Abstract
Cyclin D1 (CCND1), a crucial mediator of cell cycle progression, possesses many mutation types with different mutation frequencies in human cancers. The G870A mutation is the most common mutation in CCND1, which produces two isoforms: full-length CCND1a and divergent C-terminal CCND1b. The dysregulation of the CCND1 isoforms is associated with multiple human cancers. Exploring the molecular mechanism of CCND1 isoforms has offer new insight for cancer treatment. On this basis, the alterations of CCND1 gene are described, including amplification, overexpression, and mutation, especially the G870A mutation. Subsequently, we review the characteristics of CCND1 isoforms caused by G870A mutation. Additionally, we summarize cis-regulatory elements, trans-acting factors, and the splice mutation involved in splicing regulation of CCND1. Furthermore, we highlight the function of CCND1 isoforms in cell cycle, invasion, and metastasis in cancers. Importantly, the clinical role of CCND1 isoforms is also discussed, particularly concerning prognosis, chemotherapy, and radiotherapy. Last, emphasis is given to the corrective strategies that modulate the cancerous CCND1 isoforms. Thus, it is highlighting significance of aberrant isoforms of CCND1 as targets for cancer therapy.
Similar content being viewed by others
Facts
-
The G870A mutation is the most common mutation in CCND1, which produces two isoforms: CCND1a and CCND1b.
-
CCND1 isoforms are involved in tumor growth and progression by regulating cell cycle, invasion, and metastasis.
-
CCND1 isoforms are associated with disease risk and clinical outcome in cancers, and can be used to predict cancer risk, clinical prognosis, or treatment response.
-
Some antisense oligonucleotides or small interfering RNA can target tumor carrying CCND1b for treatment.
Open questions
-
CCND1 isoforms caused by G870A mutation that play a vital role in promoting the malignant phenotype of cancer, and what is the mechanism of its phenotype generation?
-
Can CCND1b be exploited as a prognostic marker of cancers to make more accurate diagnosis, and supervise the response to treatment in patients with cancer?
-
Various therapeutic strategies are available for targeting CCND1b, but what is the most effective therapy for cancer?
Introduction
Cyclin D1 (CCND1), a crucial mediator of cell cycle progression, is the major cyclin involved in transition of cells from the G1 to S phase and plays a vital role in the pathogenesis of cancer [1, 2]. The amplification and/or overexpression of CCND1 have frequently been found in a variety of cancers [3,4,5,6,7,8,9,10,11,12,13,14]. There are a large number of mutations in CCND1, which are closely related to the occurrence, development, prognosis, and treatment of cancers [15,16,17,18,19,20,21]. There are four major mutation types of CCND1, including missense mutation, truncating mutation, inframe mutation, and splice mutation [15]. The G870A mutation is the most common splice mutation in CCND1, and is associated with the risk, prognosis, and treatment of multiple cancers [20, 22, 23]. The conventional CCND1 consists of five distinct exons and has been studied extensively [24]. The G870A mutation produces two CCND1 isoforms by alternative splicing: full-length CCND1a and divergent C-terminal CCND1b [24, 25]. Recent studies have reported that CCND1a and CCND1b had equivalent function in regulating the cell cycle [26,27,28,29]. Zeng et al. found that both CCND1a and CCND1b can promote cell cycle progression and lead to cell proliferation, which may contribute to the potential oncogenic role of CCND1 isoforms in liver cancer Huh-7 and LO2 cell lines in vitro [26]. Interestingly, several studies have shown that CCND1b has different functions from full-length CCND1a [30, 31]. CCND1a can accelerate cell proliferation by promoting cell cycle progression, while CCND1b may inhibit cell cycle progression to prevent cell proliferation [31,32,33,34,35]. This unique activity of CCND1b is particularly significant in cancer treatment [36]. A great deal of study has indicated that dysregulated expression of CCND1 isoforms affect the multiple hallmarks of cancer [27, 37, 38]. For example, CCND1b could promote invasion and metastasis of breast cancer in a CCND1a-independent manner [38]. However, CCND1a conferred the resistance of cancer cells to DNA damage therapy by inducing DNA damage response (DDR) [39]. CCND1a and CCND1b play an essential role in modulating the switch between cell proliferation and death. Hence, this review summarizes the alterations of CCND1 gene, including amplification, overexpression, and mutation, especially the G870A mutation. Subsequently, the characteristics of CCND1 isoforms caused by G870A mutation are described. Additionally, we summarize cis-regulatory elements, trans-acting factors, and the splice mutation involved in splicing regulation of CCND1. Furthermore, we highlight the function of CCND1 isoforms in cell cycle, invasion, and metastasis in cancers. Importantly, the clinical role of CCND1 isoforms is also discussed, particularly concerning prognosis, chemotherapy, and radiotherapy. Last, emphasis is given to the corrective strategies that modulate the cancerous CCND1 isoforms. Thus, our elucidation of CCND1 isoforms from different angles will contribute to better understanding the significance of CCND1 isoforms as a biomarker for cancer and targets for future therapeutic strategies.
Alterations of CCND1 gene
Amplification and overexpression of CCND1 gene
Amplification and overexpression are the most common mechanisms of alteration [40]. Gene amplification is related to the overexpression of oncogenes in cancers [41]. CCND1 is located on chromosome 11q13, consists of 5 exons and 4 introns [42,43,44,45,46]. It was discovered in 1991 and is highly expressed in multiple cancers [43]. Its dysregulation may cause abnormal cell proliferation and contribute to the development of cancer [47, 48]. The amplification and/or overexpression of CCND1 have frequently been found in a variety of cancers [3,4,5,6,7,8,9,10,11,12,13,14]. It was reported that approximately 15–20% of breast cancer contained the amplification of CCND1 [49,50,51]. While the overexpression of CCND1 was observed in more than 50% of breast cancer [42, 52,53,54,55]. This indicates that although CCND1 amplification is closely related to the overexpression of CCND1, the overexpression of CCND1 is not always secondary to gene amplification, and other mechanisms contribute to CCND1 overexpression. Amplification and overexpression of CCND1 preferentially occurred in estrogen receptor (ER)-positive breast cancer [56,57,58,59]. Notably, the amplification of CCND1 is related to the poor prognosis of ER-positive breast cancer and ER-positive tamoxifen-treated breast cancer [4, 60,61,62]. Nevertheless, the prognostic value of CCND1 overexpression is still controversial in breast cancer. Most studies showed that CCND1 overexpression could be considered as a marker of good prognosis in breast cancer [63,64,65,66]. However, some studies suggested that CCND1 overexpression was a poor prognostic marker in breast cancer [56, 67, 68]. In head and neck squamous cell carcinoma, the amplification and overexpression of CCND1 were related to poor prognosis [69,70,71,72]. Interestingly, Kyomoto et al. found that CCND1 amplification was a more effective prognostic marker than its overexpression in human head and neck squamous cell carcinoma [72]. Moreover, Miyamoto et al. demonstrated that CCND1 amplification in oral cancer is a more reliable prognostic marker than CCND1 overexpression [73]. In addition, the amplification and overexpression of CCND1 are considered to be related to poor prognosis in multiple cancers, including gastric cancer [5], esophageal cancer [7], colorectal cancer [10], thyroid papillary cancer [12], pancreatic cancer [13], cholangiocarcinoma [14], cervical cancer [74]. Currently, studies mainly focus on the relationship between the amplification and overexpression of CCND1 and prognosis. In the future, we still need further investigation to fully clarify the role of the amplification and overexpression of CCND1 in cancers, and to make use of its prognosis and predict the value of biomarkers.
Mutation of CCND1 gene
Currently, more than 4000 mutations are discovered in CCND1 by the Catalogue of Somatic Mutations in Cancer and dbSNP databases [75, 76] (Fig. 1a). According to the mutation type of consequence, the major mutations in CCND1 include synonymous mutation, missense mutation, intron mutation, coding sequence mutation, and 3′-UTR mutation. There are four major mutation types of CCND1 based on the cBioPortal database, including missense mutation, truncating mutation, inframe mutation, and splice mutation [15]. The mutations are shown in Fig. 1b based on different mutation types. Missense mutation cause single amino acid substitutions [77]. Truncating mutation leads to protein truncation, which is closely related to many genetic diseases [78, 79]. Splice donor mutation are caused by mutations occurring at the 5′ splice site [24]. Moreover, synonymous mutation does not result in changes in amino acids due to the presence of degenerate codons [80]. Thus, different CCND1 mutations are caused by diverse mechanisms and are related to the pathogenicity and risk of diseases (Fig. 1c).
a The classification of CCND1 mutations is based on different mutation types. b CCND1 is composed of two domains (N-terminal of cyclin_N and C-terminal of cyclin_C). The main mutation types of CCND1 include missense mutations, truncating mutations, inframe mutations, and splice mutations. c Different mutation types are caused by different mechanisms and their clinical significance.
According to The cBioPortal for Cancer Genomics database analysis of samples with mutation data in pan-cancer (TCGA PanCancer Atlas Studies), the mutation frequency of CCND1 in patient is 6% [81, 82]. Plenty of studies have shown that the G870A mutation is the most common splice mutation in CCND1, and is associated with the risk, prognosis, and treatment of multiple cancers [20, 22, 23]. The G870A mutation has been determined to be related to the risk of a variety of cancers such as breast cancer [83], liver cancer [84], colorectal cancer [85], bladder cancer [86], endometrial cancer [87], esophageal and gastric cardiac cancer [88]. Table 1 summarizes the association between the G870A mutation of CCND1 and 17 different cancers. Akhter et al. performed a meta-analysis of 18 published studies to clarify the connection between the G870A mutation and breast cancer risk by increasing statistical power and ultimately found that the AA genotype of G870A mutation was connected with an increased risk of breast cancer [83]. The association between the G870A mutation and cancer risk has been found in a greater part of cancers except for cervical cancer, esophageal squamous cell carcinoma, and prostate cancer [89,90,91]. Therefore, the G870A mutation of CCND1 may be a key risk factor for cancers. In addition, other mutations of CCND1 have been studied, such as rs614367, rs498136, and rs7177 [92, 93]. In the future, we need to further study the role of these mutations in cancers.
