including commonplace water quality tests, swimming pool pH sensors, law enforcement roadside drug tests and military-issue chemical and explosive tests. These instrument-free, colorimetric kits boast simplicity, ease of use and cost-effectiveness—all of which are advantageous over conventional laboratory instrumentation. Interpretation of these colorimetric kits is not bound to absorbance measurement via conventional laboratory instrumentation. Specifically, these kits present in a variety of forms, including paper, membranes and beads, that bypass the need for specialized instruments (e.g., spectrometer), which are classically associated with colorimetry analysis via absorption, reflection and wavelength measurement. However, these field-deployable approaches also suffer from critical operational limitations and are often criticized for susceptibility to incorrect or variable user interpretation, which can lead to inaccurate results. With human vision as the detection method, these deficiencies are largely due to on-site environmental conditions (e.g., poor ambient lighting or the flashing lights of emergency vehicles) and natural, person-to-person perceptual differences4. To achieve objective analysis, the subjective nature of user-dependent visual interpretation must be mitigated.
In other words, to ameliorate the inherent uncertainties associated with human vision, an empirical model of the visible color space is required. A color space (also, model or system) is an abstract mathematical representation (coordinate system or subspace) that defines the range of perceivable colors in human vision (Fig. 1a,b). In 1931, the International Commission on Illumination (CIE) developed a mathematical model of this trichromic system (i.e., the CIE RGB color space)5,6. In 1976, the CIE elaborated on this idea by translating the CIE RGB to a new model, one that more accurately described the numerical relationship between wavelengths and human physiological response to observed color or color change. The resultant CIE L*a*b* (see Box 1—‘Color spaces’) model is now widely considered to be the gold-standard model of human color vision7. Essentially, this three-dimensional (3D) model encompasses all colors of human perception and, by design, more accurately depicts the increased spectral sensitivity of human cone cells to green wavelengths8. However, in practice, the most pervasive color system is the standard RGB (sRGB) model, which was developed in 1996 through the collaborative efforts of Hewlett-Packard and Microsoft Corporation in direct response to the growing need for standardization of color in computers and illuminated screens (https://www.w3.org/Graphics/Color/sRGB.html)9. A simplified visualization of this model is depicted in Fig. 1a. RGB models are additive tristimulus systems where each unique color in the model is represented as a tuple. More specifically, red, green and blue are the three components of the RGB color space. In many common color models, each component of the color space is assigned an 8-bit value between 0 and 255 (8-bit color component = 28 = 256). Collectively, this provides reproducible numerical representations of more than 16 million colors (24-bit RGB = 28 × 28 × 28 = 16.7 million). For example, in RGB color space, black, purple and magenta are represented by the following tuples: black = {0, 0, 0}, purple = {128, 0, 128} and magenta = {255, 0, 255}. Other color spaces (Y’UV and L*a*b*), how they relate to the RGB model and how they are used in this protocol are discussed in Box 1—‘Color Spaces’.
a, Simplified RGB and b, HSB color spaces visualized as a cube and cone, respectively. c, In HSB color space, the hue component is a continuous band of color that encompasses the circular base of a 3D cone or cylinder (simplified color band depicted here). d, By convention, the 8-bit system used in ImageJ linearizes and splits this continuous color wheel, assigning hue values ranging from 0 to 255 with pure, primary green and blue falling at ~85 and 170, respectively (G = 0.33 and B = 0.66 A.U.). However, primary red falls squarely at a hue value of 0. Thus, colors spanning the red range can exhibit ascribed hue values from ~0 to 31 and 225 to 255, effectively splitting the red spectrum. e, Rotating the color space such that the assigned location for zero falls elsewhere in the spectrum addresses the issue of split peaks when the color response spans the red range. Arrow indicates direction of rotation.
ImageJ, a freely available Java-based image processing program, was developed at the National Institutes of Health (NIH) (https://imagej.nih.gov/ij/docs/intro.html) in collaboration with the Laboratory for Optical and Computational Instrumentation (https://imagej.net/LOCI)10,11,12. Since the initial implementation of NIH Image in 1987 and the public release of ImageJ in 1997, this processing system has become a potent and pervasive tool for the scientific community. In this protocol, we create a necessary bridge between ImageJ and the biochemical/bioanalytical research community, making this technology more accessible for objective analysis of colorimetric responses across these analytical fields. Unlike many other published protocols, this procedure is not merely a simple cookbook-type, push-button approach13,14. Rather, this ‘human-in-the-loop’ approach guides users through a relatively complex process, while allowing the analyst to adapt and modify these procedures to fit a wider range of unique image sets, experimental designs and assay types. We hope that this protocol provides the impetus for further refinement, exploration, development and sharing of ImageJ-based methods across the biochemical/bioanalytical arena. Ultimately, these methods serve as a convenient, easily implemented precursor for subsequent development of more sophisticated algorithms and applications to automate image processing of bioanalytical colorimetric responses.
Development of the method and comparison with other methods
Over the last decade, necessity has dictated a heavy reliance on ImageJ as an analytical tool in our research. Consequently, we developed of a robust ‘toolkit’ of methods for enhanced procurement of empirical data from colorimetric reactions under a variety of challenging analytical conditions15,16,17,18,19. Remarkably, finding published, detailed and easy-to-implement protocols for ImageJ operation is, at best, difficult. Many of the published protocols rely on fluorescence signals and image stacks that are obtained via expensive imaging hardware (i.e., confocal microscopes and electron microscopes) coupled with specialized ImageJ plug-ins and pipelines that are not easily translated or applied to other fields of study (plug-ins are add-on software components that augment the existing ImageJ program with specific, customized features)20,21,22,23. Furthermore, many of these biomedical and biological approaches to image analysis take an individual color channel (a single slice of a color space containing a single fluorescence color signal) and convert it into a binary (black-and-white) image, disposing of a large portion of data associated with the color response in the quest for effective image segmentation.
In analytical chemistry, endpoint colorimetric response assays are used for a wide array of field-forward applications owing to simple visual readouts24. More often than not, these color-based assays are simple, lending themselves to data acquisition via inexpensive imaging technology. The measurable color changes produced by these assays contain valuable information that is used to empirically detect and quantify the presence of an analyte. Here, we delineate how to use native ImageJ tools to achieve objective, colorimetric detection and subsequently robust data analysis by exploiting a range of image manipulation techniques, including region of interest (ROI) selection and cropping; color thresholding and image masking via independent manipulation of luminance and chrominance channels in Y’UV and L*a*b* color spaces; and digital tinting for improved contrast (Figs. 2c and 3b,c). The strength of this approach is the simplicity, providing users who have little expertise or experience in image processing or computer programing with the ability to access and tailor objective, colorimetric data.
RGB color images containing colorimetry responses are opened with ImageJ. Workflow A: The Crop-and-Go approach does not require tinting or color threshold application. This approach is ideal for isolating ROIs that exhibit consistent, regular shape and size as well as uniform, homogeneous color response. Workflow B: The Crop-Threshold-and-Go approach does not require digital tinting (color balancing) before color threshold application. This workflow is preferable when the color response exhibits good contrast with the background but lacks a consistent, regular shape. Workflow C: The Crop-Tint-Threshold-and-Go approach uses color balancing to increase contrast between the background and foreground of the image, thereby enhancing ROI segmentation via subsequent color threshold application. This workflow is best suited for images depicting either colorless-to-colored or highly unsaturated colorimetric responses on a near-white/white backgrounds (i.e., when Workflows A and B do not provide adequate isolation of the color response from the background). Finally, quantitative results are exported to common analysis programs.
