Abstract
Hydrogen peroxide (HP) is a common disinfectant and antiseptic. When applied to a biofilm, it may be expected that the top layer of the biofilm would be killed by HP, the HP would penetrate further, and eventually eradicate the entire biofilm. However, using the Biofilm.jl computer model, we demonstrate a mechanism by which the biofilm can persist, and even become thicker, in the indefinite treatment with an HP solution at concentrations that are lethal to planktonic microorganisms. This surprising result is found to be dependent on the neutralization of HP by dead biomass, which provides protection for living biomass deeper within the biofilm. Practically, to control a biofilm, this result leads to the concept of treating with an HP dose exceeding a critical threshold concentration rather than a sustained, lower-concentration treatment.
Similar content being viewed by others
Introduction
The ability of microorganisms in biofilms to withstand aggressive antimicrobial treatments has been long recognized1,2,3,4,5. The mechanisms behind this recalcitrance are multifaceted and may include delayed or incomplete diffusive penetration6,7,8, dormancy or persister cell formation9,10,11, protection by extracellular polymeric substances12,13,14,15, and active adaptive responses16,17,18,19,20. Here we conduct an in silico investigation of a particularly remarkable example of tolerance: the sustained persistence of viable biofilm during continuous exposure to a biocide at concentrations sufficient to rapidly kill free-floating cells.
Hydrogen peroxide (H2O2, HP) is a commonly used disinfectant and antiseptic. One of the beneficial features of HP is that it breaks down to innocuous end products, water and oxygen gas. In many microorganisms, this deactivation is catalyzed by enzymes known as catalases. Microorganisms deploy catalases to manage intracellular oxidative stress that is inherent to aerobic respiration21,22,23,24. Microbes often contain multiple genes coding for catalase enzymes. Some catalases can be induced by oxidative stress, while others are constitutively expressed. Catalases disproportionate HP to water and diatomic oxygen, completely neutralizing the antimicrobial power of the biocide. Concentrations of HP used to study oxidative stress responses in bacteria are commonly in the range of 10−4 to 1 mM21,22,23,24, whereas concentrations of HP used to treat biofilms are typically orders of magnitude larger (0.1 to 3% or 30 to 900 mM).
In a biofilm, consumption of HP by catalases in concert with the relatively slow process of molecular diffusion can lead to concentration gradients in HP with decreasing concentration with depth into the biofilm25,26. One might anticipate that as the top layer on the “front line” of the biofilm is killed, HP would penetrate further into the biofilm and eventually eradicate it completely. We used a computer model of biofilm accumulation to analyze this interaction and discovered a mechanism by which the biofilm can persist even in the face of prolonged treatment with a flowing solution of HP.
The one-dimensional mathematical model we used has a phenomenological structure that has a long-standing history in the biofilm literature27,28,29,30,31,32,33,34. It incorporates phenomena of microbial growth, consumption of a solute to support this growth, diffusion of the solute into the biofilm, detachment of microbial biomass from the topmost layer of the biofilm, killing of microbial cells by hydrogen peroxide, consumption of hydrogen peroxide by catalase-containing cells, and diffusion of hydrogen peroxide into the biofilm. The biofilm is simulated as a uniformly thick film growing in a continuous stirred-tank reactor. A companion paper describes the equations and the code for their numerical solution in detail35. We are not aware of any prior modeling work on the problem of continuous treatment of a biofilm with hydrogen peroxide.
Results
Biofilm thickness dynamics and sustained high viability during continuous dosing with hydrogen peroxide
Simulations of untreated biofilm followed a progression of biofilm thickness increasing rapidly from the initial thickness before plateauing at a stable steady-state value. This took about two days and the biofilm attained a steady state thickness of 140 microns (Fig. 1). Biofilm treated with 500 g/m3 HP became thicker in response to the biocide challenge, an unexpected outcome, then plateaued and stably withstood ongoing continuous dosing with the biocide (Fig. 1). When the biocide was discontinued, the biofilm returned within a day to its pre-treatment steady-state thickness (Fig. 1).
Biofilms persisted despite continuous exposure to high concentrations of HP. The steady state biofilm thickness was larger than the pre-treatment thickness for HP dose concentrations up to 3,400 g/m3 (Fig. 2(a)). Only when the HP concentration was increased beyond this amount did the steady-state biofilm thickness diminish below its pre-treatment value. Concentrations of HP greater than approximately 16,600 g/m3 eventually eradicated the biofilm (Fig. 2a) Dead cells were present in the biofilm after exposure to HP, but the percentage of live cells remained quite high even for large biocide dose concentrations. The log reduction in viable cells at steady state in response to continuous treatment with 16,600 g/m3 was 0.86.
