Abstract
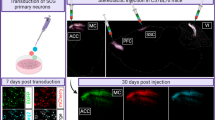
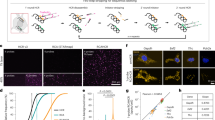
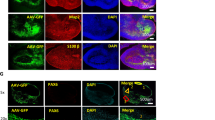
To understand and control complex tissues, the ability to genetically manipulate single cells is required. However, current delivery methods for the genetic engineering of single cells, including viral transduction, suffer from limitations that restrict their application. Here we present a protocol that describes a versatile technique that can be used for the targeted viral infection of single cells or small groups of cells in any tissue that is optically accessible. First, cells of interest are selected using optical microscopy. Second, a micropipette—loaded with magnetic nanoparticles to which viral particles are bound—is brought into proximity of the cell of interest, and a magnetic field is applied to guide the viral nanoparticles into cellular contact, leading to transduction. The protocol, exemplified here by stamping cultured neurons with adeno-associated viruses (AAVs), is completed in a few minutes and allows stable transgene expression within a few days, at success rates that approach 80%. We outline how this strategy is applied to single-cell infection in complex tissues, and is feasible both in organoids and in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data shown in this protocol are available upon reasonable request.
References
Akhtar, A. et al. A decade of molecular cell biology: achievements and challenges. Nat. Rev. Mol. Cell Biol. 12, 669–674 (2011).
Chari, R. & Church, G. M. Beyond editing to writing large genomes. Nat. Rev. Genet. 18, 749–760 (2017).
Wachsman, G. & Heidstra, R. The CRE/lox system as a tool for developmental studies at the cell and tissue level. Methods Mol. Biol. 655, 47–64 (2010).
de la Fuente-Núñez, C. & Lu, T. K. CRISPR-Cas9 technology: applications in genome engineering, development of sequence-specific antimicrobials, and future prospects. Integr. Biol. 9, 109–122 (2017).
Murrow, L. M., Weber, R. J. & Gartner, Z. J. Dissecting the stem cell niche with organoid models: an engineering-based approach. Development 144, 998–1007 (2017).
Jaenisch, R. Transgenic animals. Science 240, 1468–1474 (1988).
Weake, V. M. & Workman, J. L. Inducible gene expression: diverse regulatory mechanisms. Nat. Rev. Genet. 11, 426–437 (2010).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Maniatis, T., Goodbourn, S. & Fischer, J. A. Regulation of inducible and tissue-specific gene expression. Science 236, 1237–1245 (1987).
Mello, C. C. & Conte, D. Jr. Revealing the world of RNA interference. Nature 431, 338–342 (2004).
Ptashne, M. The chemistry of regulation of genes and other things. J. Biol. Chem. 289, 5417–5435 (2014).
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G. & Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat. Neurosci. 8, 1263–1268 (2005).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Palop, J. J. et al. Aberrant excitatory neuronal activity and compensatory remodeling of inhibitory hippocampal circuits in mouse models of Alzheimer’s disease. Neuron 55, 697–711 (2007).
Rossi, A. et al. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524, 230–233 (2015).
Berchtold, N. C. et al. Gene expression changes in the course of normal brain aging are sexually dimorphic. Proc. Natl. Acad. Sci. USA 105, 15605–15610 (2008).
Matthaei, K. I. Genetically manipulated mice: a powerful tool with unsuspected caveats. J. Physiol. 582, 481–488 (2007).
Fugger, L., Friese, M. A. & Bell, J. I. From genes to function: the next challenge to understanding multiple sclerosis. Nat. Rev. Immunol. 9, 408–417 (2009).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Zeng, H. & Sanes, J. R. Neuronal cell-type classification: challenges, opportunities and the path forward. Nat. Rev. Neurosci. 18, 530–546 (2017).
Kebschull, J. M. et al. High-throughput mapping of single-neuron projections by sequencing of barcoded RNA. Neuron 91, 975–987 (2016).
Schubert, R. et al. Virus stamping for targeted single-cell infection in vitro and in vivo. Nat. Biotechnol. 36, 81–88 (2018).
Lotze, M. T. & Kost, T. A. Viruses as gene delivery vectors: application to gene function, target validation, and assay development. Cancer Gene Ther. 9, 692–699 (2002).
Naso, M. F., Tomkowicz, B., Perry, W. L. & Strohl, W. R. Adeno-associated virus (AAV) as a vector for gene therapy. Biodrugs 31, 317–334 (2017).
