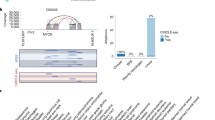
Abstract
Oncogene amplification on extrachromosomal DNA (ecDNA) is prevalent in human cancer and is associated with poor outcomes. Clonal, megabase-sized circular ecDNAs in cancer are distinct from nonclonal, small sub-kilobase-sized DNAs that may arise during normal tissue homeostasis. ecDNAs enable profound changes in gene regulation beyond copy-number gains. An emerging principle of ecDNA regulation is the formation of ecDNA hubs: micrometer-sized nuclear structures of numerous copies of ecDNAs tethered by proteins in spatial proximity. ecDNA hubs enable cooperative and intermolecular sharing of DNA regulatory elements for potent and combinatorial gene activation. The 3D context of ecDNA shapes its gene expression potential, selection for clonal heterogeneity among ecDNAs, distribution through cell division, and reintegration into chromosomes. Technologies for studying gene regulation and structure of ecDNA are starting to answer long-held questions on the distinct rules that govern cancer genes beyond chromosomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cox, D., Yuncken, C. & Spriggs, A. I. Minute chromatin bodies in malignant tumours of childhood. Lancet 286, 55–58 (1965). The original description of ecDNA in tumor cells manifesting as small chromatin bodies on chromosome spreads.
Spriggs, A. I., Boddington, M. M. & Clarke, C. M. Chromosomes of human cancer cells. Br. Med J. 2, 1431–1435 (1962).
Hoff, D. D. V., Needham-VanDevanter, D. R., Yucel, J., Windle, B. E. & Wahl, G. M. Amplified human MYC oncogenes localized to replicating submicroscopic circular DNA molecules. Proc. Natl Acad. Sci. USA 85, 4804–4808 (1988).
Turner, K. M. et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 543, 122–125 (2017). Systematic analysis of human cancer models using sequencing and cytogenetics identified ecDNAs in nearly half of human cancers and not in normal cells.
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat. Genet. 52, 891–897 (2020). Comprehensive analysis of primary tumors found increased oncogene transcription and worsened outcomes linked to ecDNA.
Kohl, N. E. et al. Transposition and amplification of oncogene-related sequences in human neuroblastomas. Cell 35, 359–367 (1983).
Benner, S., Wahl, G. & Hoff, D. V. Double minute chromosomes and homogeneously staining regions in tumors taken directly from patients versus in human tumor cell lines. Anti-cancer Drugs 2, 11–26 (1991).
Bigner, S. H., Mark, J. & Bigner, D. D. Cytogenetics of human brain tumors. Cancer Genet. Cytogenetics 47, 141–154 (1990).
Storlazzi, C. T. et al. Gene amplification as double minutes or homogeneously staining regions in solid tumors: origin and structure. Genome Res. 20, 1198–1206 (2010).
Yoshimoto, M. et al. MYCN gene amplification: identification of cell populations containing double minutes and homogeneously staining regions in neuroblastoma tumors. Am. J. Pathol. 155, 1439–1443 (1999).
Vicario, R. et al. Patterns of HER2 gene amplification and response to anti-HER2 therapies. PLoS ONE 10, e0129876 (2015).
McGill, J. R. et al. Double minutes are frequently found in ovarian carcinomas. Cancer Genet. Cytogenetics 71, 125–131 (1993).
Lin, C. C. et al. Evolution of karyotypic abnormalities and C-MYC oncogene amplification in human colonic carcinoma cell lines. Chromosoma 92, 11–15 (1985).
Wahl, G. M. The importance of circular DNA in mammalian gene amplification. Cancer Res. 49, 1333–1340 (1989).
Quinn, L. A., Moore, G. E., Morgan, R. T. & Woods, L. K. Cell lines from human colon carcinoma with unusual cell products, double minutes, and homogeneously staining regions. Cancer Res. 39, 4914–4924 (1979).
Carroll, S. M. et al. Double minute chromosomes can be produced from precursors derived from a chromosomal deletion. Mol. Cell. Biol. 8, 1525–1533 (1988).
Maurer, B. J., Lai, E., Hamkalo, B. A., Hood, L. & Attardi, G. Novel submicroscopic extrachromosomal elements containing amplified genes in human cells. Nature 327, 434–437 (1987).