Concentration gradients of glucose and HP within the biofilm result in stratified metabolic and biocidal activity
Consumption of glucose by microorganisms in the biofilm resulted in the establishment of a concentration gradient of the growth solute within the biofilm (Fig. 3a). In the untreated (no hydrogen peroxide) case, glucose penetrated about 40 microns into the biofilm (Fig. 3a). When treated with hydrogen peroxide, the glucose concentration in the bulk fluid was higher, presumably because some of the microorganisms were killed and the overall consumption of the biocide was reduced. With the increased glucose concentration in the bulk fluid, glucose penetrated about 70 microns into the biofilm. In both cases, glucose was essentially depleted in the depths of the biofilm. In the untreated case, the region of nutrient availability constituted approximately 29% (40/140) of the biofilm whereas in the treated case the zone of glucose provision was roughly 39% (70/180) of the biofilm. HP also exhibited steep concentration gradients within the biofilm during exposure to the biocide (Fig. 3b). Thus, for both glucose and hydrogen peroxide there is a chemical stratification within the biofilm that creates regions of active growth and regions of biocidal activity. Both of these zones are skewed toward the top of the biofilm.
The distribution of live and dead cells within the HP-treated biofilm was complex and non-uniform (Fig. 4a). Live cells dominated in an interior stratum whereas dead cells dominated at the base of the biofilm and in a thin zone at the very top of the biofilm. This pattern reflects the location of the growing zone in the biofilm, which is also revealed by the peak in the pattern of glucose consumption (Fig. 4b). Dead cells are relatively concentrated at the top of the biofilm because this is where HP is at highest concentrations and disinfection is most rapid. Whereas live cells constituted 100% of cells in untreated biofilm, the mean live cell volume fraction in the treated biofilms was 56%. Note that the data in this plot requires additional time to achieve steady-state due to the slow dynamics near the bottom of the biofilm. Therefore, these results are from simulations with tFinal = 2000 days, which was determined to be sufficient after analyzing the results at a range of simulations with tFinal ∈ [10, 10000] days. The volumetric consumption rate of glucose inside the biofilm was reduced in HP treated biofilms (Fig. 4b). This was not due to the lower availability of glucose as the glucose concentration in the treated biofilm was higher than that in the untreated biofilm (Fig. 3b). The reduction in glucose consumption rate was likely due to the reduced volume fraction of live, metabolically active cells in the treated biofilm.
Biofilm tolerance depends on dead cell neutralization of HP but not live cell neutralization of the biocide
When the neutralization of hydrogen peroxide mediated by dead cells was turned off, the biofilm became susceptible to killing (Fig. 5). Compared to the base case simulation that used both live and dead cell neutralization of HP, turning off dead cell neutralization resulted in the biofilm thickness decaying toward zero (Fig. 5a) as well as the elimination of live cells (Fig. 5b). Doses of HP, that in the baseline simulation were well tolerated, eliminated the biofilm in a simulation where dead cell neutralization of HP was turned off (Fig. 6). For example, whereas a biofilm in which both live and dead cells neutralize HP withstood a continuous dose of 16,600 g/m3, a dose of just 200 g/m3 fully decimated a biofilm in which dead cell neutralization was turned off (Fig. 6). Curiously, turning off the neutralization of hydrogen peroxide mediated by live cells had little influence on biofilm susceptibility (Figs. 5, 6). Therefore, it appears that the catalytic activity of dead cells versus HP is critical to robust biofilm protection, but not that of live cells.
To further analyze the impact of the neutralization of H2O2 by dead cells the biocide neutralization rate kB:D was varied from 0 to 10 gH/m3/d and the thickness of the biofilm at steady-state was computed. At low neutralization rates of less than roughly 2 gH/m3/d the biofilm thickness was small indicating the dead cells were unable to neutralize the biocide fast enough to maintain the biofilm. However, for neutralization rates larger than roughly 4 gH/m3/d, the biofilm became thicker than the case without biocide treatment indicating the result is quite insensitive to the neutralization rate provided the rate is larger than a critical value.
Biofilm tolerance to HP is modulated by the availability of glucose
When the concentration of the growth-limiting solute, glucose, was decreased, the biofilm became less tolerant (Fig. 7). Decreasing glucose from 100 g/m3 in the base case to about 16 g/m3 resulted in the elimination of the biofilm when subjected to the standard HP dose (Fig. 7). Conversely, increasing the influent concentration of glucose made the biofilm somewhat more tolerant (Fig. 7).
Discussion
A computer model of biofilm dynamics predicted robust protection of microorganisms in a biofilm from killing by hydrogen peroxide. In a base case simulation, the log reduction in viable cells after 1 day of continuous exposure of a biofilm to hydrogen peroxide at a mean bulk fluid concentration of 41.4 g/m3 over this interval was 0.063, corresponding to negligible killing. For comparison, if this concentration of HP was delivered to all of the cells in the biofilm for the same duration, the log reduction was calculated to be 9.0. The biofilm was protected because HP did not fully penetrate throughout the biofilm due to a reaction-diffusion interaction. Microbial cells in the outer regions of the biofilm consume and neutralize HP as it diffuses into the biofilm, leading to a standing gradient in biocide concentration within the biofilm (Fig. 3b).