CCND1 isoforms caused by G870A mutation
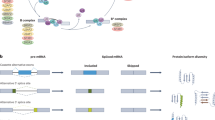
Alternative splicing produces multiple mRNA and protein isoforms by differential selection of splicing sites in precursor (pre)-mRNA (Fig. 2a) [94]. Splicing regulation plays a very important biological function, and aberrant splicing is one of the major causes of human cancer [95, 96]. A growing body of evidence has shown that aberrant splicing is widespread in cancers and plays a crucial role in cell cycle, invasion, metastasis, clinical prognosis, chemotherapy, and radiotherapy (Fig. 2a) [37, 48, 97,98,99]. The splice mutation is associated with multiple cancers and plays a crucial role in the regulation of pre-mRNA splicing [100,101,102]. The G870A mutation has been identified as a crucial splice mutation contributing to the production of CCND1a and CCND1b (Fig. 2b). Comstock et al. demonstrated that G870A mutation was associated with CCND1b by cloning the intron 4 sequences containing either the G or A allele [103]. It is generally recognized that carrying the G allele creates an optimal splicing donor site, resulting in a CCND1 transcript containing all exons (CCND1a). However, carrying the A allele may lead to abnormal splicing events, allowing partial intron 4 to remain and exon 5 to be removed, resulting in a CCND1b splice product. Several studies indicated that the A allele of G870A is preferentially related to the production of CCND1b [24, 25, 104]. CCND1a consists of five exons encoding a DNA sequence of 888 bp, which encodes a protein constituted of 295 amino acids. CCND1b is derived from a splice variant that fails to cleave at the intron 4 junction of exon 4 of CCND1 pre-mRNA to produce intron 4 retention. Since intron 4 contains a translation stop codon, CCND1b encodes a protein that produces 275 amino acids lacking exon 5 [48, 105]. There is a PEST sequence near the C-terminal of CCND1a, which is rich in Pro, Glu, Ser, and Thr residues. These residues play an important role in protein degradation [106]. In addition, the LxxLL motif is present on CCND1 and considered involving in the recruitment of SRC-1 [107]. The CCND1b protein has a completely divergent C-terminal domain, lacking the PEST motif and residues (Thr-286) that control nuclear export and protein stability [48, 108, 109]. Therefore, CCND1b was predicted to be a more stable constitutive nuclear protein with enhanced ability to modulate CDK activity and cell cycle progression. Loss of the LxxLL motif and changes in nuclear localization may alter the transcriptional activity of CCND1b [48]. We also simulate the protein structures of CCND1a and CCND1b using Swiss-Model software (Fig. 2b). It is obvious that the protein structure of CCND1b is different from CCND1a, which may lead to a special role for CCND1b in cancer. In addition, other isoforms encoded by CCND1 have been discovered in different cell types, such as CRA-a and CRA-b (Fig. 2c) [110]. Wiestner et al. have certified that CCND1 containing point mutations produced premature polyadenylation signals, leading to the production of CCND1a with truncated 3′-UTR that can significantly increase the carcinogenicity and worsen the clinical course in patients with mantle cell lymphoma [111].
a Alternative splicing and the effect of aberrant alternative splicing on cancer. The Figure shows some examples of cancer-specific alternative splicing events that contribute to the formation of distinct hallmarks of cancer. b The splice mutation of G870A occurs at the classical splice sites of exon 4 and intron 4. Carrying the G allele produces CCND1a. Carrying A allele produces CCND1b. In addition, the combination of cis-regulatory elements and trans-acting factors affects splice site selection. The protein structures of CCND1a and CCND1b are significantly different, and the mutation site is indicated by red. c General structure characteristics of CCND1 isoforms.
Regulation of CCND1 isoforms production
Cis-regulatory elements
Cis-regulatory elements are short nucleotide motifs (∼5–15 base pairs) that bind to trans-acting factors and affect pre-mRNA splicing [112]. Cis-regulatory regions can present in exons or introns and act as an enhancer or silencer of splicing, specifically controlling alternative splicing by activating or inhibiting the use of adjacent splice sites [113]. The promoter region of CCND1 has a variety of potential cis-regulatory elements that can bind to ATF/CREB, STAT5, STAT3, EGFR, EGR1, and AP1 which are important for pre-mRNA splicing and transcriptional activation of CCND1a in vitro [114,115,116,117,118]. Kang et al. found that a cis-regulatory element between −153 and −134 on CCND1 promoter could bind to the EGR1 transcription factor to regulate CCND1a transcription in vitro and in vivo [117]. Moreover, a cis-regulatory element spanning −144 to −104 of CCND1 promoter can enhance the transcription of CCND1a by binding to TGFα-induced EGR1 [119]. The cis-regulatory elements of CCND1 promoters −58 [114], −954 [118], and −674 to −261 [115] region combined with trans-acting factors are also involved in the regulation of CCND1a transcription in vitro. Therefore, the above cis-regulatory elements can be combined with the corresponding trans-acting factors to regulate CCND1 pre-mRNA splicing.
Trans-acting factors
Trans-acting factors are proteins that recognize or bind to cis-regulatory elements and participate in the formation of splicing regulatory networks, including serine/arginine-rich (SR) proteins, heterogeneous nuclear ribonucleoproteins, and some transcription factors [120]. Through the research on RNA binding protein Sam68, Paronetto et al. found that Sam68 was recruited to CCND1 and interacted with the proximal region of intron 4 to regulate its affinity for CCND1 intron 4, thereby stimulating the increase of CCND1b transcript in human prostate cancer PC3 cell line in vitro [121]. Moreover, the interaction of splice factor SRSF1 with BAF57/SMARCE1 mediated the mechanical stress-induced alternative splicing, producing CCND1b isoform in vitro [122]. Since alternative splicing is coupled with the transcription process, it is found that transcription factors also affected splice selection. Sanchez et al. demonstrated that EWS-FLI1 favored CCND1b expression by reducing the elongation rate, while EWS favored CCND1a expression in breast cancer MCF-7 and rhabdomyosarcoma A673 cell lines in vitro [28]. Additionally, BAF57 (SMARCE1) [122], MYF5 [123], SNIP1 [124], Brm (SMARCA2) [125], and other regulatory factors had also been reported to be involved in the splicing of CCND1 and summarized in Table 2.
CCND1 splice mutation
Mutation of the splice region is associated with multiple cancers and plays a crucial role in the regulation of pre-mRNA splicing [100, 101]. It is generally believed that splice mutation result in recognition of ectopic splice sites through pre-mRNA spliceosome, thereby changing splicing patterns, such as exon jumping and intron retention, and finally modulating the risks of cancer development and outcome [48, 126]. Table 3 summarizes some splice mutations of CCND1 that may regulate pre-mRNA splicing of CCND1. A total of 36 mutations are found as splice mutations of CCND1, such as rs9344 (G870A), rs367683590, rs1268871232 (G139), rs1565224976, and rs201012923. In addition, the indel, somatic single nucleotide variant, somatic insertion, and somatic deletion are found to be related to splice mutation of CCND1. At present, it has been relatively clear that the G870A mutation regulates the splicing of CCND1 pre-mRNA [71, 72]. Figure 2b illustrates the process of G870A mutation regulating the splicing of CCND1 pre-mRNA. This mutation occurs at the boundary of intron 4/exon 5, which is located at the classical splicing donor site. It can regulate the production of CCND1a and CCND1b by interacting with cis-regulatory elements and trans-acting factors. Although cells carrying the A genotype tended to increase the production of transcript b, transcript a (CCND1a) can still be detected [127, 128]. This indicates that other factors may also affect the splicing of CCND1. However, the relationship between other factors and CCND1 splicing is still unclear and needs further investigation.
The function of CCND1 isoforms in cancer
Cell cycle
Alternative splicing of CCND1 pre-mRNA is one of the oncogenic splicing events and is closely associated with the dysregulated cell cycle in cancer cells [129]. Currently, the function of CCND1 isoforms has been extensively studied, and it is generally believed that CCND1 isoforms can affect cell cycle progression through both CDK-dependent and CDK-independent mechanisms (Fig. 3). The mechanism of CCND1a and CCND1b regulating cell cycle in cancers are summarized (Table 4). In the CDK-dependent mechanism for cell cycle, CCND1a can form an active CCND1a-CDK4/6 complex by binding to and activating the G1-phase-specific CDK4/6, resulting in the phosphorylation of G1-phase cycle inhibitor protein (RB). The phosphorylated RB protein is dissociated from the bound transcription factor E2F1 to initiate transcription, which drives cells from G1 to S phase and accelerates cell proliferation (Fig. 3) [130]. Zeng et al. demonstrated that CCND1a and CCND1b can promote cell proliferation by accelerating cell cycle progression in liver cancer Huh-7 and LO2 cell lines in vitro [26]. Kim et al. investigated the effects of CCND1b small interfering RNA (siRNA) in bladder cancer SBT31A and T24 cell lines in vitro and in vivo and found that low expression of CCND1b inhibited the G1-S transition and suppressed cells proliferation [131]. This indicates that CCND1b may have the same effect as CCND1a on the CDK-dependent cell cycle mechanism, and both of them can promote cell cycle progression. However, CCND1b has revealed unexpected disparities in cell cycle regulation. Wang et al. found that overexpression of CCND1b initiated cell cycle arrest and induced apoptosis in cervical cancer HeLa cell line in vitro and in vivo, thereby inhibiting cell proliferation [129]. The reason for this difference may be related to the effect of CCND1b on RB phosphorylation. A lot of studies have shown that although CCND1b could bind to CDK4, it was significantly deficient in inducing RB phosphorylation [31, 105, 132, 133]. Thus, CCND1a and CCND1b may have different biological functions in regulating cell cycle in the CDK-dependent cell cycle mechanism. At present, the mechanism of CCND1b in cell cycle is still unclear. Therefore, how CCND1b regulates the cell cycle progression of cancer cells in a CDK-dependent manner needs to be further explored. In addition to the classical CDK-dependent cell cycle regulating activity, CCND1 itself has CDK-independent effects. CCND1 has been reported to promote cell cycle progression by regulating transcriptional factors and transcriptional coregulators involved at different levels in the cell cycle control, such as ER, androgen receptor (AR), peroxisome proliferator-activated receptor-γ (PPAR γ), SRC1, AIB1, GRIP1, STAT3 and TAFII250 [33, 134,135,136]. Zwijsen et al. revealed that CCND1a could substitute estrogen to activate ER-mediated transcription and contributed to estrogen-induced cell proliferation in estrogen-responsive tissues [34]. When ER is activated by estrogen, activation function domain-2 (AF-2) is exposed that specifically binds to steroid receptor coactivator-1 (SRC-1) [137,138,139]. The recruitment of SRC-1 to ER enhances the binding of the receptor to estrogen-responsive element (ERE), triggering transcription of target gene [107, 137]. The LxxLL motif at positions 254-259 of CCND1a has a similar structure to the AF2 of ER [35]. Therefore, CCND1a can act as a bridging factor to recruit SRC-1 into ER and activate ER-mediated transcription to promote cell cycle progression in breast cancer MCF-7 or T47D cell lines in vitro [34, 35]. However, Zhu et al. found that CCND1b could not induce ER-mediated transcription because it is unable to recruit SRC-1 to the ER [35]. Moreover, the study suggested that CCND1b could inhibit breast cancer cell proliferation by antagonizing the effect of CCND1a on ER-mediated transcription (Fig. 3). In addition, Burd et al. demonstrated that CCND1b had a different transcriptional regulation function from CCND1a in prostate cancer cell line in vitro and in vivo [140]. CCND1a was considered as a key AR corepressor [141,142,143]. CCND1a transcriptional regulation of the AR was manifested by discrete mechanisms. Transcriptional repression mediated by CCND1a binds histone deacetylase 3 (HDAC3) in the androgen-responsive element (ARE) region, thereby limiting androgen-dependent proliferation. Although CCND1b retains the function of AR binding, it selectively impairs the regulation of AR. In particular, CCND1b show impaired AR corepressor activity on the prostate-specific antigen (PSA) promoter. This defect causes CCND1b to stimulate androgen-dependent proliferation, in contrast to the inhibition of cell cycle progression mediated by CCND1a (Fig. 3) [140]. Thus, these studies suggest that the aberrant splicing isoforms play different roles in cell cycle regulation and may be therapeutic targets.
