Abstract
Precise generation of excitatory neurons and inhibitory interneurons is crucial for proper formation and function of neural circuits in the mammalian brain. Because of the size and complexity of the human brain, it is a challenge to reveal the rich diversity of interneurons. To decipher origin and diversity of interneurons in the human fetal subpallium, here we show molecular features of diverse subtypes of interneuron progenitors and precursors by conducting single-cell RNA sequencing and in situ sequencing. Interneuron precursors in the medial and lateral ganglionic eminence simultaneously procure temporal and spatial identity through expressing a combination of specific sets of RNA transcripts. Acquisition of various interneuron subtypes in adult human brains occurs even at fetal stages. Our study uncovers complex molecular signatures of interneuron progenitors and precursors in the human fetal subpallium and highlights the logic and programs in the origin and lineage specification of various interneurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw single-cell sequencing data are available in the Gene Expression Omnibus at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE165388, with the accession code GSE165388. All images of ISS are available upon request at https://github.com/yuanbell/Single-cell-sequencing-of-subpallium.
Code availability
All codes generated during this study are available upon request at https://github.com/yuanbell/Single-cell-sequencing-of-subpallium.
References
Tremblay, R., Lee, S. & Rudy, B. GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91, 260–292 (2016).
Lim, L., Mi, D., Llorca, A. & Marin, O. Development and functional diversification of cortical interneurons. Neuron 100, 294–313 (2018).
Marín, O. Interneuron dysfunction in psychiatric disorders. Nat. Rev. Neurosci. 13, 107–120 (2012).
Pauly, M. C., Dobrossy, M. D., Nikkhah, G., Winkler, C. & Piroth, T. Organization of the human fetal subpallium. Front. Neuroanat. 7, 54 (2013).
Silberberg, S. N. et al. Subpallial enhancer transgenic lines: a data and tool resource to study transcriptional regulation of GABAergic cell fate. Neuron 92, 59–74 (2016).
Nery, S., Fishell, G. & Corbin, J. G. The caudal ganglionic eminence is a source of distinct cortical and subcortical cell populations. Nat. Neurosci. 5, 1279–1287 (2002).
Hu, J. S. et al. Coup-TF1 and Coup-TF2 control subtype and laminar identity of MGE-derived neocortical interneurons. Development 144, 2837–2851 (2017).
Bandler, R. C., Mayer, C. & Fishell, G. Cortical interneuron specification: the juncture of genes, time and geometry. Curr. Opin. Neurobiol. 42, 17–24 (2017).
Xu, Q., Tam, M. & Anderson, S. A. Fate mapping Nkx2.1-lineage cells in the mouse telencephalon. J. Comp. Neurol. 506, 16–29 (2008).
Su-Feher, L. et al. Single cell enhancer activity maps neuronal lineages in embryonic mouse basal ganglia. Preprint at https://www.biorxiv.org/content/10.1101/2021.01.11.426285v1.full (2021).
Flames, N. et al. Delineation of multiple subpallial progenitor domains by the combinatorial expression of transcriptional codes. J. Neurosci. 27, 9682–9695 (2007).
Hu, J. S., Vogt, D., Sandberg, M. & Rubenstein, J. L. Cortical interneuron development: a tale of time and space. Development 144, 3867–3878 (2017).
Tosches, M. A. et al. Evolution of pallium, hippocampus, and cortical cell types revealed by single-cell transcriptomics in reptiles. Science 360, 881–888 (2018).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020).
Ma, T. et al. Subcortical origins of human and monkey neocortical interneurons. Nat. Neurosci. 16, 1588–1597 (2013).
Hansen, D. V. et al. Non-epithelial stem cells and cortical interneuron production in the human ganglionic eminences. Nat. Neurosci. 16, 1576–1587 (2013).
Boldog, E. et al. Transcriptomic and morphophysiological evidence for a specialized human cortical GABAergic cell type. Nat. Neurosci. 21, 1185–1195 (2018).
Close, J. L. et al. Single-cell profiling of an in vitro model of human interneuron development reveals temporal dynamics of cell type production and maturation. Neuron 93, 1035–1048 (2017).
Sun, T. & Hevner, R. F. Growth and folding of the mammalian cerebral cortex: from molecules to malformations. Nat. Rev. Neurosci. 15, 217–232 (2014).
Fan, X. et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res. 28, 730–745 (2018).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Zeng, Z., Miao, N. & Sun, T. Revealing cellular and molecular complexity of the central nervous system using single cell sequencing. Stem Cell Res. Ther. 9, 234 (2018).
O’Rahilly, R. & Müller, F. The Embryonic Human Brain: An Atlas of Developmental Stages, Third Edition, 1–358 (Wiley, 2005).
Clancy, B., Darlington, R. B. & Finlay, B. L. Translating developmental time across mammalian species. Neuroscience 105, 7–17 (2001).
