Abstract
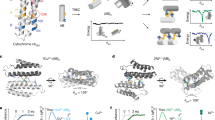
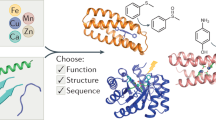
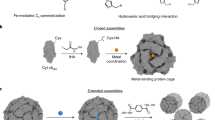
The bottom-up design and construction of functional metalloproteins remains a formidable task in biomolecular design. Although numerous strategies have been used to create new metalloproteins, pre-existing knowledge of the tertiary and quaternary protein structure is often required to generate suitable platforms for robust metal coordination and activity. Here we report an alternative and easily implemented approach (metal active sites by covalent tethering or MASCoT) in which folded protein building blocks are linked by a single disulfide bond to create diverse metal coordination environments within evolutionarily naive protein–protein interfaces. Metalloproteins generated using this strategy uniformly bind a wide array of first-row transition metal ions (MnII, FeII, CoII, NiII, CuII, ZnII and vanadyl) with physiologically relevant thermodynamic affinities (dissociation constants ranging from 700 nM for MnII to 50 fM for CuII). MASCoT readily affords coordinatively unsaturated metal centres—including a penta-His-coordinated non-haem Fe site—and well-defined binding pockets that can accommodate modifications and enable coordination of exogenous ligands such as nitric oxide to the interfacial metal centre.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The principal data supporting the findings of this work are available within the figures and the Supplementary Information. Coordinates and structure factor files for Co–H3 (6DYI), apo–CH3 (6DYB), Co–CH3 (6DYC), Fe–CH3 (6DYE), Cu–CH3 (6DYD), Fe–CH3Y (6DYG), Cu–CH3Y (6DYF), VIVO–CH3Y (6DYH), Mn–CH2E (6DY6), Fe–CH2E (6DY4), Mn–CH2EY (6DY8), Fe–CH3Y* (6DYJ), FeNO–CH3Y* (6DYK), and VIVO–CH3Y* (6DYL) have been deposited to the Protein Data Bank with the corresponding PDB ID codes. Additional data that support the findings of this study are available from the corresponding author on request.
References
Gray, H. B., Stiefel, E. I., Valentine, J. S. & Bertini, I. Biological Inorganic Chemistry: Structure and Reactivity (University Science Books, Sausalito, 2007).
Chen, A. Y. et al. Targetting metalloenzymes for therapeutic intervention. Chem. Rev. 119, 1323–1455 (2019).
Matés, J. M., Segura, J. A., Alonso, F. J. & Márques, J. Intracellular redox status and oxidative stress: implications for cell proliferation, apoptosis, and carcinogenesis. Arch. Toxicol. 82, 273–299 (2008).
Corbin, B. D. et al. Metal chelation and inhibition of bacterial growth in tissue abscesses. Science 319, 962–965 (2008).
Hayden, J. A., Brophy, M. B., Cunden, L. S. & Nolan, E. M. High-affinity manganese coordination by human calprotectin is calcium-dependent and requires a histidine-rich site formed at the dimer interface. J. Am. Chem. Soc. 135, 775–787 (2012).
Umena, Y., Kawakami, K., Shen, J. R. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 angstrom. Nature 473, 55–65 (2011).
de Montellano, P. R. O. Cytochrome P450: Structure, Mechanism, and Biochemistry (Kluwer Academic/Plenum, New York, 2005).
Yu, F. et al. Protein design: toward functional metalloenzymes. Chem. Rev. 114, 3495–3578 (2014).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Lu, Y., Yeung, N., Sieracki, N. & Marshall, N. M. Design of functional metalloproteins. Nature 460, 855–562 (2009).
Soler-Illia, G. J. A. A., Sanchez, C., Lebeau, B. & Patarin, J. Chemical strategies to design textured materials: from microporous and mesoporous oxides to nanonetworks and hierarchical structures. Chem. Rev. 102, 4093–4138 (2002).
Fürstner, A. Olefin metathesis and beyond. Angew. Chem. Int. Ed. 39, 3012–3043 (2000).
Wender, P. A., Verma, V. A., Paxton, T. J. & Pillow, T. H. Function-oriented synthesis, step economy, and drug design. Acc. Chem. Res. 41, 40–49 (2008).
Hyster, T. K., Knorr, L., Ward, T. R. & Rovis, T. Biotinylated Rh(III) complexes in engineered streptavidin for accelerated asymmetric C-H activation. Science 338, 500–503 (2012).