This protocol fully describes and clarifies previously reported Crop-and-Go15,16,17 and color profiling19 approaches. Moreover, this protocol uses native ImageJ tools to supplant the custom-written Mathematica algorithms reported in Krauss et al. and Thompson et al.17,18, which require considerable skill to write and use. That is, rather than using complex proprietary algorithms, we achieve similar color thresholding, image segmentation and color analysis via free, open-source software (Figs. 2 and 3).
Overview of the procedure
The method described in the ‘Procedure’ section offers three workflows for segmenting colorimetric responses (Fig. 3). Each workflow (A, B and C) provides the analyst with a procedural workflow that is adaptable, requiring no previous experience or knowledge of ImageJ or color science. In practice, this protocol consists of three distinct workflows that represent overlapping layers of ROI refinement and increasing segmentation complexity: Crop-and-Go, Crop-Threshold-and-Go and Crop-Tint-Threshold-and-Go (Fig. 3). The Crop-and-Go approach (Workflow A) is the most straightforward route to analysis. This workflow includes simple image cropping and image rotation (Steps 3–13), color space conversion (Step 27) and analysis (Steps 28 and 29). The Crop-Threshold-and-Go and Crop-Tint-Threshold-and-Go approaches (Workflows B and C, respectively) include optional steps for improved image segmentation and masking. Workflow B incorporates steps for color thresholding (Steps 21–26), whereas Workflow C incorporates steps for color tinting and thresholding (Steps 14–20 followed by Steps 21–26). A more detailed description of the three workflows is provided in the ‘Experimental design’ section. From a practical perspective, adaptation and implementation of this protocol is simple, requiring nothing more than image files of colorimetric experiments and a computer with the ImageJ program. All image files featured in this publication were captured with a Huawei P9 smartphone (Huawei Technologies) or an Epson Perfection V100 desktop scanner (Seiko Epson).
Advantages of the method
Application of the approach described here is simple: expertise in color science theory, software engineering, programming, app development, mathematics or physics is not required to employ this protocol. Instead, we exploit native ImageJ tools. In so doing, we leverage key underlying ImageJ design and programing principles (i.e., limiting complexity through use of simple user interfaces that a typical bench scientist or student can easily implement and understand). This protocol does not rely on fluorescent labels or signals, prepackaged image stacks or expensive imaging systems. Instead, this protocol starts with simple, raw RGB images captured using affordable imaging technology (i.e., smartphones and desktop scanners). Unlike many other methods, this workflow does not merely use color signals as a means for image segmentation; rather, it preserves raw color response data for downstream analysis. Furthermore, raw data from measured and quantified ROIs are easily exported for statistical analysis and visualization. We think that this protocol outlines invaluable methods for masking irregular shapes, analyzing inhomogeneous color changes and/or acquiring data when lighting conditions and image resolution are less than ideal.
Limitations of the method
The Crop-and-Go method described in Workflow A (Fig. 3) is simple and objective, quickly generating data for export and analysis (Steps 3–13). However, the success of this simple approach hinges entirely on uniformity of shape and homogeneity of the color response; therefore, this approach is unsuitable when the color response is inconsistent across the ROI or when shape and/or distribution of the response is complex and irregular.
Additionally, some caution is warranted when converting images into component parts of the split-spectrum HSB color space (Fig. 1b). Explicitly, color responses falling within the red region of the visible spectrum (between pink and orange) generate peaks at both the low and high extremes of the hue scale, thus confounding downstream statistical analysis and visualization (Fig. 1c,d). This is a unique and particularly troublesome feature of the otherwise robust quantitative analytical hue parameter. To elaborate, conversion from RGB color to HSB color space is essentially recasting a cubic Cartesian system into a 3D conical or cylindrical coordinate system16,26,27,28. The linear saturation and brightness components reflect the richness and intensity of the color, whereas hue is a circular variable that describes a unique color value across the visible spectrum (0–255 in ImageJ). Stated another way, in the HSB color space, hue is a continuous, circular band of visible color that spans the circumference of the cylindrical or conical base of the 3D HSB color space (Fig. 1b,c)27. By convention, the 8-bit hue system used in ImageJ assigns hue values of ~85 and 170 to pure green and blue, respectively (G = 0.33 and B = 0.66 arbitrary units (A.U.)). However, pure red falls squarely at a hue value of 0 (Fig. 1c). Thus, colors spanning the red range can exhibit ascribed hue values of ~0–31 and 225–255, effectively linearizing and splitting a continuous spectrum at 0 (Fig. 1d). In some colorimetric responses with a strong red contribution, pixels with hue values on both sides of 0 can be present within the same image. Averaging these values would result in a calculated hue value in the center of the spectrum (green to blue), resulting in a mean hue value that is not at all representative of the true color present. To ameliorate this issue, the split peaks can be eliminated by rotating the color space such that the assigned location for 0 falls outside the red range (Fig. 1e). The 3D Color Inspector plug-in (bundled with the Fiji distribution of ImageJ) can be used to both perform and visualize color rotations and shifts. After rotation, modified images can be saved for further evaluation using the protocol described here29. However, in our experience, this plugin resizes and compresses the saved image file, limiting the usefulness of the tool for sensitive analyses29. To date, we are unaware of any native ImageJ tools, Fiji plugins or published ImageJ macros that efficiently automate the rotation of hue data, and manual manipulation of the raw data is cumbersome and time-consuming. Therefore, at present, tinting and other color manipulation techniques (a priori or a posteriori) are more practical options16.
Additional challenges arise when dealing with either colorless-to-colored or highly unsaturated color responses, especially when the background is white, as is standard in paper microfluidics and paper-based lateral flow assays (Fig. 2b,c). Briefly, because white is not represented in the hue spectrum of HSB color space (Fig. 1b,c), ImageJ assigns a hue value that best represents the overall cast and tint of the background, which is often ascribed a pale yellow or blue hue value (e.g., uncoated incandescent lights produce light that is perceived as yellowish, and many fluorescent lights appear blue-greenish). This is problematic when the expected color response falls within the same hue range as the background (e.g., a colorless ammonium titanyl oxalate solution produces a yellow color change in the presence of hydrogen peroxide)16. Although digital tinting and color thresholding (Steps 14–20 and Steps 21–26, respectively) can be used to address these shortcomings, these post-processing methods bring their own inherent biases and imperfections. Specifically, because tinting parameters and color thresholds are established manually, some inherent subjective bias is expected during this procedure. However, the tradeoff is that, once the thresholds are established, they are consistently and objectively applied to all images in the set, minimizing variability between users or between experiments. Absent a machine learning approach, there are no existing ImageJ-based methods for automating the tinting and color threshold steps.
Finally, it has been our experience that the color thresholding approach (Steps 21–26) and the hue profiling method described in ref. 19 (Step 29, Option B) fail to provide adequate image segmentation under several conditions. Briefly, both methods rely on a uniformly colored background with sufficient contrast to the expected color responses. Therefore, for the same reasons discussed above, colorless-to-colored responses and very unsaturated signals can be quite difficult to segment (Fig. 2c).