Pauletti, G., Lai, E. & Attardi, G. Early appearance and long-term persistence of the submicroscopic extrachromosomal elements (amplisomes) containing the amplified DHFR genes in human cell lines. Proc. Natl Acad. Sci. USA 87, 2955–2959 (1990).
Wang, Y. et al. eccDNAs are apoptotic products with high innate immunostimulatory activity. Nature 599, 308–314 (2021).
Møller, H. D. et al. Circular DNA elements of chromosomal origin are common in healthy human somatic tissue. Nat. Commun. 9, 1069 (2018).
Møller, H. D., Parsons, L., Jørgensen, T. S., Botstein, D. & Regenberg, B. Extrachromosomal circular DNA is common in yeast. Proc. Natl Acad. Sci. USA 112, E3114–E3122 (2015).
Paulsen, T., Kumar, P., Koseoglu, M. M. & Dutta, A. Discoveries of extrachromosomal circles of DNA in normal and tumor cells. Trends Genet. 34, 270–278 (2018).
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Hung, K. L. et al. ecDNA hubs drive cooperative intermolecular oncogene expression. Nature 600, 731–736 (2021). Discovery of ecDNA hubs that enable intermolecular activation of oncogene expression through enhancer–promoter interactions.
Levan, A. & Levan, G. Have double minutes functioning centromeres? Hereditas 88, 81–92 (1978). Conclusive evidence that ecDNAs lack centromeres, which explains their distinct mode of random segregation that results in copy-number heterogeneity.
Lundberg, G. et al. Binomial mitotic segregation of MYCN-carrying double minutes in neuroblastoma illustrates the role of randomness in oncogene amplification. PLoS ONE 3, e3099 (2008).
Lange, J. T. et al. Principles of ecDNA random inheritance drive rapid genome change and therapy resistance in human cancers. Preprint at bioRxiv https://doi.org/10.1101/2021.06.11.447968 (2021).
Ståhl, F., Wettergren, Y. & Levan, G. Amplicon structure in multidrug-resistant murine cells: a nonrearranged region of genomic DNA corresponding to large circular DNA. Mol. Cell. Biol. 12, 1179–1187 (1992).
Nathanson, D. A. et al. Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science 343, 72–76 (2014).
Yu, M. & Ren, B. The three-dimensional organization of mammalian genomes. Annu. Rev. Dev. Cell Biol. 33, 265–289 (2017).
Cremer, T. & Cremer, M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2, a003889 (2010).
Spilianakis, C. G., Lalioti, M. D., Town, T., Lee, G. R. & Flavell, R. A. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005).
Apostolou, E. & Thanos, D. Virus infection induces NF-κB-dependent interchromosomal associations mediating monoallelic IFN-β gene expression. Cell 134, 85–96 (2008).
Maass, P. G., Barutcu, A. R. & Rinn, J. L. Interchromosomal interactions: a genomic love story of kissing chromosomes. J. Cell Biol. 218, 27–38 (2019).
Yi, E. et al. Live-cell imaging shows uneven segregation of extrachromosomal DNA elements and transcriptionally active extrachromosomal DNA hubs in cancer. Cancer Discov. 12, 468–483 (2021).
Itoh, N. & Shimizu, N. DNA replication-dependent intranuclear relocation of double minute chromatin. J. Cell Sci. 111, 3275–3285 (1998).
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007).
Kanda, T., Sullivan, K. F. & Wahl, G. M. Histone-GFP fusion protein enables sensitive analysis of chromosome dynamics in living mammalian cells. Curr. Biol. 8, 377–385 (1998).
Oobatake, Y. & Shimizu, N. Double-strand breakage in the extrachromosomal double minutes triggers their aggregation in the nucleus, micronucleation, and morphological transformation. Genes Chromosomes Cancer 59, 133–143 (2020).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Gibson, B. A. et al. Organization of chromatin by intrinsic and regulated phase separation. Cell 179, 470–484 (2019).
Shin, Y. et al. Liquid nuclear condensates mechanically sense and restructure the genome. Cell 175, 1481–1491 (2018).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Henssen, A. et al. Targeting MYCN-driven transcription by BET-bromodomain inhibition. Clin. Cancer Res. 22, 2470–2481 (2016).
