Abstract
The coronavirus SARS-CoV-2 causes the severe disease COVID-19. SARS-CoV-2 infection is initiated by interaction of the viral spike protein and host receptor angiotensin-converting enzyme 2 (ACE2). We report an improved bright and reversible fluorogenic reporter, named SURF (split UnaG-based reversible and fluorogenic protein–protein interaction reporter), that we apply to monitor real-time interactions between spike and ACE2 in living cells. SURF has a large dynamic range with a dark-to-bright fluorescence signal that requires no exogenous cofactors. Utilizing this reporter, we carried out a high-throughput screening of small-molecule libraries. We identified three natural compounds that block replication of SARS-CoV-2 in both Vero cells and human primary nasal and bronchial epithelial cells. Cell biological and biochemical experiments validated all three compounds and showed that they block the early stages of viral infection. Two of the inhibitors, bruceine A and gamabufotalin, were also found to block replication of the Delta and Omicron variants of SARS-CoV-2. Both bruceine A and gamabufotalin exhibited potent antiviral activity in K18-hACE2 and wild-type C57BL6/J mice, as evidenced by reduced viral titres in the lung and brain, and protection from alveolar and peribronchial inflammation in the lung, thereby limiting disease progression. We propose that our fluorescent assay can be applied to identify antiviral compounds with potential as therapeutic treatment for COVID-19 and other respiratory diseases.
Similar content being viewed by others
Main
The worldwide pandemic of the respiratory disease COVID-19, caused by the coronavirus SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2), has produced huge health challenges and economic loss. While effective vaccines and therapeutic monoclonal antibodies are available, a large part of the global population remains unvaccinated and without access to monoclonal therapies due to their cost and limited supply1,2. Moreover, immune escape mutations in the spike (S) protein may weaken the efficacy of current interventions3. Thus, small-molecule drugs are urgently needed to curb progression of the COVID-19 pandemic.
The first step in the entry of SARS-CoV-2 into human cells is viral binding to the host cell, which is achieved by protein–protein interaction (PPI) between the virus surface-anchored S protein and the host cell receptor angiotensin-converting enzyme 2 (ACE2)4,5,6,7. The trimeric S protein is cleaved into S1 and S2 subunits; subunit S1 contains a receptor-binding domain (RBD) that binds the extracellular peptidase domain of ACE2. The coronavirus enters the host cells8,9 and viral genomic RNA is released into the cytoplasm. After viral RNA replication, mature virions are assembled and released via budding from host cells.
Inhibition of the PPI between the S protein and ACE2 may therefore block the entry of SARS-CoV-2 into human cells, prevent infection and ultimately remedy COVID-19 disease. Indeed, this PPI is a major target for therapeutic intervention and researchers have developed antibodies, de novo designed proteins, peptides and engineered ACE2 to bind the S protein and inhibit viral binding and infection10,11,12,13. Here, we report the identification of small-molecule inhibitors, including natural compounds, that block the interaction of S with ACE2, inhibit binding of SARS-CoV-2 to cells and potently suppress replication of coronavirus in host cells.
We identified the inhibitors using a reversible and fluorogenic PPI reporter that enables direct visualization of spike and ACE2 interactions in living cells. This PPI reporter has a large dynamic range, with a dark-to-bright fluorescence signal that requires no exogenous cofactors. It visualizes the interaction between S protein and its receptor ACE2 and is generalizable to imaging the interaction of spike with other receptors such as neuropillin-1. Two of the natural products identified using this method potently inhibit replication of SARS-CoV-2 in vivo and could be developed into antiviral therapeutics for treatment of COVID-19.
Results
Engineering reporters for imaging of spike–ACE2 interactions
To design a fluorogenic reporter for detecting interaction between the S protein and ACE2, we improved a green fluorescent PPI reporter that we previously engineered from UnaG14,15. We named this PPI reporter SURF (split UnaG-based reversible and fluorogenic PPI reporter). Like its parent protein UnaG, SURF incorporates endogenous bilirubin as the chromophore and thus its genetically encoded fluorescence does not require an exogenous cofactor. To visualize interaction of two proteins, we fuse each fragment of SURF (carboxyl-terminal fragment, cSURF; amino-terminal fragment, nSURF) to the proteins of interest (Fig. 1a). When interactions of the two proteins bring the two fragments into close proximity, the two parts of SURF reconstitute and become fluorescent (Fig. 1a and Supplementary Fig. 1). Because SURF is reversible, inhibition of PPI leads to dissociation of the two fragments of SURF, resulting in the absence of fluorescence.
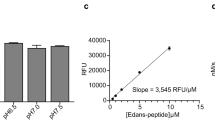
a, Schematic of the PPI reporter SURF based on a split fluorescent protein. X and Y represent two proteins of interest. b, Schematic of structure-based design of an initial SURF reporter for imaging of spike–ACE2 interaction. c,d, Fluorescence images (c) and quantification (d) of the initial and improved SURF reporters in imaging of spike–ACE2 interaction. SURF brightness was improved 270-fold via protein engineering by ACE2 truncation and mCherry insertion. c, Scale bars, 20 µm. d, Norm. SURF Fluo.: normalized SURF fluorescence. SRBD (residues 319–541) truncations refer (left to right) to: SRBD (333–541) with ACE2 (1–615, 10–615 and 17–615) and SRBD (333–518) with ACE2 (1–615, 10–615 and 17–615). Data presented as mean ± s.d. (n = 5). e, The final SURF reporter, SRBD::ΔACE2 SURF, is bright and reversible. ΔACE2, truncated ACE2 (residues 17–615); IFP2, a near-infrared fluorescent protein. Experiments were repeated three times independently, with similar results.
Next, we applied SURF to the design of PPI reporters of S–ACE2 interaction by structure-based protein design and engineering. Crystal structures of complexes of the S protein, ACE2 and B0AT1, revealed that the PPI of S and ACE2 is mediated by the receptor-binding domain of the S protein (SRBD)4,5,6,7. Thus, we fused C- and N-terminal components of SURF—that is, cSURF and nSURF—to SRBD and ACE2, respectively (Fig. 1b). The interaction of SRBD and ACE2 brought the two fragments of SURF into close proximity so that SURF reconstituted and became fluorescent. Disruption of SRBD–ACE2 interaction led to dissociation of the two SURF fragments, resulting in the loss of fluorescence. To design the two parts of the SURF reporter into one construct, we utilized the ‘self-cleaving’ 2 A peptide16 (Fig. 1c, upper left). We expressed this initial SURF reporter in human embryonic kidney 293 T (HEK293T) cells (Fig. 1c, lower left), which showed green fluorescence as expected although fluorescence was dim.
To make the reporter more suitable for high-throughput screening (HTS) we used structure-guided protein engineering, which increased the level of brightness by ~270-fold (Fig. 1c, right, 1d). Specifically, we engineered ACE2 by truncating the C-terminal fragment of ACE2 and found that removal of residues 616–727 improved brightness 2.2-fold (Fig. 1d). We then shortened the N-terminal fragment of ACE2 by removal of residues 1–16, which further increased brightness 3.5-fold. The truncated ACE2, including residues 17–615, was named ΔACE2. Truncation of SRBD did not improve brightness. We next incorporated a red fluorescent protein mCherry17,18,19 into our reporter to normalize the green fluorescence of SURF. We placed mCherry between SRBD and cSURF, which surprisingly improved SURF brightness 30-fold (Fig. 1d). mCherry probably functions as a spacer between SRBD and cSURF that may enhance SURF reconstitution following association of SRBD and ΔACE2. A second mCherry between nSURF and ΔACE2 further improved brightness, leading to a SURF reporter 270-fold brighter than the initial reporter. We named this variant of SURF reporter SRBD::ΔACE2 SURF (for simplicity, the two incorporated mCherry moieties are not shown).
Next, we examined reversibility of the SRBD::ΔACE2 SURF system. SRBD or ΔACE2 (each fused to near-infrared fluorescence protein IFP2)20,21 was co-expressed to compete against SRBD-cSURF or ΔACE2-nSURF, respectively, which would inhibit SURF reconstitution. SURF fluorescence was abolished, whereas mCherry fluorescence was stable (Fig. 1e and Supplementary Fig. 2). As a control, IFP2 expression did not perturb SURF fluorescence. Quantification of the results and calculation of a z-factor22
gave us ~0.7 with IFP2 and IFP2 fusion to SRBD (or ΔACE2) as positive (+) and negative (–) controls of the S and ACE2 PPI, respectively (here, σ is standard deviation, μ is mean). This indicates that SRBD::ΔACE2 SURF is a highly robust and versatile assay for HTS of PPI inhibitors that block S–ACE2 interaction in HEK293T cells. Thus, our SURF-based approach enabled us to conduct HTS in living cells at biosafety level 2.