Applications of the method
In large part, this protocol is based on analytical pipelines mentioned but not fully described in refs. 15,16,17,18,19,30. Similar ImageJ-based image analysis techniques have been used to evaluate assays for 19-nortestosterone31, Salmonella32, Yersinia pestis33, Escherichia coli and Enterococcus sp.34 and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) (Coronavirus Disease 2019)35,36. To date, we have also used this protocol to evaluate colorimetric responses for the detection of a variety of analytical targets, namely narcotics4, explosives15,16 and organophosphates (nerve agent and pesticide analogs—not yet published), as well as forensically and clinically relevant targets (e.g. semen19 and bilirubin18). We are also currently using this image analysis approach for colorimetric detection of several Tier 1 viral and bacterial agents (Centers for Disease Control and Prevention Select Agents and Toxins List). Specifically, we are developing a loop-mediated isothermal amplification method with colorimetric detection of SARS-CoV-2 N1 and immunochromatographic assays for Ebola virus-like particles and Y. pestis F1 antigen (https://www.selectagents.gov/SelectAgentsandToxinsList.html). The images for these colorimetric assessments were obtained from a variety of analytical platforms and formats, including liquid volumes within microcentrifuge tubes or microdevice chambers, along with color responses on solid, cellulosic and silica-based substrates. In view of this, we present a simple-to-use, robust analytical tool that will provide the user with the bandwidth necessary to segment and analyze a wide range of color responses.
Experimental design
Image capture and file type
Performing the colorimetric experiments and capturing raw image files of the results are the first, and perhaps most critical, steps in all three processing workflows described in this protocol (Steps 1 and 2). We routinely use smartphones and desktop scanners to acquire digital images. However, before acquiring any pictures, thoughtful deliberation is warranted when considering file type and storage options. Many imaging platforms default to lossy compression formats, such as JPEG. Lossless file formats, such as PNG, GIF and TIFF, are recommended for sensitive quantitative analyses. Scanning images at higher resolutions (dpi) improves overall image quality. But higher-resolution image capture is more time-consuming and generates appreciably larger file sizes. In our experience, scanning images between 600 and 1200 dpi provides a reasonable balance among acquisition time, file size and resolution. Natively, the ImageJ program opens a limited number of common lossy and lossless file formats, including JPEG, TIFF, PNG, GIF and BMP. Other formats (*.jp2, *.ome.tif, *.mov, *.avi, etc) are readable only via plugins that facilitate the importation of non-native file formats (https://imagej.nih.gov/ij/features.html).
Most commercial imaging devices (e.g., scanners and digital cameras) incorporate Bayer-type color filter arrays (CFAs). Although an in-depth discussion of these CFAs and in situ color calibration are well beyond the scope of this protocol, the potential variability introduced by manufacturer-to-manufacturer differences and the associated color mapping software are noteworthy. CFA-induced variability stands to substantially affect imaging and subsequent analysis. As such, we recommend using the same device for all image capture within a given set of experiments to ensure homogeneous application of image segmentation within the dataset.
ImageJ interface
Upon launching the ImageJ program, you will notice a single window consisting entirely of a menu bar, tool bar and status bar (https://imagej.nih.gov/ij/docs/guide/146-Part-IV.html#par:User-Interface1) (Fig. 4a). The File, Edit, Image and Analyze menus provide access to all submenu and command paths used in this protocol (https://imagej.nih.gov/ij/docs/guide/146-Part-V.html). The protocol relies heavily on the straight-line, rectangle and oval selection tools, all of which are accessible via icons on the tool bar (Fig. 4b). Right-clicking or double-clicking on a selection tool icon provides access to alternative selection modes and brushes. Other selection tools are available within the ImageJ workspace (i.e., the polygon, freehand and wand tracing tools provide additional selection options but are not discussed or used here). A recent protocol by Boudaoud et al. offers several examples of the polygon and straight-line tools in action14. A detailed overview of all selection tools is available at https://imagej.nih.gov/ij/docs/tools.html.
This figure features an image of 2.5-mm laser cutouts of plastic-backed, silica thin-layer chromatography (TLC) plates. The circular, TLC plate cuttings were treated with malachite green and placed in individual microfluidic chambers for colorimetric, organophosphate detection and differentiation. Subgroups of organophosphates, namely chemical warfare agents and pesticides, are highly deleterious to human health either through terroristic use or accidental consumption. Shown here, we demonstrate how to select, refine and crop an ROI using the ImageJ GUI. a, The ImageJ GUI consists entirely of the menu, tool and status bars. Selection tool icons are accessible directly from the tool bar (red box). b, Dragging-and-dropping an image file directly onto the status bar opens a new window containing the active image. Similarly, images can be opened via the File > Open… submenu. Clicking the oval selection tool icon allows the user to define a circular/ovoid ROI by clicking-and-dragging on the active image (yellow outline). Clicking-and-dragging the handles on the selection permits course refinement of the ROI. c, The Edit > Selection > Specify submenu permits further refinement and modification of the current ROI selection. d. After refining the ROI selection, the active image is cropped via keyboard shortcuts (Ctrl + Shift + X) or the Image > Crop submenu. Notice that ImageJ crops the image to a rectangle that completely bounds the active selection. e, The area outside the ROI selection is removed via the Edit > Clear Outside submenu.
Workflow A: Crop-and-Go
Regardless of ROI shape and complexity, almost all of our analyses begin with simple cropping and image rotation. Figures 4 and 5 provide a synopsis of the ImageJ graphical user interface (GUI), area selection tools, cropping and image rotation. In many cases, it is possible to proceed directly from these cropping steps (Steps 3–13) to image-type conversion and data analysis (Steps 27–29), effectively precluding the need for tinting and color threshold application (Steps 14–20 and Steps 21–26). In our experience, this workflow is ideal when the ROI is clearly defined with uniform shape (e.g,. rectangular or ovoid) and size that do not change between images or trials within a set. In fact, poor segmentation results with this Crop-and-Go method-motivated exploration and incorporation of the tinting and color thresholding steps that follow.
This figure features an image of six prototype lateral flow immunoassay strips for the detection of a biological agent that the Centers for Disease Control and Prevention has listed as a Tier 1 Select agent (i.e., great risk for potential misuse and significant probability for mass casualties). Shown here, we demonstrate how to rotate, align and crop ROIs from the image before downstream analysis with the Plot Profile tool. a, For easier visualization and alignment of the image, temporary gridlines are applied to the active image via the Image > Transform > Rotate… submenu (red arrows). b, The Rotate dialogue window allows the user to specify, preview and apply rotational image transformations (integer values of ± 90° only). In this figure, the active image is rotated 87° (red arrow). c, As seen in the previous figure, clicking the rectangle selection tool icon allows the user to define a square/rectangular ROI by clicking-and-dragging on the active image (not shown). Again, the Edit > Selection > Specify submenu is used for ROI refinement before cropping.
Workflow B: Crop-Threshold-and-Go
Suboptimal illumination, image capture and experimental execution produce artifacts, anomalies (e.g., shadows, glares and bubbles) and inhomogeneous color responses that confound and muddle downstream analysis when using the simple Crop-and-Go method. The ImageJ color thresholding module allows the user to define and mask irregular, asymmetrical shapes associated with the color response (Steps 21–26) by clearly demarcating ROI boundaries and more effectively removing troublesome background color(s) and artifacts (Fig. 6). Similarly to the Crop-and-Go workflow, this color thresholding approach begins with simple ROI selection and cropping of the original, true-color RGB image (Steps 1–13).
