Abstract
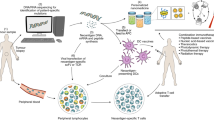
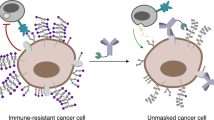
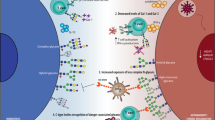
Cancer cell glycocalyx is a major line of defence against immune surveillance. However, how specific physical properties of the glycocalyx are regulated on a molecular level, contribute to immune evasion and may be overcome through immunoengineering must be resolved. Here we report how cancer-associated mucins and their glycosylation contribute to the nanoscale material thickness of the glycocalyx and consequently modulate the functional interactions with cytotoxic immune cells. Natural-killer-cell-mediated cytotoxicity is inversely correlated with the glycocalyx thickness of the target cells. Changes in glycocalyx thickness of approximately 10 nm can alter the susceptibility to immune cell attack. Enhanced stimulation of natural killer and T cells through equipment with chimeric antigen receptors can improve the cytotoxicity against mucin-bearing target cells. Alternatively, cytotoxicity can be enhanced through engineering effector cells to display glycocalyx-editing enzymes, including mucinases and sialidases. Together, our results motivate the development of immunoengineering strategies that overcome the glycocalyx armour of cancer cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw data are available from the corresponding author upon request. Source data are provided with this paper.
Code availability
Packages for fitting the SAIM image sequences with the above model have been implemented in C++ and Julia and are available via GitHub at https://github.com/mjc449/SAIMscannerV3 and https://github.com/paszeklab/SAIMFitKit.jl.
References
Shimasaki, N., Jain, A. & Campana, D. NK cells for cancer immunotherapy. Nat. Rev. Drug Discov. 19, 200–218 (2020).
Heczey, A. et al. Anti-GD2 CAR-NKT cells in relapsed or refractory neuroblastoma: updated phase 1 trial interim results. Nat. Med. 29, 1379–1388 (2023).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Ghasempour, S. & Freeman, S. A. The glycocalyx and immune evasion in cancer. FEBS J. 290, 55–65 (2021).
Möckl, L. The emerging role of the mammalian glycocalyx in functional membrane organization and immune system regulation. Front. Cell Dev. Biol. 8, 253 (2020).
Zhou, J. Y. & Cobb, B. A. Glycans in immunologic health and disease. Annu. Rev. Immunol. 39, 511–536 (2021).
Fernandes, Â. et al. Glycans as shapers of tumour microenvironment: a sweet driver of T‐cell‐mediated anti‐tumour immune response. Immunology 168, 217–232 (2023).
van de Wiel-van Kemenade, E. et al. Episialin (MUC1) inhibits cytotoxic lymphocyte-target cell interaction. J. Immunol. 151, 767–776 (1993).
Hollingsworth, M. A. & Swanson, B. J. Mucins in cancer: protection and control of the cell surface. Nat. Rev. Cancer 4, 45–60 (2004).
Shurer, C. R. et al. Physical principles of membrane shape regulation by the glycocalyx. Cell 177, 1757–1770.e21 (2019).
Paszek, M. J. et al. The cancer glycocalyx mechanically primes integrin-mediated growth and survival. Nature 511, 319–325 (2014).
Kuo, J. C.-H., Gandhi, J. G., Zia, R. N. & Paszek, M. J. Physical biology of the cancer cell glycocalyx. Nat. Phys. 14, 658–669 (2018).
Bell, G. I., Dembo, M. & Bongrand, P. Cell adhesion. Competition between nonspecific repulsion and specific bonding. Biophys. J. 45, 1051–1064 (1984).
Suzuki, Y., Sutoh, M., Hatakeyama, S. & Mori, K. MUC1 carrying core 2 O-glycans functions as a molecular shield against NK cell attack, promoting bladder tumor metastasis. Int. J. Oncol. 40, 1831–1838 (2012).