The ability of aggregated microorganisms to neutralize hydrogen peroxide and prevent its penetration into the depths of a biofilm has been experimentally demonstrated. Hydrogen peroxide is biologically inactivated by enzymes, known as catalases, that convert the biocide to water and diatomic oxygen (O2). In a biofilm treated with sufficient HP, the evolution of gaseous oxygen can be visibly observed as effervescence emanating from the biofilm. Incomplete penetration of HP into biofilm was first experimentally demonstrated by Lu et al.25 using microelectrode technology. During 2 h of treatment with 3000 g/m3 of HP, gradients in the concentration of the biocide within the biofilm persisted. Stewart et al.26 used the same approach to demonstrate the failure of HP to penetrate to the bottom of a P. aeruginosa biofilm that was approximately 130 microns thick. In the same study, biofilms formed by a P. aeruginosa katA mutant that was deficient in catalase activity were fully penetrated by the same application of HP26. This result demonstrates that catalase activity alone provides sufficient reactive neutralization of hydrogen peroxide to prevent its effective penetration into a biofilm. Catalase activity has been shown to contribute to biofilm protection from HP in various microorganisms16,36,37,38,39,40.
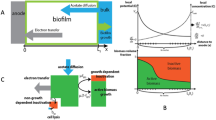
We propose a simplified conceptual model to explain the remarkable tolerance of biofilm to HP and the ability of the biofilm to maintain this defense indefinitely (Fig. 8). This model conceptualizes a zone at the top layer of the biofilm that is predominantly dead cells and an active zone beneath this layer that is predominantly living cells. In reality, the biofilm is not sharply stratified but harbors gradients in dead and live cells. The dead zone contains microbial cells, that while non-viable and unable to grow, nevertheless retain their catalase activity. Catalase does not require ATP or any regenerated co-factor and so this enzyme can continue to operate at full activity within a completely dead cell. The function of the cells in the outer dead layer is to provide a reactive shield that limits the penetration and access of HP to the viable cells in the interior of the biofilm. In the relocated active zone, viable bacteria continue to receive solute (glucose) and are able to grow. Indeed, because the biocide reduces viable bacteria at the surface of the biofilm, less glucose is consumed and the glucose concentration in the bulk fluid increases. This increases the concentration of glucose available to bacteria in the relocated active zone. The function of the growing cells in the relocated active zone is to provide a continuous supply of catalase-containing cells to replenish the cells in the dead zone. Dead cells at the very top of the biofilm detach and are released into the bulk fluid, but they are also continuously replaced by fresh cells coming from the growing layer beneath. There is an essential cohesive function of the biofilm matrix or extracellular polymeric substances in retaining dead cells and their associated catalase activity37. We hypothesize that the mechanistic features described above allow the biofilm to persist even in the face of long-term continuous exposure to HP.
A Prior to biocide treatment, a metabolically active zone (green) resides at the top of the biofilm with metabolically inactive cells in the interior of the biofilm (white). B During HP exposure, the top layer of the biofilm is composed mostly of dead cells (magenta). The metabolically active zone (green) relocates beneath the dead zone. Inactive cells (white) are located at the bottom of the biofilm. Growth of cells in the relocated active zone pushes a continuous supply of catalase-containing cells into the dead zone. Even though many of these cells die as they encounter HP in the upper layer of the biofilm, they retain their catalase activity and are able to consume HP and shield more deeply embedded cells from the biocide.
The conceptual model outlined in the preceding paragraph is sufficient to provide enduring protection from HP. Phenomena that were not included in the computer model are therefore not essential to the prolonged tolerance we have simulated. The model did not incorporate the possibility of an adaptive response, i.e., induction of increased catalase activity upon exposure to HP. The ability of biofilm bacteria to induce catalase in response to oxidative stress is known16,41. If this effect were included in the computational model, one would expect that it would further enhance tolerance. However, it is not necessary to include an adaptive stress response to predict robust protection in the biofilm state. Similarly, the phenomena of dormancy or a shift in metabolic state that makes cells less vulnerable to HP was not included in our simulations. Because there are regions of the biofilm in which cellular metabolic activity is much reduced as a consequence of local deprivation for glucose, such a mechanism is plausible. If this effect were added to the computational model, it would also be expected to increase tolerance.
Most experimental data in the literature involve static, short-term challenge of biofilm with HP15,42,43,44,45,46,47. These are not appropriate comparisons for the simulations we have conducted that incorporate continuous provision of both growth-supporting nutrients as well as the biocide. Our predictions of increased biofilm thickness upon continuous exposure to moderate concentrations of hydrogen peroxide and complete elimination of biofilm at a distinct threshold high concentration have not been experimentally observed or tested. This represents an opportunity for future work by experimenters.