This figure features an image of a five-layer, polymeric microfluidic device. a, Twenty of the 25 chambers contain dye volumes (100 mM tartrazine) ranging from 2 to 10 µl in 2-µl increments (n = 4 each). Shown here, we demonstrate image segmentation via color threshold application for the acquisition of raw ROI area data. As seen in Fig. 5, the Rectangle selection tool and the Image > Transform > Rotate… submenu are used for ROI selection, refinement, rotation and cropping. b, Color thresholds are applied via the Image > Adjust> Color Threshold… submenu. Specifically, the Y’UV threshold windows for this analysis are Y’ = 50–255, U = 0–100 and V= 130–255. Panel c shows representative cropped and thresholded images across the five dye volumes used in this experiment. The red threshold color shows the proposed ROI selection based on the current threshold settings (2-µl example). After applying the color thresholds (Select button), the ROI selection is bounded by a yellow outline. Pixel area measurements provide the necessary ROI data to plot a calibration curve for future microfluidic recovery studies (Image > Type > HSB Stack and Analyze > Measure).
At a fundamental level, color thresholding is the independent manipulation and application of filters to the component channels within a color space, effectively defining a range of values that further isolate the pixels of interest. Explicitly, each of the previously discussed color spaces (RGB, HSB, Y’UV and L*a*b*; Box 1) are trichromic, characterizing each pixel within an image as a combination of three constituent slices or channels (e.g., the three component color channels of the RGB color model are red, green and blue) (Fig. 6b). User-specified color thresholds individually partition pixels into non-overlapping zones that clearly delimit the boundaries between the background and foreground of an image, refining the original ROI (Fig. 6c)27,37.
From the menu bar, the ImageJ color threshold command is nested within the Image > Adjust submenu. Upon opening the color threshold window, the user will be presented with three histograms corresponding to the pixel distributions within each of the component channels of the color space (Fig. 6b). Each histogram has interactive sliders for independent manipulation and thresholding. By default, ImageJ presents the user with HSB histograms that, as discussed, are a good starting point. However, we have found Y’UV and L*a*b* color spaces to be more visually intuitive and more user-friendly for color threshold determination and application (Fig. 6b,c). Numerical values (0–255) on the x axis of each histogram ease reproducibility between iterations. By default, pixels within the bounds of the lower threshold (LT) and upper threshold (UT) are shown in red (i.e., LT ≤ values ≤ UT) (Fig. 6c). The user will need to define these LT and UT values for each channel within the color space. The ImageJ-generated histograms offer a path to more objective segmentation, ensuring that setting threshold values is not completely arbitrary or random. Using Fig. 6 as an example, by comparing histograms of empty microfluidic chambers (not shown) with those of dye-filled chambers, peaks corresponding to the background and to the dye solution were identified, allowing us to set threshold values for each of the Y’UV color channels (Y’ = 50–255, U = 0–100 and V = 130–255). A closer look at the histograms reveals that our user-defined threshold ranges include dark pixels in the Y’ channel, yellow pixels in the U channel and red pixels in the V channel, effectively removing the background and isolating the yellow fluid within the microfluidic chamber.
If application of color thresholds (Step 21–26) generates adequate segmentation of the images, one can forgo digital tinting (Step 14–20) and proceed directly to image-type conversion and data analysis (Steps 27–29). However, if ‘adequate’ segmentation remains elusive, digital tinting described in the next section (Workflow C) might improve demarcation of the ROIs (Fig. 7). Here, the term ‘adequate’ is intended to suggest satisfactory or acceptable segmentation both in quality and quantity. The user will need to define what is adequate for each application or assay in question. It should be noted that digital tinting and color thresholding are not ‘magic bullets’ and will not work in all situations. The success or failure of these techniques depends largely on the results and images in question. As an example, attempts to tint and color threshold the immunochromatographic images featured in Fig. 5 failed to mask known, weak-positive results (i.e., lowest dilution in the dilution series). However, in those instances, the native ImageJ Plot Profile tool was capable of differentiating this weak-positive from negative samples and, therefore, provided one possible path to analysis.
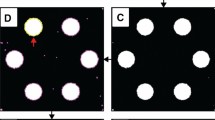
a, This figure features images of 2.5-mm laser cutouts of plastic-backed, silica thin-layer chromatography plates. The circular cuttings were treated with magnesium phthalocyanine and exposed to diethyl 1-phenylethyl phosphonate, a nontoxic chemical warfare agent organophosphate analog. Unsaturated, inhomogeneous, colorless-to-colored chromogenesis was observed, mandating digital tinting for increased contrast between the color response and the image background. b–d, As seen in Fig. 4, the Oval selection tool and the Edit > Selection > Specify submenu are used for ROI selection, refinement and cropping. e, A magenta color adjustment (tint) is applied to the active image via the Image > Adjust > Color Balance… submenu. f, Color thresholds are applied to the tinted image via the Image > Adjust> Color Threshold… submenu. Specifically, the Y’UV threshold windows are Y’ = 174–255, U = 138–255 and V = 0–140. g, Clicking Select in the color threshold window applies the user-defined thresholds, effectively partitioning pixels into ‘object’ and ‘nonobject’ regions (yellow selection outline). h, Removing the background outside the selection permits clear visualization of the selected ROI (Edit > Clear Outside).
Workflow C: Crop-Tint-Threshold-and-Go
For more complex segmentation tasks (e.g., monochromatic color responses containing a subset of images that are very unsaturated or difficult to mask), we often take an iterative approach to setting color thresholds. Briefly, we establish minimal, initial threshold values for masking around a strong positive color response, shown as peaks in the color threshold histograms, while excluding the background color of a clean, negative response, as described in the previous section. Those initial threshold values are applied to a representative range of images within the set, and the resultant ROIs are evaluated (Fig. 7). If necessary, threshold values are adjusted, and the ROI evaluation is repeated. For example, poorly defined threshold values for the images in Fig. 6 can generate ROIs that over-select background pixels or the eliminate the yellow pixels of interest. As discussed in the ‘Limitations’ section, some user bias is inherent in this visual thresholding method. However, once established, final threshold values can be used universally for a given application, guaranteeing consistent, uniform application across image sets and experiments and between users.
As mentioned, when processing images that depict either colorless-to-colored or highly unsaturated colorimetric responses (Fig. 7a), ImageJ can assign color values to the background that are similar to the expected, native color response of the assay16. In these cases, digital tinting via color balance and contrast adjustment is particularly helpful. The ImageJ ‘Brightness/Contrast’ and ‘Color Balance…’ modules enhance the segmentation process by improving image contrast and giving the background a more saturated, non-white color, which can be more easily distinguished from the targeted color response. Adjustments via these modules (Steps 14–20) (Fig. 7e) should be applied before application of color thresholds and image segmentation (Steps 21–26) (Fig. 7f). As a starting point, we recommend making initial adjustments in a color channel that is farthest from the native color response associated with the assay (e.g., on the RGB color wheel, green is complementary to magenta, and red is complementary to cyan). The rationale supporting this recommendation is discussed in Krauss et al.16.
Furthermore, brightness and contrast are both attributes of a color’s luminance. In simple terms, brightness is the overall lightness or darkness of an object, whereas contrast is often associated with visual acuity, or sharpness, and is related to how readily objects or details can be distinguished from other features in the field of view (FOV). As an analogy, consider visual acuity on sunny versus foggy days. On foggy days, objects and details in a person’s FOV are more difficult to distinguish than on sunny days due to lower contrast and similar light intensity. Within the ImageJ menu bar, both the Brightness/Contrast and Color Balance… modules are nested under the Image > Adjust submenu. From the ‘Brightness/Contrast’ window, global brightness and contrast adjustments are applied to the active image via interactive sliders. Narrowing the display range with the ‘Contrast’ slider increases the luminance difference between neighboring pixels, ideally making objects within the image easier to distinguish and isolate. Conversely, expanding the display range decreases contrast. Shifting the display range with the ‘Brightness’ slider alters the overall lightness or darkness of the image. Pixels that fall outside the range defined by the LT and UT of the histogram are displayed as black or white, respectively. Similarly, the ‘Color Balance…’ module (Fig. 7e) allows the user to make adjustments to a single-color channel of the RGB color space. In addition to the expected R, G and B color channels, this module also permits independent manipulation of cyan, yellow and magenta channels (C, Y and M). It is important to note that adjustment of single C, Y or M channels is equivalent to simultaneous manipulation of the blue-green, blue-red and red-green color channels, respectively.