Okamoto, T. et al. Core2 O-glycan-expressing prostate cancer cells are resistant to NK cell immunity. Mol. Med. Rep. 7, 359–364 (2013).
Madsen, C. B. et al. Glycan elongation beyond the mucin associated Tn antigen protects tumor cells from immune-mediated killing. PLoS ONE 8, e72413 (2013).
Tsuboi, S. et al. A novel strategy for evasion of NK cell immunity by tumours expressing core2 O-glycans. EMBO J. 30, 3173–3185 (2011).
Ajo-Franklin, C. M., Ganesan, P. V. & Boxer, S. G. Variable incidence angle fluorescence interference contrast microscopy for z-imaging single objects. Biophys. J. 89, 2759–2769 (2005).
Lambacher, A. & Fromherz, P. Fluorescence interference-contrast microscopy on oxidized silicon using a monomolecular dye layer. Appl. Phys. A 63, 207–216 (1996).
Paszek, M. J. et al. Scanning angle interference microscopy reveals cell dynamics at the nanoscale. Nat. Methods 9, 825–827 (2012).
Colville, M. J., Park, S., Zipfel, W. R. & Paszek, M. J. High-speed device synchronization in optical microscopy with an open-source hardware control platform. Sci. Rep. 9, 12188 (2019).
Colville, M., Park, S., Singh, A., Paszek, M. & Zipfel, W. R. Azimuthal beam scanning microscope design and implementation for axial localization with scanning angle interference microscopy. in Biomedical Engineering Technologies. Methods in Molecular Biology Vol. 2393, 127–152 (Springer, 2022).
Nason, R. et al. Display of the human mucinome with defined O-glycans by gene engineered cells. Nat. Commun. 12, 4070 (2021).
Lan, Y., Ni, W. & Tai, G. Expression of MUC1 in different tumours and its clinical significance (review). Mol. Clin. Oncol. 17, 161 (2022).
Carson, D. D. The cytoplasmic tail of MUC1: a very busy place. Sci. Signal. 1, pe35 (2008).
Tang, X. et al. First-in-man clinical trial of CAR NK-92 cells: safety test of CD33-CAR NK-92 cells in patients with relapsed and refractory acute myeloid leukemia. Am. J. Cancer Res. 8, 1083–1089 (2018).
Arai, S. et al. Infusion of the allogeneic cell line NK-92 in patients with advanced renal cell cancer or melanoma: a phase I trial. Cytotherapy 10, 625–632 (2008).
Williams, B. A. et al. A phase I trial of NK-92 cells for refractory hematological malignancies relapsing after autologous hematopoietic cell transplantation shows safety and evidence of efficacy. Oncotarget 8, 89256–89268 (2017).
Yu, A. C. Y., Worrall, L. J. & Strynadka, N. C. J. Structural insight into the bacterial mucinase StcE essential to adhesion and immune evasion during enterohemorrhagic E. coli infection. Structure 20, 707–717 (2012).
Walsh, M. D., Luckie, S. M., Cummings, M. C., Antalis, T. M. & McGuckin, M. A. Heterogeneity of MUC1 expression by human breast carcinoma cell lines in vivo and in vitro. Breast Cancer Res. Treat. 58, 255–266 (1999).
van de Wall, S., Santegoets, K. C. M., van Houtum, E. J. H., Büll, C. & Adema, G. J. Sialoglycans and siglecs can shape the tumor immune microenvironment. Trends Immunol. 41, 274–285 (2020).
Hudak, J. E., Canham, S. M. & Bertozzi, C. R. Glycocalyx engineering reveals a Siglec-based mechanism for NK cell immunoevasion. Nat. Chem. Biol. 10, 69–75 (2014).
Rosenstock, P., Horstkorte, R., Gnanapragassam, V. S., Harth, J. & Kielstein, H. Siglec-7 expression is reduced on a natural killer (NK) cell subset of obese humans. Immunol. Res 65, 1017–1024 (2017).