Morton, A. R. et al. Functional enhancers shape extrachromosomal oncogene amplifications. Cell 179, 1330–1341 (2019). Description of oncogene-enhancer co-amplification on ecDNAs, showing hijacking of both cognate and ectopic enhancers to drive oncogene expression.
Tanabe, H. et al. Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc. Natl Acad. Sci. USA 99, 4424–4429 (2002).
Helmsauer, K. et al. Enhancer hijacking determines extrachromosomal circular MYCN amplicon architecture in neuroblastoma. Nat. Commun. 11, 5823 (2020).
Nikolaev, S. et al. Extrachromosomal driver mutations in glioblastoma and low-grade glioma. Nat. Commun. 5, 5690 (2014).
Risca, V. I., Denny, S. K., Straight, A. F. & Greenleaf, W. J. Variable chromatin structure revealed by in situ spatially correlated DNA cleavage mapping. Nature 541, 237–241 (2017).
Fragkos, M., Ganier, O., Coulombe, P. & Méchali, M. DNA replication origin activation in space and time. Nat. Rev. Mol. Cell Biol. 16, 360–374 (2015).
Carré-Simon, À. & Fabre, E. 3D genome organization: causes and consequences for DNA damage and repair. Genes 13, 7 (2022).
Zhu, Y. et al. Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription. Cancer Cell 39, 694–707 (2021).
Deshpande, V. et al. Exploring the landscape of focal amplifications in cancer using AmpliconArchitect. Nat. Commun. 10, 392 (2019).
Luebeck, J. et al. AmpliconReconstructor integrates NGS and optical mapping to resolve the complex structures of focal amplifications. Nat. Commun. 11, 4374 (2020).
Xu, K. et al. Structure and evolution of double minutes in diagnosis and relapse brain tumors. Acta Neuropathol. 137, 123–137 (2019).
L’Abbate, A. et al. Genomic organization and evolution of double minutes/homogeneously staining regions with MYC amplification in human cancer. Nucleic Acids Res. 42, 9131–9145 (2014).
Shoshani, O. et al. Chromothripsis drives the evolution of gene amplification in cancer. Nature 591, 137–141 (2021).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Uslu, V. V. et al. Long-range enhancers regulating Myc expression are required for normal facial morphogenesis. Nat. Genet. 46, 753–758 (2014).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Schoenfelder, S. & Fraser, P. Long-range enhancer-promoter contacts in gene expression control. Nat. Rev. Genet. 20, 437–455 (2019).
Pang, J. et al. Extrachromosomal DNA in HPV mediated oropharyngeal cancer drives diverse oncogene transcription. Clin. Cancer Res. 27, 6772–6786 (2021).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Weischenfeldt, J. et al. Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking. Nat. Genet. 49, 65–74 (2017).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Schwab, M., Klempnauer, K. H., Alitalo, K., Varmus, H. & Bishop, M. Rearrangement at the 5′ end of amplified c-myc in human COLO 320 cells is associated with abnormal transcription. Mol. Cell. Biol. 6, 2752–2755 (1986).
Cho, S. W. et al. Promoter of lncRNA gene PVT1 is a tumor-suppressor DNA boundary element. Cell 173, 1398–1412.e22 (2018).
Northcott, P. A. et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488, 49–56 (2012).
Tolomeo, D., Agostini, A., Visci, G., Traversa, D. & Storlazzi, C. T. PVT1: a long non-coding RNA recurrently involved in neoplasia-associated fusion transcripts. Gene 779, 145497 (2021).
Kalyana-Sundaram, S. et al. Gene fusions associated with recurrent amplicons represent a class of passenger aberrations in breast cancer. Neoplasia 14, 702–708 (2012).
Robinson, D. R. et al. Integrative clinical genomics of metastatic cancer. Nature 548, 297–303 (2017).