Screening PPI inhibitors of spike and ACE2 using SURF
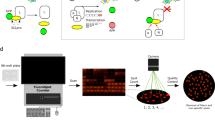
Using the SRBD::ΔACE2 SURF assay, we screened libraries of 1,516 US Food and Drug Administration (FDA)-approved drugs and 2,592 natural compounds (Fig. 2a and Extended Data Fig. 1a). First, the reporter was transfected into HEK293T cells followed by addition of the compounds with incubation for 24 h. Next, we imaged the SURF and mCherry fluorescence of the cells. The degree of PPI inhibition was calculated based on SURF fluorescence normalized by mCherry. From the FDA-approved drug library, 15 drugs showed >40% PPI inhibition with P < 0.01 (Extended Data Fig. 1b); for the natural compound library, 52 compounds showed >50% PPI inhibition with P < 0.01 (Fig. 2b).
a, Schematic showing experimental procedure. b, Volcano plot of ~2,600 natural compounds involved in inhibition of PPI of spike and ACE2. HEK293T cells were transfected with SURF reporter; SURF fluorescence was normalized by mCherry. Cells were incubated with each compound at 10 µM for 24 h before imaging. Dashed horizontal and vertical red lines represent P = 0.01 and 50% PPI inhibition, respectively. P values, two-sided nonpaired t-test between compound- and DMSO-treated groups (n = 5). c, Normalized SURF fluorescence of cells preincubated with 25 compounds (10 µM) for 6 h. Dashed red line represents 50% PPI inhibition. d, Inhibition kinetics of the identified natural compounds against spike–ACE2 interaction. Each compound (10 µM) was added to HEK293T cells expressing the SURF reporter, followed by time-lapse imaging. Estimated PPI half-life (𝜏) following inhibition is shown for each compound. c,d, Data presented as mean ± s.d. (n = 5).
To confirm the inhibitory activity of the FDA-approved drugs, an independent imaging of 15 drugs was conducted under the same conditions. We selected the top ten drugs (Extended Data Fig. 1c) and characterized their inhibition kinetics by time-lapse imaging using SURF. Seven drugs that inhibited the PPI within 6 h (Extended Data Fig. 1d) were selected for antiviral activity measurement (see below). Finally, the drugs showed little effect on a SURF control where cSURF and nSURF were directly linked, suggesting that they have little effect on the reporter itself (Supplementary Fig. 3).
For the identified natural compounds, their activity was confirmed by a second round of imaging (Supplementary Fig. 4). Next, we narrowed the number of hits by shortening incubation time to 6 h: 25 compounds exhibited >50% PPI inhibition (Fig. 2c) while the other 27 either displayed little inhibition (19 compounds) or strong cytotoxicity based on morphological changes (Supplementary Fig. 5). We next characterized inhibition kinetics: 19 compounds inhibited the PPI within 2 h (Fig. 2d). Among these, bruceine D, bruceine A, homoharrington, anisomycin and gamabufotalin blocked the PPI with τ ~0.5–0.7 h. The remaining compounds showed slower inhibition (Supplementary Fig. 6). Finally, the top 19 compounds had little effect on the SURF control (Supplementary Fig. 7).
Antiviral activity against SARS-CoV-2 in cells
We next selected the top-performing drugs and natural compounds against the PPI and determined their antiviral activity using SARS-CoV-2 in Vero-E6 cells (Fig. 3a). Six of these drugs inhibited SARS-CoV-2 at 5 μM (Fig. 3b), with the natural compounds inhibiting SARS-CoV-2 replication at 1 or 2 μM (Fig. 3c). Strikingly, gamabufotalin, anisomycin and bruceine A exhibited >6 log reduction in viral titre at 1 µM while at 0.1 μM concentration they inhibited SARS-CoV-2 by 10–1,000-fold (Fig. 3d). Interestingly, bruceine D, a compound structurally related to bruceine A, exhibited no inhibition at 0.1 μM (Supplementary Fig. 8).
a, Schematic showing the experimental design (Methods). b–d, Quantitative analysis of viral titre from plaque assays on Vero-E6 cells treated with drugs at 5 μM (b) or natural compounds at 2 or 1 μM (c) or at 0.1 μM (d). b–d, MOI = 0.5. Data presented as mean + s.d. (n = 3). P values, two-sided nonpaired t-test between compound- and DMSO-treated groups. *P < 0.05. **P < 0.01, ***P < 0.001. Exact P values: b, starting from ciclopirox: 0.14, 0.00096, 0.00096, 0.0013, 0.00099, 0.00096, 0.00096; c, starting from oleandrin: 0.00083, 0.0011, 0.00088, 0.0011, 0.00090, 0.0026, 0.00087, 0.00089, 0.0011, 0.00085, 0.0025, 0.00087, 0.00080, 0.00080, 0.00080, 0.00081, 0.00080; d, starting from bruceine D: 0.22, 0.00093, 0.00093, 0.0010, 0.0020. e, Dose–response and cell toxicity curve of each compound against SARS-CoV-2 by plaque assay (MOI = 0.5). Percentage of relative infection was determined by the ratio of infection rate of SARS-CoV-2 treated with each compound divided by that of DMSO control. EC50 and CC50 are presented as mean ± s.d. (n = 3). f,g, Immunofluorescence staining (f) of SARS-CoV-2 N and S protein in Vero-E6 cells and quantification (g) of number of stained cells at 16 h.p.i. (MOI = 0.5), treated with 200 nM anisomycin, bruceine A, gamabufotalin or remdesivir. Scale bars, 100 μm. Data presented as mean ± s.d. (n = 3). P values, two-sided nonpaired t-test between compound- and DMSO-treated groups. ***P < 0.001. Exact P values: starting from gamabufotalin anti-N: 0.00023, 0.00028, 0.00029, 0.82; starting from gamabufotalin anti-S: 0.00016, 0.00019, 0.00021, 0.63. NS, not significant; ND, not detectable.
Next, we determined dose–response curves (Fig. 3e). Gamabufotalin showed the most potent antiviral activity, with half-maximal effective concentration (EC50) ~0.003 μM, followed by anisomycin (0.016 μM) and bruceine A (0.054 μM). Interestingly, bruceine A showed fourfold higher potency than bruceine D. These compounds were largely not cytotoxic in Vero cells at EC50, with selectivity index (50% cytotoxicity concentration (CC50)/EC50) >10. Other natural compounds and FDA-approved drugs showed less potent antiviral activity, with EC50 at submicromolar (Fig. 3e) or micromolar range (Supplementary Fig. 9). The top three inhibitors were further confirmed using antibodies against the nucleocapsid (N) or spike. At 200 nM, gamabufotalin, anisomycin and bruceine A potently blocked SARS-CoV-2 (Fig. 3f,g); these compounds also blocked SARS-CoV-2 variants Delta and Omicron (Extended Data Fig. 2).
Antiviral activity during SARS-CoV-2 infection
To further understand mode of action, we examined the effects of the natural compounds at three time points before or after SARS-CoV-2 infection (Fig. 4a). For the first condition we preincubated Vero cells with inhibitors for 3 h, followed by the addition of SARS-CoV-2 (multiplicity of infection (MOI) = 5) for adsorption at 4 °C for 1 h. We then removed the solution and added fresh medium containing the compounds (Fig. 4a (i), −4 h). For the other two conditions, compounds were added at 1- or 4-h post-infection (h.p.i.) (Fig. 4a (ii), +1 h and (iii), +4 h samples). Cells were then incubated at 37 °C; 16 h later both supernatant and cells were collected for plaque assay and quantitative PCR with reverse transcription (RT–qPCR).
a, Schematic of the staging experiment. SARS-CoV-2 (MOI = 5) was added to Vero-E6 cells, then natural compounds (1 μM) were added at various time points as indicated. b,c, Samples were characterized by plaque assay (b) and RT–qPCR (c). Total RNAs were extracted from Vero cells with Trizol reagent. vRNA genomes were measured by RT–qPCR with primers targeting the nucleocapsid gene (N) of SARS-CoV-2, normalized to internal control HRT1. d, SARS-CoV-2 binding assay with immunofluorescence staining. Vero cells were preincubated with natural compounds (2 μM), followed by the addition of SARS-CoV-2 (MOI = 100) for adsorption at 4 °C for 1 h. Left, experimental procedures; middle, immunofluorescence images; right, quantification of immunofluorescence against N protein. b–d, Data presented as mean ± s.d. (n = 3). P values, two-sided nonpaired t-test between compound- and DMSO-treated groups. *P < 0.05 considered significant. **P < 0.01, ***P < 0.001, ****P < 0.0001. Exact P values <0.0001 (b,c); starting from anisomycin: 0.00020, 0.00021, 0.00020 (d). Final concentration of natural compounds was 2 μM for the above experiments. d, Scale bars, 40 μm. e, Schematic of the rVSV-eGFP-SARS-CoV-2 pseudovirus infection assay. The rVSV-eGFP-SARS-CoV-2 pseudovirus comprises replication-competent rVSVs encoding the spike protein of SARS-CoV-2 in place of the original G glycoprotein. A549 cells stably expressing hACE2 were preincubated with natural compounds for 3 h, followed by incubation with rVSV-eGFP-SARS-CoV-2 (MOI = 1) at 37 °C for 1 h and washing with fresh medium containing compounds. Cell eGFP fluorescence was imaged at 16 h.p.i. f, Dose–response curves of inhibitors against rVSV-eGFP-SARS-CoV-2 pseudovirus infection assay. Relative infection was calculated based on eGFP fluorescence. Data presented as mean ± s.d. (n = 3).