Invasion and metastasis
The isoforms of CCND1 had been deemed to contribute to the invasion and metastasis of various human cancers [38, 144, 145]. The mechanism of CCND1b isoform regulating cell invasion and metastasis in cancers is summarized in Table 4. CCND1b promotes invasion and metastasis in a manner independent of CCND1a in breast cancer MCF-7 cell line in vivo [38, 146]. Specifically, CCND1b can modulate the metastatic phenotype characterized by αvβ3 expression and synergize with HOXD3 to enhance the invasive and metastasis potential of breast cancer cells (Fig. 4a). αvβ3 is a key integrin mediating the invasive and migration of cancer cells [147, 148]. Interestingly, αvβ3 is overexpressed only in metastatic tumor cells, while it is very low expressed in non-metastatic cancer cells [148, 149]. It has been found that even if NF-κB is effectively activated by TLR4 ligand, the expression of αvβ3 cannot be effectively increased by NF-κB in non-metastatic MCF-7 cells [149]. In addition, CCND1b has also been found to promote cell invasion via Erk or RB phosphorylation in bladder cancer and rectal tumorigenesis [144, 145]. By introducing the CCND1b siRNA, Kim et al. demonstrated that the CCND1b siRNA significantly inhibited the invasiveness of bladder cancer cells in vivo [131]. Since CCND1b promotes invasion and metastasis of cancer cells, CCND1b can be used as a new clinical target for anti-metastasis therapy. However, different from the mechanism of CCND1b, CCND1a may participate in the process of cell invasion and metastasis through various mechanisms [1]. Studies have reported the mechanism of a new regulatory axis composed of CCND1-CDK4/Paxilin-Rac1 in the invasion and metastasis of cancer cells [1, 150, 151]. Paxillin (Pxn) is a key component of local adhesion for the monitoring of Rho GTPases [152]. Pxn has many phosphorylation sites, and the CCND1a-CDK4 complex may promote the activation of small GTPase RAC1 by regulating the phosphorylation of Pxn to induce cell invasion and metastasis (Fig. 4b) [1, 151]. Although a lot of studies have shown that CCND1 subtype is related to the invasion and metastasis of cancer cells, how CCND1 isoforms regulate invasion and metastasis independently from the cell cycle still needs to be further explored.
CCND1B transgenic mice models
The CCND1 transgenic mice model has been widely used to find new biological functions of CCND1 and contribute to better understand the role of CCND1 in tumorigenesis in vivo [153, 154]. A large body of literature has shown that genetic alterations of CCND1 are extremely common in human cancers [155]. However, in vitro functional studies have not exhibited transforming activity and CCND1 overexpression has shown weak or no carcinogenic activity in vivo transgenic models [156,157,158,159]. Rodriguez-Puebla et al. confirmed that CCND1 overexpression in mice epidermis can increase cell proliferation and the activity of the cyclin-dependent kinase in vivo, but it does not affect the development of skin tumor by transgenic mice (K5D1 mice) overexpressing CCND1 [157]. Robles et al. also agreed with this view, believing that the expression of CCND1 in epithelial tissues of transgenic mice resulted in epidermal hyperproliferation and severe thymic hyperplasia, which were not related to the development of skin tumors [158]. Yet, studies with CCND1-null mice (D1KO) or CCND1 deficient cells have indicated that CCND1 is necessary for tumor development [160,161,162,163]. This paradox may be related to the existence of alternative splice product of CCND1. By exploring transgenic mice expressing human CCND1b under the control of the bovine K5 promoter (K5D1b mice), Rojas et al. found that K5D1b mice basically had no macroscopic or microscopic phenotype. Interestingly, the skin carcinogenesis of K5D1b mice was enhanced and lack of thymus hyperplasia [164]. The lack of thymus phenotype in K5D1b mice may be related to the functional loss of exon 5 in CCND1b [165]. To elucidate the carcinogenic potential of CCND1b, Kim et al. developed CCND1b transgenic mice and indicated that CCND1b expression was conducive to female-specific rectal carcinogenesis. In addition, Augello et al. described the first-in-field model for switching from CCND1a to CCND1b using a new genetically engineered mice model. It provided the first genetic evidence for CCND1b as an oncogene [166]. Notably, this study not only confirmed the first preclinical evidence for the method of specifically targeting CCND1b-expressing tumors, but also provided the basic principle for developing CCND1b expression as a new biomarker of therapeutic response. Thus, the transgenic mice model of CCND1b is a favorable model for studying the role of CCND1b in cancer, which is essential for preclinical research.
Clinical impaction of CCND1 isoforms in cancer
Clinical prognosis
CCND1 isoforms are associated with disease risk and/or clinical outcome in cancers, and can be used to some extent to predict cancer risk, clinical prognosis, or therapeutic response (Table 4). Multiple clinical studies have suggested that CCND1b overexpression could be used as a prognostic marker in breast cancer, non-small cell lung cancer, and thyroid cancer [30, 167, 168]. Millar et al. showed that high expression of CCND1b was independently relevant to adverse outcomes in breast cancer, including recurrence, metastasis, and decreased survival [30]. However, the high expression of CCND1a was inversely associated with Ki67 markers and not correlated with clinical prognosis [30]. Since CCND1b functions independently of CCND1a, the association between CCND1b and disease outcome is not regulated by the state of CCND1a. Abramson et al. analysed CCND1a and CCND1b expression in primary human breast cancer and found that patients who co-expressed CCND1a and CCND1b had a higher risk of recurrence than the expression of either alone [37]. Additionally, the expression of nuclear CCND1b in papillary thyroid cancer was associated with aggressive clinicopathological features, including lymph node metastasis, risk of recurrence, and advanced stage, while the expression of cytoplasmic CCND1b was related to lymph node metastasis and high risk for cancer recurrence [167]. In non-small cell lung cancer, CCND1b expression was associated with prognosis, whereas CCND1a expression was not relevant to prognosis [168]. In cervical cancer, CCND1a expression was connected to tumor size and degree of differentiation, and CCND1b expression was associated with lymph node metastasis, but it was not related to the prognosis of cervical cancer [169]. Moreover, Gupta et al. indicated that although increased total CCND1 expression was correlated with survival in esophageal adenocarcinoma, the expression of CCND1a and CCND1b was not associated with overall survival [170]. In colorectal cancer, the expression of CCND1a and CCND1b was also not significantly related to prognosis [171]. These studies indicate that CCND1b plays a vital role as a clinical prognosis marker in cancers and warrants further investigation.
Chemotherapy
Chemoresistance is currently obstructing the success of chemotherapy in cancer. CCND1 isoforms have been shown to contribute to clinical responses and provide therapeutic targets for chemotherapeutic drugs [39, 172]. Plenty of evidence suggested that the expression of CCND1a and CCND1b was related to chemoresistance in cancers (Fig. 5) [39, 173]. In Table 4, we summarize the role of two CCND1 isoforms in modulating chemoresistance in cancers. A study on colon cancer revealed that CCND1a increased the phosphorylation of γH2AX induced by chemotherapy agents (doxorubicin or 5-fluorouracil) and recruited Rad51 to local chromatin in response to DNA damage, triggering DDR characterized by phosphorylation of γH2AX in colon cancer HCT116 cell line in vitro. However, CCND1b failed to recruit Rad51 even in response to DNA damage [39]. Regarding breast cancer, the CCND1b siRNA has been demonstrated to synergistically enhance the cell killing effect of doxorubicin, thereby inhibiting tumor growth in mice model [173]. CCND1a was also found to mediate paclitaxel resistance of breast cancer cells through the pRB/E2F1 pathway and AKT phosphorylation [172]. Additionally, Myklebust et al. identified the upregulation of CCND1a expression as a positive predictor of adjuvant 5-fluorouracil and levamisole therapy for colon cancer, especially in stage III colon cancer [171]. Upregulated CCND1a expression was also proved to be involved in chemoresistance to temozolomide in human malignant glioma cells [174]. Consistently, induction of CCND1a silencing in liver cancer HepG2 and SMMC-7721 cell lines significantly increases susceptibility to 5-fluorouracil in vitro [175]. Similarly, this phenomenon had also been observed in gastric cancer AGS cell line [176]. Therefore, the expression of CCND1a and CCND1b during cancer treatment is crucial for the proper selection of chemotherapeutic drugs.
Radiotherapy
The splicing favor of CCND1a is associated with radiosensitivity (Table 4). Fu et al. analysed the expression of CCND1a in biopsy specimens of nasopharyngeal carcinoma patients by immunohistochemical method and found that the expression level of CCND1a was negatively associated with the radiosensitivity of nasopharyngeal carcinoma [177]. Additionally, the high expression of CCND1a improved the sensitivity of radiotherapy for oral squamous cell carcinoma and esophageal cancer in vitro [178, 179]. Moreover, Choi et al. demonstrated that low expression of CCND1a increased proton radiosensitivity in triple-negative breast cancer cells in vitro, possibly because depletion of CCND1a prevents Rad51 recruitment to double-strand break sites [180]. It has been reported that DNA repair induced by upregulated CCND1a expression demonstrated a potential radioresistant mechanism in ependymoma 293T cell line in vitro [181]. A recent study also showed that upregulated CCND1a expression could promote DNA repair and enhance radioresistance in lung cancer cells in vitro [182]. However, no association of CCND1b expression with radiosensitivity was observed. Therefore, these findings suggested that CCND1a is closely related to the radiosensitivity of cancer cells and may play a vital role in the regulation of cellular radiosensitivity.