Gandhi, J. G., Koch, D. L. & Paszek, M. J. Equilibrium modeling of the mechanics and structure of the cancer glycocalyx. Biophys. J. 116, 694–708 (2019).
de Gennes, P. G. Polymers at an interface; a simplified view. Adv. Colloid Interface Sci. 27, 189–209 (1987).
Paturej, J., Sheiko, S. S., Panyukov, S. & Rubinstein, M. Molecular structure of bottlebrush polymers in melts. Sci. Adv. 2, e1601478 (2016).
Kudelka, M. R. et al. Cellular O-glycome reporter/amplification to explore O-glycans of living cells. Nat. Methods 13, 81–86 (2016).
Argüeso, P. et al. Association of cell surface mucins with galectin-3 contributes to the ocular surface epithelial barrier. J. Biol. Chem. 284, 23037–23045 (2009).
Lang, M., Hoffmann, M., Dockhorn, R., Werner, M. & Sommer, J.-U. Fluctuation driven height reduction of crosslinked polymer brushes: a Monte Carlo study. J. Chem. Phys. 139, 164903 (2013).
Olguin-Olguin, A. et al. Chemokine-biased robust self-organizing polarization of migrating cells in vivo. Proc. Natl Acad. Sci. USA 118, e2018480118 (2021).
Zhang, C. et al. Chimeric antigen an off-the-shelf cellular therapeutic for targeted elimination of cancer cells and induction of protective antitumor immunity. Front. Immunol. 8, 533 (2017).
Malaker, S. A. et al. The mucin-selective protease StcE enables molecular and functional analysis of human cancer-associated mucins. Proc. Natl Acad. Sci. USA 116, 7278–7287 (2019).
Wang, L. et al. Monoclonal antibody targeting MUC1 and increasing sensitivity to docetaxel as a novel strategy in treating human epithelial ovarian cancer. Cancer Lett. 300, 122–133 (2011).
Cipollone, J. A. et al. The anti-adhesive mucin podocalyxin may help initiate the transperitoneal metastasis of high grade serous ovarian carcinoma. Clin. Exp. Metastasis 29, 239–252 (2012).
Singh, R. et al. Target-specific cytotoxic activity of recombinant immunotoxin scFv(MUC1)-ETA on breast carcinoma cells and primary breast tumors. Mol. Cancer Ther. 6, 562–569 (2007).
Cho, J. H., Collins, J. J. & Wong, W. W. Universal chimeric antigen receptors for multiplexed and logical control of T cell responses. Cell 173, 1426–1438.e11 (2018).
Pedram, K. et al. Design of a mucin-selective protease for targeted degradation of cancer-associated mucins. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01840-6 (2023).
Zhou, H.-X., Rivas, G. & Minton, A. P. Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu. Rev. Biophys. 37, 375–397 (2008).
Pan, H., Colville, M. J., Supekar, N. T., Azadi, P. & Paszek, M. J. Sequence-specific mucins for glycocalyx engineering. ACS Synth. Biol. 8, 2315–2326 (2019).
Schönfeld, K. et al. Selective inhibition of tumor growth by clonal NK cells expressing an ErbB2/HER2-specific chimeric antigen receptor. Mol. Ther. 23, 330–338 (2015).
Xiao, Q. et al. Size-dependent activation of CAR-T cells. Sci. Immunol. 7, eabl3995 (2022).
Shurer, C. R. et al. Genetically encoded toolbox for glycocalyx engineering: tunable control of cell adhesion, survival, and cancer cell behaviors. ACS Biomater. Sci. Eng. 4, 388–399 (2018).
Chitirala, P. et al. Studying the biology of cytotoxic T lymphocytes in vivo with a fluorescent granzyme B-mTFP knock-in mouse. eLife 9, e58065 (2020).
Shinoda, H. et al. Acid-tolerant monomeric GFP from olindias FORMOSA. Cell Chem. Biol. 25, 330–338.e7 (2018).