Otis, E. M. & Brent, R. Equivalent ages in mouse and human embryos. Anat. Rec. 120, 33–63 (1954).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318 (2017).
Mayer, C. et al. Developmental diversification of cortical inhibitory interneurons. Nature 555, 457–462 (2018).
Mi, D. et al. Early emergence of cortical interneuron diversity in the mouse embryo. Science 360, 81–85 (2018).
Harkin, L. F. et al. Distinct expression patterns for type II topoisomerases IIA and IIB in the early foetal human telencephalon. J. Anat. 228, 452–463 (2016).
Bansod, S., Kageyama, R. & Ohtsuka, T. Hes5 regulates the transition timing of neurogenesis and gliogenesis in mammalian neocortical development. Development 144, 3156–3167 (2017).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Zhao, Q. et al. Single-cell transcriptome analyses reveal endothelial cell heterogeneity in tumors and changes following antiangiogenic treatment. Cancer Res. 78, 2370–2382 (2018).
Petros, T. J., Bultje, R. S., Ross, M. E., Fishell, G. & Anderson, S. A. Apical versus basal neurogenesis directs cortical interneuron subclass fate. Cell Rep. 13, 1090–1095 (2015).
Lindtner, S. et al. Genomic resolution of DLX-orchestrated transcriptional circuits driving development of forebrain GABAergic neurons. Cell Rep. 28, 2048–2063 (2019).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Alzu’bi, A. et al. The transcription factors COUP-TFI and COUP-TFII have distinct roles in arealisation and GABAergic interneuron specification in the early human fetal telencephalon. Cereb. Cortex 27, 4971–4987 (2017).
Butt, S. J. et al. The temporal and spatial origins of cortical interneurons predict their physiological subtype. Neuron 48, 591–604 (2005).
Kanatani, S., Yozu, M., Tabata, H. & Nakajima, K. COUP-TFII is preferentially expressed in the caudal ganglionic eminence and is involved in the caudal migratory stream. J. Neurosci. 28, 13582–13591 (2008).
Rudy, B., Fishell, G., Lee, S. & Hjerling-Leffler, J. Three groups of interneurons account for nearly 100% of neocortical GABAergic neurons. Dev. Neurobiol. 71, 45–61 (2011).
Kepecs, A. & Fishell, G. Interneuron cell types are fit to function. Nature 505, 318–326 (2014).
Turrero Garcia, M. & Harwell, C. C. Radial glia in the ventral telencephalon. FEBS Lett. 591, 3942–3959 (2017).
Silva, T. P. et al. Transcriptome profiling of human pluripotent stem cell-derived cerebellar organoids reveals faster commitment under dynamic conditions. Biotechnol. Bioeng. 118, 2781–2803 (2021).
Enterría-Morales, D. et al. Molecular targets for endogenous glial cell line-derived neurotrophic factor modulation in striatal parvalbumin interneurons. Brain Commun. 2, fcaa105 (2020).
Flandin, P., Kimura, S. & Rubenstein, J. L. The progenitor zone of the ventral medial ganglionic eminence requires Nkx2-1 to generate most of the globus pallidus but few neocortical interneurons. J. Neurosci. 30, 2812–2823 (2010).
Tiveron, M. C. et al. Zic-proteins are repressors of dopaminergic forebrain fate in mice and C. elegans. J. Neurosci. 37, 10611–10623 (2017).
Sousa, V. H., Miyoshi, G., Hjerling-Leffler, J., Karayannis, T. & Fishell, G. Characterization of Nkx6-2-derived neocortical interneuron lineages. Cereb. Cortex 19 i1–i10 (2009).
Lu, K. M., Evans, S. M., Hirano, S. & Liu, F. C. Dual role for Islet-1 in promoting striatonigral and repressing striatopallidal genetic programs to specify striatonigral cell identity. Proc. Natl Acad. Sci. USA 111, E168–E177 (2014).
Li, J. et al. Transcription factors Sp8 and Sp9 coordinately regulate olfactory bulb interneuron development. Cereb. Cortex 28, 3278–3294 (2018).
O’Leary, D. D. & Nakagawa, Y. Patterning centers, regulatory genes and extrinsic mechanisms controlling arealization of the neocortex. Curr. Opin. Neurobiol. 12, 14–25 (2002).
Xu, Q. et al. Sonic hedgehog signaling confers ventral telencephalic progenitors with distinct cortical interneuron fates. Neuron 65, 328–340 (2010).
Ma, T. et al. A subpopulation of dorsal lateral/caudal ganglionic eminence-derived neocortical interneurons expresses the transcription factor Sp8. Cereb. Cortex 22, 2120–2130 (2012).