Plaque assay-based measurement of viral particles in the supernatant showed that anisomycin and bruceine A reduced viral titre by between four and five logs and gamabufotalin by between two and three (Fig. 4b (i), −4 h). For the +1 h samples, anisomycin and bruceine A reduced viral titre by four logs and gamabufotalin by between one and two. The inhibitory effect was further reduced in the +4 h regimen. Anisomycin and bruceine A showed viral titre reduction by two logs and gamabufotalin by less than one.
Consistent with virus production measured by plaque assay, viral genome copy number measured by RT–qPCR in infected cells showed that, for −4 h samples, anisomycin, bruceine A and gamabufotalin reduced vRNA load by between five and six, between four and five and between two and three logs, respectively. Inhibitory activity was reduced in +1 h samples and further reduced in +4 h samples (Fig. 4c). These results indicate that inhibitors act on the early stages of the viral infection.
To test whether these natural compounds inhibit cellular binding of SARS-CoV-2, we conducted an immunofluorescence-based binding assay. We preincubated Vero cells with each inhibitor (2 μM) for 3 h, followed by the addition of SARS-CoV-2 (MOI = 100) for adsorption at 4 °C for 1 h (Fig. 4d). We then removed the solution, washed the cells and paraformaldehyde (PFA) fixed and stained the samples with antibodies against the N protein. Immunofluorescence of the N protein was decreased by around tenfold, suggesting that SARS-CoV-2 viral particle binding to cells is largely reduced in the presence of inhibitors (Fig. 4d). These results indicate that inhibitors act by blockage of SARS-CoV-2 binding to host cells.
To further examine whether inhibitors block the PPI of S and ACE2, we utilized the replication-competent recombinant vesicular stomatitis virus (rVSV) encoding the spike in place of the original G glycoprotein (rVSV-enhanced green fluorescent protein (eGFP)-SARS-CoV-2)23. We examined the effects of the inhibitors on pseudovirus in ACE2-expressing A549 cells (Fig. 4e). Bruceine A, anisomycin and gamabufotalin showed strong inhibition, with an EC50 of 0.018, 0.018 and 0.005 μM, respectively. Finally, we showed that our compounds had no effect on a control virus, a Sindbis virus (SINV; Supplementary Fig. 10). The above results suggest that these compounds exert their activity by blockage of the PPI of S and ACE2.
Anti-SARS-CoV-2 activity in primary human cells
Next, we determined the antiviral potential of the natural compounds gamabufotalin, bruceine A and anisomycin in human primary cells relevant to SARS-CoV-2 infection in humans24. We used primary human nasal epithelial cells (HNEpCs) and primary human bronchial epithelial cells (HBEpCs), both of which were cultured in an air–liquid interface. We preincubated HNEpCs with 2 μM natural compounds for 3 h before infection (Fig. 5a). Next, we infected HNE cells with SARS-CoV-2 for 1 h at 37 °C and incubation for 48 h (both stages in the presence of the 2 μM inhibitors). Following viral infection, cells were fixed and stained with anti-N and anti-S antibodies. Immunofluorescence imaging showed that the natural compounds largely reduced SARS-CoV-2 infection in HNEpCs (Fig. 5b,d). We then conducted similar experiments in HBEpCs with the compounds at 15, 50 and 200 nM and 1 μM (Fig. 5c and Supplementary Fig. 11a). Quantification of HBEpC imaging data showed that all three compounds reduced infection even at 15 nM (Fig. 5e and Supplementary Fig. 11b). We also confirmed the inhibitory effect of gamabufotalin and anisomycin in HNE cells using plaque assays and RT–qPCR (Extended Data Fig. 3). These inhibitors therefore have potent antiviral activity against SARS-CoV-2 infection in primary human airway epithelial cells.
a, Experimental procedure. b,c, Immunofluorescence staining of SARS-CoV-2 in primary HNEpCs (b) and HBEpCs (c) treated with gamabufotalin, bruceine A or anisomycin at 2 μM (b) or 50 nM (c). MOI = 3 for HNEpCs, MOI = 1 for HBEpCs. Scale bars, 50 μm (b) and 100 μm (c). d,e, Quantification of nucleocapsid-positive (N+) or spike-positive (S+) cells. HNEpCs (d) and HBEpCs (e) were immunostained with N or S, respectively. Numbers of N+ or S+ cells per 100 cells were quantified. Rectangles bordered by dashed grey lines represent samples treated with the same compound at different concentrations. Data presented as mean ± s.d. (n = 3). P values, two-sided nonpaired t-test between compound- and DMSO-treated groups. *P < 0.05, **P < 0.01. Exact P values: d, starting from gamabufotalin anti-N: 0.0023, 0.0042, 0.0027; starting from gamabufotalin anti-S: 0.0016, 0.0030, 0.0019; e, starting from gamabufotalin anti-N: 0.0045, 0.0030, 0.0030, 0.018, 0.0056, 0.0030, 0.013, 0.0035, 0.0030; starting from gamabufotalin anti-S: 0.0030, 0.0020, 0.0020, 0.013, 0.0040, 0.0020, 0.0088, 0.0023, 0.0019.
We also examined the effects of the natural compounds when given before or after infection of SARS-CoV-2 in HBEpCs (Extended Data Fig. 4). First we preincubated HBEpCs with inhibitors for 3 h, followed by the addition of SARS-CoV-2 for adsorption for 1 h. We then removed the solution and added fresh medium containing the compounds (Extended Data Fig. 4a (i), −4 h). For the other two conditions, compounds were added at 1 or 4 h.p.i. (Extended Data Fig. 4a (ii), +1 h and (iii) +4 h samples). Cells were incubated at 37 °C for 72 h and fixed for immunofluorescence staining against SARS-CoV-2 N and S proteins. Our data showed that, for all three compounds, −4 h samples showed complete inhibition of SARS-CoV-2 infection, +1 h samples showed ~90% inhibition but +4 h samples showed no significant inhibition (Extended Data Fig. 4b,c). These results are consistent with those for Vero cells. Our data indicate that these inhibitors act on the early stages of viral infection in HBEpCs. We also examined the toxicity of the natural compounds in HBEpCs. Following incubation with gamabufotalin, bruceine A or anisomycin at 2 μM for 72 h, no cytotoxicity was observed in HBEpCs (Supplementary Fig. 12).
Anti-SARS-CoV-2 activity in mouse models of infection
We next determined antiviral activity of the potent inhibitors gamabufotalin and bruceine A in mouse models of SARS-CoV-2 infection. K18-hACE2 mice25 were infected intranasally with SARS-CoV-2 (103 plaque-forming units (PFU) per mouse) in the absence (vehicle control, solvent) or presence of inhibitor. Treatment was repeated on days 1 and 2 (Fig. 6a). We directly examined the effects of bruceine A and gamabufotalin on virus replication. At day 3 post-infection, lungs and brain were harvested for analysis. Both compounds reduced viral titre by between two and three logs in the lung (Fig. 6b). Bruceine A treatment reduced SARS-CoV-2 titre to undetectable levels in brain tissues (<10 PFU g–1; n = 6 animals). Similarly, we were unable to detect virus in the brains of four out of six animals treated with gamabufotalin and we observed a reduction of between one and two logs in viral titre for the remaining two animals. Immunofluorescence imaging analysis revealed similar antiviral activity for bruceine A and gamabufotalin in lung sections. Immunofluorescence of anti-N and anti-S indicated that the lungs of both bruceine A- and gamabufotalin-treated animals showed markedly fewer N- and S-positive cells than vehicle controls (Fig. 6c,d).
a, Schematic of the experimental design. K18-hACE2 mice were infected with SARS-CoV-2 (1,000 PFU) intranasally with either inhibitor or vehicle control (solvent) at day 0. Inhibitors were then delivered intranasally once per day on days 1 and 2. At 3 d.p.i., lung and brain were harvested for analysis. b, Virus titre measurement by plaque assay for lung and brain. n = 6 mice per group. Bruceine A and gamabufotalin were administered at 3.25 mg kg–1 by intranasal inoculation. P values, two-sided nonpaired Welch’s t-test between vehicle and treated groups. c, Immunofluorescence images of lung tissues. Slides were stained with antibodies against N, S, acetylated tubulin (AcTub) and DAPI. Immunofluorescence of N, S, AcTub and DAPI denoted by pseudocolours green, red, grey and blue, respectively. Lower panels correspond to boxed areas of upper panels. Scale bars: 100 μm (upper), 200 μm (lower). d, Quantification of immunofluorescence. Immunofluorescence samples were acquired from two mice for each group. Data presented as as mean ± s.d. P values, two-sided nonpaired t-test between vehicle and treated groups. ****P < 0.0001. Exact P values: anti-N: bruceine A 7.8 × 10–7, gamabufotalin 5.4 × 10–7; anti-S: bruceine A 8.8 × 10–7, gamabufotalin 6.0 × 10–7.