Key, H. M., Dydio, P., Clark, D. S. & Hartwig, J. F. Abiological catalysis by artificial haem proteins containing nobel metals in place of iron. Nature. 534, 534–537 (2016).
Bos, J., Fusetti, F., Diressen, A. J. M. & Roelfes, G. Enantioselective artificial metalloenzymes by creation of a novel active site at the protein dimer interface. Angew. Chem. Int. Ed. 51, 7472–7475 (2012).
Cavazza, C. et al. Crystallographic snapshots of the reaction of aromatic C–H with O2 catalyzed by a protein-bound iron complex. Nat. Chem. 2, 1069–1076 (2010).
Hayashi, T., Hilvert, D. & Green, A. P. Engineered metalloenzymes with non-canonical coordination environments. Chem. Eur. J. 24, 1–11 (2018).
Jeschek, M. et al. Directed evolution of artificial metalloenzymes for in vivo metathesis. Nature 537, 661–665 (2016).
Yang, H. et al. Evolving artificial metalloenzymes via random mutagenesis. Nat. Chem. 10, 318–324 (2018).
Mann, S. I., Heinisch, T., Ward, T. R. & Borovik, A. S. Peroxide activation regulated by hydrogen bonds within artificial Cu proteins. J. Am. Chem. Soc. 139, 17289–17292 (2017).
Yeung, N. et al. Rational design of a structural and functional nitric oxide reductase. Nature 462, 1079–1082 (2009).
Toscano, M. D., Woycechowsky, K. J. & Hilvert, D. Minimalist active-site redesign: teaching old enzymes new tricks. Angew. Chem. Int. Ed. 46, 3212–3236 (2007).
Lombardi, A. et al. Retrostructural analysis of metalloproteins: application to the design of a minimal model for diiron proteins. Proc. Natl Acad. Sci. USA 97, 6298–6305 (2000).
Reig, A. J. et al. Alteration of the oxygen-dependent reactivity of de novo Due Ferri proteins. Nat. Chem. 4, 900–906 (2012).
Koder, R. L. et al. Design and engineering of an O2 transport protein. Nature 458, 305–310 (2009).
Lombardi, A. et al. Miniaturized metalloproteins: applications to iron-sulfur proteins. Proc. Natl Acad. Sci. USA 97, 11922–11927 (2000).
Zastrow, M. L., Peacock, A. F. A., Stuckey, J. A. & Pecoraro, V. L. Hydrolytic catalysis and structural stabilization in a designed metalloprotein. Nat. Chem. 4, 118–123 (2012).
Der, B. S., Edwards, D. R. & Kuhlman, B. Catalysis by a de novo zinc-mediated protein interface: implications for natural enzyme evolution and rational enzyme engineering. Biochemistry 51, 3933–3940 (2012).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143–4148 (2018).
Reetz, M. T. Combinatorial and evolution-based methods in the creation of enantioselective catalysts. Angew. Chem. Int. Ed. 40, 284–310 (2001).
Rufo, C. M. et al. Short peptides self-assemble to produce catalytic amyloids. Nat. Chem. 6, 303–309 (2014).
Salgado, E. N., Radford, R. J. & Tezcan, F. A. Metal-directed protein self-assembly. Acc. Chem. Res. 43, 661–672 (2010).
Churchfield, L. A., Medina-Morales, A., Brodin, J. D., Perez, A. & Tezcan, F. A. De novo design of an allosteric metalloprotein assembly with strained disulfide bonds. J. Am. Chem. Soc. 138, 13163–13166 (2016).
Churchfield, L. A., Alberstein, R. G., Williamson, L. M. & Tezcan, F. A. Determining the structural and energetic basis of allostery in a de novo designed metalloprotein assembly. J. Am. Chem. Soc. 140, 10043–10053 (2018).
Song, W. J. & Tezcan, F. A. A designed supramolecular protein assembly with in vivo enzymatic activity. Science 346, 1525–1528 (2014).
Song, W. J., Yu, J. & Tezcan, F. A. Importance of scaffold flexibility/rigidity in the design and directed evolution of artificial metallo-β-lactamases. J. Am. Chem. Soc. 139, 16772–16779 (2017).
Eijsink, V. G. H. et al. Rational engineering of enzyme stability. J. Biotech. 113, 105–120 (2004).
Dombkowski, A. A., Sultana, K. Z. & Craig, D. B. Protein disulfide engineering. FEBS Lett. 588, 206–212 (2014).
Robertson, D. E. et al. Design and synthesis of multi-haem proteins. Nature 368, 425–432 (1994).