Bryceson, Y. et al. Functional analysis of human NK cells by flow cytometry. in Natural Killer Cell Protocols 612, 335–352 Humana Press, (2010).
Wang, S.-S. et al. Efficient inhibition of O-glycan biosynthesis using the hexosamine analog Ac5GalNTGc. Cell Chem. Biol. 28, 699–710.e5 (2021).
Acknowledgements
This investigation was supported by the National Science Foundation (NSF) grant 1752226 (M.J.P.), the Breast Cancer Coalition of Rochester predoctoral fellowship (S.P.) and the following National Institutes of Health (NIH) grants: CA276398 (M.J.P.), GM229133 (M.J.P.), CA210184 (M.J.P. and C.F.), CA193043 (M.J.P. and W.R.Z.), GM138692 (M.J.P.), GM137314 (M.J.P and M.D.), AI147362 (M.K.), T32AI007285 (C.J.S.) and CA273349 (S.N.). NGM138692 supported the glycoengineered cell-line development and glycocalyx material characterization. Work was performed at the Cornell Nanoscale Facility (NSF NNCI-2025233), Biotechnology Resource Center (RRID:SCR_021740) and Imaging Facility (RRID:SCR_021741) with NYSTEM (C029155) and NIH (S10OD018516) funding the ZEISS LSM880 instrument. The SAIM instrument development was supported by the Kavli Institute at Cornell for Nanoscale Science and project no. CA193043. We thank X. Su for providing the CD19 CAR cDNA and S. Malaker for helpful insights.
Author information
Authors and Affiliations
Contributions
All authors contributed to the design of experiments, interpretation of results and preparation of the manuscript. Construction of plasmids, preparation of cell lines and execution of experiments were performed by S.P., M.J.C., J.H.P., C.R.S., A.S., E.J.S., C.J.S., L.-T.H., J.C.-H.K, M.C.G. and J.S. M.J.C. built the SAIM microscope. C.J.S. and M.K. generated the CAR T cells. A.S. and S.N. conducted the CORA analysis. S.P. and M.J.P. wrote the manuscript with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
Cornell’s Center for Technology Licensing has submitted a patent related to the findings of this work with two authors (S.P. and M.J.P.) listed as the inventors (PCT/US2022/080937). All other authors declare no competing interests.
Peer review
Peer review information
Nature Materials thanks Kamil Godula, Heinz Läubli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional analysis of Muc1-mediated protection against NK cell cytotoxic function.
a,b, Representative confocal and brightfield images of two clonally expanded MCF10A cell lines expressing Muc1-GFP with 42 tandem repeats (TRs) under the control of a tetracycline inducible promoter at the indicated doxycycline induction level. Scale bars, 10 µm. c, Representative confocal and brightfield images of Muc1-expresssing B2 clonal cells labelled with anti-Human Muc1 Janelia Fluor 549; the doxycycline induction level is indicated. Scale bars, 10 µm. d, NK-92 cell-mediated cytotoxicity against the polyclonal Muc1-GFP and three Muc1-GFP expressing clonal cell lines (1E7, 2E4, and 2G9) expressing MCF10A line from which the clonal lines were isolated, and a clonal MCF10A cell line expressing the Muc1 construct without the GFP reporter (B2 clone); the doxycycline induction levels are indicated. Results are the mean ± s.d of n =3 experimental replicates. NK cell to target cell ratio is 5:1. e. Mean fluorescence intensity of cell-surface Muc1-GFP in 1E7 cells treated with the indicated concentration of StcE mucinase; cell-surface Muc1-GFP probed with Alexa Fluor 647 conjugated GFP nanobody. Results are shown for induction of Muc1-GFP at 100 ng/ml and 1000 ng/ml doxycycline. Results are the mean ± s.d. of at least 14 cells from one representative of 3 independent experiments. Statistical analysis by one-way ANOVA with multiplicity-adjusted P values from Tukey’s multiple comparisons test.