Strategies modulating CCND1 isoforms in cancer
Antisense oligonucleotides
Small molecule modulators of pre-mRNA splicing represent an attractive option for establishing novel therapeutic strategy in cancer treatment [183]. To date, antisense oligonucleotides (ASO) have been widely used to modulate the splicing mode of pre-mRNA. ASO, typically 15–30 nucleotides, is a short, synthetic, antisense, and modified single-stranded deoxyribonucleotide analogue [184]. It is designed to base pairs in an antisense orientation to a specific pre-mRNA sequence and create a steric hindrance to the binding of splicing factors to the pre-mRNA [185]. This binding alters the recognition of splice sites by the spliceosome, ultimately leading to splicing isoforms switching. Morpholino antisense oligonucleotides (MAOs) are the third-generation ASO, whose ribose is replaced by a morpholino ring and the negatively charged phosphodiester bond is substituted with a neutral phosphoramidate linkage (Fig. 6a) [186]. It has been extensively used to modify splicing and can completely and specifically block splicing events [187,188,189,190]. In general, MAO can base pairs with alternative splice site of CCND1 pre-mRNA to block the aggregation of spliceosome and the binding of splicing factors to their target sequences, which lead to the splicing favorable switch [191]. Since the CCND1b transcript is generated due to a failure to splice at the exon 4/intron 4 boundary of CCND1 pre-mRNA, Augello et al. inhibited splicing at the exon 4/intron 4 boundary by designing an MAO that specifically binds to CCND1 mRNA [191]. Although MAO treatment did not affect overall CCND1 levels, the introduction of MAO into prostate cancer cells effectively suppressed the splicing events of CCND1a and upregulated CCND1b transcripts in a dose-dependent manner, and eventually led to increased prostate cancer cell proliferation and invasion (Fig. 6b). In addition, the study has confirmed that the level of CCND1a is down-regulated by miR-195 overexpression. Zhang et al. designed an ASO against miR-195, which could reduce the level of miR-195 by binding to endogenous miR-195, thereby upregulating CCND1a expression and eventually leading to the proliferation of colorectal cancer cells in vitro [192]. Moreover, a related study found that Bcl-2 silencing was associated with decreased CCND1a expression but not with CCND1b in mantle cell lymphoma cell lines in vitro [193]. Therefore, the use of ASO against Bcl-2 (oblimersen) can also indirectly inhibit the transcription of CCND1a through the interaction between Bcl-2 and CCND1a, resulting in higher proliferation and invasive of mantle cell lymphoma. Figure 6c summarizes some sequences of ASO related to the regulation of CCND1 pre-mRNA splicing. In a word, ASO is an effective strategy to correct the expression of cancer-related CCND1 isoforms through redirection of splicing and rebalancing the ratio of CCND1a/CCND1b.
a Chemical formulae of morpholino and oblimersen. b Morpholino that binds to the exon 4/intron 4 boundary prevents the binding of spliceosome, resulting in a splicing shift to the short isoform CCND1b. siRNA targeting CCND1 isoforms can down-regulate the expression of CCND1 isoforms by binding to the mRNA of target gene. c Sequences of ASO and siRNA used to modulate CCND1 isoforms.
Small interfering RNA
In addition to ASO, RNA interference (RNAi) is the most common transcript-targeted therapy tool that can be used to modulate gene expression [194]. RNAi, the process of machining double-stranded RNA into short siRNA, has been used to target mRNA to downregulate gene expression via degradation of Dicer-RNA induced silencing complex pathway (RISC) [195]. Compared to ASO technologies, RNAi relies on a catalytic mechanism, because after cleavage of the target mRNA, siRNA-loaded RISC can isolate and link to another mRNA molecule, which mainly depends on 100% complementarity of the bind [196, 197]. Therefore, extremely low siRNA concentrations are able to induce efficient target gene knockdown [198]. siRNAs are also widely used to modulate the expression of aberrant isoforms. Some siRNA sequences targeting CCND1b used in studies have been summarized in Fig. 6c. Sanchez et al. efficiently decreased the expression of CCND1b by using the siRNAs that specifically bind to CCND1b, resulting in reduced tumorigenesis in Ewing sarcoma cells in vitro [28]. In general, after the siRNA targeting CCND1a and CCND1b enters the cell, it is integrated to form RISC under the guidance of its antisense strand. The endogenous mRNA with homologous sequences is cleaved by binding to RISC, ultimately leading to the silencing of CCND1a and CCND1b (Fig. 6b). In addition, Kim et al. demonstrated the use of siRNA targeting CCND1b would be a novel therapy for CCND1b-expressing in bladder cancer cell line in vitro. It suppresses the malignant phenotypes of bladder cancer by inducing apoptosis, inhibiting the cancer cell stemness, and epithelial-mesenchymal transition [131]. Moreover, CCND1b can promote the invasion and metastasis of breast cancer cells in vitro, and the use of siRNA targeting CCND1b may provide a new target for the treatment of metastatic breast cancer [27]. Currently, siRNAs have been identified as therapeutic tools for the treatment of cancers [199]. Therefore, siRNA targeting CCND1b is also expected to develop into new anti-cancer drugs.
Conclusions
CCND1 has many mutation types with different mutation frequencies in human cancers. The G870A mutation has an extremely high probability of mutation in clinical tumor and is closely related to the risk, treatment, and prognosis of multiple cancer. The G870A mutation is widely regarded as an attractive target for clinical prediction in cancers. However, how the G870A mutation is generated and its molecular mechanism in cancer need to be further explored. Gene expression disorder caused by abnormal RNA splicing is also an important reason for the genesis and development of cancer [200, 201]. Currently, the regulatory of splicing has become a potent therapeutic strategy for cancer. Studies have proved that the production of CCND1a and CCND1b by alternative splicing of G870A mutation plays a key role in the occurrence and development of cancer. It is generally believed that the imbalanced CCND1a/b ratio can cause cancer, and the high expression of CCND1b is closely related to carcinogenicity. In recent years, in vitro and in vivo studies have revealed the new effect of CCND1b in cell cycle, cell invasion, and metastasis. Moreover, the internal relationship between CCND1b-regulated cell invasion and metastasis has been discovered, providing a new clinical viewpoint for targeting CCND1b anti-metastasis therapy. However, the biological mechanism of preferential splicing of CCND1b results in its high expression in cells is not explicit. Therefore, it is still urgent to further study the mechanisms of CCND1 isoforms in cells. Correction of CCND1 splicing by ASO and small molecule modulators has been shown efficacy in cancer therapy. The development of splicing regulatory drugs targeting CCND1b is expected to become a new option for cancer treatment. Nevertheless, an inhibitor of specific splicing factor for CCND1 splice correction needs to be identified. Besides CCND1, the cyclin family members including CCNA [202], CCNB [203], and CCNE [204] also have multiple splice isoforms, but it is not clear that the multiple splice isoforms of the other family members play a finely coordinated biological role. Thus, there still exists much work in future research on this gene.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Montalto FI, De Amicis F. Cyclin D1 in cancer: a molecular connection for cell cycle control, adhesion and invasion in tumor and stroma. Cells. 2020;9:2648.
Baldin V, Lukas J, Marcote MJ, Pagano M, Draetta G. Cyclin D1 is a nuclear protein required for cell cycle progression in G1. Genes Dev. 1993;7:812–21.
Kaminagakura E, Werneck da Cunha I, Soares FA, Nishimoto IN, Kowalski LP. CCND1 amplification and protein overexpression in oral squamous cell carcinoma of young patients. Head Neck. 2011;33:1413–9.
Jirström K, Stendahl M, Rydén L, Kronblad A, Bendahl PO, Stål O, et al. Adverse effect of adjuvant tamoxifen in premenopausal breast cancer with cyclin D1 gene amplification. Cancer Res. 2005;65:8009–16.
Gao P, Zhou GY, Liu Y, Li JS, Zhen JH, Yuan YP. Alteration of cyclin D1 in gastric carcinoma and its clinicopathologic significance. World J Gastroenterol. 2004;10:2936–9.
Sheyn I, Noffsinger AE, Heffelfinger S, Davis B, Miller MA, Fenoglio-Preiser CM. Amplification and expression of the cyclin D1 gene in anal and esophageal squamous cell carcinomas. Hum Pathol. 1997;28:270–6.
Inomata M, Uchino S, Tanimura H, Shiraishi N, Adachi Y, Kitano S. Amplification and overexpression of cyclin D1 in aggressive human esophageal cancer. Oncol Rep. 1998;5:171–6.
Zhang YJ, Jiang W, Chen CJ, Lee CS, Kahn SM, Santella RM, et al. Amplification and overexpression of cyclin D1 in human hepatocellular carcinoma. Biochem Biophys Res Commun. 1993;196:1010–6.
Reissmann PT, Koga H, Figlin RA, Holmes EC, Slamon DJ. Amplification and overexpression of the cyclin D1 and epidermal growth factor receptor genes in non-small-cell lung cancer. Lung Cancer Study Group. J Cancer Res Clin Oncol 1999;125:61–70.
Bahnassy AA, Zekri AR, El-Houssini S, El-Shehaby AM, Mahmoud MR, Abdallah S, et al. Cyclin A and cyclin D1 as significant prognostic markers in colorectal cancer patients. BMC Gastroenterol. 2004;4:22.
Smalley KS, Lioni M, Dalla Palma M, Xiao M, Desai B, Egyhazi S, et al. Increased cyclin D1 expression can mediate BRAF inhibitor resistance in BRAF V600E-mutated melanomas. Mol Cancer Ther. 2008;7:2876–83.
Várkondi E, Gyory F, Nagy A, Kiss I, Ember I, Kozma L. Oncogene amplification and overexpression of oncoproteins in thyroid papillary cancer. In Vivo. 2005;19:465–70.
Gansauge S, Gansauge F, Ramadani M, Stobbe H, Rau B, Harada N, et al. Overexpression of cyclin D1 in human pancreatic carcinoma is associated with poor prognosis. Cancer Res. 1997;57:1634–7.
Sugimachi K, Aishima S, Taguchi K, Tanaka S, Shimada M, Kajiyama K, et al. The role of overexpression and gene amplification of cyclin D1 in intrahepatic cholangiocarcinoma. J Hepatol. 2001;35:74–9.
Xu J, Lin DI. Oncogenic c-terminal cyclin D1 (CCND1) mutations are enriched in endometrioid endometrial adenocarcinomas. PLoS ONE. 2018;13:e0199688.
Moreno-Bueno G, Rodríguez-Perales S, Sánchez-Estévez C, Hardisson D, Sarrió D, Prat J, et al. Cyclin D1 gene (CCND1) mutations in endometrial cancer. Oncogene. 2003;22:6115–8.
Ikeda Y, Oda K, Hiraike-Wada O, Koso T, Miyasaka A, Kashiyama T, et al. Cyclin D1 harboring the T286I mutation promotes oncogenic activation in endometrial cancer. Oncol Rep. 2013;30:584–8.
Benzeno S, Lu F, Guo M, Barbash O, Zhang F, Herman JG, et al. Identification of mutations that disrupt phosphorylation-dependent nuclear export of cyclin D1. Oncogene. 2006;25:6291–303.
Kowalczyk MM, Barańska M, Fendler W, Borkowska EM, Kobos J, Borowiec M, et al. G870A polymorphic variants of CCND1 gene and cyclin D1 protein expression as prognostic markers in laryngeal lesions. Diagnostics. 2022;12:1059.
El Menshawy N, El Marghany AB, Sarhan MM, Aladle DA. Cyclin D1 G870A polymorphism: relation to the risk of ALL development, prognosis impact, and methotrexate cytotoxicity. Asian Pac J Cancer Prev. 2020;21:2941–7.