We next examined histopathological lung changes by staining sections with haematoxylin and eosin (H&E). In vehicle-treated samples we detected both alveolar and peribronchial inflammation. Bruceine A-treated lung showed little or no inflammation in alveolar and peribronchial areas, similar to that from animals not infected with SARS-CoV-2 (Extended Data Figs. 5 and 6). Gamabufotalin-treated lung also revealed little alveolar inflammation and largely reduced peribronchial inflammation.
We also examined whether inhibitors could block coronavirus after viral infection in mice. We first infected wild-type C57BL6 mice with another strain of SARS-CoV-2 (B.1.1.7) via intranasal inoculation without co-application of inhibitor. Treatment with bruceine A and gamabufotalin were then given twice, at 24 and 48 h.p.i., via intranasal inoculation-elicited protection (Extended Data Fig. 7a). Lungs were harvested for analysis at 72 h.p.i. Bruceine A and gamabufotalin reduced lung virus titre by 0.8 and one log, respectively (Extended Data Fig. 7b). Immunofluorescence of anti-N and anti-S showed that both drugs largely reduced N- and S-positive cells in lung (Extended Data Fig. 7c,d).
Histopathological changes in the C57BL6/B.1.1.7 model were mild, but perivascular and alveolar inflammation was still detected in the vehicle-treated group (Extended Data Fig. 8a). With five mice per group analysed, significant differences were shown between the histopathology score of the vehicle-treated group (1.2 out of 16) and the bruceine A-treated (0.2 out of 16) and gamabufotalin-treated (0.2 out of 16) groups (Extended Data Fig. 8b). These results suggest that the two natural compounds may provide therapeutic treatment for COVID-19.
Discussion
Several small-molecule inhibitors interfere with aspects of the life cycle of SARS-CoV-2, including the main protease Mpro26,27,28,29 and the SARS-CoV-2 replicase RdRp30. However, no potent small molecules at nanomolar range have been reported to inhibit the interaction of the viral S protein and host receptor ACE2. In the present study, we designed a brightly fluorescent PPI reporter that visualizes the interaction between spike protein and its receptor. This reporter is fluorogenic and reversible. Its large dynamic range and bright signal enable robust HTS of small-molecule inhibitors in living cells.
Using fluorogenic PPI reporters, we identified potent inhibitors that block the interaction of the two proteins in living cells at nanomolar concentrations, with verified inhibition of SARS-CoV-2 infection in both Vero cells and primary human airway cells at 15 nM. Additionally, the reversibility of our reporter allowed us to characterize the inhibition kinetics of the identified compounds which, interestingly, are proportional to their efficacy in cells. Our work demonstrates that this innovative reporter and structure-based engineering strategy can be applied to the future design of reporters for imaging key PPIs that mediate other virus–host interactions, and to screen for small-molecule inhibitors, following an approach similar to that developed here.
Our results suggest that the potent compounds bruceine A and gamabufotalin inhibit SARS-CoV-2 infection by blocking the S protein- and ACE2-mediated early stages of viral infection. First, inhibitors exert their activity on the early stages of viral infection because their antiviral activities are largely reduced when introduced at the later stages. Second, the SARS-CoV-2 binding assay showed that inhibitors block binding of the virus to host cells. Finally, using the pseudovirus, we further determined that the compounds exert their activity by blocking interaction of the S protein and ACE2 because the pseudovirus expresses only one of the SARS-CoV-2 proteins, the S protein. Furthermore, these compounds have no activity against SINV, which does not have the spike protein of SARS-CoV-2.
To gain insight into how these compounds inhibit S–ACE2 interaction, we docked both into potential pockets of the S–ACE2 protein complex and found that they were able to dock into the active site of ACE2 (Extended Data Fig. 9). An ACE2 inhibitor bound to the same site was reported to be effective in blocking SARS-CoV spike protein-mediated cell fusion31, demonstrating that the ACE2 inhibitor induces marked conformational changes in the active site that impacts surrounding residues, including those involved in binding to the spike protein. Considering the high sequence similarity between the two spike proteins (~75% homologous) of SARS-CoV and SARS-CoV-2, our compounds may also inhibit S–ACE2 interaction because the ACE2 inhibitor, possibly by conformational change of ACE2, blocks interaction with the spike protein.
Considering that the SARS-CoV-2 virus constantly changes through mutations, especially in the spike RBD domain, we tested the antiviral effect of bruceine A and gamabufotalin on Delta and Omicron variants. Both compounds maintained similar or slightly weaker inhibition effect on Delta and Omicron infection in Vero cells.
We demonstrate here that two natural compounds (gamabufotalin and bruceine A) exhibit potent antiviral efficacy in vivo when administered together with the virus as a prophylactic treatment. We chose these two inhibitors because their analogues showed largely reduced antiviral activity, indicating a structure–activity relationship. For example, cinobufotalin, which is structurally related to gamabufotalin, showed fivefold lower antiviral activity while bruceine D had fourfold lower antiviral activity than bruceine A (Fig. 3e). Both bruceine A and gamabufotalin reduced viral titre in the lung by between two and three logs at 3 days post-infection (d.p.i.) using K18-hACE2 mouse models of SARS-CoV-2 infection. These results were verified by immunofluorescence imaging of lung tissues. Furthermore, bruceine A eliminated coronavirus in mouse brain while gamabufotalin also significantly blocked SARS-CoV-2 replication in the brain. Histopathological analysis of lung showed a significant reduction in both alveolar and peribronchial inflammation. These findings demonstrate that both inhibitors significantly reduced SARS-CoV-2 replication in lung and brain, and prevented lung inflammation in the mouse model when the inhibitors were administered together with the virus. On the other hand, when the inhibitors were provided 24 h.p.i. they showed mild activity in blocking viral replication. This is consistent with results in cultured cells in which the inhibitors mainly blocked the early stages of viral infection. Our work suggests that the two compounds may be used as therapeutic treatment for COVID-19 via nasal application.
The PPI reporter we developed in this work visualizes the interaction between spike protein and its receptor ACE2. Our engineering strategy improved the reporter’s brightness by ~270-fold. The improved PPI reporter visualizes spike–ACE2 interaction with a wide dynamic range, which enabled HTS of small-molecule inhibitors of this PPI. This reporter strategy will have important applications in the future regarding the study of spike–receptor interactions, and it will also guide future engineering of reporters in imaging spike interaction with other receptors such as neuropillin-1 (NRP1). The spike protein of SARS-CoV-2 has been shown to interact with NRP1 (ref. 32), which facilitates infection of SARS-CoV-2 in neurons such as the NRP1-positive olfactory neurons facing the nasal cavity. We further show that the reporter is generalizable because it can be used to visualize the interaction between spike and NRP1 (Extended Data Fig. 10). This PPI reporter will be valuable in the study of spike–NRP1 interaction, as well as in identification of small-molecule inhibitors by HTS. Therefore, our work provides a general strategy in the engineering of PPI reporters for imaging interaction between a viral protein and its host cell receptor, and in screening for inhibitors that block viral interaction with host cells. Our PPI reporter, its engineering strategy and the screening approach will become a valuable resource in effectively dealing with any future pandemic by SARS-CoV-2 variants and other viruses.
Methods
Plasmid construction
To create the SRBD::ΔACE2 SURF in pcDNA3.1, we used different truncations of the S protein and ACE2 and fused them to cSURF and nSURF, respectively. FKBP::Frb SURF was also cloned into pcDNA3.1, using full-length human FKBP CDS and human mTOR Frb domain (mTOR2025–2114). pcDNA3.1-SRBD-IFP2 and pcDNA3.1-IFP2-ΔACE2 were created by fusion of either SRBD or ΔACE2 to IFP2 with a ten-amino-acid linker. All constructs were made using standard enzyme digestion and ligation methods.
Cell culture, transfection and live cell imaging
HEK293T cells (ATCC, no. 3216) were cultured in DMEM (Gibco, no. 11965-092) supplemented with 10% fetal bovine serum (FBS; Gibco, no. 16000044) and 1% penicillin/streptomycin/glutamine (Gibco Invitrogen). Vero-E6 cells (ATCC, no. 1586) were maintained in MEM (Gibco Invitrogen) supplemented with 10% FBS and 1% penicillin/streptomycin/glutamine. A549-ACE2 cells were a kind gift from P. Jackson’s laboratory at Stanford University. All cells were cultured at 37 °C in humidified 5% CO2 incubators.