Color spaces: conversion
To date, we have exclusively used RGB and HSB color spaces for reporting colorimetric results. As discussed in Box 1, we use Y’UV and L*a*b* color models for color threshold application and image segmentation, not for analysis and reporting; i.e., the Y’UV and L*a*b* color spaces are used only as a means for enhanced visualization and segmentation. Thus, the active image is still a 24-bit RGB file. As such, before ROI measurement and data acquisition, you will need to convert each active image into a three-slice stack. Selecting ‘RGB Stack’ or ‘HSB Stack’ from the Image > Type submenu converts the active image from 24-bit RGB into three layers that are stacked in a single window, with each slice or layer representing a component channel of the chosen color space. Interacting with the slider bar at the bottom of the window scrolls between layers of the stack (Step 12) (Fig. 2a,c).
Materials
Equipment
-
Image capture device; examples include Huawei P9 smartphone and Epson Perfection V100 desktop scanner
Critical
The image capture device should be able to create raw RGB image files that are readable by ImageJ (see ‘Experimental design’—‘Image capture and file type’ section).
-
Computer with ImageJ (https://imagej.nih.gov/ij/download.html) or the Fiji distribution of ImageJ (https://imagej.net/Fiji/Downloads)
-
Example images: source image files for all featured figures are available via open access at zendo.org, which can be found at https://doi.org/10.5281/zenodo.3976070
Procedure
Critical
As discussed in the ‘Overview of the procedure’ section, this protocol consists of three distinct workflows that represent overlapping layers of ROI refinement and increasing segmentation complexity (Fig. 3). Workflow A, the most elementary route to analysis, includes all steps of the protocol except the more complex portions of the protocol (e.g. Steps 14–20 and Steps 21–26, which encompass tinting, color manipulation and thresholding). Both extended workflows (B and C) fit within, and rely on, the foundational elements outlined in Workflow A. Many of the tools, options and modules detailed in the ‘Procedure‘ section will already be familiar to users with previous experience using ImageJ. However, for users who are unfamiliar with the software, we understand that a period of acclimation and learning is necessary. As such, we recommend familiarization with the steps and tools used in Workflow A before progressing to the more advanced steps featured in Workflows B and C (Steps 14–20 and Steps 21–26).
Critical
It should be noted that the ImageJ Macro Recorder can be used to track, record and save user actions as macros and scripts that can then be used to batch process large numbers of images. Details on ImageJ batch processing are available at https://imagej.nih.gov/ij/docs/pdfs/ImageJBatchProcessing.pdf. In our experience, some familiarity with Java, Python and/or Jython scripting is necessary for full understanding and use of this Macro Recorder scripting feature.
Launching ImageJ and opening the image file
Timing 1 min
-
1
Launch the ImageJ application (Fig. 4a).
-
2
Open your image file. This can be done by selecting the File > Open command path, using keyboard shortcuts Ctrl + O or ⌘ + O or dragging-and-dropping the file directly onto the status bar within the ImageJ GUI.
Selecting and refining an ROI
Timing 30 s per ROI
-
3
(Optional) Use the Image > Duplicate command path to spawn a duplicate of the active image in a new window. Rename the duplicate copy via the popup window that appears. Click OK when finished. Working from a copy prevents inadvertent overwriting of the original image file.
-
4
Select an ROI using one of the following native selection tools (Fig. 4a):
-
Activate the Straight Line selection tool by clicking the icon on the toolbar. Click and drag on the active image to draw a line that transects the image. Holding the Shift key constrains the selection to a horizontal or vertical straight line.
-
Activate the Oval or Rectangle selection tool by clicking the appropriate icon on the toolbar. Click and drag on the active image to define a rough ROI. Holding the Shift key down constrains these rectangular and ovoid selections to square and circular shapes, respectively. While dragging, values related to the selections location and dimensions will appear in the status bar (x, y, w and h). Composite selections (multiple overlapping selections) can be created by holding the Shift or Alt keys (i.e., holding one of these keys while creating a second, overlapping selection adds to or subtracts from a previous selection, respectively (Fig. 4b)).
-
Using the Edit > Selection > Specify command path permits the creation of rectangular and ovoid selections with predefined dimensions and coordinate locations within the active image (Fig. 4c).
Critical step
It might be helpful to change the apparent magnification (Zoom level) of the active image. To access the Zoom In/Out commands, do one of the following: (i) Access the In and Out commands via the Image > Zoom submenu; (ii) Use keyboard shortcuts + or – or Shift + Mouse Wheel; (iii) Restore the active image to original size by clicking the Magnifying glass icon on the ImageJ tool bar.
Critical step
This protocol focuses exclusively on the rectangle or oval selection tools. However, other selection tools are available within ImageJ, including polygon, freehand, wand tracing and straight-line tools. Right-clicking or doubling-clicking the rectangle, oval and straight-line tool icons provides access to other alternative selection modes and brushes. An overview of these tools is available at https://imagej.nih.gov/ij/docs/tools.html. Use of the polygon and straight-line tools is described in Boudaoud et al.14.
-
-
5
(Optional) Rotational transformation (Steps 5–9): To perform rotational transformation of the image, first click Image > Transform > Rotate to open a popup dialogue for rotational transformations (Fig. 5a).
-
6
Click the Preview option to superimpose gridlines onto the active image and to preview the transformation.
-
7
Specify the desired number of gridlines by entering an integer value into the dialogue box (Fig. 5b).
-
8
Rotate the active image. There are several ways to specify the degree of rotation:
-
Enter an integer value directly into the dialogue.
-
Click the arrow buttons to make ±1° rotational adjustments incrementally.
-
Click directly on the slider bar to make ±10° rotational adjustments (−90° to 90°). For rotations <−90° and >90°, apply multiple, sequential rotational transformations via the slider bar or enter larger integer values directly into the dialogue (e.g., −360° to 360°) (Fig. 5b).
Critical step
The Image > Transform submenu and dialogue windows contains additional image transformation options that are not discussed or used here (e.g., flips, rotations and translations).
-
-
9
Click OK to apply the transformation and to exit the Rotate dialogue.
-
10
Refine the selected ROI:
-
For coarse adjustments of the ROI, click and drag the handles that decorate the boundaries of the selection (Fig. 4b).
-
Use keyboard shortcuts: For single-pixel refinement of the selection, press Alt + Arrow. Note: This command executes strict vertical and/or horizontal amendments to the ROI. Use the arrow keys to adjust the position of the selection on the active image.
-
Use the Edit > Selection > Specify command path for swift, precise adjustment of the selection (Figs. 4c and 5c). Clicking through this command path opens a popup window containing fields for width, height and x–y coordinate values. Ticking the appropriate toggle fields constrains rectangular and ovoid selections into squares or circles, respectively. For example, a 163 × 122-pixel ovoid selection (Fig. 4c) can be converted into a constrained to a 100-pixel-wide circular selection (Fig. 4c).