Shastry BS. SNPs: impact on gene function and phenotype. Methods Mol Biol. 2009;578:3–22.
Garcia-Aguilar J, Chen Z, Smith DD, Li W, Madoff RD, Cataldo P, et al. Identification of a biomarker profile associated with resistance to neoadjuvant chemoradiation therapy in rectal cancer. Ann Surg. 2011;254:486–92. discussion 92-3
Thakur N, Kumari S, Mehrotra R. Association between Cyclin D1 G870A (rs9344) polymorphism and cancer risk in Indian population: meta-analysis and trial sequential analysis. Biosci Rep. 2018;38:BSR20180694.
Betticher DC, Thatcher N, Altermatt HJ, Hoban P, Ryder WD, Heighway J. Alternate splicing produces a novel cyclin D1 transcript. Oncogene. 1995;11:1005–11.
Howe D, Lynas C. The cyclin D1 alternative transcripts [a] and [b] are expressed in normal and malignant lymphocytes and their relative levels are influenced by the polymorphism at codon 241. Haematologica. 2001;86:563–9.
Zeng Z, Tu J, Cheng J, Yao M, Wu Y, Huang X, et al. Influence of CCND1 G870A polymorphism on the risk of HBV-related HCC and cyclin D1 splicing variant expression in Chinese population. Tumour Biol. 2015;36:6891–900.
Wang Y, Liu Y, Xiang L, Han L, Yao X, Hu Y, et al. Cyclin D1b induces changes in the macrophage phenotype resulting in promotion of tumor metastasis. Exp Biol Med. 2021;246:2559–69.
Sanchez G, Bittencourt D, Laud K, Barbier J, Delattre O, Auboeuf D, et al. Alteration of cyclin D1 transcript elongation by a mutated transcription factor up-regulates the oncogenic D1b splice isoform in cancer. Proc Natl Acad Sci USA. 2008;105:6004–9.
Sanchez G, Delattre O, Auboeuf D, Dutertre M. Coupled alteration of transcription and splicing by a single oncogene: boosting the effect on cyclin D1 activity. Cell Cycle. 2008;7:2299–305.
Millar EK, Dean JL, McNeil CM, O’Toole SA, Henshall SM, Tran T, et al. Cyclin D1b protein expression in breast cancer is independent of cyclin D1a and associated with poor disease outcome. Oncogene. 2009;28:1812–20.
Lévêque C, Marsaud V, Renoir JM, Sola B. Alternative cyclin D1 forms a and b have different biological functions in the cell cycle of B lymphocytes. Exp Cell Res. 2007;313:2719–29.
Lukas J, Parry D, Aagaard L, Mann DJ, Bartkova J, Strauss M, et al. Retinoblastoma-protein-dependent cell-cycle inhibition by the tumour suppressor p16. Nature. 1995;375:503–6.
Chen S, Li L. Degradation strategy of cyclin D1 in cancer cells and the potential clinical application. Front Oncol. 2022;12:949688.
Zwijsen RM, Wientjens E, Klompmaker R, van der Sman J, Bernards R, Michalides RJ. CDK-independent activation of estrogen receptor by cyclin D1. Cell. 1997;88:405–15.
Zhu J, Sen S, Wei C, Frazier ML. Cyclin D1b represses breast cancer cell growth by antagonizing the action of cyclin D1a on estrogen receptor alpha-mediated transcription. Int J Oncol. 2010;36:39–48.
Wang Y, Dean JL, Millar EK, Tran TH, McNeil CM, Burd CJ, et al. Cyclin D1b is aberrantly regulated in response to therapeutic challenge and promotes resistance to estrogen antagonists. Cancer Res. 2008;68:5628–38.
Abramson VG, Troxel AB, Feldman M, Mies C, Wang Y, Sherman L, et al. Cyclin D1b in human breast carcinoma and coexpression with cyclin D1a is associated with poor outcome. Anticancer Res. 2010;30:1279–85.
Wu FH, Luo LQ, Liu Y, Zhan QX, Luo C, Luo J, et al. Cyclin D1b splice variant promotes αvβ3-mediated adhesion and invasive migration of breast cancer cells. Cancer Lett. 2014;355:159–67.
Li Z, Jiao X, Wang C, Shirley LA, Elsaleh H, Dahl O, et al. Alternative cyclin D1 splice forms differentially regulate the DNA damage response. Cancer Res. 2010;70:8802–11.
Nasser MM, Mehdipour P. Exploration of involved key genes and signaling diversity in brain tumors. Cell Mol Neurobiol. 2018;38:393–419.
Mohammedi L, Doula FD, Mesli F, Senhadji R. Cyclin D1 overexpression in Algerian breast cancer women: correlation with CCND1 amplification and clinicopathological parameters. Afr Health Sci. 2019;19:2140–6.
Bartkova J, Lukas J, Müller H, Lützhøft D, Strauss MJB. Cyclin D1 protein expression and function in human breast cancer. Int J Cancer. 1994;57:353–61.
Rosenberg CL, Kim HG, Shows TB, Kronenberg HM, Arnold A. Rearrangement and overexpression of D11S287E, a candidate oncogene on chromosome 11q13 in benign parathyroid tumors. Oncogene. 1991;6:449–53.
Xiong Y, Connolly T, Futcher B, Beach D. Human D-type cyclin. Cell. 1991;65:691–9.
Arnold A, Kim HG, Gaz RD, Eddy RL, Fukushima Y, Byers MG, et al. Molecular cloning and chromosomal mapping of DNA rearranged with the parathyroid hormone gene in a parathyroid adenoma. J Clin Investig. 1989;83:2034–40.
Motokura T, Bloom T, Kim HG, Jüppner H, Ruderman JV, Kronenberg HM, et al. A novel cyclin encoded by a bcl1-linked candidate oncogene. Nature. 1991;350:512–5.
Biliran H Jr., Wang Y, Banerjee S, Xu H, Heng H, Thakur A, et al. Overexpression of cyclin D1 promotes tumor cell growth and confers resistance to cisplatin-mediated apoptosis in an elastase-myc transgene-expressing pancreatic tumor cell line. Clin Cancer Res. 2005;11:6075–86.
Knudsen KE, Diehl JA, Haiman CA, Knudsen ES. Cyclin D1: polymorphism, aberrant splicing and cancer risk. Oncogene. 2006;25:1620–8.
Buckley MF, Sweeney KJ, Hamilton JA, Sini RL, Manning DL, Nicholson RI, et al. Expression and amplification of cyclin genes in human breast cancer. Oncogene. 1993;8:2127–33.
Dickson C, Fantl V, Gillett C, Brookes S, Bartek J, Smith R, et al. Amplification of chromosome band 11q13 and a role for cyclin D1 in human breast cancer. Cancer Lett. 1995;90:43–50.
Lammie GA, Fantl V, Smith R, Schuuring E, Brookes S, Michalides R, et al. D11S287, a putative oncogene on chromosome 11q13, is amplified and expressed in squamous cell and mammary carcinomas and linked to BCL-1. Oncogene. 1991;6:439–44.
Gillett C, Smith P, Gregory W, Richards M, Millis R, Peters G, et al. Cyclin D1 and prognosis in human breast cancer. Int J Cancer. 1996;69:92–9.
McIntosh GG, Anderson JJ, Milton I, Steward M, Parr AH, Thomas MD, et al. Determination of the prognostic value of cyclin D1 overexpression in breast cancer. Oncogene. 1995;11:885–91.
Gillett C, Fantl V, Smith R, Fisher C, Bartek J, Dickson C, et al. Amplification and overexpression of cyclin D1 in breast cancer detected by immunohistochemical staining. Cancer Res. 1994;54:1812–7.
Bartkova J, Lukas J, Strauss M, Bartek J. Cyclin D1 oncoprotein aberrantly accumulates in malignancies of diverse histogenesis. Oncogene. 1995;10:775–8.
Tobin NP, Bergh J. Analysis of cyclin D1 in breast cancer: a call to arms. Curr Breast Cancer Rep. 2012;4:171–3.
Holm K, Staaf J, Jönsson G, Vallon-Christersson J, Gunnarsson H, Arason A, et al. Characterisation of amplification patterns and target genes at chromosome 11q13 in CCND1-amplified sporadic and familial breast tumours. Breast Cancer Res Treat. 2012;133:583–94.
Hui R, Ball JR, Macmillan RD, Kenny FS, Prall OW, Campbell DH, et al. EMS1 gene expression in primary breast cancer: relationship to cyclin D1 and oestrogen receptor expression and patient survival. Oncogene. 1998;17:1053–9.
Hui R, Campbell DH, Lee CS, McCaul K, Horsfall DJ, Musgrove EA, et al. EMS1 amplification can occur independently of CCND1 or INT-2 amplification at 11q13 and may identify different phenotypes in primary breast cancer. Oncogene. 1997;15:1617–23.
Aaltonen K, Amini RM, Landberg G, Eerola H, Aittomäki K, Heikkilä P, et al. Cyclin D1 expression is associated with poor prognostic features in estrogen receptor positive breast cancer. Breast Cancer Res Treat. 2009;113:75–82.
Roy PG, Pratt N, Purdie CA, Baker L, Ashfield A, Quinlan P, et al. High CCND1 amplification identifies a group of poor prognosis women with estrogen receptor positive breast cancer. Int J Cancer. 2010;127:355–60.
Bilal E, Vassallo K, Toppmeyer D, Barnard N, Rye IH, Almendro V, et al. Amplified loci on chromosomes 8 and 17 predict early relapse in ER-positive breast cancers. PLoS ONE. 2012;7:e38575.
Hwang TS, Han HS, Hong YC, Lee HJ, Paik NS. Prognostic value of combined analysis of cyclin D1 and estrogen receptor status in breast cancer patients. Pathol Int. 2003;53:74–80.
Naidu R, Wahab NA, Yadav MM, Kutty MK. Expression and amplification of cyclin D1 in primary breast carcinomas: relationship with histopathological types and clinico-pathological parameters. Oncol Rep. 2002;9:409–16.
Reis-Filho JS, Savage K, Lambros MB, James M, Steele D, Jones RL, et al. Cyclin D1 protein overexpression and CCND1 amplification in breast carcinomas: an immunohistochemical and chromogenic in situ hybridisation analysis. Mod Pathol. 2006;19:999–1009.
Quintayo MA, Munro AF, Thomas J, Kunkler IH, Jack W, Kerr GR, et al. GSK3β and cyclin D1 expression predicts outcome in early breast cancer patients. Breast Cancer Res Treat. 2012;136:161–8.
Guo LL, Gao P, Wu YG, Jian WC, Hao CY, Li H, et al. Alteration of cyclin D1 in Chinese patients with breast carcinoma and its correlation with Ki-67, pRb, and p53. Arch Med Res. 2007;38:846–52.