While the comparison to experimental results is therefore very limited, we examine two published studies that provide clues. In one study using P. aeruginosa biofilms grown in the drip-flow biofilm reactor, biofilms were exposed to a continuous flow of 50 mM HP (1700 g/m3) in a glucose-minimal medium for 1 h16,26 This resulted in removal of approximately 25% of the biofilm while viability of the remaining attached cells was reduced to approximately 80%. A quantitative comparison is not possible, but these effects are qualitatively consistent with our simulations (Fig. 2).
An interesting recent study, Zhang et al.48 used bacteria growing in microfluidic devices, interrogated via fluorescence microscopy, to investigate mechanisms of Escherichia coli biofilm response to treatment with HP under continuous cultivation. Biofilms treated with 300 g/m3 HP continued to grow and the katG catalase gene was expressed locally in a narrow region near, but interior, to the top of the biofilm. The HP caused oxidative stress in cells at the periphery of the biofilm as visualized by a reactive oxygen species sensitive fluorescent probe. During HP exposure, the zone of growth was located deeper within the biofilm. These observations are qualitatively consistent with the conceptual model diagrammed in Fig. 8.
The theory presented here should be applicable to any antimicrobial agent that can be enzymatically deactivated in a reaction that is not dependent on the viability of the cell. An example would be the class of β-lactam antibiotics49. These drugs, of which penicillin is a familiar example, contain a β-lactam ring that is cleaved by enzymes known as β-lactamases. This cleavage deactivates the antibiotic and is a primary mechanism of the spread of antibiotic resistance among pathogenic bacteria. In biofilms, β-lactamases have been shown to confer community protection50,51,52 Our model predicts that, just as for catalase-positive biofilms treated with HP, a biofilm formed by a β-lactamase-positive bacterium could exhibit sustained viability and even expansion when continuously treated with a penicillin-class antibiotic.
This work extends prior mathematical and computational models of biofilm inactivation. Previous reports have investigated the dynamics of antimicrobial penetration into biofilms30,53, the contribution of slow-growing or nutrient-limited microbes to antibiotic tolerance29,32,54,55,56, and the phenomenon of persister cells34,55,57. The current study makes a new contribution to understanding biofilm resilience by integrating the reaction-diffusion limited penetration of an antimicrobial with the continual regeneration of catalytic activity within a localized growing zone.
A practical prediction of this set of simulations is that the dose-response behavior of a biofilm treated with HP under continuous flow is highly non-linear. The biofilm will persist despite increasing the dose concentration well above concentrations that are lethal to planktonic microorganisms. This leads to the concept of a threshold treatment concentration that is needed to achieve biofilm control rather than a progressive dose response. Consider for example a biofouling control scenario for which biofilm thickness may be the most appropriate measure of success (rather than viability). HP dose concentrations up to 16,600 g/m3 only reduce the biofilm thickness from the untreated baseline by about 20%. Increasing the HP dose to 16,700 g/m3 results in the complete elimination of the biofilm and zero fouling. Thus, if HP is to be used in a biofouling control capacity, it would be important to establish the threshold concentration for biofilm elimination.
The results of this study also support the biofouling control strategy of reducing nutrients available to support biofilm growth. Reducing nutrient availability not only reduced innate fouling potential prior to biocide treatment, it also reduces the ability of the biofilm to persist during continuous HP exposure (Fig. 7).
Methods
Simulations are conducted with Biofilm.jl58, which models a one-dimensional biofilm within a stirred tank reactor. Details are provided in35 on the governing equations, numerical methods, and application of the solver to a number of test cases. In this paper, the governing equations are described followed by a high-level description of the numerical methods.
The model solves for the concentrations of particulates and solutes within the tank (Xt and St) and biofilm (Xb and Sb) as well as the thickness of the biofilm (Lf). For the biofilm studied in this paper, the particulates are live (L) and dead (D) biomass, and the particulates concentrations can be written as Xt = [Xt:L, Xt:D] within the tank and Xb = [Xb:L, Xb:D] within the biofilm. The solutes in this problem are glucose (G) and hydrogen peroxide (H), and the tank solute concentrations can be written as St = [St:G, St:H] and the biofilm solute concentrations can be written as Sb = [Sb:G, Sb:H].
Tank liquid volume
The concentrations of particulates and solutes are solved for within the tank using
for j ∈ {L, D} and k ∈ {G, H} with initial conditions
The first term on the right-hand-side (RHS) of Eqns. (1) and (2) describes the flow at a flow rate Q into and out of the tank with volume V. The flow of hydrogen peroxide into the tank controls the dosing and is given as
The second terms on the RHS of Eqns (1) and (2) are the flux from the biofilm with surface area A. The flux of particulates is due to detachment from the top of the biofilm \({J}_{\det :j}={v}_{\det }{\left.{X}_{b:j}\right\vert }_{z = {L}_{f}}\) where \({v}_{\det }\) is the detachment velocity that is modeled with
where \({K}_{\det }\) is the detachment coefficient. Particulate attachment is neglected as this process is expected to be important in the early stages of biofilm initiation but not in the long-term persistence of mature biofilms as is the focus in this investigation. The flux of solutes is due to diffusion between the tank and biofilm and is defined below in Eq. (12).