Packer, A. M., Roska, B. & Häusser, M. Targeting neurons and photons for optogenetics. Nat. Neurosci. 16, 805–815 (2013).
Judkewitz, B., Rizzi, M., Kitamura, K. & Häusser, M. Targeted single-cell electroporation of mammalian neurons in vivo. Nat. Protoc. 4, 862–869 (2009).
Wertz, A. et al. Single-cell-initiated monosynaptic tracing reveals layer-specific cortical network modules. Science 349, 70–74 (2015).
Alsteens, D. et al. Atomic force microscopy-based characterization and design of biointerfaces. Nat. Rev. Mater. 2, 17008 (2017).
Rancz, E. A. et al. Transfection via whole-cell recording in vivo: bridging single-cell physiology, genetics and connectomics. Nat. Neurosci. 14, 527–532 (2011).
Marshel, J. H., Mori, T., Nielsen, K. J. & Callaway, E. M. Targeting single neuronal networks for gene expression and cell labeling in vivo. Neuron 67, 562–574 (2010).
Martínez-Martín, D. et al. Inertial picobalance reveals fast mass fluctuations in mammalian cells. Nature 550, 500–505 (2017).
Stiefel, P. et al. Cooperative vaccinia infection demonstrated at the single-cell level using FluidFM. Nano Lett. 12, 4219–4227 (2012).
Nguyen, T. D. et al. Targeted single-neuron infection with rabies virus for transneuronal multisynaptic tracing. J. Neurosci. Methods 209, 367–370 (2012).
Rancz, E. A. & Schaefer, A. T. Viruses leave their stamp on single cells. Nat. Biotechnol. 36, 42–44 (2018).
Vogt, N. Putting a stamp on single cells. Nat. Methods 15, 95 (2018).
Muñoz, W., Tremblay, R. & Rudy, B. Channelrhodopsin-assisted patching: in vivo recording of genetically and morphologically identified neurons throughout the brain. Cell Rep. 9, 2304–2316 (2014).
Komazin-Meredith, G. et al. The positively charged surface of herpes simplex virus UL42 mediates DNA binding. J. Biol. Chem. 283, 6154–6161 (2008).
Alsteens, D. et al. Nanomechanical mapping of first binding steps of a virus to animal cells. Nat. Nanotechnol. 12, 177–183 (2017).
Ginger, M., Haberl, M., Conzelmann, K. K., Schwarz, M. K. & Frick, A. Revealing the secrets of neuronal circuits with recombinant rabies virus technology. Front. Neural Circuits 7, 2 (2013).
Rothermel, M., Brunert, D., Zabawa, C., Díaz -Quesada, M. & Wachowiak, M. Transgene expression in target-defined neuron populations mediated by retrograde infection with adeno-associated viral vectors. J. Neurosci. 33, 15195–15206 (2013).
Jurgens, H. A., Amancherla, K. & Johnson, R. W. Influenza infection induces neuroinflammation, alters hippocampal neuron morphology, and impairs cognition in adult mice. J. Neurosci. 32, 3958–3968 (2012).
Cronin, J., Zhang, X. Y. & Reiser, J. Altering the tropism of lentiviral vectors through pseudotyping. Curr. Gene Ther. 5, 387–398 (2005).
Walther, W. & Stein, U. Cell type specific and inducible promoters for vectors in gene therapy as an approach for cell targeting. J. Mol. Med. 74, 379–392 (1996).
Conzelmann, K. K. Nonsegmented negative-strand RNA viruses: genetics and manipulation of viral genomes. Annu. Rev. Genet. 32, 123–162 (1998).
Goldsmith, C. S. & Miller, S. E. Modern uses of electron microscopy for detection of viruses. Clin. Microbiol. Rev. 22, 552–563 (2009).
Uchida, E. et al. Optimization of the virus concentration method using polyethyleneimine-conjugated magnetic beads and its application to the detection of human hepatitis A, B and C viruses. J. Virol. Methods 143, 95–103 (2007).
Phelan, K. & May, K. M. Basic techniques in mammalian cell tissue culture. Curr. Protoc. Cell Biol. 66, 1.1.1–1.1.22 (2015).
Pavlov, T. S. et al. Implementing patch clamp and live fluorescence microscopy to monitor functional properties of freshly isolated PKD epithelium. J. Vis. Exp. 103, e53035 (2015).