Torigoe, M., Yamauchi, K., Kimura, T., Uemura, Y. & Murakami, F. Evidence that the laminar fate of LGE/CGE-derived neocortical interneurons is dependent on their progenitor domains. J. Neurosci. 36, 2044–2056 (2016).
Rubin, A. N. et al. The germinal zones of the basal ganglia but not the septum generate GABAergic interneurons for the cortex. J. Neurosci. 30, 12050–12062 (2010).
Hern, W. M. Correlation of fetal age and measurements between 10 and 26 weeks of gestation. Obstet. Gynecol. 63, 26–32 (1984).
Gayoso, A., Shor, J., Carr, A. J., Sharma, R. & Pe’er, D. JonathanShor/DoubletDetection: HOTFIX: correct setup.py installation. Zenodo (2019). https://doi.org/10.5281/zenodo.3376859
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Stevant, I. et al. Dissecting cell lineage specification and sex fate determination in gonadal somatic cells using single-cell transcriptomics. Cell Rep. 26, 3272–3283 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Korotkevich, G., Sukhov, V. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at https://www.biorxiv.org/content/10.1101/060012v3 (2021).
Acknowledgements
This study was supported by the Scientific Research Funds of Huaqiao University (Z16Y0017 to T.S.), the Fundamental Research Funds for the Central Universities (ZQN-715 to Y.S.), the Innovation Awards of Quanzhou Talents (2018C057R to Y.C.), the Natural Science Foundation of Fujian Province of China (2018J01585 to S.H.), the National Key Research and Development Program of China (2017YFA0106800 to R.K.) and the National Natural Science Foundation of China (31771141 to T.S., 11802096 to Y.S. and 31770927 to R.K.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Y.Y. and T.S. conceived and designed the study. Y.Y. performed the single-cell sequencing experiments. Y.Y., Z.Z., R.C. and S.H. analyzed sequencing data. Y.Y., D.X. and R.K. performed the ISS experiment. Y.S., W.J.C., W.H.C. and W.L. prepared brain tissues. Y.Y. and T.S. wrote the paper. T.S. supervised the entire study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-cell transcriptomic maps of the human fetal subpallium.
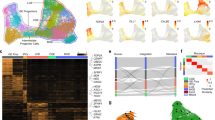
a, Cell clustering of human fetal subpallial samples at gestational weeks (GW) 9 to 12 integrated using harmony and depicted using t-SNE. b, The distribution of each subclusters in the subpallium at four developing stages. Colors indicate cell clusters as shown in a. c, Clustering of all cells from four developing stages after batch correction visualized using t-SNE. d, Heatmap illustrating differentially expressed genes (DEGs) in five major clusters and 17 subclusters. e, Assessment of 71 genes from 2,922 DEGs using Random Forest Classifier from scikit-learn of the remaining 16 subclusters after removing excitatory lineages. ‘True label’ indicates the manual annotation based on the 71 genes.
Extended Data Fig. 2 Expression patterns of canonical genes from each cell types.
a-f, Representative genes expressed in cell clusters from human fetal subpallium visualized using t-SNE. Each dot represents one cell. Major clusters include excitatory lineages (a), neural progenitor cells (NPCs, b) and interneuron precursors (INPs, c) derived from the medial ganglionic eminence (MGE_INPs, d), caudal ganglionic eminence (CGE_INPs, e) and lateral ganglionic eminence (LGE_INPs, f).
Extended Data Fig. 3 Heterogeneity of interneuron progenitors in the human fetal subpallium.
a-c, Expression of DLX2 and GAD2 at gestational weeks (GW) 9 (a), GW11 (b), GW12 (c), visualized by the t-SNE plot. d, Violin plots of expression patterns of progenitor markers expressed at the ventricular zone (VZ) and subventricular zone (SVZ) in the subpallium at GW12. e, Violin plots of expression patterns of ganglionic eminence regional markers in the progenitors from GW9 to GW12.
Extended Data Fig. 4 Expression patterns of subpallial genes in the ganglionic eminence of the human fetal brain as detected by in situ sequencing (ISS).
a and b, Expression patterns of 6 subpallial genes in the medial ganglionic eminence (MGE) (a), lateral ganglionic eminence (LGE) (a) and caudal ganglionic eminence (CGE) (b) at gestational week 12 (GW12). c-e, Boxed areas in a and b are shown in high power views. Scale bars: 1 mm in a and b; 100 µm in c-e.
Extended Data Fig. 5 Expression patterns of genes in the subpallium of the human fetal brain as detected by in situ sequencing (ISS).
a-d, Expression patterns of genes expressed in the ventricular zone (VZ) and subventricular zone (SVZ) at gestational week 12 (GW12) are shown in green and purple pseudo-colors, respectively, in the medial ganglionic eminence (MGE), lateral ganglionic eminence (LGE) and caudal ganglionic eminence (CGE). Scale bars: 1 mm.