The third term is the rates defined for particulates as
which describes 1) the growth of live biomass using the Monod equation59 with maximum growth rate \({\mu }_{\max }\) and half-saturation constant KM and 2) the death of live biomass due to the presence of hydrogen peroxide with kdis being the disinfection neutralization rate coefficient. The rates for solutes are
which describes the consumption of glucose due to the growth of biomass with the yield coefficient of glucose on biomass YGL and the neutralization of hydrogen peroxide due to live and dead biomass with neutralization rate coefficients of kB:L and kB:D, respectively.
Biofilm volume
The concentration of particulates and solutes are solved for as a function of time t and location z within the biofilm using
for j ∈ {L, D} and k ∈ {G, H} with initial conditions
and boundary conditions
Note that the particulate concentration is related to the volume fraction Pb:j by Pb:j = Xb:j/ρj The first term on the RHS of Eq. (9) describes the vertical transport of particulates due to changes at deeper depths in the biofilm characterized by the growth velocity v(z), i.e.,
where \({P}_{{{{\rm{tot}}}}}=\mathop{\sum }\nolimits_{j = 1}^{{N}_{x}}{X}_{b:j}/{\rho }_{j}\) is the total volume fraction of particulates and ρj is the density of the particulates. The first term on the RHS of Eq. (10) is the diffusion of solutes through the biofilm with a diffusion coefficient Db:k. The rate terms, RX:j and RS:k use the expressions in Eqns. (5) to (8).
The boundary conditions at z = 0, the bottom of the biofilm on the solid surface, enforce a zero-flux condition. The diffusion flux boundary condition at the top of the biofilm (z = Lf) defines Jdif:k and enforces conservation of mass at the surface of the biofilm, i.e., the flux into the biofilm (left-term) matches the flux through a mass transfer boundary layer within the tank on the surface of the biofilm of thickness LL with a diffusion coefficient of Dt:k (middle-term).
Biofilm thickness
The thickness of the biofilm is described by
with initial condition
The thickness depends on the growth velocity at the top of the biofilm (see Eq. (13)) and the detachment velocity (see Eq. (4)).
Numerical methods
The equations are solved using Biofilm.jl which is written in Julia. The main workhorse of Biofilm.jl is the differential equation solver that is part of the DifferentialEquations.jl library60. To solve the equations, the partial differential equations (Eqs. (9) and (10)) are discretized on a one-dimensional grid within the biofilm with Nz = 50 grid cells and solved from t = 0 to tFinal = 100 days. Additional details on how the equations are discretized and solved are provided in Owkes et al.35. The source code used to run the simulations and produce the results is available at https://github.com/markowkes/Biofilm.jl/blob/HydrogenPeroxidePaper/HydrogenPeroxideDosing.jl.
Simulation cases
The solver was run with a number of different cases. The standard parameters are provided in Table 1, which describes the dosing off case. Parameters for system geometry (V, A, Q) were arbitrarily chosen while values of diffusion coefficients61,62, biomass yield coefficient and maximum specific growth rate63, disinfection rate coefficients64 and biocide neutralization rate coefficients65 were based on typical values from literature. The detachment rate coefficient was selected to produce a steady-state biofilm thickness in the dosing off case of approximately 150 microns. Modifications of these parameters for the cases are in Table 2. The dosing on case (B) has a H2O2 dosing after 2 days. Case C has the dosing turn on at 2 days and then off after 6 days. Case D is a series of simulations with varying H2O2 dosing concentrations. Cases E and F explore the influence of the H2O2 neutralization by the dead and live biomass, respectively. Finally, Case G is a series of simulations that vary the influent glucose concentration.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
No data is used in this study. The simulated results are produced with Biofilm.jl, which is described in the subsequent section.
Code availability
The results in this study are produced with the Biofilm.jl software, which is freely available with an MIT license58. To run Biofilm.jl with the appropriate inputs and perform postprocessing to generate the figures in this manuscript can be done with a file called HydrogenPeroxideDosing.jl. This file and the version of Biofilm.jl used with this manuscript is available on the HydrogenPeroxidePaper branch of the git repository https://github.com/markowkes/Biofilm.jl/tree/HydrogenPeroxidePaper. A Code Ocean capsule is also provided that reproduces all the figures in this manuscript https://doi.org/10.24433/CO.8353351.v2.