HEK293T cells were transfected using calcium phosphate or polyethylenimine. All live cell imaging was performed using a Nikon Eclipse Ti inverted microscope equipped with a Yokogawa CSU-W1 confocal scanner unit (Andor) and a CO2/temperature control unit, using Nikon NIS-elements software.
For determination of inhibition kinetics of the identified drugs, we added each drug (10 μM) to HEK293T cells expressing SRBD::ΔACE2 SURF, performed time-lapse imaging and analysed fluorescence changes over time. The inhibition kinetics of each drug were determined from normalized SURF fluorescence (by the red fluorescence of mCherry) over time. Seven drugs inhibited the PPI within 6 h, three of which (mitoxantrone (2 HCl), digoxin and cinobufotalin) inhibited it relatively rapidly with an inhibition half-life (τ) of ≤1.5 h.
HTS
A library of natural compounds containing 2,592 compounds (TargetMol, no. L6000) and an FDA-approved drug library (MedChemExpress, no. HY-L022) containing 1,516 compounds were used for HTS. For each screening, HEK293T cells were transfected with SRBD::ΔACE2 SURF using polyethylenimine and seeded onto 96-well plates at ~5,000 cells per well, 6 h after transfection. One day after seeding, compounds were added to cells (5 or 10 μM final concentration for natural compounds or FDA-approved drugs, respectively). After 16 h of incubation with the compounds, images were taken for each well in the 488 and 561 channels. Total cell fluorescence was calculated using ImageJ. SURF fluorescence intensity was normalized by mCherry intensity and then compared with that of control wells incubated with DMSO to acquire fold-change values.
Cytotoxicity assay
Cytotoxicity of the identified compounds on Vero-E6 cells and HBEpCs was determined with WST-1 cell proliferation assays (Roche, no. 5015944001). For Vero-E6 cells, about 3,000 per well were seeded into 96-well plates and cultured for 1 day at 37 °C, followed by the addition of 1 μl of each compound at decreasing concentrations. After 18 or 72 h of incubation at 37 °C, WST-1 reagents were added to each well with incubation for 1 h. HBEpCs were cultured on 6.5 mm transwell inserts, compounds were added to the culture medium for 72 h then WST-1 reagent was added with incubation for 4 h. Absorbance at 450 and 650 nm was measured using a Tecan Infinite M1000 plate reader with iControl 1.8 software. All experiments were performed in three biological independent repeats.
SARS-CoV-2 strains including Delta and Omicron
A clinical isolate of SARS-CoV-2 (USA-WA1/2020, BEI NR-52281) was propagated in Vero-E6 cells and used for testing of drugs in this cell culture. A clinical isolate, SARS-CoV-2/human/USA/CA-UCSF-0001C/2020 (ref. 33), was propagated in A549 cells expressing ACE2 receptor under control of the CMV promoter and was used for both mouse and primary cell experiments.
Omicron and Delta variants were isolated in Vero-hACE2-TMPRSS2 cells (BEI NR-54970): 200 μl of nasal specimen was added to one well of a 96-well plate and serially diluted 1:1 over five additional wells; 100 μl of freshly trypsinized cells, resuspended in infection medium (DMEM with 10% FBS, 200 μg ml–1 penicillin/streptomycin and 5 μg ml–1 amphotericin B) at 2.5 × 105 cells ml–1, was then added to each sample dilution. Cells were cultured at 37 °C with 5% CO2 and checked for cytopathic effects at 2–3 d.p.i. Supernatants were harvested on day 3 after inoculation; 200 μl of P0 was used to infect a confluent T25 flask to generate a P1 culture then harvested after 3 days. Viruses were passaged three times after isolation to obtain a sufficiently high titre (>107 PFU ml–1). Viral titres were quantified by plaque assay. All infection experiments were performed at biosafety level 3.
Antiviral assay on Vero-E6 cells
For assessment of initial antiviral activity and EC50 of drugs, 70% confluent monolayers of Vero-E6 cells (2 × 105 per well in 24-well plates) were pretreated with drug for 3 h (pretreatment) and then infected with SARS-CoV-2 (MOI = 0.5) at 37 °C for 1 h. Virus solution was removed, then cells were washed twice with PBS and added to fresh medium containing drug/natural compound or DMSO to continue culture. At 16 h.p.i., viral titres of supernatants were determined by plaque assay.
Plaque assay
Confluent monolayers of Vero-E6 cells grown in six-well plates were incubated with serial dilutions of virus samples (250 μl per well) at 37 °C for 1 h. Next, cells were overlaid with 1% agarose (Invitrogen) prepared with MEM supplemented with 2% FBS and 1% penicillin/streptomycin/glutamine (100x, Gibco Invitrogen). After 3 days cells were fixed with 4% formaldehyde for 2 h, the overlay was discarded and samples stained with crystal violet dye.
Adsorption assay
Vero-E6 cell monolayers at 70% confluence grown on coverslips were preincubated with natural compounds at 37 °C for 3 h. Cells were then added to SARS-CoV-2 virus at MOI = 100 and immediately transferred to 4 °C for 1 h. The virus suspension was quickly removed and cells were washed three times with ice-cold PBS. Cells were immediately fixed with 4% PFA for 30 min at room temperature. PFA was then washed away with PBS and quenched with 1 M glycine in PBS.
Drug staging experiment
Drugs were added at different stages of SARS-CoV-2 infection. For −4 h samples, Vero-E6 cells (~2 × 105 per well in 24-well plates) were pretreated with 2 μM compound for 3 h (preincubation) and infected with SARS-CoV-2 at MOI = 5. After 1 h of incubation at 37 °C, cells were washed twice with PBS to remove virus and then incubated with fresh medium containing drug; for +1 h and +4 h samples, there was no preincubation, and drugs were added 1 or 4 h.p.i., respectively. At 16 h.p.i., supernatant was collected for plaque assay and cells harvested using Trizol reagent (Ambion) for RT–qPCR.
Pseudovirus rVSV-eGFP-SARS-CoV-2
We preincubated ACE2-expressing A549 cells with compounds for 3 h, infected the cells with rVSV-eGFP-SARS-CoV-2 (MOI = 1) at 37 °C for 1 h then removed the pseudovirus, added compounds again and cultured the cells for 16 h. Cells were then imaged and infection rates calculated based on percentage of cells with eGFP signal.
SINV–eGFP generation
The eGFP expressing Sindbis virus (SINV–eGFP) was cloned by insertion of the eGFP coding sequence into the XbaI site of the double-subgenomic Sindbis vector pTE392J (provided by C. Rice). Viral genomic RNA was obtained by in vitro transcription using MEGAscript SP6 kit (Invitrogen) with transfection into BHK-21 cells. The initial virus stock was collected from the supernatant of transfected cells and amplified once by infecting a new culture of BHK-21 cells (passage 1, P1). Finally, P1 viral stock titration was performed by plaque assay in BHK-21 cells. A procedure similar to that for SARS-CoV-2 was followed here to examine effects of the natural compounds on SINV.
Molecular docking
The structure of SARS-CoV-2 S protein complexed with human ACE2 (PDB ID: 6VW1) was used in molecular docking calculations, with calculations performed as previously described34. Protonation states of residues in the ACE2 active site were visually checked. Atom charges were assigned to the structure using the AMBER united atom force field. Energy grids were pregenerated with CHEMGRID for van der Waals potential, with QNIFFT for Poisson–Boltzmann-based electrostatic potential and with SOLVMAP for ligand desolvation penalty. Presampling three-dimensional conformer libraries were generated by ZINC. Compounds were docked against the ACE2 active site using DOCK3.7. Docking poses were outputted and examined visually. Figures were generated using UCSF Chimera.
Infection of primary cells
Primary human nasal epithelial cells (PromoCell) were pretreated with compound for 3 h, infected with virus at MOI = 3, incubated with medium containing 2 μM inhibitor for 48 h, then fixed with 4% paraformaldehyde in PBS.
Primary HBEpCs were cultured on 6.5 mm transwell inserts (Corning) for 4 weeks at an air–liquid interface to obtain well-differentiated epithelium. Mucus was removed immediately before viral infection by washing the apical surface with a prewarmed solution of 10 mM dithiothreitol (Thermo) in PBS for 10 min, then washed twice with PBS without dithioreitol. Cells were inoculated by the addition of virus to the apical surface at MOI = 1. After 1 h, the apical surface was washed twice with PBS and cells were returned to the incubator. Cells were fixed at 72 h.p.i. in 4% paraformaldehyde in PBS.
Immunofluorescence
Both Vero-E6 and primary cells were fixed in 4% paraformaldehyde, permeabilized by PBST (PBS with 0.1% Triton X-100), blocked with 2% BSA and 10% goat serum and incubated overnight at 4 °C with antinucleocapsid antibody (1:500; Genetex, no. GTX135357) and/or antispike antibody (1:600; Genetex, no. GTX632604). After washing three times with PBST, cells were incubated with goat antirabbit IgG H&L (1:800, Alexa Fluor 488; Abcam, no. ab150077) or goat antimouse IgG H&L (1:800, Alexa Fluor 555; Abcam, no. ab150114).