Extended Data Fig. 6 Expression patterns of genes expressed in the ventricular zone (VZ) in the subpallium of the human fetal brain as detected by in situ sequencing (ISS).
a, Merged expression of CLU, LIX1, PTN and SPARC in the medial ganglionic eminence (MGE), lateral ganglionic eminence (LGE) and caudal ganglionic eminence (CGE) at gestational week 12 (GW12). b-e, Expression of CLU (b), LIX1 (c), PTN (d) and SPARC (e) in the MGE, LGE and CGE. Scale bars: 100 µm.
Extended Data Fig. 7 Specification of medial ganglionic eminence (MGE)-derived interneuron precursors.
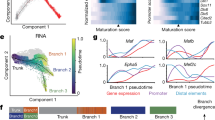
a, Clustering of MGE-derived interneuron precursors from GW9 to GW12 visualized by t-SNE embedding. Cluster 5, 1, 2, 4 and 7 were similar to ANGPT2+/CRABP1+, ZEB2+/MAF+, POU3F2+/CNTNAP2+, NR2F1+/MEIS2+ and LHX8+/NKX2-1+, respectively, in the Fig. 4a, cluster 3, 6, 8 were raised from the GW12. b, Heatmap of differentially expressed genes from subclusters in a. c, Diffusion map of the most variable genes and reconstruction of the cell lineages. Dots represent cells, and black lines represent predicted cell lineage-1 and -2. As examples, the cluster 6 was predicted to the lineage 1 and the cluster 3 to the lineage 2. d. Expression patterns of genes highly expressed in the dorsal (MEIS2 and NR2F1) and ventral (LHX8 and ZIC4) MGE in human fetal brains at gestational week 12 (GW12) as detected using in situ sequencing (ISS). e, Box plots representing relative numbers of positive dots of MGE marker genes in boxed areas in the dorsal and ventral MGE in d. Box: 25–75th percentiles, whiskers: 10–90th percentiles, horizontal line in box: median. Scale bars: 1 mm.
Extended Data Fig. 8 Expression patterns of medial ganglionic eminence (MGE) specific genes of the human fetal brain as detected by in situ sequencing (ISS).
a, Coronal sections labeled with DAPI to illustrate cell nucleoli of a human fetal brain at gestational week 12 (GW12). Boxed areas highlight the MGE. The dorsal (D), ventral (V), medial (M) and lateral (L) orientations of the sections are labeled. b, High expression of CNTNAP2 and MAF at the root of lineage-1 and -2 based on the diffusion map in lineage analyses. c-e, Expression patterns of 10 MGE-precursor marker genes in coronal sections of human fetal brains. The dorsal and ventral MGEs are labeled as dMGE (c) and vMGE (d). Scale bars: 1 mm.
Extended Data Fig. 9 Lateral ganglionic eminence (LGE)-derived interneuron precursors.
a and b, Expression patterns of 8 LGE specific genes in coronal sections of the human fetal brain at gestational week 12 (GW12) as detected by in situ sequencing (ISS). The ventral (green box) and dorsal (blue box) LGE are shown as vLGE and dLGE. The boxed areas also are shown in high power views. Scale bars: 1 mm and 100 µm.
Extended Data Fig. 10 Caudal ganglionic eminence (CGE)-derived interneuron precursors.
a, The transcription network regulated by CGE-specific transcription factors in interneuron precursors. b, Gene ontology (GO) enrichment analysis of target genes for CGE-specific transcription factors. c, A coronal section labeled with DAPI to illustrate cell nucleoli of one human fetal brain at gestational week 12 (GW12). d, Expression patterns of CALB2, NPAS3, ST18 and SP9 in the CGE in coronal sections of the human fetal brain at GW12, as detected by in situ sequencing (ISS). Scale bar: 1 mm.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5
Supplementary Tables
Supplementary Tables 1–18
Rights and permissions
About this article
Cite this article
Yu, Y., Zeng, Z., Xie, D. et al. Interneuron origin and molecular diversity in the human fetal brain. Nat Neurosci 24, 1745–1756 (2021). https://doi.org/10.1038/s41593-021-00940-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00940-3
This article is cited by
-
Drug targeting in psychiatric disorders — how to overcome the loss in translation?
Nature Reviews Drug Discovery (2024)
-
Deciphering Oligodendrocyte Lineages in the Human Fetal Central Nervous System Using Single-Cell RNA Sequencing
Molecular Neurobiology (2024)
-
Integrated neural tracing and in-situ barcoded sequencing reveals the logic of SCN efferent circuits in regulating circadian behaviors
Science China Life Sciences (2024)
-
Identifying foetal forebrain interneurons as a target for monogenic autism risk factors and the polygenic 16p11.2 microdeletion
BMC Neuroscience (2023)
-
Insights into Alzheimer’s disease from single-cell genomic approaches
Nature Neuroscience (2023)