References
Stewart, P. S. Antimicrobial Tolerance in Biofilms. Microbiol. Spectr. 3(3), 3.07 (2015).
Hall, C. W. & Mah, T.-F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 41, 276–301 (2017).
Yan, J. & Bassler, B. L. Surviving as a Community: Antibiotic Tolerance and Persistence in Bacterial Biofilms. Cell Host Microbe 26, 15–21 (2019).
Ciofu, O., Moser, C., Jensen, P. Ø. & Høiby, N. Tolerance and resistance of microbial biofilms. Nat. Rev. Microbiol. 20, 621–635 (2022).
Sandvik, E. L., Borgogna, T. R. & Stewart, P. S. Antimicrobial and Innate Immune Tolerance Mechanisms in Biofilms. In Richter, K. & Kragh, K. N. (eds.) Antibiofilm Strategies: Current and Future Applications to Prevent, Control and Eradicate Biofilms, 17–35 (Springer International Publishing, Cham, 2022). https://doi.org/10.1007/978-3-031-10992-8_2.
De Beer, D., Srinivasan, R. & Stewart, P. S. Direct measurement of chlorine penetration into biofilms during disinfection. Appl. Environ. Microbiol. 60, 4339–4344 (1994).
Stewart, P. S. & Raquepas, J. B. Implications of reaction-diffusion theory for the disinfection of microbial biofilms by reactive antimicrobial agents. Chem. Eng. Sci. 50, 3099–3104 (1995).
Davison, W. M., Pitts, B. & Stewart, P. S. Spatial and Temporal Patterns of Biocide Action against Staphylococcus epidermidis Biofilms. Antimicrob. Agents Chemother. 54, 2920–2927 (2010).
Anderl, J. N., Zahller, J., Roe, F. & Stewart, P. S. Role of Nutrient Limitation and Stationary-Phase Existence in Klebsiella pneumoniae Biofilm Resistance to Ampicillin and Ciprofloxacin. Antimicrob. Agents Chemother. 47, 1251–1256 (2003).
Borriello, G. et al. Oxygen Limitation Contributes to Antibiotic Tolerance of Pseudomonas aeruginosa in Biofilms. Antimicrob. Agents. Chemother. 48, 2659–2664 (2004).
Keren, I., Kaldalu, N., Spoering, A., Wang, Y. & Lewis, K. Persister cells and tolerance to antimicrobials. FEMS Microbiol. Lett. 230, 13–18 (2004).
Colvin, K. M. et al. The Pel Polysaccharide Can Serve a Structural and Protective Role in the Biofilm Matrix of Pseudomonas aeruginosa. PLOS Pathog. 7, 1–13 (2011).
Billings, N. et al. The Extracellular Matrix Component Psl Provides Fast-Acting Antibiotic Defense in Pseudomonas aeruginosa Biofilms. PLOS Pathog. 9, 1–12 (2013).
Goltermann, L. & Tolker-Nielsen, T. Importance of the Exopolysaccharide Matrix in Antimicrobial Tolerance of Pseudomonas aeruginosa Aggregates. Antimicrob. Agents Chemother. 61, e02696–16 (2017).
Stewart, P. S. et al. Conceptual Model of Biofilm Antibiotic Tolerance That Integrates Phenomena of Diffusion, Metabolism, Gene Expression, and Physiology. J. Bacteriol. 201, e00307–19 (2019).
Elkins, J. G., Hassett, D. J., Stewart, P. S., Schweizer, H. P. & McDermott, T. R. Protective Role of Catalase in Pseudomonas aeruginosa Biofilm Resistance to Hydrogen Peroxide. Appl. Environ. Microbiol. 65, 4594–4600 (1999).
Pamp, S. J., Gjermansen, M., Johansen, H. K. & Tolker-Nielsen, T. Tolerance to the antimicrobial peptide colistin in Pseudomonas aeruginosa biofilms is linked to metabolically active cells, and depends on the pmr and mexAB-oprM genes. Mol. Microbiol. 68, 223–240 (2008).
Nguyen, D. et al. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 334, 982–986 (2011).
Stewart, P. S. et al. Contribution of stress responses to antibiotic tolerance in Pseudomonas aeruginosa Biofilms. Antimicrob. Agents Chemother. 59, 3838–3847 (2015).
Crabbé, A., Jensen, P. Ø., Bjarnsholt, T. & Coenye, T. Antimicrobial tolerance and metabolic adaptations in microbial biofilms. Trends Microbiol. 27, 850–863 (2019).
Farr, S. B. & Kogoma, T. Oxidative stress responses in Escherichia coli and Salmonella typhimurium. Microbiol. Rev. 55, 561–585 (1991).
Belkin, S., Smulski, D. R., Vollmer, A. C., Van Dyk, T. K. & LaRossa, R. A. Oxidative stress detection with Escherichia coli harboring a katG’::lux fusion. Appl. Environ. Microbiol. 62, 2252–2256 (1996).