Lundgren K, Brown M, Pineda S, Cuzick J, Salter J, Zabaglo L, et al. Effects of cyclin D1 gene amplification and protein expression on time to recurrence in postmenopausal breast cancer patients treated with anastrozole or tamoxifen: a TransATAC study. Breast Cancer Res. 2012;14:R57.
Akervall JA, Michalides RJ, Mineta H, Balm A, Borg A, Dictor MR, et al. Amplification of cyclin D1 in squamous cell carcinoma of the head and neck and the prognostic value of chromosomal abnormalities and cyclin D1 overexpression. Cancer. 1997;79:380–9.
Michalides R, van Veelen N, Hart A, Loftus B, Wientjens E, Balm A. Overexpression of cyclin D1 correlates with recurrence in a group of forty-seven operable squamous cell carcinomas of the head and neck. Cancer Res. 1995;55:975–8.
Namazie A, Alavi S, Olopade OI, Pauletti G, Aghamohammadi N, Aghamohammadi M, et al. Cyclin D1 amplification and p16(MTS1/CDK4I) deletion correlate with poor prognosis in head and neck tumors. Laryngoscope. 2002;112:472–81.
Kyomoto R, Kumazawa H, Toda Y, Sakaida N, Okamura A, Iwanaga M, et al. Cyclin-D1-gene amplification is a more potent prognostic factor than its protein over-expression in human head-and-neck squamous-cell carcinoma. Int J Cancer. 1997;74:576–81.
Miyamoto R, Uzawa N, Nagaoka S, Hirata Y, Amagasa T. Prognostic significance of cyclin D1 amplification and overexpression in oral squamous cell carcinomas. Oral Oncol. 2003;39:610–8.
Cheung TH, Yu MM, Lo KW, Yim SF, Chung TK, Wong YF. Alteration of cyclin D1 and CDK4 gene in carcinoma of uterine cervix. Cancer Lett. 2001;166:199–206.
Tate JG, Bamford S, Jubb HC, Sondka Z, Beare DM, Bindal N, et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 2019;47:D941–D7.
Smigielski EM, Sirotkin K, Ward M, Sherry ST. dbSNP: a database of single nucleotide polymorphisms. Nucleic Acids Res. 2000;28:352–5.
Stefl S, Nishi H, Petukh M, Panchenko AR, Alexov E. Molecular mechanisms of disease-causing missense mutations. J Mol Biol. 2013;425:3919–36.
Mihaylova V, Chablais F, Herenger Y, Spiegel R, Heinrich Jung H. Novel truncating mutations of MYO18B causing congenital myopathy in a Swiss patient. Neurol Genet. 2020;6:e458.
Farazi Fard MA, Rebelo AP, Buglo E, Nemati H, Dastsooz H, Gehweiler I, et al. Truncating mutations in UBAP1 cause hereditary spastic paraplegia. Am J Hum Genet. 2019;104:767–73.
Hunt RC, Simhadri VL, Iandoli M, Sauna ZE, Kimchi-Sarfaty C. Exposing synonymous mutations. Trends Genet: TIG. 2014;30:308–21.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Akhter N, Alzahrani FA, Dar SA, Wahid M, Sattar RSA, Hussain S, et al. AA genotype of cyclin D1 G870A polymorphism increases breast cancer risk: findings of a case-control study and meta-analysis. J Cell Biochem. 2019;120:16452–66.
Wang DD, Duan WD, Zhu ZM, Tu YL, Dou CQ, Han MM, et al. CCND1 rs9344 polymorphism is associated with the risk of hepatocellular carcinoma in Caucasian population. J Cancer Res Ther. 2018;14:S516–S8.
Xie M, Zhao F, Zou X, Jin S, Xiong S. The association between CCND1 G870A polymorphism and colorectal cancer risk: a meta-analysis. Medicine. 2017;96:e8269.
Wang Y, Li Z, Liu N, Zhang G. Association between CCND1 and XPC polymorphisms and bladder cancer risk: a meta-analysis based on 15 case-control studies. Tumour Biol. 2014;35:3155–65.
Kang S, Kim JW, Park NH, Song YS, Kang SB, Lee HP. Cyclin D1 polymorphism and the risk of endometrial cancer. Gynecol Oncol. 2005;97:431–5.
Zhang J, Li Y, Wang R, Wen D, Sarbia M, Kuang G, et al. Association of cyclin D1 (G870A) polymorphism with susceptibility to esophageal and gastric cardiac carcinoma in a northern Chinese population. Int J Cancer. 2003;105:281–4.
Yang M, Zhu H, Hu T, Liu S, Wang H. Association of CCND1 gene polymorphism with cervical cancer susceptibility in Caucasian population: a meta-analysis. Int J Clin Exp Med. 2015;8:12983–8.
Tang W, Yu P, Wang Y, Kang M, Sun B, Yin J, et al. Lack of association between cyclin D1 A870G (rs9344) polymorphism and esophageal squamous cell carcinoma risk: case-control study and meta-analysis. Int J Clin Exp Med. 2015;8:12685–95.
Zheng M, Wan L, He X, Qi X, Liu F, Zhang DH. Effect of the CCND1 A870G polymorphism on prostate cancer risk: a meta-analysis of 3,820 cases and 3,825 controls. World J Surg Oncol. 2015;13:55.
Naqvi AZ, Mahjabeen I, Ameen S, Ahmed MW, Khan AU, Akram Z, et al. Genetic and expression variations of cell cycle pathway genes in brain tumor patients. Biosci Rep. 2020;40:BSR20190629.
Petkevicius V, Salteniene V, Juzenas S, Wex T, Link A, Leja M, et al. Polymorphisms of microRNA target genes IL12B, INSR, CCND1 and IL10 in gastric cancer. World J Gastroenterol. 2017;23:3480–7.
Irimia M, Blencowe BJ. Alternative splicing: decoding an expansive regulatory layer. Curr Opin Cell Biol. 2012;24:323–32.
Montes M, Sanford BL, Comiskey DF, Chandler DS. RNA splicing and disease: animal models to therapies. Trends Genet. 2019;35:68–87.
Duan Y, Jia Y, Wang J, Liu T, Cheng Z, Sang M, et al. Long noncoding RNA DGCR5 involves in tumorigenesis of esophageal squamous cell carcinoma via SRSF1-mediated alternative splicing of Mcl-1. Cell Death Dis. 2021;12:587.
Bonnal SC, López-Oreja I, Valcárcel J. Roles and mechanisms of alternative splicing in cancer - implications for care. Nat Rev Clin Oncol. 2020;17:457–74.
Lee SC, Abdel-Wahab O. Therapeutic targeting of splicing in cancer. Nat Med. 2016;22:976–86.
He C, Li A, Lai Q, Ding J, Yan Q, Liu S, et al. The DDX39B/FUT3/TGFβR-I axis promotes tumor metastasis and EMT in colorectal cancer. Cell Death Dis. 2021;12:74.
Busslinger M, Moschonas N, Flavell RA. Beta + thalassemia: aberrant splicing results from a single point mutation in an intron. Cell. 1981;27:289–98.
Sarkar A, Panati K, Narala VR. Code inside the codon: The role of synonymous mutations in regulating splicing machinery and its impact on disease. Mutat Res Rev Mutat Res. 2022;790:108444.
Jin Z, Feng H, Liang J, Jing X, Zhao Q, Zhan L, et al. FGFR3(△7-9) promotes tumor progression via the phosphorylation and destabilization of ten-eleven translocation-2 in human hepatocellular carcinoma. Cell Death Dis. 2020;11:903.
Comstock CE, Augello MA, Benito RP, Karch J, Tran TH, Utama FE, et al. Cyclin D1 splice variants: polymorphism, risk, and isoform-specific regulation in prostate cancer. Clin Cancer Res. 2009;15:5338–49.
Holley SL, Parkes G, Matthias C, Bockmühl U, Jahnke V, Leder K, et al. Cyclin D1 polymorphism and expression in patients with squamous cell carcinoma of the head and neck. Am J Pathol. 2001;159:1917–24.
Lu F, Gladden AB, An JAD. alternatively spliced cyclin D1 isoform, cyclin D1b, is a nuclear oncogene. Cancer Res. 2003;63:7056–61.
Langenfeld J, Kiyokawa H, Sekula D, Boyle J, Dmitrovsky E. Posttranslational regulation of cyclin D1 by retinoic acid: a chemoprevention mechanism. Proc Natl Acad Sci USA. 1997;94:12070–4.
Zwijsen RM, Buckle RS, Hijmans EM, Loomans CJ, Bernards R. Ligand-independent recruitment of steroid receptor coactivators to estrogen receptor by cyclin D1. Genes Dev. 1998;12:3488–98.
Alt JR, Cleveland JL, Hannink M, Diehl JA. Phosphorylation-dependent regulation of cyclin D1 nuclear export and cyclin D1-dependent cellular transformation. Genes Dev. 2000;14:3102–14.
Diehl JA, Cheng M, Roussel MF, Sherr CJ. Glycogen synthase kinase-3beta regulates cyclin D1 proteolysis and subcellular localization. Genes Dev. 1998;12:3499–511.
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, et al. The sequence of the human genome. Science. 2001;291:1304–51.
Wiestner A, Tehrani M, Chiorazzi M, Wright G, Gibellini F, Nakayama K, et al. Point mutations and genomic deletions in CCND1 create stable truncated cyclin D1 mRNAs that are associated with increased proliferation rate and shorter survival. Blood. 2007;109:4599–606.
Yocca AE, Edger PP. Current status and future perspectives on the evolution of cis-regulatory elements in plants. Curr Opin Plant Biol. 2022;65:102139.
Wang Z, Burge CB. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA. 2008;14:802–13.
Lee RJ, Albanese C, Stenger RJ, Watanabe G, Inghirami G, Haines GK 3rd, et al. pp60(v-src) induction of cyclin D1 requires collaborative interactions between the extracellular signal-regulated kinase, p38, and Jun kinase pathways. A role for cAMP response element-binding protein and activating transcription factor-2 in pp60(v-src) signaling in breast cancer cells. J Biol Chem. 1999;274:7341–50.
Matsumura I, Kitamura T, Wakao H, Tanaka H, Hashimoto K, Albanese C, et al. Transcriptional regulation of the cyclin D1 promoter by STAT5: its involvement in cytokine-dependent growth of hematopoietic cells. EMBO J. 1999;18:1367–77.
Xu Y, Shi Y, Yuan Q, Liu X, Yan B, Chen L, et al. Epstein-Barr Virus encoded LMP1 regulates cyclin D1 promoter activity by nuclear EGFR and STAT3 in CNE1 cells. J Exp Clin Cancer Res. 2013;32:90.
Kang JH, Kim MJ, Ko SH, Jeong IK, Koh KH, Rhie DJ, et al. Upregulation of rat Ccnd1 gene by exendin-4 in pancreatic beta cell line INS-1: interaction of early growth response-1 with cis-regulatory element. Diabetologia. 2006;49:969–79.