Lichtenstein, B. R. et al. eE Engineering oxidoreductases: maquette proteins designed from scratch. Biochem. Soc. Trans. 40, 561–566 (2012).
Gibney, B. R. et al. Self-assembly of heme A and heme B in a designed four-helix bundle: implications for a cytochrome c oxidase maquette. Biochemistry 39, 11041–11049 (2000).
Faraone-Mennella, J., Tezcan, F. A., Gray, H. B. & Winkler, J. R. Stability and folding kinetics of structurally characterized cytochrome c-b 562. Biochemistry 45, 10504–10511 (2006).
Black, D. S. C. & Hartshorn, A. J. Ligand design and synthesis. Coord. Chem. Rev. 9, 219–274 (1972).
Goncharenko, K. V., Vit, A., Blankenfeldt, W. & Seebeck, F. P. Structure of the sulfoxide synthase EgtB from the ergothioneine biosynthetic pathway. Ang. Chem. Int. Ed. 54, 2821–2824 (2015).
Krissinel, E. & Henrick, K. Interface of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Addison, A. W., Rao, T. N., Reedijk, J., van Rijn, J. & Verschoor, G. C. Synthesis, structure, and spectroscopic properties of copper(II) compounds containing nitrogen–sulphur donor ligands; the crystal and molecular structure of aqua[1,7-bis(N-methylbenzimidazol-2’-yl)-2,6-dithiaheptane]copper(II) perchlorate. J. Chem. Soc. Dalton. Trans. 1984, 1349–1356 (1984).
Smith, T. S., LoBrutto, R. & Pecoraro, V. L. Paramagnetic spectroscopy of vanadyl complexes and its applications to biological systems. Coord. Chem. Rev. 228, 1–18 (2002).
Shi, R. et al. Crystal structures of apo and metal-bound forms of the UreE protein from Helicobacter pylori: role of multiple metal binding sites. Biochemistry 49, 7080–7088 (2010).
Zhu, G., Koszelak-Rosenblum, M., Connelly, S. M., Dumont, M. E. & Malkowski, M. G. The crystal structure of an integral membrane fatty acid α-hydroxylase. J. Biol. Chem. 290, 29820–29833 (2015).
Parmar, A. S., Pike, D. & Nanda, V. in Protein Design Methods and Applications (ed. Köhler, V.) Ch. 12 (Humana, New York, 2014).
Grynkiewicz, G., Poenie, M. & Tsien, R. Y. A New generation of Ca2+ indicators with greatly improved fluorescence properties. J. Biol. Chem. 260, 3440–3450 (1985).
Zastrow, M. L. & Pecoraro, V. L. Designing hydrolytic zinc metalloenzymes. Biochemistry 53, 957–978 (2014).
Vita, C., Roumestand, C., Toma, F. & Ménez, A. Scorpion toxins as natural scaffolds for protein engineering. Proc. Natl Acad. Sci. USA 92, 6404–6408 (1995).
Brodin, J. D. et al. Evolution of metal selectivity in templated protein interfaces. J. Am. Chem. Soc. 132, 8610–8617 (2010).
Hosseinzadeh, P. et al. Enhancing Mn(II)-binding and manganese peroxidase activity in a designed cytochrome c peroxidase through fine-tuning secondary-sphere interactions. Biochemistry 55, 1494–1502 (2016).
Golynskiy, M. V., Gunderson, W. A., Hendrich, M. P. & Cohen, S. M. Metal binding studies and EPR spectroscopy of the manganese transport regulator MntR. Biochemistry 45, 15359–15372 (2006).
Cotruvo, J. A. & Stubbe, J. Metallation and mismetallation of iron and manganese proteins in vitro and in vivo: the class I ribonucleotide reductases as a case study. Metallomics. 4, 1020–1036 (2012).
Koepke, J. et al. pH modulates the quinone position in the photosynthetic reaction center from Rhodobacter sphaeroides in the neutral and charge separated states. J. Mol. Biol. 371, 396–409 (2007).
Chu, R. et al. Redesign of a four-helix bundle protein by phage display coupled with proteolysis and structural characterization by NMR and X-ray crystallography. J. Mol. Biol. 323, 253–262 (2002).
Pasternak, A., Kaplan, J., Lear, J. D. & DeGrado, W. F. Proton and metal ion-dependent assembly of a model diiron protein. Protein Sci. 10, 958–969 (2001).