Extended Data Fig. 2 Additional data supporting Siglec-independent protection by the mucin layer.
a, b, Surface expression levels of CD3 (a) and CD56 (b) in NK-92 cells and primary NK cells analyzed by flow cytometry, validating isolation of CD56+/CD3- NK cells. c, d, Surface levels of Siglec-7 (c) and Siglec-9 (d) on NK-92 and primary human NK cells evaluated by flow cytometry, demonstrating minimal to no detectable levels of the receptors on NK-92. e, Western blot analysis confirming UDP-N-acetylglucosamine 2-epimerase (GNE) knockout (KO) in 1E7 cells to ablate sialylation. Flow cytometry data for primary NK cells are representative results for one of n =3 human blood donors.
Extended Data Fig. 3 Measurement of the nanoscale glycocalyx thickness with Ring Scanning Angle Interference Microscopy (Ring-SAIM).
a, Muc1-GFP size standards with biopolymer domains comprised of 0, 10, 21, and 42 tandem repeats (TRs). b, Western blot analysis of Muc1-GFP constructs stably expressed in MCF10A epithelial cells; arrows indicate fully glycosylated Muc1 with 10, 21, or 42 TRs; the primary antibody reacts with the Muc1 TRs and does not probe the 0TR Muc1. c, Flow cytometry analysis of cell-surface Muc1-GFP probed with Alexa Fluor 647 conjugated GFP nanobody. d, Examples of the pixelwise SAIM interferograms for Muc1-GFP constructs labelled with Alexa Fluor 647 conjugated GFP nanobody. e, Representative widefield image and glycocalyx thickness map of live MCF10A epithelial cells expressing the indicated Muc1 constructs labeled with Alexa Fluor 647 conjugated GFP nanobody; the Muc1 x42 mutant construct is comprised of 42 TRs of the mutated Muc1 tandem repeat – PDARPAPGATAPPAHGVTAA – which has three of the five serine/threonine O-glycosylation sites mutated to alanine. Scale bars, 10 µm. f, g, Quantification of glycocalyx thickness in cells expressing the indicated Muc1 constructs (f) and the Muc1-expressing B2 clonal line induced at the indicated doxycycline levels (b). Boxes and whiskers show the first and third quartiles (boxes), median, and range of the data. Each condition includes a minimum of 19 cells from a representative experiment (n=3 independent experiments). Statistical analysis by one-way ANOVA with Tukey’s post-hoc tests. h, Log of the mean glycocalyx thickness versus log of the mucin biopolymer length from (f); best-fit line to a power-law model with slope = 0.16 and R2= 0.62. i, Log of the mean glycocalyx thickness in 1E7 cells expressing Muc1-GFP at varying doxycycline induction levels versus the log of the mean GFP nanobody signal, a proportionate measure of Muc1 surface density; plotted data from Fig. 2f and Fig. 1d; best-fit line to a power-law model has slope = 0.06 and R2 = 0.79. j, Quantification of glycocalyx thickness in cells expressing the Muc1 x42 and the Muc1 x42 mutant construct. Boxes and whiskers show the first and third quartiles (boxes), median, and range of the data. Each condition includes a minimum of 23 cells from a representative experiment (n=3 independent experiments). Statistical analysis by two-tailed t tests. Unless otherwise indicated, doxycycline induction of mucin expression is at 1,000 ng/ml.
Extended Data Fig. 4 Validation of glycoengineered cell lines.
