Abstract
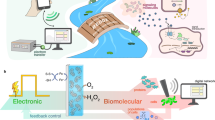
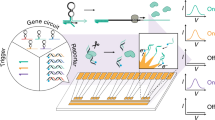
We developed a bioelectronic communication system that is enabled by a redox signal transduction modality to exchange information between a living cell-embedded bioelectronics interface and an engineered microbial network. A naturally communicating three-member microbial network is ‘plugged into’ an external electronic system that interrogates and controls biological function in real time. First, electrode-generated redox molecules are programmed to activate gene expression in an engineered population of electrode-attached bacterial cells, effectively creating a living transducer electrode. These cells interpret and translate electronic signals and then transmit this information biologically by producing quorum sensing molecules that are, in turn, interpreted by a planktonic coculture. The propagated molecular communication drives expression and secretion of a therapeutic peptide from one strain and simultaneously enables direct electronic feedback from the second strain, thus enabling real-time electronic verification of biological signal propagation. Overall, we show how this multifunctional bioelectronic platform, termed a BioLAN, reliably facilitates on-demand bioelectronic communication and concurrently performs programmed tasks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets that support the findings of this study are available at https://figshare.com/s/30bcc0241826827d12f4. Source data are provided with this paper.
Code availability
The MATLAB code for the models used in this study is available from the corresponding author upon request.
References
Gubbi, J., Buyya, R., Marusic, S. & Palaniswami, M. Internet of Things (IoT): a vision, architectural elements, and future directions. Future Gener. Comput. Syst. 29, 1645–1660 (2013).
Tian, B. et al. Roadmap on semiconductor–cell biointerfaces. Phys. Biol. 15, 031002 (2018).
Atkinson, J. T. et al. Metalloprotein switches that display chemical-dependent electron transfer in cells. Nat. Chem. Biol. 15, 189–195 (2018).
Bird, L. J. et al. Engineered living conductive biofilms as functional materials. MRS Commun. 9, 505–517 (2019).
Shao, J. et al. Smartphone-controlled optogenetically engineered cells enable semiautomatic glucose homeostasis in diabetic mice. Sci. Transl. Med. 9, eaal2298 (2017).
Mimee, M. et al. An ingestible bacterial–electronic system to monitor gastrointestinal health. Science 360, 915–918 (2018).
Yang, X. et al. Bioinspired neuron-like electronics. Nat. Mater. 18, 510–517 (2019).
Tschirhart, T. et al. Electronic control of gene expression and cell behaviour in Escherichia coli through redox signalling. Nat. Commun. 8, 14030 (2017).
Bhokisham, N. et al. A redox-based electrogenetic CRISPR system to connect with and control biological information networks. Nat. Commun. 11, 2427 (2020).
Sadat Mousavi, P. et al. A multiplexed, electrochemical interface for gene-circuit-based sensors. Nat. Chem. 12, 48–55 (2020).
Liu, Y. et al. Connecting biology to electronics: molecular communication via redox modality. Adv. Healthc. Mater. 6, 1700789 (2017).
Lee, H. et al. Wearable/disposable sweat-based glucose monitoring device with multistage transdermal drug delivery module. Sci. Adv. 3, e1601314 (2017).
Wang, S., Payne, G. F. & Bentley, W. E. in Gene Expression and Control (ed. Uchiumi, F.) Ch. 9 (IntechOpen, 2019).
Green, J. & Paget, M. S. Bacterial redox sensors. Nat. Rev. Microbiol. 2, 954–966 (2004).
Hirose, A., Kouzuma, A. & Watanabe, K. Towards development of electrogenetics using electrochemically active bacteria. Biotechnol. Adv. 37, 107351 (2019).
McCarty, N. S. & Ledesma-Amaro, R. Synthetic biology tools to engineer microbial communities for biotechnology. Trends Biotechnol. 37, 181–197 (2019).
Lindemann, S. R. et al. Engineering microbial consortia for controllable outputs. ISME J. 10, 2077–2084 (2016).
Pomposiello, P. J. & Demple, B. Redox-operated genetic switches: the SoxR and OxyR transcription factors. Trends Biotechnol. 19, 109–114 (2001).
Liu, Y. et al. Using a redox modality to connect synthetic biology to electronics: hydrogel-based chemo-electro signal transduction for molecular communication. Adv. Healthc. Mater. 6, 1600908 (2017).