Albanese C, Johnson J, Watanabe G, Eklund N, Vu D, Arnold A, et al. Transforming p21ras mutants and c-Ets-2 activate the cyclin D1 promoter through distinguishable regions. J Biol Chem. 1995;270:23589–97.
Yan YX, Nakagawa H, Lee MH, Rustgi AK. Transforming growth factor-alpha enhances cyclin D1 transcription through the binding of early growth response protein to a cis-regulatory element in the cyclin D1 promoter. J Biol Chem. 1997;272:33181–90.
Cáceres JF, Kornblihtt AR. Alternative splicing: multiple control mechanisms and involvement in human disease. Trends Genet. 2002;18:186–93.
Paronetto MP, Cappellari M, Busà R, Pedrotti S, Vitali R, Comstock C, et al. Alternative splicing of the cyclin D1 proto-oncogene is regulated by the RNA-binding protein Sam68. Cancer Res. 2010;70:229–39.
Feng J, Xu X, Fan X, Yi Q, Tang L. BAF57/SMARCE1 interacting with splicing factor SRSF1 regulates mechanical stress-induced alternative splicing of cyclin D1. Genes. 2021;12:306.
Panda AC, Abdelmohsen K, Martindale JL, Di Germanio C, Yang X, Grammatikakis I, et al. Novel RNA-binding activity of MYF5 enhances Ccnd1/Cyclin D1 mRNA translation during myogenesis. Nucleic Acids Res. 2016;44:2393–408.
Bracken CP, Wall SJ, Barré B, Panov KI, Ajuh PM, Perkins ND. Regulation of cyclin D1 RNA stability by SNIP1. Cancer Res. 2008;68:7621–8.
Batsché E, Yaniv M, Muchardt C. The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nat Struct Mol Biol. 2006;13:22–9.
Kitamura K, Nimura K. Regulation of RNA splicing: aberrant splicing regulation and therapeutic targets in cancer. Cells. 2021;10:923.
Sp1 B. Androgen receptor as a target in androgen-independent prostate cancer. Urology. 2002;60:132–8.
Trapman J, Brinkmann AO. The androgen receptor in prostate cancer. Pathol Res Pract. 1996;192:752–60.
Wang N, Wei H, Yin D, Lu Y, Zhang Y, Jiang D, et al. Cyclin D1b overexpression inhibits cell proliferation and induces cell apoptosis in cervical cancer cells in vitro and in vivo. Int J Clin Exp Pathol. 2014;7:4016–23.
Sherr CJ. D-type cyclins. Trends Biochem Sci. 1995;20:187–90.
Kim CJ, Terado T, Tambe Y, Mukaisho KI, Sugihara H, Kawauchi A, et al. Anti-oncogenic activities of cyclin D1b siRNA on human bladder cancer cells via induction of apoptosis and suppression of cancer cell stemness and invasiveness. Int J Oncol. 2018;52:231–40.
Marzec M, Kasprzycka M, Lai R, Gladden AB, Wlodarski P, Tomczak E, et al. Mantle cell lymphoma cells express predominantly cyclin D1a isoform and are highly sensitive to selective inhibition of CDK4 kinase activity. Blood. 2006;108:1744–50.
Solomon DA, Wang Y, Fox SR, Lambeck TC, Giesting S, Lan Z, et al. Cyclin D1 splice variants. Differential effects on localization, RB phosphorylation, and cellular transformation. The. J Biol Chem. 2003;278:30339–47.
Musgrove EA, Caldon CE, Barraclough J, Stone A, Sutherland RL. Cyclin D as a therapeutic target in cancer. Nat Rev Cancer. 2011;11:558–72.
Coqueret O. Linking cyclins to transcriptional control. Gene. 2002;299:35–55.
Fu M, Wang C, Li Z, Sakamaki T, Pestell RG. Minireview: Cyclin D1: normal and abnormal functions. Endocrinology. 2004;145:5439–47.
Beato M. Gene regulation by steroid hormones. Cell. 1989;56:335–44.
Evans RM. The steroid and thyroid hormone receptor superfamily. Science. 1988;240:889–95.
Kumar V, Chambon P. The estrogen receptor binds tightly to its responsive element as a ligand-induced homodimer. Cell. 1988;55:145–56.
Burd CJ, Petre CE, Morey LM, Wang Y, Revelo MP, Haiman CA, et al. Cyclin D1b variant influences prostate cancer growth through aberrant androgen receptor regulation. Proc Natl Acad Sci USA. 2006;103:2190–5.
Reutens AT, Fu M, Wang C, Albanese C, McPhaul MJ, Sun Z, et al. Cyclin D1 binds the androgen receptor and regulates hormone-dependent signaling in a p300/CBP-associated factor (P/CAF)-dependent manner. Mol Endocrinol. 2001;15:797–811.
Petre CE, Wetherill YB, Danielsen M, Knudsen KE. Cyclin D1: mechanism and consequence of androgen receptor co-repressor activity. J Biol Chem. 2002;277:2207–15.
Knudsen KE, Cavenee WK, Arden KC. D-type cyclins complex with the androgen receptor and inhibit its transcriptional transactivation ability. Cancer Res. 1999;59:2297–301.
Kim CJ, Tambe Y, Mukaisho KI, Sugihara H, Kawauchi A, Inoue H. Akt-dependent activation of Erk by cyclin D1b contributes to cell invasiveness and tumorigenicity. Oncol Lett. 2016;12:4850–6.
Kim CJ, Nishi K, Isono T, Okuyama Y, Tambe Y, Okada Y, et al. Cyclin D1b variant promotes cell invasiveness independent of binding to CDK4 in human bladder cancer cells. Mol Carcinog. 2009;48:953–64.
Luo BP, Luo J, Hu YB, Yao XW, Wu FH. Cyclin D1b splice variant promotes αvβ3-mediated EMT induced by LPS in breast cancer cells. Curr Med Sci. 2018;38:467–72.
Lai TH, Fong YC, Fu WM, Yang RS, Tang CH. Stromal cell-derived factor-1 increase alphavbeta3 integrin expression and invasion in human chondrosarcoma cells. J Cell Physiol. 2009;218:334–42.
Rolli M, Fransvea E, Pilch J, Saven A, Felding-Habermann B. Activated integrin alphavbeta3 cooperates with metalloproteinase MMP-9 in regulating migration of metastatic breast cancer cells. Proc Natl Acad Sci USA. 2003;100:9482–7.
Liao SJ, Zhou YH, Yuan Y, Li D, Wu FH, Wang Q, et al. Triggering of Toll-like receptor 4 on metastatic breast cancer cells promotes αvβ3-mediated adhesion and invasive migration. Breast Cancer Res Treat. 2012;133:853–63.
Fusté NP, Ferrezuelo F, Garí E. Cyclin D1 promotes tumor cell invasion and metastasis by cytoplasmic mechanisms. Mol Cell Oncol. 2016;3:e1203471.
Fusté NP, Fernández-Hernández R, Cemeli T, Mirantes C, Pedraza N, Rafel M, et al. Cytoplasmic cyclin D1 regulates cell invasion and metastasis through the phosphorylation of paxillin. Nat Commun. 2016;7:11581.
Tang K, Boudreau CG, Brown CM, Khadra A. Paxillin phosphorylation at serine 273 and its effects on Rac, Rho and adhesion dynamics. PLoS Comput Biol. 2018;14:e1006303.
Hutchinson JN, Muller WJ. Transgenic mouse models of human breast cancer. Oncogene. 2000;19:6130–7.
Sutherland RL, Musgrove EA. Cyclin D1 and mammary carcinoma: new insights from transgenic mouse models. Breast Cancer Res. 2002;4:14–7.
Deshpande A, Sicinski P, Hinds PW. Cyclins and cdks in development and cancer: a perspective. Oncogene. 2005;24:2909–15.
Yamamoto H, Ochiya T, Takeshita F, Toriyama-Baba H, Hirai K, Sasaki H, et al. Enhanced skin carcinogenesis in cyclin D1-conditional transgenic mice: cyclin D1 alters keratinocyte response to calcium-induced terminal differentiation. Cancer Res. 2002;62:1641–7.
Rodriguez-Puebla ML, LaCava M, Conti CJ. Cyclin D1 overexpression in mouse epidermis increases cyclin-dependent kinase activity and cell proliferation in vivo but does not affect skin tumor development. Cell Growth Differ. 1999;10:467–72.
Robles AI, Larcher F, Whalin RB, Murillas R, Richie E, Gimenez-Conti IB, et al. Expression of cyclin D1 in epithelial tissues of transgenic mice results in epidermal hyperproliferation and severe thymic hyperplasia. Proc Natl Acad Sci USA. 1996;93:7634–8.
Gladden AB, Diehl JA. Location, location, location: the role of cyclin D1 nuclear localization in cancer. J Cell Biochem. 2005;96:906–13.
Yu Q, Geng Y, Sicinski P. Specific protection against breast cancers by cyclin D1 ablation. Nature. 2001;411:1017–21.
Lee RJ, Albanese C, Fu M, D’Amico M, Lin B, Watanabe G, et al. Cyclin D1 is required for transformation by activated Neu and is induced through an E2F-dependent signaling pathway. Mol Cell Biol. 2000;20:672–83.
Perez-Roger I, Kim SH, Griffiths B, Sewing A, Land H. Cyclins D1 and D2 mediate myc-induced proliferation via sequestration of p27(Kip1) and p21(Cip1). EMBO J. 1999;18:5310–20.
Robles AI, Rodriguez-Puebla ML, Glick AB, Trempus C, Hansen L, Sicinski P, et al. Reduced skin tumor development in cyclin D1-deficient mice highlights the oncogenic ras pathway in vivo. Genes Dev. 1998;12:2469–74.
Rojas P, Benavides F, Blando J, Perez C, Cardenas K, Richie E, et al. Enhanced skin carcinogenesis and lack of thymus hyperplasia in transgenic mice expressing human cyclin D1b (CCND1b). Mol Carcinog. 2009;48:508–16.
Lu F, Gladden AB, Diehl JA. An alternatively spliced cyclin D1 isoform, cyclin D1b, is a nuclear oncogene. Cancer Res. 2003;63:7056–61.
Augello MA, Berman-Booty LD, Carr R 3rd, Yoshida A, Dean JL, Schiewer MJ, et al. Consequence of the tumor-associated conversion to cyclin D1b. EMBO Mol Med. 2015;7:628–47.
Jeon S, Kim Y, Jeong YM, Bae JS, Jung CK. CCND1 splice variant as A novel diagnostic and predictive biomarker for thyroid cancer. Cancers 2018;10:437.
Li R, An SJ, Chen ZH, Zhang GC, Zhu JQ, Nie Q, et al. Expression of cyclin D1 splice variants is differentially associated with outcome in non-small cell lung cancer patients. Hum Pathol. 2008;39:1792–801.