a, Agarose gel electrophoresis of PCR-amplified and EcoRI-digested segment of C1GALT1 showing successful homozygous knockout (KO) in 1E7 cells (See 2A7 sub-clone of 1E7); the homology directed repair (HDR) template for CRISPR/Cas9 mediated C1GALT1 KO introduces a unique EcoRI site. b, Sanger-sequencing results of a PCR-amplified segment of C1GALT1 with insertion of EcoRI restriction site (GAATTC) and stop codon (TAA) in the 1E7 C1GALT1 KO clone, verifying homozygous KO. (Top: 1E7; bottom: 1E7 C1GALT1 KO) c, Fluorescence images of Glucosaminyl (N-Acetyl) Transferase 1 (GCNT1) in wild-type and GCNT1 overexpressing (GCNT1 OE) 1E7 cells with Muc1-GFP expression induced at 1,000 ng/ml of doxycycline; GCNT1 is localized in the Golgi. d, Relative composition of Core O-glycans in wild-type, 1E7 GCNT1 OE, and 1E7 GNE KO cells measured with the Cellular O-glycome Reporter (CORA) method e, Flow cytometry analysis of wild-type and glycoengineered 1E7 cells. Results for the parental MCF10A epithelial cells that do not express Muc1-GFP (-Muc1-GFP) are included as a control. f, Western blot showing the relative size of Muc1 in the wild-type and glycoengineered 1E7 cells. g, sWGA lectin blot after extended SDS-PAGE run time showing increased molecular weight of Muc1-GFP in 1E7 GCNT1 OE cells compared to wild-type 1E7 cells. h, Rate of proliferation of wild-type and glycoengineered 1E7 cells. Results are the mean of 2 independent experiment replicates. i, Flow cytometry analysis of 1E7 cells and their glycoengineered progeny induced at 0 ng/ml and 1,000 ng/ml of doxycycline.
Extended Data Fig. 5 Galectin interactions with Muc1 O-glycans.
a, Quantification of recombinant human galectin-3 (rhGal-3) binding to wild-type, C1GALT1 knockout (KO), and GCNT1 overexpressing (OE) 1E7 cells; data normalized to the Muc1 signal intensity and presented as the mean and ± s.d. of 20 cells from one representative of n = 3 independent experiments. b, Same as in a, with 5,000 U/mL PNGase F treatment to selectively remove N-glycans prior to analysis. c, Quantification of endogenous cell-surface galectin-3 (Gal-3) and Muc1 before and after treatment with 200 nM StcE mucinase. Results are the mean ± s.d. of at least 11 cells from one representative of n = 3 independent experiments. Statistical analysis by two-tailed t tests. d, Representative confocal images of Muc1-GFP and immuno-labelled Gal-3 in wild-type and LGALS3 overexpressing (LGALS3 OE) 1E7 cells. Scale bars, 10 µm. e, Quantification of endogenous Gal-3 on the 1E7 cell surface following treatment with the indicated concentration of galectin inhibitor, TD139; data normalized to the Muc1 signal intensity and presented as the mean ± s.d. of 10 cells from one representative of n = 3 independent experiments. f, Representative confocal images of endogenous galectin-3 on the cell membrane in presence and absence of 50µM TD139. Scale bars, 10 µm. g, Quantification of recombinant human galectin-1 (rhGal-1) binding to wild-type and glycoengineered 1E7 cells; data normalized to the Muc1 signal intensity and presented as the mean ± s.d. of 20 cells from one representative of n = 3 independent experiments. h, Same as in g with 5,000 U/mL PNGase F treatment to selectively remove N-glycans prior to analysis. i, FRAP analysis of Muc1-GFP in wild-type and 1E7 GCNT1 OE cells treated with control buffer, as well as 1E7 GCNT1 OE cells treated with 10 µM rhGal-1. Results are the mean ± s.d. of intensity profiles of n= 11, 10, 9 cells, respectively. j, k, Quantification of mobile fraction (j) and half recovery time (k) from (i). In all experiments, Muc1-GFP expression was induced with 1,000 ng/ml of doxycycline. Unless otherwise indicated, statistical analysis by one-way ANOVA with multiplicity-adjusted P values from Tukey’s multiple comparisons test.
Extended Data Fig. 6 Apoptosis is not inhibited by Muc1-GFP expression.
a, Quantification of 1E7 cell death resulting from treatment with the indicated concentrations of recombinant perforin for 30 minutes; Muc1-GFP expression induced at 1, 100, 1,000 ng/ml doxycycline. Results are the mean ± s.d. of n = 3 technical replicates for one representative of three independent experiments. b, Quantification of cell viability (%) of uninduced (0 ng/ml) and doxycycline-induced (1,000 ng/ml) 1E7 cells following treatment with indicated concentration of doxorubicin for 48 hours. Results are the mean ± s.d. of n = 3 technical replicates.