-
-
11
Crop the image to the fit selected ROI (Figs. 4d and 5c). It should be noted that ImageJ generates rectangular crops, regardless of the size, shape and complexity of the user-defined selection. That is, ImageJ crops the image to the smallest rectangular shape that fully bounds the currently selected ROI. To crop, use either the Image > Crop command path or keyboard shortcuts Ctrl +Shift + X or ⌘ + Shift + X.
-
12
(Optional) To fully isolate the selected ROI and remove the background (from default rectangular selection) from the cropped ROI, click through the Edit > Clear Outside or Edit > Clear command paths (Fig. 4e). An ROI must be selected for this step to work properly.
-
13
Save and rename a copy of this cropped image using the File > Save As > TIFF… command path. Saving in the lossless TIFF format preserves the active ROI overlay, preventing unnecessary replication of this work later.
(Optional, for Workflow C only) Color manipulation for enhanced contrast
Timing 30 s per ROI
-
14
Use the Image > Adjust > Color Balance… to open the Color Balance popup window. This popup dialogue contains an intensity histogram of the active image as well as interactive sliders for shifting the histogram display range (brightness) and amending the lower and upper limits of the display range (minimum and maximum) (Fig. 7e).
-
15
Select a color channel for adjustment. The displayed histogram will change to reflect the intensity distribution of the selected color channel.
-
16
Click and drag the Brightness slider to make global changes to the color channel.
Critical step
The previous changes can be cancelled either by clicking Reset without clicking Apply or by selecting another color channel without clicking Apply.
-
17
Close the Color Balance popup window without clicking Apply.
-
18
Click Apply to preserve the changes and to continue.
-
19
(Optional) Repeat Steps 15–18 to perform multiple, sequential color balance adjustments.
-
20
Close the Color Balance popup window.
(Optional, for Workflows B and C only) Color thresholding for image segmentation and ROI refinement
Timing 30 s per ROI
Critical
The color thresholding steps in this section will remove pixels from downstream analysis. As such, these steps should be performed after digital tinting (Steps 14–20).
-
21
Click through the Image > Adjust > Color Threshold… command path to open the Threshold Color popup window (Figs. 6b and 7f).
-
22
Select a color space for threshold adjustment. As previously discussed, RGB is a logical starting point, but L*a*b* or Y’UV might be more visually intuitive workspaces choices for color thresholding (Box 1). The displayed histograms will change to reflect the three component parts of the chosen trichromic color space.
-
23
Click and drag the sliders to adjust the lower and upper limits for each of the component color channels within the selected color space. By default, pixels that fall within the user-defined threshold window are shown in red (Fig. 6c).
Critical step
The previous changes can be cancelled either by clicking the Original button without clicking Select or by closing the Threshold Color Popup window without clicking Select.
-
24
Click Select to apply and preserve the newly selected ROI. The newly selected ROI is displayed with a yellow outline (Fig. 6c).
-
25
Close the Threshold Color popup window.
-
26
Save the newly segmented image by using the File > Save As > Tiff… submenu to save and rename the active image file with the associated ROI overlay. Alternatively, navigate to the File > Save As > Selection… submenu to save the active ROI selection as a stand-alone ROI file.
Caution
Saving in the JPG format will not preserve the active ROI overlay.
Image-type conversion
Timing 15 s per ROI
-
27
Convert the active image into a three-slice RGB or HSB stack (Figs. 8c,9 and 10) by using the Image > Type > RGB Stack or Image > Type > HSB Stack submenu, respectively.
Fig. 8: Workflow A—The Crop-and-Go method of segmentation and analysis. a, This figure features an image of 12 commercially available lateral flow assays (ABAcard, Abacus Diagnostics) for the detection of p30 prostate-specific antigen. These ABAcards were used to detect the presence of semen in mock forensic sexual assault samples. Briefly, after the manufacturer’s protocol, cell lysate was loaded onto each assay card. The image presented here was captured with a desktop scanner after 10 min of incubation/runtime at room temperature. b, As seen in Fig. 5, the Rectangle selection tool and the Image > Transform > Rotate… submenu are used for ROI selection, refinement, rotation and cropping. c, The Analyze >Plot Profile tool is used to evaluate the raw RGB images as well as HSB stacks. c,d, Notice that image type (raw RGB or three-slice HSB stack) and selection method (line or area) affect the output and the consistency of the profiling analysis. Explicitly, for very faint, visually ambiguous color responses, line selections fail to detect a color shift (false negatives). Conversely, area selections correctly reveal positive test results (red arrows).
Fig. 9: Workflow A—The Crop-and-Go method of segmentation and analysis. This figure features images of malachite green before (left) and after (right) exposure to hydrochloric acid vapor, a known hydrolysis product of some organophosphates (Fig. 4 describes this setup). This color change is strongly bichromatic and homogeneous, making it amenable to simple analysis using the Crop-and-Go method depicted here. As seen in Fig. 4, the Oval selection tool and the Edit > Selection > Specify submenu are used for ROI selection, refinemen, and cropping (100 × 100-pixel circular crop). All cropped ROIs are converted to three-slice HSB color space stacks via the Image > Type > HSB stack submenu. Raw hue (yellow highlight) and saturation (blue highlight) data are extracted (Analyze > Measure), copied (Ctrl + A and Ctrl + C or File > Save As…) and exported/pasted to another program (e.g. Microsoft Excel, Origin Pro, R studio, etc) for analysis and bar plot construction.
Fig. 10: Workflow C—The Crop-Tint-Threshold-and-Go method of segmentation and analysis. This figure features images of magnesium phthalocyanine before (left) and after (right) exposure to diethyl 1-phenylethyl phosphonate (Fig. 7 describes this setup). As previously mentioned, this unsaturated, inhomogeneous, colorless-to-colored chromogenesis dictates more involved image processing. Here we demonstrate application of the Crop-Tint-Threshold-and-Go workflow. As seen in Fig. 4, the Oval selection tool and the Edit > Selection > Specify submenu are used for ROI selection, refinement and cropping (100 × 100-pixel circular crop). Digital tinting and color thresholds are applied via the Image > Adjust> Color Balance… and Image > Adjust> Color Threshold… submenus, respectively. All cropped ROIs are converted to three-slice HSB color space stacks via the Image > Type > HSB stack submenu. Raw hue (yellow highlight), saturation (blue highlight) and pixel area data are extracted (Analyze > Measure), copied (Ctrl + A and Ctrl + C or File > Save As…) and exported/pasted to another program (e.g. Microsoft Excel, Origin Pro, R studio, etc) for analysis and bar plot construction. By default, ImageJ analyzed the entire 35 × 35-pixel square (area = 1,225 pixels) when the color threshold eliminated all pixels for the ROI (negatives or before images)—red strikethrough added to raw data window. Thus, pixel area becomes an additional metric for evaluation of the color response.
Analysis
Timing 15 s per analysis tool
-
28
(Optional) Click through the Analyze > Set Measurements submenu to specify what area statistics are to be included in subsequent measurements (e.g., area, minimum, maximum, mean, decimal, places, etc.).
-
29
Analyze the data. Follow Option A for analyzing the histograms of the active ROI selection (Fig. 2a). Follow Option B for analyzing the plot profiles of the active ROI selection (Figs. 2b and 8). Follow Option C for directly analyzing and measuring the active ROI (Fig. 2a).
-
(A)
Option A: Analyzing and measuring the histograms of the active ROI selection:
-
(i)
Click Analyze > Histogram or Ctrl + H to open the histogram popup window. This window consists of a two-dimensional (2D) histogram of the ROI, five buttons (List, Copy, Log, Live and RGB) and an interactive slider. Adjusting the current ROI (Step 6) while the Live button is active modifies the histogram and area statistics in real time.
