Abstract
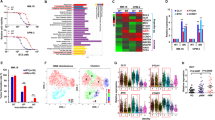
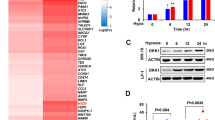
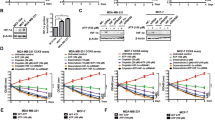
The bone marrow microenvironment in multiple myeloma (MM) is hypoxic and provides multi-advantages for the initiation of chemoresistance, but the underlying mechanisms and key regulators are still indistinct. In the current study, we found that hypoxia stimulus easily induced chemoresistance to proteasome inhibitors (PIs), and the steroid receptor coactivator 3 (SRC-3) expression was remarkably augmented at posttranslational level. Protein interactome analysis identified SENP1 as a key modifier of SRC-3 stability, as SENP1-mediated deSUMOylation attenuated the K11-linked polyubiquitination of SRC-3. SENP1 depletion in the SENP1fl/flCD19Cre/+ B cells showed impaired SRC3 stability, and knockdown of SENP1 in MM cells by CRISPR/cas9 sgRNA accelerated the degradation of SRC-3 and remarkably overcame the resistance to PIs. In the Vk*Myc and 5TGM1 mouse models as well as patient-derived xenograft (PDX) of myeloma, SENP1 inhibitor Momordin Ιc (Mc) increased the sensitivity to PIs in MM cells. Importantly, SENP1 level was positively correlated with SRC-3 level in the tissues from refractory/relapsed MM, as well as in xenograft tissues from mice treated with bortezomib and Mc. Taken together, our findings suggest that hypoxia-induced SENP1 is a crucial regulator of chemoresistance to PIs, and shed light on developing therapeutic strategies to overcome chemoresistance by using small molecules targeting SENP1 or SRC-3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials. Source data are provided with this paper. Requests for any materials in this study should be directed to Zhiqiang Liu and obtained through an MTA.
References
van de Donk N, Pawlyn C, Yong KL. Multiple myeloma. Lancet 2021;397:410–27.
Moser-Katz T, Joseph NS, Dhodapkar MV, Lee KP, Boise LH. Game of bones: how myeloma manipulates its microenvironment. Front Oncol. 2020;10:625199.
Kumar SK, Rajkumar V, Kyle RA, van Duin M, Sonneveld P, Mateos MV, et al. Multiple myeloma. Nat Rev Dis Prim. 2017;3:17046.
From the American Association of Neurological Surgeons ASoNC, Interventional Radiology Society of Europe CIRACoNSESoMINTESoNESOSfCA, Interventions SoIRSoNS, World Stroke O, Sacks D, Baxter B, et al. Multisociety consensus quality improvement revised consensus statement for endovascular therapy of acute ischemic stroke. Int J Stroke. 2018;13:612–32.
Ikeda S, Tagawa H. Impact of hypoxia on the pathogenesis and therapy resistance in multiple myeloma. Cancer Sci. 2021;112:3995–4004.
Ria R, Vacca A. Bone marrow stromal cells-induced drug resistance in multiple myeloma. Int J Mol Sci. 2020;21:613.
Kawano Y, Moschetta M, Manier S, Glavey S, Gorgun GT, Roccaro AM, et al. Targeting the bone marrow microenvironment in multiple myeloma. Immunol Rev. 2015;263:160–72.
Yen CH, Hsiao HH. NRF2 is one of the players involved in bone marrow-mediated drug resistance in multiple myeloma. Int J Mol Sci. 2018;19:3503.
Pennathur-Das R, Levitt L. Augmentation of in vitro human marrow erythropoiesis under physiological oxygen tensions is mediated by monocytes and T lymphocytes. Blood 1987;69:899–907.
Danet GH, Pan Y, Luongo JL, Bonnet DA, Simon MC. Expansion of human SCID-repopulating cells under hypoxic conditions. J Clin Invest. 2003;112:126–35.
Hu J, Handisides DR, Van Valckenborgh E, De Raeve H, Menu E, Vande Broek I, et al. Targeting the multiple myeloma hypoxic niche with TH-302, a hypoxia-activated prodrug. Blood 2010;116:1524–7.
Martin SK, Diamond P, Gronthos S, Peet DJ, Zannettino AC. The emerging role of hypoxia, HIF-1 and HIF-2 in multiple myeloma. Leukemia 2011;25:1533–42.
Xu Y, Guo J, Liu J, Xie Y, Li X, Jiang H, et al. Hypoxia-induced CREB cooperates MMSET to modify chromatin and promote DKK1 expression in multiple myeloma. Oncogene 2021;40:1231–41.
Yuen VW, Wong CC. Hypoxia-inducible factors and innate immunity in liver cancer. J Clin Invest. 2020;130:5052–62.
de Heer EC, Jalving M, Harris AL. HIFs, angiogenesis, and metabolism: elusive enemies in breast cancer. J Clin Invest. 2020;130:5074–87.
Mani DR, Krug K, Zhang B, Satpathy S, Clauser KR, Ding L, et al. Cancer proteogenomics: current impact and future prospects. Nat Rev Cancer. 2022;22:298–313.
Lothrop AP, Torres MP, Fuchs SM. Deciphering post-translational modification codes. FEBS Lett. 2013;587:1247–57.
Wirth M, Schick M, Keller U, Kronke J. Ubiquitination and ubiquitin-like modifications in multiple myeloma: biology and therapy. Cancers. 2020;12:3764.
Liu J, Xie Y, Guo J, Li X, Wang J, Jiang H, et al. Targeting NSD2-mediated SRC-3 liquid-liquid phase separation sensitizes bortezomib treatment in multiple myeloma. Nat Commun. 2021;12:1022.