Imlay, J. A. Cellular defenses against superoxide and hydrogen peroxide. Ann. Rev. Biochem. 77, 755–776 (2008).
Hébrard, M., Viala, J. P. M., Méresse, S., Barras, F. & Aussel, L. Redundant hydrogen peroxide scavengers contribute to Salmonella virulence and oxidative stress resistance. J. Bacteriol. 191, 4605–4614 (2009).
Lu, X., Roe, F., Jesaitis, A. & Lewandowski, Z. Resistance of biofilms to the catalase inhibitor 3-amino-1,2, 4-triazole. Biotechnol. Bioeng. 59, 156–162 (1998).
Stewart, P. S. et al. Effect of catalase on hydrogen peroxide penetration into Pseudomonas aeruginosa biofilms. Appl. Environ. Microbiol. 66, 836–838 (2000).
Horn, H. & Lackner, S. Modeling of biofilm systems: a review. Adv. Biochem. Eng. /Biotechnol. 146, 53–76 (2014).
Wanner, O. & Gujer, W. A multispecies biofilm model. Biotechnol. Bioeng. 28, 314–328 (1986).
Stewart, P. S. Biofilm accumulation model that predicts antibiotic resistance of Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 38, 1052–1058 (1994).
Stewart, P. S., Hamilton, M. A., Goldstein, B. R. & Schneider, B. T. Modeling biocide action against biofilms. Biotechnol. Bioeng. 49, 445–455 (1996).
Wanner, O. & Reichert, P. Mathematical modeling of mixed-culture biofilms. Biotechnol. Bioeng. 49, 172–184 (1996).
Roberts, M. E. & Stewart, P. S. Modeling antibiotic tolerance in biofilms by accounting for nutrient limitation. Antimicrob. Agents Chemother. 48, 48–52 (2004).
Wanner, O. & Morgenroth, E. Biofilm modeling with AQUASIM. Water Sci. Technol. J. Int. Asso. Water Poll. Res. 49, 137–144 (2004).
Roberts, M. E. & Stewart, P. S. Modelling protection from antimicrobial agents in biofilms through the formation of persister cells. Microbiol. (Read., Engl.) 151, 75–80 (2005).
Owkes, M., Coblentza, K., Erikssona, A., Kammerzella, T. & Stewart, P. S. Biofilm.jl: a fast solver for one-dimensional biofilm chemistry and ecology. http://arxiv.org/abs/2307.06153 (2023).
Hassett, D. J. et al. Quorum sensing in Pseudomonas aeruginosa controls expression of catalase and superoxide dismutase genes and mediates biofilm susceptibility to hydrogen peroxide. Mol. Microbiol. 34, 1082–1093 (1999).
Hahn, M. M., González, J. F. & Gunn, J. S. Salmonella Biofilms Tolerate Hydrogen Peroxide by a Combination of Extracellular Polymeric Substance Barrier Function and Catalase Enzymes. Front. Cell. Infec. Microbiol. 11. https://www.frontiersin.org/articles/10.3389/fcimb.2021.683081 (2021).
Fortuna, A. et al. The Pseudomonas aeruginosa DksA1 protein is involved in H2O2 tolerance and within-macrophages survival and can be replaced by DksA2. Sci. Rep. 12, 10404 (2022).
Fernandez, N. L. & Waters, C. M. Cyclic di-GMP increases catalase production and hydrogen peroxide tolerance in Vibrio cholerae. Appl. Environ. Microbiol. 85, e01043–19 (2019).
Khakimova, M., Ahlgren, H. G., Harrison, J. J., English, A. M. & Nguyen, D. The stringent response controls catalases in Pseudomonas aeruginosa and is required for hydrogen peroxide and antibiotic tolerance. J. Bacteriol. 195, 2011–2020 (2013).
Peeters, E., Sass, A., Mahenthiralingam, E., Nelis, H. & Coenye, T. Transcriptional response of Burkholderia cenocepacia J2315 sessile cells to treatments with high doses of hydrogen peroxide and sodium hypochlorite. BMC Genomics 11, 90 (2010).
Wood, P., Jones, M., Bhakoo, M. & Gilbert, P. A novel strategy for control of microbial biofilms through generation of biocide at the Biofilm-Surface Interface. Appl. Environ. Microbiol. 62, 2598–2602 (1996).
Liao, J. & Sauer, K. The MerR-Like Transcriptional Regulator BrlR Contributes to Pseudomonas aeruginosa Biofilm Tolerance. J. Bacteriol. 194, 4823–4836 (2012).
DeQueiroz, G. A. & Day, D. F. Antimicrobial activity and effectiveness of a combination of sodium hypochlorite and hydrogen peroxide in killing and removing Pseudomonas aeruginosa biofilms from surfaces. J. Appl. Microbiol. 103, 794–802 (2007).