-
(ii)
Click the Live button to view a histogram of the active RGB and HSB stacks. Manipulate the interactive slider in the stack window to switch between the slices of the stack and the associated histograms. For original 24-bit RGB images that are not yet converted to RGB or HSB stacks, click the RGB button to scroll through the six different histogram options.
-
(iii)
Export the active histogram and associated data.
-
(iv)
Save the active histogram as an image file by clicking through the File > Save As… command path on the main ImageJ menu bar.
-
(v)
Save this histogram data as a *.csv file by clicking List, followed by the File > Save As… command path.
-
(vi)
Add the histogram data to your system clipboard by clicking Edit > Cut, Edit > Copy or the Copy button. This option permits cut, copy and paste data transfers from ImageJ to other widgets or applications (i.e. text editors, spreadsheet programs or other software).
-
(i)
-
(B)
Option B: Analyzing the Plot Profiles of the active ROI selection
-
(i)
Clicking Analyze > Plot Profile or Ctrl + K opens the Plot Profile popup window. This window consists of a 2D intensity plot of the ROI and four interactive buttons (List, Data>>, More>> and Live). Customize the appearance of the plot by clicking the More >> button followed by any of the nested options in the More >> submenu (e.g. Set range…, Axis options…, Legend…, etc). Modifying the current ROI (Step 10), while the Live button is active, adjusts the 2D plot profile in real time.
-
(ii)
Export the plot profile and associated data. Save the active plot profile view as an image file by clicking through the File > Save As.. command path on the main ImageJ menu bar.
-
(iii)
Export and save the plot profile x–y coordinate data as a *.csv file by clicking List > File > Save As… or Data >> Save As..
-
(iv)
Add the plot profile x–y data to your system clipboard by clicking the List button followed by Ctrl + A and Ctrl + C (Windows) or ⌘ + O and ⌘ + O (Mac).
-
(i)
-
(C)
Option C: Directly analyzing and measuring the active ROI
-
(i)
Calculate and display area statistics for the current ROI and active stack slice by clicking Analyze > Measure, Ctrl + M (Windows) or ⌘ + M (Mac). To switch between slices of the color space stack, manipulate the interactive slider in the active stack window. Measurements are added to and displayed in a Results popup window.
-
(ii)
Save the results by clicking through the File > Save As.. command path in the Results window.
-
(iii)
Add the measurements data to your system clipboard by clicking Ctrl + A and Ctrl + C (Windows) or ⌘ + O and ⌘ + O (Mac).
-
(i)
-
(A)
Troubleshooting
Troubleshooting advice can be found in Table 1.
Timing
The suggested timing information below is for experienced users. New users who are unfamiliar with ImageJ will find that familiarization with these steps takes much longer, and they should anticipate at least double the timing reported in this section.
-
Steps 1 and 2, launching ImageJ and Opening the Image File: 1 min
-
Steps 3–13, selecting and refining an ROI: 30 s per ROI (initial crop size and threshold determination might take longer)
-
Steps 14–20, (Optional, for Workflow C only) Color manipulation for enhanced contrast: 30 s per ROI
-
Steps 21–26, (Optional, for Workflows B and C only) Color thresholding for image segmentation and ROI refinement: 30 s per ROI
-
Step 27, image-type conversion: 15 s per ROI
-
Steps 28 and 29, analysis: 15 s per analysis tool
Anticipated results
In addition to raw numerical data, successful implementation of this protocol will allow the analyst to generate results in several formats, namely histograms, plot profiles, bar plots and scatter plots.
Histograms
The Analyze > Histogram command path generates a histogram and calculates area statistics for the current ROI (Fig. 2). Step 29, Option A of the procedure details saving histograms as image files and exporting histogram data as a *.csv file. For three-slice image stacks, the displayed histogram represents pixel distributions (y axis) across all intensity values of the selected color space slice (x axis = 0–255). Likewise, individual RGB histograms depict pixel distributions (y axis) across all intensity values of the selected color channel of an RGB image (x axis = 0–255). For the unweighted and weighted grayscale histograms, ImageJ converts individual RGB pixels into grayscale intensities with these formulas,
or
Weighted conversions produce grayscale images with brightness that is perceptually most similar to the original color image. However, although visually darker (closer to black or 0 on a 0–255 grayscale), the unweighted grayscale conversion produces images that are directly equivalent to the original color image and are not biased toward the green component. As such, for the duration of this protocol, default unweighted grayscale conversions are implemented. You can browse the histograms by activating the RGB button (RGB images) or the Live button (converted image stacks) located in the histogram popup window. Switching between color space channels while the RGB or Live buttons are active forces real-time changes to the area statics and the channel histogram.
Plot profiles
The Analyze > Plot Profile command path generates 2D plots depicting grayscale pixel intensities across a line or rectangular area selection (Fig. 2). Step 29, Option B of the ‘Procedure’ section details saving plot profiles as image files and exporting profile data as a *.csv file. For line selections, the plot profile displays grayscale pixel intensity (y axis) along the horizontal distance of the line through the active image (x axis). For area selections, the plot profile displays a ‘column averaged plot’, where the x and y axes represent the horizontal distance through the image and the vertically averaged grayscale pixel intensity, respectively. Activating the Live button forces dynamic, real-time changes to the plot profile as modifications are made to the current ROI selection (Step 10).
Bar and scatter lots
Exported ImageJ datasets can be used to construct bar and scatter plots via any number of third-party widgets, programs and applications (e.g., Microsoft Excel, R Studio, OriginPro and GraphPad Prism) (Fig. 2). Step 29, Option C of the ‘Procedure’ section details acquiring ROI area measurements and exporting the raw data for downstream visualization and statistical analysis.
Data availability
All figures and any associated data have not been previously published. The source image files used to generate the primary data underlying the figures featured in this protocol are available via open access at zenodo.org, which can be found at https://doi.org/10.5281/zenodo.3976070.
Code availability
This protocol requires no custom scripts or algorithms. Rather, this protocol uses free, publicly available software packages (e.g., ImageJ 2.0.0-rc-69/1.52p bundled with Java 1.8.0_172 (32-bit) (https://imagej.nih.gov/ij/download.html) and the Fiji distribution of ImageJ (https://imagej.net/Fiji/Downloads))10,11,12 as well as available commercial software (e.g., Microsoft Excel for Microsoft 365).
References
Lee, B. B. The evolution of concepts of color vision. Neurociencias 4, 209–224 (2008).
Molday, R. S. & Moritz, O. L. Photoreceptors at a glance. J. Cell Sci. 128, 4039–4045 (2015).
Jameson, K. A. in The Oxford Companion to Consciousness 155–158 (Oxford University Press, 2009).
Krauss, S. T. et al. Objective method for presumptive field-testing of illicit drug possession using centrifugal microdevices and smartphone analysis. Anal. Chem. 88, 8689–8697 (2016).
CIE. Commission internationale de l’eclairage proceedings, 1931 (Cambridge University, 1932).
Smith, T. & Guild, J. The CIE colorimetric standards and their use. Trans. Opt. Soc. 33, 73 (1931).
CIE. Colorimetry-Part 4: CIE 1976 L* a* b* colour space. International Organization for Standardization https://www.iso.org/standard/74166.html (2008).
Schnapf, J., Kraft, T. & Baylor, D. Spectral sensitivity of human cone photoreceptors. Nature 325, 439–441 (1987).