For both Vero-E6 and primary HBEpCs, images were acquired using a Nikon Eclipse Ti-E Spinning Disk microscope with a ×60 numerical aperture (NA) 1.49 Plan-Apo objective. The software used was NIS-Element AR 4.20. For primary HNEpCs, images were acquired on an Everest deconvolution workstation (Intelligent Imaging Innovations) equipped with a Zeiss AxioImager Z1 microscope, CoolSnapHQ cooled CCD camera (Roper Scientific) and a ×40 NA 1.3 Plan-Apochromat objective lens (Zeiss, no. 420762-9800). The software used for acquisition was SlideBook (v.6). All images were processed in ImageJ. Fluorescence signals were calculated by pixel intensity using Analyze Particle function in ImageJ, and cell numbers calculated using Cell Counter plugin in ImageJ.
Three wells of either Vero-E6 or primary cell cultures were infected for each condition. Ten field views for each well of Vero-E6, three for each HNE culture and ten for each HBE culture were imaged. In total, 1,500 cells were analysed for each treatment for Vero-E6 or HBEpCs, and 500 for each treatment for HNEpCs. Total N- and S-positive cell numbers per 100 cells were used for statistical analysis.
vRNA isolation and qPCR
Genomic RNA was extracted from cell pellets using Trizol (Ambion) according to the manufacturer’s instructions. RNA was treated with DNAse I (NEB) and used as a template to reverse transcribe complementary DNA with the Iscript kit (NEB). qPCR was performed using the Luna Universal qPCR Master Mix (NEB) and a CFX connect qPCR Detection System (Bio-Rad). To determine the number of vRNA copies per millilitre, plasmids containing the nucleocapsid gene of SARS-CoV-2 (cloned from the USA-WA1/2020 isolate) were used as standards and diluted serially tenfold to determine target copy numbers. Threshold cycle values were plotted against the number of target copies, with the resultant standard curve used to determine the number of genome equivalents of vRNA in samples. For cell pellet samples, vRNA copy number was normalized to the housekeeping gene HRT1. All samples were within the range of linearity of a standard curve, and primer efficiency was 100 ± 5%. The primer sequences targeting nucleocapsid were 5′-TCCTGGTGATTCTTCTTCAGG-3′ and 5′-TCTGAGAGAGGGTCAAGTGC-3′; HRT1 primer sequences were 5′-GGTCCTTTTCACCAGCAAGCT-3′ and 5′-TGACACTGGCAAAACAATGCA-3′.
Inhibitor preparation for in vivo study
Natural products were obtained from MedChemExpress (bruceine A, no. HY-N0841 and gamabufotalin, no. HY-N0883). Natural products were prepared at 100 mg ml–1 in DMSO, then each was diluted in solvent ((5% Tween 80; Sigma, no. P4780) and 40% PEG300 (MedChemExpress, no. HY-Y0873, 55% PBS)).
Toxicity testing in animals
Natural products were delivered intranasally. Briefly, C57BL6/J mice were anaesthetized with isoflurane and inoculated with solvent and compounds at different diluted concentrations in 50 μl (6.50, 3.25 and 1.63 mg kg–1). Mice were treated every 24 h for 3 days. No obvious toxicity was observed at dosages up to 6.5 mg kg–1 per mouse by intranasal inoculation.
Antiviral effect testing in mouse models
All antiviral studies were performed in an animal biosafety level 3 facility at the Department of Immunology and Microbiology, University of California, San Francisco (UCSF). SARS-CoV-2 experiments were performed in 8–10-week-old K18-hACE2 mice (The Jackson Laboratory, no. 034860, B6.Cg-Tg (K18-hACE2) 2Prlmn/J, hemizygous) and C57BL6/J mice (The Jackson Laboratory, no. 000664, RRID:IMSR_JAX:000664). Mice were housed and bred in a specific-pathogen-free, AAALAC-certified animal facility at UCSF at ~23 °C and 50% humidity under a 12/12-h light/dark cycle, with standard chow diet provided. All procedures were performed in accordance with the guidelines of the Laboratory Animal Center of the National Institutes of Health. The Institutional Animal Care and Use Committee of the UCSF approved all animal protocols (approved protocol no. AN178420-02B).
For K18-hACE2 mice, on day 0 we administered solvent, bruceine A and gamabufotalin and, simultaneously, 1,000 PFU of SARS-CoV-2 in 50 μl (25 µl of virus solution in PBS plus 25 µl of drug solution or solvent) intranasally. Bruceine A and gamabufotalin were then administrated intranasally once per day for 2 days. Mice were anaesthetized with isoflurane before intranasal infection. This experiment was repeated twice independently; a total of eight mice were used in each control or drug-treated group.
For C57BL6/J mice, on day 0 we administered 105 PFU of SARS-CoV-2 (B.1.1.7) intranasally. Bruceine A and gamabufotalin were then administrated intranasally at 24 and 48 h.p.i. A total of five mice were used in each control or drug-treated group.
For virus tissue distribution, at 3 d.p.i. animals were humanely euthanized and lungs and brains harvested. Half of the lungs were fixed in 4% PFA for 72 h, transferred to 70% ethyl alcohol, embedded in paraffin and processed for H&E and immunofluorescence staining. The other half of the lungs, and brains, were weighted and homogenized in 1 ml of 2% FBS MEM medium with gentleMACS C Tubes (Miltenyi Biotec, no. 130-093-237). Samples were centrifuged at 2,500g for 10 min and the supernatant transferred to a new set of tubes, then frozen at −80 °C for viral titration by plaque assay. Infectious titres were quantified by plaque assay titration using Vero-E6 cells as described above. Unpaired t-testing was used for tissue distribution analysis.
Mouse lung histological analysis
Tissue samples were embedded in paraffin. Lung tissue blocks were cut into 5 μm sections in the Gladstone pathology and light microscope core facility. Sections were stained with H&E, imaged under digital light microscopy in the UCSF Nikon imaging center and then analysed by ImageJ. H&E-stained lung sections were examined using a semiquantitative method previously described35. The severity of four histopathological parameters—(1) perivascular inflammation, (2) bronchial or bronchiolar epithelial degeneration or necrosis, (3) bronchial or bronchiolar inflammation and (4) alveolar inflammation)—was defined and a score assigned for each parameter (0, none; 1, <25%; 2, 26–50%; 3, 51–75%; and 4, >75%). Unpaired t-testing was used for statistical analysis.
Immunofluorescence examination of lung sections
Deparaffinization, rehydration and heat-induced epitope retrieval were performed on an ST4020 small linear stainer (Leica Biosystems). For deparaffinization, slides were baked at 70 °C for 1.0–1.5 h followed by rehydration in descending concentrations of ethanol (100% twice, 95% twice and 80%, 70% and ddH2O twice; each step for 30 s). Washes were performed using a Leica ST4020 Linear Stainer (Leica Biosystems) programmed to three dips per wash for 30 s each. Heat-induced epitope retrieval was performed in a Lab VisionTM PT module (Thermo Fisher) using Dako Target Retrieval Solution pH 9.0 (no. S236784-2, DAKO Agilent) at 97 °C for 10 min and cooled to 65 °C. After further cooling to room temperature for 30 min, slides were washed three times (10 min each) in Tris-buffered saline (TBS) containing 0.1% Tween 20 (TBST, Cell Marque). Sections were then blocked in 5% normal donkey serum in TBST at room temperature for 1 h, followed by incubation with primary antibodies in the blocking solution. After overnight incubation of primary antibodies at 4 °C, sections were washed three times with TBST and stained with the appropriate secondary antibodies in PBS plus 3% bovine serum albumin, 0.4% saponin and 0.02% sodium azide at room temperature for 1 h. Next, sections were washed three times with TBST and mounted with ProLong Gold Antifade mounting medium containing DAPI (Invitrogen). The primary antibodies and final titrations used were mouse antiacetylated α-tubulin (1:300; Santa Cruz, no. sc-23950), rabbit anti-SARS-CoV-2 nucleocapsid (1:1,000; GeneTex, no. GTX135361) and mouse anti-SARS-CoV-2 spike (1:600; GeneTex, no. GTX632604). Secondary antibodies included highly cross-adsorbed donkey antirabbit Alexa Fluor Plus 647 (1:500; Thermo, no. A32795) and highly cross-adsorbed donkey antimouse Alexa Fluor Plus 555 (1:500; Thermo, no. A32773). Fluorescence-immunolabelled images were acquired using either a Zeiss AxioImager Z1 or Keyence BZ-X710 fluorescent microscope. Post-imaging processing was performed using the FIJI package ImageJ 1.52t.