Zhuang, J. et al. Design of a five-coordinate heme protein maquette: a spectroscopic model of deoxymyoglobin. Inorg. Chem. 43, 8218–8220 (2004).
McLaughlin, M. P. et al. Azurin as a protein scaffold for a low-coordinate non-heme iron site with a small-molecule binding pocket. J. Am. Chem. Soc. 134, 19746–19757 (2012).
Chakraborty, S. et al. Spectroscopic and computational study of a nonheme iron nitrosyl center in a biosynthetic model of nitric oxide reductase. Angew. Chem. Int. Ed. 126, 2449–2453 (2014).
Orville, A. M. & Lipscomb, J. D. Simultaneous binding of nitric oxide and isotopically-labeled substrates of inhibitors by reduced protocatechuate 3,4-dioxygenase. J. Biol. Chem. 268, 8596–8607 (1993).
Martini, R. J. et al. Experimental correlation of substrate position with reaction outcome in the aliphatic halogenase, SyrB2. J. Am. Chem. Soc. 137, 6912–6919 (2015).
Li, M. et al. Tuning the electronic structure of octahedral iron complexes [FeL(X)] (L = 1-alkyl-4,7-bis(4-tert-butyl-2-mercaptobenzyl)-1,4,7-triazacyclononane, X = Cl, CH3O, CN, NO). The S = 1/2, S = 3/2 spin equilibrium of [FeLPr(NO)]. Inorg. Chem. 41, 3444–3456 (2002).
Münck, E. in Physical Methods in Bioinorganic Chemistry (ed. Que, L. Jr) Ch. 6 (University Science Books, Sausalito, 2000).
Ray, M. et al. Structure and magnetic properties of trigonal bipyramidal iron nitrosyl complexes. Inorg. Chem. 38, 3110–3115 (1999).
Speelman, A. L. et al. Non-heme high-spin {FeNO}6-8 complexes: one ligand platform can do it all. J. Am. Chem. Soc. 140, 11341–11359 (2018).
Serres, R. G. et al. Structural, spectroscopic, and computational study of an octahedral, non-haem {Fe-NO}6-8 Series: [Fe(NO)(cyclam-ac)]2+/+/0. J. Am. Chem. Soc. 126, 5138–5153 (2004).
Mbughuni, M. M. et al. Trapping and spectroscopic characterization of an FeIII-superoxo intermediate from a nonheme mononuclear iron-containing enzyme. Proc. Natl Acad. Sci. USA 107, 16788–16793 (2010).
Nakashige, T. G. & Nolan, E. M. Human calprotectin affects the redox speciation of iron. Metallomics 9, 1086–1095 (2017).
Acknowledgements
This work was supported by the National Science Foundation (grant no. CHE1607145 to F.A.T.). J.R. was supported by a postdoctoral fellowship from the National Institute of General Medical Sciences of the National Institute of Health (grant no. F32GM120981). Portions of this research were carried out at the Stanford Synchrotron Radiation Lightsource at the Stanford Linear Accelerator Center and the Advanced Light Source at the Lawrence Berkeley National Laboratory, which are supported by the DOE, Office of Science, and Office of Basic Energy Sciences under contracts DE-AC02-76SF00515 and DE-AC02-05CH11231, respectively. We thank C. Moore and M. Gembicky for assistance with XRD experiments.
Author information
Authors and Affiliations
Contributions
J.R. co-conceived the project, designed and performed experiments, analysed data and co-wrote the paper. M.J.F. and M.T.G. performed EPR and Mössbauer experiments. F.A.T. conceived and directed the project and wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
The Supplementary Information File that accompanies this manuscript includes detailed experimental protocols and a description of the materials and methods employed. Supplementary tables and figures include data of interest to specialists and that support the arguments found in the main text.
Rights and permissions
About this article
Cite this article
Rittle, J., Field, M.J., Green, M.T. et al. An efficient, step-economical strategy for the design of functional metalloproteins. Nat. Chem. 11, 434–441 (2019). https://doi.org/10.1038/s41557-019-0218-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0218-9
This article is cited by
-
Supramolecular assembling systems of hemoproteins using chemical modifications
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2023)
-
Overcoming universal restrictions on metal selectivity by protein design
Nature (2022)
-
Detection of environmental pollutant cadmium in water using a visual bacterial biosensor
Scientific Reports (2022)
-
Recent advances in bacterial biosensing and bioremediation of cadmium pollution: a mini-review
World Journal of Microbiology and Biotechnology (2022)
-
Engineering and emerging applications of artificial metalloenzymes with whole cells
Nature Catalysis (2021)