Chen SL, Wang SC, Hosking B, Muscat GE. Subcellular localization of the steroid receptor coactivators (SRCs) and MEF2 in muscle and rhabdomyosarcoma cells. Mol Endocrinol. 2001;15:783–96.
Wu H, Sun L, Zhang Y, Chen Y, Shi B, Li R, et al. Coordinated regulation of AIB1 transcriptional activity by sumoylation and phosphorylation. J Biol Chem. 2006;281:21848–56.
Chesi M, Matthews GM, Garbitt VM, Palmer SE, Shortt J, Lefebure M, et al. Drug response in a genetically engineered mouse model of multiple myeloma is predictive of clinical efficacy. Blood 2012;120:376–85.
Wang J, Zhu X, Dang L, Jiang H, Xie Y, Li X, et al. Epigenomic reprogramming via HRP2-MINA dictates response to proteasome inhibitors in multiple myeloma with t(4;14) translocation. J Clin Invest. 2022;132:e149526.
Song X, Chen J, Zhao M, Zhang C, Yu Y, Lonard DM, et al. Development of potent small-molecule inhibitors to drug the undruggable steroid receptor coactivator-3. Proc Natl Acad Sci USA. 2016;113:4970–5.
Kumar R, Sabapathy K. RNF4-A Paradigm for SUMOylation-mediated ubiquitination. Proteomics 2019;19:e1900185.
Li L, Deng CX, Chen Q. SRC-3, a steroid receptor coactivator: implication in cancer. Int J Mol Sci. 2021;22:4760.
Geng C, Rajapakshe K, Shah SS, Shou J, Eedunuri VK, Foley C, et al. Androgen receptor is the key transcriptional mediator of the tumor suppressor SPOP in prostate cancer. Cancer Res. 2014;74:5631–43.
Lanz RB, Bulynko Y, Malovannaya A, Labhart P, Wang L, Li W, et al. Global characterization of transcriptional impact of the SRC-3 coregulator. Mol Endocrinol. 2010;24:859–72.
Xu J, Wu RC, O’Malley BW. Normal and cancer-related functions of the p160 steroid receptor co-activator (SRC) family. Nat Rev Cancer. 2009;9:615–30.
Raninga PV, Di Trapani G, Vuckovic S, Tonissen KF. TrxR1 inhibition overcomes both hypoxia-induced and acquired bortezomib resistance in multiple myeloma through NF-small ka, Cyrillicbeta inhibition. Cell Cycle. 2016;15:559–72.
Murray MY, Zaitseva L, Auger MJ, Craig JI, MacEwan DJ, Rushworth SA, et al. Ibrutinib inhibits BTK-driven NF-kappaB p65 activity to overcome bortezomib-resistance in multiple myeloma. Cell Cycle. 2015;14:2367–75.
Mougenot P, Bressolle F, Culine S, Solassol I, Poujol S, Pinguet F. In vitro cytotoxic effect of melphalan and pilot phase II study in hormone-refractory prostate cancer. Anticancer Res. 2006;26:2197–203.
Kunz K, Piller T, Muller S. SUMO-specific proteases and isopeptidases of the SENP family at a glance. J Cell Sci. 2018;131:jcs211904.
Ma C, Wu B, Huang X, Yuan Z, Nong K, Dong B, et al. SUMO-specific protease 1 regulates pancreatic cancer cell proliferation and invasion by targeting MMP-9. Tumour Biol. 2014;35:12729–35.
Xiang-Ming Y, Zhi-Qiang X, Ting Z, Jian W, Jian P, Li-Qun Y, et al. SENP1 regulates cell migration and invasion in neuroblastoma. Biotechnol Appl Biochem. 2016;63:435–40.
Chang HM, Yeh ETH. SUMO: From bench to bedside. Physiol Rev. 2020;100:1599–619.
Scanlon SE, Glazer PM. Multifaceted control of DNA repair pathways by the hypoxic tumor microenvironment. DNA Repair. 2015;32:180–9.
Rohwer N, Cramer T. Hypoxia-mediated drug resistance: novel insights on the functional interaction of HIFs and cell death pathways. Drug Resist Update. 2011;14:191–201.
Acknowledgements
This work was supported by Cancer Biobank of Tianjin Medical University Cancer Institute and Hospital. We thank Dr. Leif Bergsagel at the Mayo Clinic for kindly providing the Vk*Myc mouse spleen cells. This work was supported by the Beijing Natural Science Foundation of China (Z200020, ZQL), the National Natural Science Foundation of China (81870161, 82070221, ZQL; 81900215, JYW; 81870150, ZGZ), the Tianjin Research Innovation Project for Postgraduate Students (2020YJSB162, YX), and Tianjin Key Medical Discipline (Specialty) Construction Project (TJYXZDXK-009A).
Author information
Authors and Affiliations
Contributions
LG and ZQL contributed to writing the manuscript; JG, YYL, SW, YX, HMJ, XL, ZY.P., YXW, and JPM contributed to performing the experiments and statistical analyses; JG, SW, MLH, and MQW were in charge of the animal studies; ZGZ and XKC provided the patient samples and clinical statistics; ZQL and ZGZ contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All authors concur with the submission and publication of this research article.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, J., Lv, Y., Wang, S. et al. Hypoxia induces chemoresistance to proteasome inhibitors through orchestrating deSUMOylation and ubiquitination of SRC-3 in multiple myeloma. Oncogene 41, 4971–4979 (2022). https://doi.org/10.1038/s41388-022-02494-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02494-5