Anderson, M., Motta, R., Chandrasekar, S. & Stokes, M. in Color and Imaging Conference 238–245 (Society for Imaging Science and Technology, 1996).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18, 529 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676 (2012).
Roels, J. et al. An interactive ImageJ plugin for semi-automated image denoising in electron microscopy. Nat. Commun. 11, 1–13 (2020).
Boudaoud, A. et al. FibrilTool, an ImageJ plug-in to quantify fibrillar structures in raw microscopy images. Nat. Protoc. 9, 457 (2014).
Krauss, S. T., Holt, V. C. & Landers, J. P. Simple reagent storage in polyester-paper hybrid microdevices for colorimetric detection. Sens. Actuators B Chem. 246, 740–747 (2017).
Krauss, S. T., Nauman, A. Q., Garner, G. T. & Landers, J. P. Color manipulation through microchip tinting for colorimetric detection using hue image analysis. Lab Chip 17, 4089–4096 (2017).
Krauss, S. T. et al. Centrifugal microfluidic devices using low-volume reagent storage and inward fluid displacement for presumptive drug detection. Sens. Actuators B Chem. 284, 704–710 (2019).
Thompson, B. L., Wyckoff, S. L., Haverstick, D. M. & Landers, J. P. Simple, reagentless quantification of total bilirubin in blood via microfluidic phototreatment and image analysis. Anal. Chem. 89, 3228–3234 (2017).
Jackson, K. R. et al. A novel loop-mediated isothermal amplification method for identification of four body fluids with smartphone detection. Forensic Sci. Int. Genet. 45, 102195 (2020).
Russell, R. A. et al. Segmentation of fluorescence microscopy images for quantitative analysis of cell nuclear architecture. Biophys. J. 96, 3379–3389 (2009).
Balsam, J., Bruck, H. A., Kostov, Y. & Rasooly, A. Image stacking approach to increase sensitivity of fluorescence detection using a low cost complementary metal-oxide-semiconductor (CMOS) webcam. Sens. Actuators B Chem. 171, 141–147 (2012).
Perez, A. J. et al. A workflow for the automatic segmentation of organelles in electron microscopy image stacks. Front. Neuroanat. 8, 126 (2014).
Stegmaier, J. et al. Fast segmentation of stained nuclei in terabyte-scale, time resolved 3D microscopy image stacks. PLoS ONE 9, e90036 (2014).
Capitan-Vallvey, L. F., Lopez-Ruiz, N., Martinez-Olmos, A., Erenas, M. M. & Palma, A. J. Recent developments in computer vision-based analytical chemistry: a tutorial review. Anal. Chim. Acta 899, 23–56 (2015).
Cao, S., Huang, D., Wang, Y. & Li, G. in Advances in Mechanical and Electronic Engineering 381–386 (Springer, 2012).
Cantrell, K., Erenas, M., de Orbe-Payá, I. & Capitán-Vallvey, L. Use of the hue parameter of the hue, saturation, value color space as a quantitative analytical parameter for bitonal optical sensors. Anal. Chem. 82, 531–542 (2010).
Ibraheem, N. A., Hasan, M. M., Khan, R. Z. & Mishra, P. K. Understanding color models: a review. ARPN J. Sci. Technol. 2, 265–275 (2012).
Hanbury, A. The taming of the hue, saturation and brightness colour space. Proc. 7th Computer Vision Winter Workshop http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.3.2574 (2002).
Barthel, K. U. 3D-data representation with ImageJ. Proc. 1st ImageJ User & Developer Conference http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.414.725 (2006).
Clark, C. P. et al. Closable valves and channels for polymeric microfluidic devices. Micromachines 11, 627 (2020).
Tian, Z., Liu, L. Q., Peng, C., Chen, Z. & Xu, C. A new development of measurement of 19-Nortestosterone by combining immunochromatographic strip assay and ImageJ software. Food Agric. Immunol. 20, 1–10 (2009).
Hwang, J., Kwon, D., Lee, S. & Jeon, S. Detection of Salmonella bacteria in milk using gold-coated magnetic nanoparticle clusters and lateral flow filters. RSC Adv. 6, 48445–48448 (2016).
Kortli, S. et al. Yersinia pestis detection using biotinylated dNTPs for signal enhancement in lateral flow assays. Anal. Chim. Acta 1112, 54–61 (2020).
Adkins, J. A. et al. Colorimetric and electrochemical bacteria detection using printed paper-and transparency-based analytic devices. Anal. Chem. 89, 3613–3621 (2017).
Tanner, N. A., Zhang, Y. & Evans, T. C. Jr Visual detection of isothermal nucleic acid amplification using pH-sensitive dyes. Biotechniques 58, 59–68 (2015).
Zhang, Y. et al. Rapid molecular detection of SARS-CoV-2 (COVID-19) virus RNA using colorimetric LAMP. Preprint at https://www.medrxiv.org/content/10.1101/2020.02.26.20028373v1 (2020).
Singh, H. K., Tomar, S. K. & Maurya, P. K. Thresholding techniques applied for segmentation of RGB and multispectral images. https://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.735.9363&rep=rep1&type=pdf (2012).
Acknowledgements
We gratefully acknowledge K. Siller (UVA Research Computing, University of Virginia) for his advice and instruction provided in ImageJ-Fiji workshops given at the University of Virginia (attended by M.S.W. and A.T.S. in 2018). We also recognize the initial foundational image analysis work of previous Landers lab members, including C. P. Clark, S. T. Krauss, K. R. Jackson, J. Li, D. A. Nelson, O. N. Scott and B. L. Thompson.
Author information
Authors and Affiliations
Contributions
All author contributions are based on CRediT standards. Writing—original draft: M.S.W. Writing—review and editing: all authors. Conceptualization, methodology, investigation, data curation, formal analysis and visualization of the image segmentation procedure: M.S.W. Conceptualization, methodology, investigation and image acquisition for all colorimetric experiments: M.S.W., L.M.D. and A.T.S. Project administration and funding acquisition: J.P.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Bahram Hemmateenejad and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Clark, C. P. et al. Micromachines 11, 627 (2020): https://doi.org/10.3390/mi11070627
Jackson, K. R. et al. Forensic Sci. Int. Genet. 45, 102195 (2020): https://doi.org/10.1016/j.fsigen.2019.102195
Krauss, S. T. et al. Sens. Actuators B Chem. 284, 704−710 (2019): https://doi.org/10.1016/j.snb.2018.12.113
Krauss, S.T. et al. Lab Chip 17, 4089−4096 (2017): https://doi.org/10.1039/C7LC00796E
Rights and permissions
About this article
Cite this article
Woolf, M.S., Dignan, L.M., Scott, A.T. et al. Digital postprocessing and image segmentation for objective analysis of colorimetric reactions. Nat Protoc 16, 218–238 (2021). https://doi.org/10.1038/s41596-020-00413-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00413-0
This article is cited by
-
Application of machine learning algorithms for accurate determination of bilirubin level on in vitro engineered tissue phantom images
Scientific Reports (2024)
-
Smartphone-based imaging colorimetric assay for monitoring the quality of curcumin in turmeric powder
Analytical Sciences (2024)
-
Smartphone-based platforms implementing microfluidic detection with image-based artificial intelligence
Nature Communications (2023)
-
Computer vision meets microfluidics: a label-free method for high-throughput cell analysis
Microsystems & Nanoengineering (2023)
-
An android smartphone-based digital image colorimeter for detecting acid fuchsine dye in aqueous solutions
Journal of the Iranian Chemical Society (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.