Gu J, Zhang X, Yang Z, Wang N. Expression of cyclin D1 protein isoforms and its prognostic significance in cervical cancer. Cancer Manag Res. 2019;11:9073–83.
Gupta VK, Feber A, Xi L, Pennathur A, Wu M, Luketich JD, et al. Association between CCND1 G/A870 polymorphism, allele-specific amplification, cyclin D1 expression, and survival in esophageal and lung carcinoma. Clinical cancer research: an official journal of the American Association for. Cancer Res. 2008;14:7804–12.
Myklebust MP, Li Z, Tran TH, Rui H, Knudsen ES, Elsaleh H, et al. Expression of cyclin D1a and D1b as predictive factors for treatment response in colorectal cancer. Br J Cancer. 2012;107:1684–91.
Bao C, Chen J, Chen D, Lu Y, Lou W, Ding B, et al. MiR-93 suppresses tumorigenesis and enhances chemosensitivity of breast cancer via dual targeting E2F1 and CCND1. Cell Death Dis. 2020;11:618.
Wei M, Zhu L, Li Y, Chen W, Han B, Wang Z, et al. Knocking down cyclin D1b inhibits breast cancer cell growth and suppresses tumor development in a breast cancer model. Cancer Sci. 2011;102:1537–44.
Zhang D, Dai D, Zhou M, Li Z, Wang C, Lu Y, et al. Inhibition of cyclin D1 expression in human glioblastoma cells is associated with increased temozolomide chemosensitivity. Cell Physiol Biochem. 2018;51:2496–508.
Ding H, Wang Y, Zhang H. CCND1 silencing suppresses liver cancer stem cell differentiation and overcomes 5-Fluorouracil resistance in hepatocellular carcinoma. J Pharmacol Sci. 2020;143:219–25.
Seo JH, Jeong ES, Lee KS, Heo SH, Jeong DG, Choi YK. Lentivirus-mediated shRNA targeting of cyclin D1 enhances the chemosensitivity of human gastric cancer to 5-fluorouracil. Int J Oncol. 2013;43:2007–14.
Fu SM, Xu MX, Lin SM, Liang Z, Cai JH. Association of cyclin D1 and survivin expression with sensitivity to radiotherapy in patients with nasopharyngeal carcinoma. Genet Mol Res. 2014;13:3502–9.
Jin J, Guo Y, Dong X, Liu J, He Y. Methylation-associated silencing of miR-193b improves the radiotherapy sensitivity of esophageal cancer cells by targeting cyclin D1 in areas with zinc deficiency. Radiother Oncol. 2020;150:104–13.
Shintani S, Mihara M, Ueyama Y, Matsumura T, Wong DT. Cyclin D1 overexpression associates with radiosensitivity in oral squamous cell carcinoma. Int J Cancer. 2001;96:159–65.
Choi C, Park S, Cho WK, Choi DH. Cyclin D1 is associated with radiosensitivity of triple-negative breast cancer cells to proton beam irradiation. Int J Mol Sci. 2019;20:4943.
Liang ML, Hsieh TH, Liu YR, Chen YW, Lee YY, Chang FC, et al. Significance of cyclin D1 overexpression in progression and radio-resistance of pediatric ependymomas. Oncotarget. 2018;9:2527–42.
Yao G, Tang J, Yang X, Zhao Y, Zhou R, Meng R, et al. Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer. Theranostics. 2020;10:11144–58.
Salton M, Misteli T. Small molecule modulators of pre-mRNA splicing in cancer therapy. Trends Mol Med. 2016;22:28–37.
Xue K, MacLaren RE. Antisense oligonucleotide therapeutics in clinical trials for the treatment of inherited retinal diseases. Expert Opin Investig Drugs. 2020;29:1163–70.
Havens MA, Hastings ML. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 2016;44:6549–63.
Amantana A, Iversen PL. Pharmacokinetics and biodistribution of phosphorodiamidate morpholino antisense oligomers. Curr Opin Pharmacol. 2005;5:550–5.
Schmajuk G, Sierakowska H, Kole R. Antisense oligonucleotides with different backbones. Modification of splicing pathways and efficacy of uptake. J Biol Chem. 1999;274:21783–9.
Morcos PA. Gene switching: analyzing a broad range of mutations using steric block antisense oligonucleotides. Methods Enzymol. 2000;313:174–89.
Gebski BL, Mann CJ, Fletcher S, Wilton SD. Morpholino antisense oligonucleotide induced dystrophin exon 23 skipping in mdx mouse muscle. Hum Mol Genet. 2003;12:1801–11.
Draper BW, Morcos PA, Kimmel CB. Inhibition of zebrafish fgf8 pre-mRNA splicing with morpholino oligos: a quantifiable method for gene knockdown. Genesis. 2001;30:154–6.
Augello MA, Burd CJ, Birbe R, McNair C, Ertel A, Magee MS, et al. Convergence of oncogenic and hormone receptor pathways promotes metastatic phenotypes. J Clin Investig. 2013;123:493–508.
Zhang X, Xu J, Jiang T, Liu G, Wang D, Lu Y. MicroRNA-195 suppresses colorectal cancer cells proliferation via targeting FGF2 and regulating Wnt/β-catenin pathway. Am J Cancer Res. 2016;6:2631–40.
Tucker CA, Kapanen AI, Chikh G, Hoffman BG, Kyle AH, Wilson IM, et al. Silencing Bcl-2 in models of mantle cell lymphoma is associated with decreases in cyclin D1, nuclear factor-kappaB, p53, bax, and p27 levels. Mol Cancer Ther. 2008;7:749–58.
Barresi V, Musmeci C, Rinaldi A, Condorelli DF. Transcript-targeted therapy based on RNA interference and antisense oligonucleotides: current applications and novel molecular targets. Int J Mol Sci. 2022;23:8875.
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998;391:806–11.
Rivas FV, Tolia NH, Song JJ, Aragon JP, Liu J, Hannon GJ, et al. Purified Argonaute2 and an siRNA form recombinant human RISC. Nat Struct Mol Biol. 2005;12:340–9.
Zamore PD, Tuschl T, Sharp PA, Bartel DP. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 2000;101:25–33.
Wittrup A, Ai A, Liu X, Hamar P, Trifonova R, Charisse K, et al. Visualizing lipid-formulated siRNA release from endosomes and target gene knockdown. Nat Biotechnol. 2015;33:870–6.
Friedrich M, Aigner A. Therapeutic siRNA: state-of-the-art and future perspectives. BioDrugs 2022;36:549–71.
Stanley RF, Abdel-Wahab O. Dysregulation and therapeutic targeting of RNA splicing in cancer. Nat Cancer. 2022;3:536–46.
Wang E, Aifantis I. RNA splicing and cancer. Trends Cancer. 2020;6:631–44.
Honda A, Valogne Y, Bou Nader M, Bréchot C, Faivre J. An intron-retaining splice variant of human cyclin A2, expressed in adult differentiated tissues, induces a G1/S cell cycle arrest in vitro. PLoS ONE. 2012;7:e39249.
Lozano JC, Schatt P, Marquès F, Peaucellier G, Fort P, Féral JP, et al. A presumptive developmental role for a sea urchin cyclin B splice variant. J Cell Biol. 1998;140:283–93.
Sewing A, Rönicke V, Bürger C, Funk M, Müller R. Alternative splicing of human cyclin E. J Cell Sci. 1994;107:581–8.
Zhao Y, He HR, Wang MY, Ren XD, Zhang L, Dong YL, et al. Cyclin D1 G870A gene polymorphism and risk of leukemia and hepatocellular carcinoma: a meta-analysis. Genet Mol Res. 2015;14:5171–80.
Lan J, Li M, Wang H. CCDN1 rs603965 polymorphism may serve as a genetic biomarker of brain tumor: A meta-analysis of 5,769 subjects. Mol Genet Genom Med. 2019;7:e00655.
Liao D, Wu Y, Pu X, Chen H, Luo S, Li B, et al. Cyclin D1 G870A polymorphism and risk of nasopharyngeal carcinoma: a case-control study and meta-analysis. PLoS ONE. 2014;9:e113299.
Meng Y, Zhang C, Zhou X. Association between the Cyclin D1 G870A polymorphism and the susceptibility to and prognosis of upper aerodigestive tract squamous cell carcinomas: an updated meta-analysis. Onco Targets Ther. 2016;9:367–76.
Zong H, Cao L, Ma C, Zhao J, Ming X, Shang M, et al. Association between the G870A polymorphism of Cyclin D1 gene and glioma risk. Tumour Biol. 2014;35:8095–101.
Wen L, Hu YY, Yang GL, Liu DX. CCND1 G870A polymorphism contributes to the risk of esophageal cancer: An updated systematic review and cumulative meta-analysis. Biomed Rep. 2014;2:549–54.
Wang L, Wang Z, Gao X, Li X, Sun X, Liu C. Association between Cyclin D1 polymorphism and oral cancer susceptibility: a meta-analysis. Tumour Biol. 2014;35:1149–55.
Lin H, Fang L, Lin D. Association of cyclin D1 variants with head and neck cancer susceptibility: evidence from a meta-analysis. Asian Pac J Cancer Prev. 2014;15:5645–51.
Qin LY, Zhao LG, Chen X, Yang Z, Mo WN. The CCND1 G870A gene polymorphism and leukemia or non-Hodgkin lymphoma risk: a meta-analysis. Asian Pac J Cancer Prev. 2014;15:6923–8.
Zhou C, An H, Hu M, Liu Q, Geng P, Xu J, et al. The cyclin D1 (CCND1) G870A polymorphism and lung cancer susceptibility: a meta-analysis. Tumour Biol. 2013;34:3831–7.
Xu Y, Chen SY, Ross KN, Balk SP. Androgens induce prostate cancer cell proliferation through mammalian target of rapamycin activation and post-transcriptional increases in cyclin D proteins. Cancer Res. 2006;66:7783–92.
Kim CJ, Tambe Y, Mukaisho K, Sugihara H, Isono T, Sonoda H, et al. Female-specific rectal carcinogenesis in cyclin D1b transgenic mice. Carcinogenesis. 2014;35:227–36.
Funding
This work was supported by grants from the National Key R&D project of the Chinese Ministry of Science and Technology [2018YFE0205100]; the Gansu Province Science and Technology Innovation Service Platform Construction Project [18JR2TA024]; the Key Program of the National Natural Science Foundation of China [U1632270]; the National Natural Science Foundation of China [11675234].
Author information
Authors and Affiliations
Contributions
JW collected the related papers and drafted the manuscript. XX and CD participated in the design of the review. WS, TZ, SZ, HL, FM, MS, and WS initiated the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Nickolai Barlev
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Su, W., Zhang, T. et al. Aberrant Cyclin D1 splicing in cancer: from molecular mechanism to therapeutic modulation. Cell Death Dis 14, 244 (2023). https://doi.org/10.1038/s41419-023-05763-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-023-05763-7