Han, Q. et al. Regrowth of microcosm biofilms on titanium surfaces after various antimicrobial treatments. Front. Microbiol. 10, 2693 (2019).
Govaert, M., Smet, C., Verheyen, D., Walsh, J. L. & Van Impe, J. F. M. Combined Effect of Cold Atmospheric Plasma and Hydrogen Peroxide Treatment on Mature Listeria monocytogenes and Salmonella Typhimurium Biofilms. Front. Microbiol. 10, 2674 (2019).
Bjarnsholt, T. et al. Pseudomonas aeruginosa tolerance to tobramycin, hydrogen peroxide and polymorphonuclear leukocytes is quorum-sensing dependent. Microbiology (Reading, England) 151, 373–383 (2005).
Zhang, Y., Cai, Y. & Chen, Z. Community-specific diffusion characteristics determine resistance of biofilms to oxidative stress. Sci. Adv. 9, eade2610 (2023).
Bush, K. & Bradford, P. A. β-Lactams and β-Lactamase Inhibitors: An Overview. Cold Spring Harbor Persp. Med. 6, a025247 (2016).
Budhani, R. K. & Struthers, J. K. Interaction of Streptococcus pneumoniae and Moraxella catarrhalis: Investigation of the Indirect Pathogenic Role of β-Lactamase-Producing Moraxellae by Use of a Continuous-Culture Biofilm System. Antimicrob. Agents. Chemother. 42, 2521–2526 (1998).
Anderl, J. N., Franklin, M. J. & Stewart, P. S. Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 44, 1818–1824 (2000).
Sorg, R. A. et al. Collective Resistance in Microbial Communities by Intracellular Antibiotic Deactivation. PLOS Biol. 14, e2000631 (2016).
Dibdin, G. H., Assinder, S. J., Nichols, W. W. & Lambert, P. A. Mathematical model of beta-lactam penetration into a biofilm of Pseudomonas aeruginosa while undergoing simultaneous inactivation by released beta-lactamases. J. Antimicrob. Chemother. 38, 757–769 (1996).
Cogan, N. G., Cortez, R. & Fauci, L. Modeling physiological resistance in bacterial biofilms. Bull. Math. Biol. 67, 831–853 (2005).
Chambless, J. D., Hunt, S. M. & Stewart, P. S. A Three-Dimensional Computer Model of Four Hypothetical Mechanisms Protecting Biofilms from Antimicrobials. Appl. Environ. Microbiol. 72, 2005–2013 (2006).
Gade, P. A. V. et al. Modelling of ciprofloxacin killing enhanced by hyperbaric oxygen treatment in Pseudomonas aeruginosa PAO1 biofilms. PloS One 13, e0198909 (2018).
De Leenheer, P. & Cogan, N. G. Failure of antibiotic treatment in microbial populations. J. Math. Biol. 59, 563–579 (2009).
Owkes, M. Biofilm.jl. https://github.com/markowkes/Biofilm.jl.
Monod, J. The Growth of Bacterial Cultures. Ann. Rev. Microbiol. 3, 371–394 (1949).
Rackauckas, C. & NIE, Q. DifferentialEquations.jl - A Performant and Feature-Rich Ecosystem for Solving Differential Equations in Julia. J. Open Res. Softw. 5, 15 (2017).
Stewart, P. S. Diffusion in Biofilms. J. Bacteriol. 185, 1485–1491 (2003).
Stewart, P. S. A review of experimental measurements of effective diffusive permeabilities and effective diffusion coefficients in biofilms. Biotechnol. Bioeng. 59, 261–272 (1998).
Robinson, J. A., Trulear, M. G. & Characklis, W. G. Cellular reporoduction and extracellular polymer formation by Pseudomonas aeruginosa in continuous culture. Biotechnol. Bioeng. 26, 1409–1417 (1984).
Stewart, P. S. & Parker, A. E. Measuring Antimicrobial Efficacy against Biofilms: a Meta-analysis. Antimicrob. Agents Chemother. 63, e00020–19 (2019).
Brown, S. M., Howell, M. L., Vasil, M. L., Anderson, A. J. & Hassett, D. J. Cloning and characterization of the katB gene of Pseudomonas aeruginosa encoding a hydrogen peroxide-inducible catalase: purification of KatB, cellular localization, and demonstration that it is essential for optimal resistance to hydrogen peroxide. J. Bacteriol. 177, 6536–6544 (1995).
Author information
Authors and Affiliations
Contributions
P.S. conceptualized the biofilm and was a contributor to writing the manuscript. M.O. ran the simulations, performed post-processing, made the figures, and contributed to writing the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stewart, P.S., Owkes, M. Simulation of catalase-dependent tolerance of microbial biofilm to hydrogen peroxide with a biofilm computer model. npj Biofilms Microbiomes 9, 60 (2023). https://doi.org/10.1038/s41522-023-00426-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-023-00426-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.