VanArsdale, E. et al. Redox-based synthetic biology enables electrochemical detection of the herbicides dicamba and Roundup via rewired Escherichia coli. ACS Sens. 4, 1180–1184 (2019).
McKay, R. et al. A platform of genetically engineered bacteria as vehicles for localized delivery of therapeutics: toward applications for Crohn’s disease. Bioeng. Transl. Med. 3, 209–221 (2018).
Zheng, M., Aslund, F. & Storz, G. Activation of the OxyR transcription factor by reversible disulfide bond formation. Science 279, 1718–1721 (1998).
Kim, S. O. et al. OxyR: a molecular code for redox-related signaling. Cell 109, 383–396 (2002).
Aslund, F., Zheng, M., Beckwith, J. & Storz, G. Regulation of the OxyR transcription factor by hydrogen peroxide and the cellular thiol-disulfide status. Proc. Natl Acad. Sci. USA 96, 6161–6165 (1999).
Demple, B. & Halbrook, J. Inducible repair of oxidative DNA damage in Escherichia coli. Nature 304, 466–468 (1983).
Sultana, S. T. et al. Electrochemical scaffold generates localized, low concentration of hydrogen peroxide that inhibits bacterial pathogens and biofilms. Sci. Rep. 5, 14908 (2015).
Rubens, J. R., Selvaggio, G. & Lu, T. K. Synthetic mixed-signal computation in living cells. Nat. Commun. 7, 11658 (2016).
Virgile, C. et al. Engineering bacterial motility towards hydrogen-peroxide. PLoS ONE 13, e0196999 (2018).
Hornstrom, D., Larsson, G., van Maris, A. J. A. & Gustavsson, M. Molecular optimization of autotransporter-based tyrosinase surface display. Biochim. Biophys. Acta Biomembr. 1861, 486–494 (2019).
Terrell, J. L. et al. Nano-guided cell networks as conveyors of molecular communication. Nat. Commun. 6, 8500 (2015).
Brown, S. Metal-recognition by repeating polypeptides. Nat. Biotechnol. 15, 269–272 (1997).
Tamerler, C. et al. Materials specificity and directed assembly of a gold-binding peptide. Small 2, 1372–1378 (2006).
Verde, A. V., Acres, J. M. & Maranas, J. K. Investigating the specificity of peptide adsorption on gold using molecular dynamics simulations. Biomacromolecules 10, 2118–2128 (2009).
Medintz, I. L. et al. Quantum-dot/dopamine bioconjugates function as redox coupled assemblies for in vitro and intracellular pH sensing. Nat. Mater. 9, 676–684 (2010).
Tschirhart, T. et al. Electrochemical measurement of the beta-galactosidase reporter from live cells: a comparison to the Miller assay. ACS Synth. Biol. 5, 28–35 (2016).
DeLisa, M. P., Wu, C. F., Wang, L., Valdes, J. J. & Bentley, W. E. DNA microarray-based identification of genes controlled by autoinducer 2-stimulated quorum sensing in Escherichia coli. J. Bacteriol. 183, 5239–5247 (2001).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Servinsky, M. D. et al. Directed assembly of a bacterial quorum. ISME J. 10, 158–169 (2016).
Shang, W. et al. Selective assembly and functionalization of miniaturized redox capacitor inside microdevices for microbial toxin and mammalian cell cytotoxicity analyses. Lab Chip 18, 3578–3587 (2018).
Pennacchio, F. A., Garma, L. D., Matino, L. & Santoro, F. Bioelectronics goes 3D: new trends in cell–chip interface engineering. J. Mater. Chem. B 6, 7096–7101 (2018).
Yuk, H., Lu, B. & Zhao, X. Hydrogel bioelectronics. Chem. Soc. Rev. 48, 1642–1667 (2019).
Kang, M. et al. Signal processing approach to probe chemical space for discriminating redox signatures. Biosens. Bioelectron. 112, 127–135 (2018).
Hwang, I. Y. et al. Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models. Nat. Commun. 8, 15028 (2017).
Sun, F., Zhang, W.-B., Mahdavi, A., Arnold, F. H. & Tirrell, D. A. Synthesis of bioactive protein hydrogels by genetically encoded SpyTag–SpyCatcher chemistry. Proc. Natl Acad. Sci. USA 111, 11269 (2014).