Statistics and reproducibility
All statistics were performed in GraphPad Prism or Microsoft Excel. The IC50 or EC50 of drug inhibition, cytotoxicity and antiviral activity was calculated using the nonlinear fit function (variable slope). All SURF reporter testing and cell immunofluorescence staining were repeated at least three times. Mouse lung section immunofluorescence staining was performed in two mice per group, with three slides per mouse analysed. H&E staining was performed in two mice per group for K18-hACE2 mice, and five mice per group for C57BL6 mice; three slides per mouse were prepared. No statistical methods were used to predetermine sample size, but our sample sizes are similar to those reported in previous publications29,35. Cell cultures and animals were all randomly grouped. Library screening and initial antiviral tests (Figs. 2 and 3) were carried out blindly, labelled with numbers but not compound names. The following experiments were not blind, since differences between compound and control groups were readily observed from imaging data or antiviral tests. No more than two data points, if any, for one drug in drug library screening were excluded due to low imaging quality. One outlier data point in the RT–qPCR assay for the anti-Omicron variant test, DMSO group was excluded due to low RNA yield.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available within the paper and its Supplementary information. The complete screening results and raw data for mouse experiments can be found in the Source Data section.
References
Taylor, P. C. et al. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat. Rev. Immunol. 21, 382–393 (2021).
Dai, L. & Gao, G. F. Viral targets for vaccines against COVID-19. Nat. Rev. Immunol. 21, 73–82 (2021).
Annavajhala, M. K. et al. Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York. Nature 597, 703–708 (2021).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Wang, Q. et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 181, 894–904 (2020).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Yan, R. et al. Structural basis for the recognition of the SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).
Glebov, O. O. Understanding SARS‐CoV‐2 endocytosis for COVID‐19 drug repurposing. FEBS J. 92, 207–208 (2020).
Shang, J. et al. Cell entry mechanisms of SARS-CoV-2. Proc. Natl Acad. Sci. USA 117, 11727–11734 (2020).
Cao, L. et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science 370, 426–431 (2020).
Chan, K. K. et al. Engineering human ACE2 to optimize binding to the spike protein of SARS coronavirus 2. Science 369, 1261–1265 (2020).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Zhang, G., Pomplun, S., Loftis, A. R., Loas, A. & Pentelute, B. L. The first-in-class peptide binder to the SARS-CoV-2 spike protein. Preprint at bioRxiv https://doi.org/10.1101/2020.03.19.999318 (2020).
Kumagai, A. et al. A bilirubin-inducible fluorescent protein from eel muscle. Cell 153, 1602–1611 (2013).
To, T.-L., Zhang, Q. & Shu, X. Structure-guided design of a reversible fluorogenic reporter of protein-protein interactions. Protein Sci. 25, 748–753 (2016).
Szymczak, A. L. et al. Correction of multi-gene deficiency in vivo using a single ‘self-cleaving’ 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Campbell, R. E. et al. A monomeric red fluorescent protein. Proc. Natl Acad. Sci. USA 99, 7877–7882 (2002).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Shu, X., Shaner, N. C., Yarbrough, C. A., Tsien, R. Y. & Remington, S. J. Novel chromophores and buried charges control color in mFruits. Biochemistry 45, 9639–9647 (2006).
Shu, X. et al. Mammalian expression of infrared fluorescent proteins engineered from a bacterial phytochrome. Science 324, 804–807 (2009).
Yu, D. et al. An improved monomeric infrared fluorescent protein for neuronal and tumour brain imaging. Nat. Commun. 5, 3626 (2014).
Zhang, J., Chung, T. & Oldenburg, K. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Case, J. B. et al. Neutralizing antibody and soluble ACE2 inhibition of a replication-competent VSV-SARS-CoV-2 and a clinical isolate of SARS-CoV-2. Cell Host Microbe 28, 475–485 (2020).
Ryu, G. & Shin, H.-W. SARS-CoV-2 infection of airway epithelial cells. Immune Netw. 21, e3 (2021).
Winkler, E. S. et al. SARS-CoV-2 infection of human ACE2-transgenic mice causes severe lung inflammation and impaired function. Nat. Immunol. 21, 1327–1335 (2020).
Dai, W. et al. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science 368, 1331–1335 (2020).
Jin, Z. et al. Structure of Mpro from COVID-19 virus and discovery of its inhibitors. Nature 582, 289–293 (2020).
Zhang, L. et al. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 368, 409–412 (2020).
Li, X. et al. Ethacridine inhibits SARS-CoV-2 by inactivating viral particles. PLoS Pathog. 17, e1009898 (2021).
Yin, W. et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368, 1499–1504 (2020).
Huentelman, M. J. et al. Structure-based discovery of a novel angiotensin-converting enzyme 2 inhibitor. Hypertension 44, 903–906 (2004).
Daly, J. L. et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 370, 861–865 (2020).
Samuel, R. M. et al. Androgen signaling regulates SARS-CoV-2 receptor levels and is associated with severe COVID-19 symptoms in men. Cell Stem Cell 27, 876–889 (2020).
Bender, B. J. et al. A practical guide to large-scale docking. Nat. Protoc. 16, 4799–4832 (2021).
White, K. M. et al. Plitidepsin has potent preclinical efficacy against SARS-CoV-2 by targeting the host protein eEF1A. Science 371, 926–931 (2021).
Acknowledgements
We thank D. Gordon and N. Krogan for sharing the cDNA of SARS-CoV-2 spike protein; S. P. J. Whelan for sharing the rVSV-eGFP-SARS-CoV-2 pseudovirus; S. G. Remesh, K. Leung and J. Wells for help with the the PPI assay; and L. Rodriguez for help with the plaque assay for HBEpCs. This work was supported by the UCSF Program for Breakthrough Biomedical Research (funded in part by the Sandler Foundation), NIH (no. R35GM131766 to X.S. and nos. R01 AI36178, AI40085 and P01 AI091575), the Bill and Melinda Gates Foundation and the DARPA Intercept programme (contract no. HR0011-17-2-0027 to R.A.) and NIH (nos. R35 HL145235 and U19 AI077439 to D.J.E.)
Author information
Authors and Affiliations
Contributions
J.Y. and X.S. initiated the project and designed fluorescent reporters and assays. J.Y., C-I.C. and X.L. prepared the drug and natural compound library. J.Y. performed screens and characterized the kinetics of PPI inhibition in cells. Y.X., P.V.L. and R.A. designed SARS-CoV-2 biosafety level 3 experiments in cells and animals and interpreted data. Y.X. and P.V.L. performed plaque assays and calculated plaque numbers. J.Y., Y.X. and P.V.L. performed quantitative analysis and plotted data. M.A.G.-K. isolated virus variants, performed RT–qPCR and analysed data. J.Y. performed toxicity assays. P.V.L. and Y.X. infected Vero cells. J.Y. performed immunostaining and analysed immunofluorescence. Y.X., P.V.L., T.N., J.V.N., C.-T.W., P.K.J. and R.A. designed HNEpC infection experiments. Y.X. and P.L. performed viral infection. C.-T.W., T.N. and I.T.L. performed cell culturing. C.-T.W. conducted immunostaining and imaging. J.Y. and X.L. performed quantitative analysis. J.Y. and L.R.B. designed HBEpC infection experiments, L.R.B. and K.G. performed HBEpC culture. P.V.L. and M.T. performed viral infection. J.Y. performed immunostaining, imaging and quantitative analysis. S.P. and B.K.S. performed and analysed molecular docking. X.S., J.Y. and Y.X. wrote the manuscript. J.Y., Y.X., P.V.L., M.K., C.-T.W., S.P. and M.T. wrote the methods used. All authors edited and contributed to the final draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interest: J.Y. and X.S. have filed a patent on the imaging assay.
Peer review
Peer review information
Nature Microbiology thanks Anne Moscona and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 High throughput screening of FDA-approved drugs that inhibits spike and ACE2 interaction using the SURF reporter in living cells.
(a) Schematic showing experimental procedure. (b) Volcano plot of ~1600 drugs in inhibiting the PPI of spike and ACE2. HEK293T cells were transfected with the SURF reporter. SURF fluorescence was normalized by mCherry. Cells were incubated with compounds at 10 𝜇M concentration for 24 h before imaging. The dotted horizontal and vertical red lines represent p-value = 0.01 and 40% PPI inhibition, respectively. P value, two-sided non-paired t-test between compound- and DMSO-treated groups (n = 5). (c) Normalized SURF fluorescence of cells incubated with 15 compounds by a second round of imaging. The dotted red line represents 50% PPI inhibition. Data are mean ± s.d. (n = 5). (d) Inhibition kinetics of the identified drugs against spike and ACE2 interaction. Each clinical drug (10 𝜇M) was added to HEK293T cells expressing the SURF reporter, followed by time-lapse imaging. Estimated PPI half-time (𝜏) upon inhibition is shown for each drug. Data are mean ± s.d. (n = 5).
Extended Data Fig. 2 The identified natural compounds inhibit infection of SARS-CoV-2 Delta and Omicron variants.