Extended Data Fig. 7 Validation of endogenous labeling of Granzyme B (GzmB) in NK-92 cells.
Using CRISPR/Cas9 and an appropriate homology directed repair template, the gene sequence for a flexible GGSGGSGGS linker and the pH-tolerant green fluorescent protein, Gamillus, was inserted immediately before the endogenous GZMB stop codon in NK-92 cells. a, Representative brightfield and fluorescence images of NK-92 GzmB-Gamillus cells stained with Lysotracker Deep Red. Scale bar, 10µm. b, Pearson’s coefficients of correlation between GzmB-Gamillus and Lysotracker. Results are the mean ± s.d. of 20 cells. c, d, NK-92 GzmB-Gamillus mediated cytotoxicity against 1E7 with Muc1-GFP expression induced at the indicated doxycycline concentration. Results are the mean ± s.d. of n =3 independent experimental replicates. NK cell to target cell ratio is 10:1. Statistical analysis by one-way ANOVA with multiplicity-adjusted P values from Tukey’s multiple comparisons test. e, Western blot showing the relative expression level of Granzyme B (GzmB) in the NK-92 cells and NK-92 GzmB-Gamillus knock-in (KI) cells. Note that the acid-labile linker between GzmB and Gamillus is expected to undergo cleavage in the low pH of lysosomes and cytolytic granules.
Extended Data Fig. 8 Supporting characterization for StcE NK-92 and Zip-NK-92 cells.
a, Representative confocal images of Muc1-GFP on 1E7 cells treated with control buffer or 100 nM StcE, StcE ΔX409, StcE ΔX409 fused to the EE leucine zipper (StcE-EE); Alexa Fluor 647 conjugated GFP nanobody is used to probe cell-surface Muc1-GFP constructs. Scale bar, 10 µm. b, Viability of NK-92 and Zip-NK-92 after treatment with indicated concentration of StcE-EE. Propidium iodide (20 μg/ml) was used to detect dead cell population by flow cytometry. c, Cytotoxicity mediated by Zip-NK-92 cells coupled with the indicated concentration of Sialidase-EE zipper fusion protein (Sialidase-EE) against B2 cells at the indicated doxycycline induction level; NK cell to target cell ratio is 5:1. Results are the mean ± s.d. of n = 3 technical replicates for each doxycycline concentration.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Tables 1–3, Figs. 1–3 and References.
Source data
Source Data Fig. 1
Source data for Fig. 1.
Source Data Fig. 2
Source data for Fig. 2.
Source Data Fig. 3
Source data for Fig. 3.
Source Data Fig. 4
Source data for Fig. 4.
Source Data Fig. 5
Source data for Fig. 5.
Source Data Fig. 3
Uncropped gels and blots for Fig. 3d.
Source Data Extended Data Fig. 2
Uncropped gels and blots for Extended Data Fig. 2i.
Source Data Extended Data Fig. 3
Uncropped gels and blots for Extended Data Fig. 3b.
Source Data Extended Data Fig. 4a
Uncropped gels and blots for Extended Data Fig. 4a.
Source Data Extended Data Fig. 4f
Uncropped gels and blots for Extended Data Fig. 4f.
Source Data Extended Data Fig. 4g
Uncropped gels and blots for Extended Data Fig. 4g.
Source Data Extended Data Fig. 7
Uncropped gels and blots for Extended Data Fig. 7e.
Source Data Extended Data Fig. 1
Source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Park, S., Colville, M.J., Paek, J.H. et al. Immunoengineering can overcome the glycocalyx armour of cancer cells. Nat. Mater. 23, 429–438 (2024). https://doi.org/10.1038/s41563-024-01808-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-024-01808-0