Booker, S. A., Song, J. & Vida, I. Whole-cell patch-clamp recordings from morphologically- and neurochemically-identified hippocampal interneurons. J. Vis. Exp. 91, e51706 (2014).
Stuart, G. J., Dodt, H. U. & Sakmann, B. Patch-clamp recordings from the soma and dendrites of neurons in brain slices using infrared video microscopy. Pflug. Arch. 423, 511–518 (1993).
Kitamura, K., Judkewitz, B., Kano, M., Denk, W. & Häusser, M. Targeted patch-clamp recordings and single-cell electroporation of unlabeled neurons in vivo. Nat. Methods 5, 61–67 (2008).
Grutzendler, J., Kasthuri, N. & Gan, W. B. Long-term dendritic spine stability in the adult cortex. Nature 420, 812–816 (2002).
Chou, S. J. et al. Analysis of spatial-temporal gene expression patterns reveals dynamics and regionalization in developing mouse brain. Sci. Rep. 6, 19274 (2016).
Ran, Y. F. et al. Rapid, highly sensitive detection of herpes simplex virus-1 using multiple antigenic peptide-coated superparamagnetic beads. Analyst 139, 6126–6134 (2014).
Kendall, C. G. et al. Amine analysis using AlexaFluor 488 succinimidyl ester and capillary electrophoresis with laser-induced fluorescence. J. Anal. Methods Chem. 2015, 368362 (2015).
Acknowledgements
We thank P. Buchmann and P. Argast for building our magnet Supplementary Figure 1. The study was supported by an European Union grant (FP7/211800 to D.J.M.), a Canada Research Chair (Tier II to S.T.), Swiss National Science Foundation grants (310030B_160225 to D.J.M. and 3100330B_163457 to B.R.), the National Center of Competence in Research (NCCR) Molecular Systems Engineering (to D.J.M. and B.R.), the European Research Council (669157, RETMUS to B.R.) and a DARPA grant (HR0011-17-C-0038, Cortical Sight to B.R.).
Author information
Authors and Affiliations
Contributions
All authors discussed the protocol and contributed to the writing of the protocol. R.S., S.H. and S.T. designed and optimized the experiments described. Figures and videos were created by R.S., S.H. and S.T.
Corresponding authors
Ethics declarations
Competing interests
R.S., S.T., D.J.M. and B.R. applied for a patent related to the virus-stamping approach. The remaining author declares no competing interests.
Additional information
Peer review information Nature Protocols thanks Martina Anton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Schubert, R. et al. Nat. Biotechnol. 36, 81–88 (2018): https://doi.org/10.1038/nbt.4034
Wertz, A. et al. Science 349, 70–74 (2015): https://doi.org/10.1126/science.aab1687
Alsteens, D. et al. Nat. Nanotechnol. 12, 177–183 (2017): https://doi.org/10.1038/nnano.2016.228
Integrated supplementary information
Supplementary Fig. 1 Schematic of setup used for shielded virus stamping for experiments.
To maintain health of cells during the stamping procedure and allowing for glass capillary (pipette), optical and magnet access we built a custom chamber made of two concentric compartments31. The inner compartment is a glass-bottomed dish in which cells are cultured at 37 °C. The surrounding compartment is continuously filled via an inlet with humidified (<98% relative humidity) synthetic air (80% N2 and 20% O2) supplemented with 5% CO2. The top of the chamber is left open to access the cells by the glass capillary pipette and objective (top-down dipping objective). A 35 mm diameter opening of the outer chamber (chamber lid) allows optical access from the top. The angle between pipette and glass-bottom dish, which is usually ~ 45° can be adjusted according to the experimental needs. The bottom of the chamber can be optically assessed as well as indicated by the condenser. Depending on the angle of the pipette, the electromagnet (or permanent magnet) is aligned from below without blocking the light pathway. If experiments are to be conducted over extended time periods, the chambers can be covered with silicone sealant to enable gas exchange between compartments and to prevent evaporation of the cell-culture media.
Supplementary information
Supplementary Information
Supplementary Fig. 1
Supplementary Video 1
Nanoparticle mobility in the glass capillary pipette in the presence of a magnetic field.
Supplementary Video 2
Virus stamping of a cultured rat cortical neuron.
Rights and permissions
About this article
Cite this article
Schubert, R., Herzog, S., Trenholm, S. et al. Magnetically guided virus stamping for the targeted infection of single cells or groups of cells. Nat Protoc 14, 3205–3219 (2019). https://doi.org/10.1038/s41596-019-0221-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0221-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.