(a) Schematic showing the experimental design, see methods for details.(b,c) Quantitative analysis of viral titer by plaque assays (B) and viral RNA genome copies by qRT-PCR (C) on Vero E6 cells treated with drugs at 1 𝜇M. qRT-PCR was performed with primers targeting the nucleocapsid gene (N) of SARS-CoV-2, normalized to internal control HRT1. Green bars indicate results of the Delta variant, red bars indicate results of the Omicron variant. The data are shown as mean ± s.d. (n = 3). P value, two-sided non-paired t-test between compound- and DMSO-treated groups. **P < 0.01. ***P < 0.001. Exact P values: (B) starting from bruceine A for Delta: 0.00048, 0.00048, 0.00034, 0.00034. (C) starting from bruceine A for Delta: 0.00052, 0.00052, 0.0022, 0.0021.
Extended Data Fig. 3 Plaque assays and qRT-PCR confirm gamabufotalin and bruceine A inhibit SARS-CoV-2 infection in primary human nasal epithelial cells.
(a) Experimental procedure. (b) Plaque assays using culture medium on top of the transwells. (c) Plaque assays using culture medium under the transwells. (d) qRT-PCR using HBE cells. The data are shown as mean ± s.d. (n = 3). P value, two-sided non-paired t-test between compound- and DMSO-treated groups.
Extended Data Fig. 4 The identified natural compounds inhibit early stages of SARS-CoV-2 infection in primary human bronchial epithelial cells.
(a) Experimental procedure. (b) Immunofluorescence staining of SARS-CoV-2 in the primary human bronchial (HBEpC) epithelial cells. Gamabufotalin, bruceine A or anisomycin were added at different timepoints, at 200 nM, MOI = 1. Nucleocapsid is shown in green. Spike is shown in red. Scale bar: 50 μm. (c) Quantification of the immunofluorescence images in HBEpC. The data are shown as mean ± s.d. (n = 3). P value, two-sided non-paired t-test between compound- and DMSO-treated groups. **P < 0.01. NS: not significant. Exact P values: starting from gamabufotalin anti-N: 0.0036, 0.0057, 0.69, 0.0036, 0.0046, 0.21, 0.0036, 0.0043, 0.11; starting from gamabufotalin anti-S: 0.0039, 0.0059, 0.86, 0.0039, 0.0050, 0.12, 0.0039, 0.0047, 0.083.
Extended Data Fig. 5 Histopathological analysis of the SARS-CoV-2 infected mice.
H & E analysis of lung sections from the infected mice. K18-hACE2 mice were infected with SARS-CoV-2 (1000 PFU) intranasally with the natural compounds bruceine A, gamabufotalin or vehicle control (solvent) at day 0. Then the inhibitors were delivered intranasally once per day at day 1 and day 2. At day 3 (that is, 3 days post-infection), the lungs were harvested and fixed in 4% PFA and embedding into paraffin and cut at 5 μm slides, stained with hematoxylin and eosin (H & E). Black boxes point to regions of the lung anatomy where inflammation was assessed. The magnified images corresponding to the boxed areas are shown on the right panels. Regions where inflammation was detected are indicated by arrows. Scale bar: 200 μm. The H & E staining was performed in two mice per group and 3 slides per mouse were analyzed.
Extended Data Fig. 6 Pathological scores of lung sections.
The semi-quantitative method is described in the methods. For the histology slides, we selected two representative mice out of the 6 mice based on the plaque assay data (Fig. 7B). The dotted black line represents pathology score = 0. Data are mean ± s.d. N = 2 independent animals for each treatment group. 3 slides per animal were examined. P value, two-sided non-paired t-test between treated and vehicle groups. ****p < 0.0001. Exact P values: bruceine A 1.4E-07, gamabufotalin 2.8E-07.
Extended Data Fig. 7 Bruceine A and gamabufotalin exhibit mild activity in inhibiting infection of SARS-CoV-2 B.1.1.7 strain in the C57BL6/J mice.
(a) Schematic of the experimental design. C57BL6/J mice were infected by the intranasal route with 105 pfu of SARS-CoV-2 (B.1.1.7). Bruceine A and gamabufotalin were used at 3.25 mg/kg by intranasal inoculation at both 24- and 48-h post-infection. (b) Virus loads in the lungs. Lung tissues were collected at 3 days post-infection, homogenized, and supernatants were tittered by plaque assay in Vero-TMPRSS2 cells. Data are shown as mean ± s.d. N = 4 independent animals for compound group, and N = 5 independent animals for vehicle group. 2 plaque assays per animal were conducted. Two-tailed Mann-Whitney test was used for statistical analysis. Significance is noted with asterisks as follow: * p < 0.05. Exact P values: bruceine A 0.0412, gamabufotalin 0.0412. (c) Immunofluorescence images of the C57BL6/J mice lung tissues. The lung slides were stained with antibodies against nucleocapsid protein (N), spike (S) and DAPI. Immunofluorescence of N, S and DAPI are shown with pseudo-colors green, red and blue, respectively. Lower panels correspond to the boxed areas of the upper panels. Scale bar, 100 μm (upper panels); 50 μm (lower panels). (d) Quantification of the immunofluorescence. Data are shown as mean ± s.d. N = 2 independent animals for each treatment group. Three slides per animal were examined. P value, two-sided non-paired t-test between compound- and DMSO-treated groups. ****P < 0.0001. Exact P values: anti-N: bruceine A 1.1E-07, gamabufotalin 1.0E-07; anti-S: bruceine A 1.1E-07, gamabufotalin 5.8E-08.
Extended Data Fig. 8 Histopathological analysis of the infection of SARS-CoV-2 B.1.1.7 strain in C57BL6 mice.
(a) H & E staining of lung sections from the infected mice. C57BL6 mice were infected by the intranasal route with 105 pfu of SARS-CoV-2 (B.1.1.7). Bruceine A and gamabufotalin were used as 3.25 mg/kg by intranasal inoculation at both 24- and 48-h post-infection. At day 3 (that is, 3 days post-infection), lungs were harvested and fixed in 4% PFA and embedding into paraffin and cut at 5 μm slides, stained with hematoxylin and eosin (H & E). Black boxes point to regions of the lung anatomy where inflammation was assessed. The magnified images corresponding to the boxed areas are shown on the right panels. Regions where inflammation was detected are indicated by arrows. Scale bar: 200 μm. (b) Pathological scores of the lung sections. Five mice in each group were analyzed using a semi-quantitative method described in the methods. The dotted black line represents pathology score = 0. Data are shown as mean ± s.d. P value, two-sided non-paired t-test between treated and vehicle groups. *P < 0.05. Exact P values: bruceine A 0.046, gamabufotalin 0.046.
Extended Data Fig. 9 Models of binding poses of Bruceine A (left) and Gamabufotalin right) in the ACE2 active site.
The compounds are colored by orange. The residues in ACE2 active pocket are colored by light green. The hydrogen bonds are illustrated with black solid lines. The co-ordinations with zinc ion are shown with black dashed lines. For both bruceine A and gamabufotalin, the poses with highest possibility were generated via docking the compounds into the active site of ACE2. Bruceine A forms good polar and non-polar interactions with protein residues. The carbonyl and hydroxyl groups coordinate with the zinc ion in the ACE2 protein and another carbonyl group forms decent hydrogen bonds with Arg 518. Bruceine A also has good hydrophobic contacts with Tyr 515, Phe 274, Ile 446 and Thr 371. Gamabufotalin also coordinates with the zinc ion via its carbonyl group and shows hydrophobic contacts with ACE2. These models show possible binding poses of bruceine A and gamabufotalin in human ACE2 protein.
Extended Data Fig. 10 The PPI reporter SURF can be used to image interaction between spike and NRP1.
Spike (529-685aa) and NRP1 (273-429aa) are fused to each part of SURF. The SURF reporter was transiently co-transfected into HEK293T with mApple_T2A_H2B-mIFP. Green: SURF fluorescence. Red: mCherry fluorescence. Blue: mIFP fluorescence. Scale bar: 20 μm. The experiments were repeated for three times independently with similar results.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 with legends.
Supplementary Data
Statistical Source Data for Supplementary Figures.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Fig. 1c
Image Source Data.
Source Data Fig. 1e
Image Source Data.
Source Data Fig. 3f
Image Source Data.
Source Data Fig. 4d
Image Source Data.
Source Data Fig. 5b
Image Source Data.
Source Data Fig. 5c
Image Source Data.
Source Data Fig. 6c
Image Source Data.
Source Data Extended Data Fig. 4b
Image Source Data.
Source Data Extended Data Fig. 5
Image Source Data.
Source Data Extended Data Fig. 7c
Image Source Data.
Source Data Extended Data Fig. 8a
Image Source Data.
Source Data Extended Data Fig. 10
Image Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, J., Xiao, Y., Lidsky, P.V. et al. Fluorogenic reporter enables identification of compounds that inhibit SARS-CoV-2. Nat Microbiol 8, 121–134 (2023). https://doi.org/10.1038/s41564-022-01288-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01288-5
This article is cited by
-
SURFing SARS-CoV-2 inhibition
Nature Reviews Microbiology (2023)