Chapman, O. S. et al. The landscape of extrachromosomal circular DNA in medulloblastoma. Preprint at bioRxiv https://doi.org/10.1101/2021.10.18.464907 (2021).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Terakawa, T. et al. The condensin complex is a mechanochemical motor that translocates along DNA. Science 358, 672–676 (2017).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Hamkalo, B. A., Farnham, P. J., Johnston, R. & Schimke, R. T. Ultrastructural features of minute chromosomes in a methotrexate-resistant mouse 3T3 cell line. Proc. Natl Acad. Sci. USA 82, 1126–1130 (1985).
van der Bliek, A. M., Lincke, C. R. & Borst, P. Circular DNA of 3T6R50 double minute chromosomes. Nucleic Acids Res. 16, 4841–4851 (1988).
Rattner, J. B. & Lin, C. C. Ultrastructural organization of double minute chromosomes and HSR regions in human colon carcinoma cells. Cytogenetic Genome Res. 38, 176–181 (1984).
VanDevanter, D. R., Piaskowski, V. D., Casper, J. T., Douglass, E. C. & Von Hoff, D. D. Ability of circular extrachromosomal DNA molecules to carry amplified MYCN protooncogenes in human neuroblastomas in vivo. J. Natl Cancer Inst. 82, 1815–1821 (1990).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Hung, K. L. et al. Targeted profiling of human extrachromosomal DNA by CRISPR–CATCH. Preprint at bioRxiv https://doi.org/10.1101/2021.11.28.470285 (2022).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602–1612 (2017).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Zheng, M. et al. Multiplex chromatin interactions with single-molecule precision. Nature 566, 558–562 (2019).
Lareau, C. A. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol. 37, 916–924 (2019).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183, 1103–1116 (2020).
Koche, R. P. et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat. Genet. 52, 29–34 (2019).
Jiang, W. et al. Cas9-assisted targeting of chromosome segments CATCH enables one-step targeted cloning of large gene clusters. Nat. Commun. 6, 1–8 (2015).
Overhauser, J. in Pulsed-Field Gel Electrophoresis: Protocols, Methods, and Theories (eds. Burmeister, M. & Ulanovsky, L.) 129–134 (Humana Press, 1992).
Cao, H. et al. Rapid detection of structural variation in a human genome using nanochannel-based genome mapping technology. GigaScience 3, 34 (2014).
Baskin, F., Rosenberg, R. N. & Dev, V. Correlation of double-minute chromosomes with unstable multidrug cross-resistance in uptake mutants of neuroblastoma cells. Proc. Natl Acad. Sci. USA 78, 3654–3658 (1981).
Møller, H. D., Parsons, L., Jørgensen, T. S., Botstein, D. & Regenberg, B. Extrachromosomal circular DNA is common in yeast. Proc. Natl Acad. Sci. USA 112, E3114–E3122 (2015).
Shoura, M. J. et al. Intricate and cell type-specific populations of endogenous circular DNA (eccDNA) in Caenorhabditis elegans and Homo sapiens. G3 7, 3295–3303 (2017).
Smith, C. A. & Vinograd, J. Small polydisperse circular DNA of HeLa cells. J. Mol. Biol. 69, 163–178 (1972).
Acknowledgements
H.Y.C. is supported by US National Institutes of Health grant R35-CA209919 and is an Investigator of the Howard Hughes Medical Institute. P.S.M. is supported in part by grants U24CA264379 and R01 CA238249 from the US National Institutes of Health. H.Y.C. and P.S.M. are supported by Cancer Grand Challenges CGCSDF-2021\100007 with support from Cancer Research UK and the National Cancer Institute. K.L.H. is supported by a Stanford Graduate Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Anton Henssen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Beth Moorefield, Tiago Faial and Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hung, K.L., Mischel, P.S. & Chang, H.Y. Gene regulation on extrachromosomal DNA. Nat Struct Mol Biol 29, 736–744 (2022). https://doi.org/10.1038/s41594-022-00806-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00806-7
This article is cited by
-
Extrachromosomal DNA in cancer
Nature Reviews Cancer (2024)
-
Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
Nature Communications (2024)
-
ATACAmp: a tool for detecting ecDNA/HSRs from bulk and single-cell ATAC-seq data
BMC Genomics (2023)
-
Extrachromosomal circular DNA in colorectal cancer: biogenesis, function and potential as therapeutic target
Oncogene (2023)
-
Circlehunter: a tool to identify extrachromosomal circular DNA from ATAC-Seq data
Oncogenesis (2023)