French, K. E., Zhou, Z. & Terry, N. Horizontal ‘gene drives’ harness indigenous bacteria for bioremediation. Sci. Rep. 10, 15091 (2020).
Jiang, Y., Dong, W., Xin, F. & Jiang, M. Designing synthetic microbial consortia for biofuel production. Trends Biotechnol. 38, 828–831 (2020).
Jawed, K., Yazdani, S. S. & Koffas, M. A. G. Advances in the development and application of microbial consortia for metabolic engineering. Metab. Eng. Commun. 9, e00095 (2019).
Alper, H. S. & Avalos, J. L. Metabolic pathway engineering. Synth. Syst. Biotechnol. 3, 1–2 (2018).
Stephens, K., Pozo, M., Tsao, C. Y., Hauk, P. & Bentley, W. E. Bacterial co-culture with cell signaling translator and growth controller modules for autonomously regulated culture composition. Nat. Commun. 10, 4129 (2019).
Beardslee, L. A. et al. Ingestible sensors and sensing systems for minimally invasive diagnosis and monitoring: the next frontier in minimally invasive screening. ACS Sens. 5, 891–910 (2020).
Windmiller, J. R. et al. Electrochemical sensing based on printable temporary transfer tattoos. Chem. Commun. 48, 6794–6796 (2012).
Akyildiz, I. F., Pierobon, M., Balasubramaniam, S. & Koucheryavy, Y. The Internet of Bio-Nano Things. IEEE Commun. Mag. 53, 32–40 (2015).
Hall, B. G., Acar, H., Nandipati, A. & Barlow, M. Growth rates made easy. Mol. Biol. Evol. 31, 232–238 (2014).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Gawarzewski, I. et al. Crystal structure of the transport unit of the autotransporter adhesin involved in diffuse adherence from Escherichia coli. J. Struct. Biol. 187, 20–29 (2014).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Dong, H. et al. Living bacteria–nanoparticle hybrids mediated through surface-displayed peptides. Langmuir 34, 5837–5848 (2018).
Acknowledgements
Partial support of this work was provided by DTRA (HDTRA1-19-0021), NSF (DMREF 1435957, ECCS 1807604, CBET 1805274), the National Institutes of Health (R21EB024102) and the Office of Naval Research (N0001417WX01318, N0001418WX01042). This work was also supported by the Office of the Under Secretary of Defense for Research and Engineering (USD(R&E)) through the Applied Research for Advancement of S&T Priorities (ARAP) Program on Synthetic Biology for Military Environments (SBME).
Author information
Authors and Affiliations
Contributions
J.L.T., T.T., Y.L., C.Y.T., H.-C.W., G.V., G.F.P., D.N.S.-C. and W.E.B. were involved with the conception and design of the work. J.L.T., T.T., K.S., R.M. and M.P. were involved with engineering strains used in this work. J.L.T., T.T., J.P.J., K.S. and H.D. were involved with data acquisition and interpretation. J.P.J. and M.M.H. were involved with computational kinetic studies and protein modelling, respectively. J.L.T., T.T., J.P.J., G.F.P., D.N.S.-C. and W.E.B. were involved with writing and documentation of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Nanotechnology thanks Ada Poon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Figs. 1–15, Tables 1–8 and references.
Source data
Source Data Fig. 2
Statistical source data and R code.
Source Data Fig. 3
Model data and corresponding MATLAB-generated figures.
Source Data Fig. 4
Protein sequence, unprocessed images, statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Terrell, J.L., Tschirhart, T., Jahnke, J.P. et al. Bioelectronic control of a microbial community using surface-assembled electrogenetic cells to route signals. Nat. Nanotechnol. 16, 688–697 (2021). https://doi.org/10.1038/s41565-021-00878-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-021-00878-4
This article is cited by
-
Synthetic microbiology in sustainability applications
Nature Reviews Microbiology (2024)
-
Redox-enabled electronic interrogation and feedback control of hierarchical and networked biological systems
Nature Communications (2023)
-
A versatile bioelectronic interface programmed for hormone sensing
Nature Communications (2023)
-
An electrogenetic interface to program mammalian gene expression by direct current
Nature Metabolism (2023)
-
Redox signaling-driven modulation of microbial biosynthesis and biocatalysis
Nature Communications (2023)