Abstract
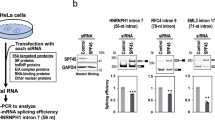
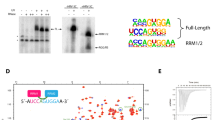
The U2AF-homology motif (UHM) mediates protein-protein interactions between factors involved in constitutive RNA splicing. Here we report that the splicing factor SPF45 regulates alternative splicing of the apoptosis regulatory gene FAS (also called CD95). The SPF45 UHM is necessary for this activity and binds UHM-ligand motifs (ULMs) present in the 3′ splice site–recognizing factors U2AF65, SF1 and SF3b155. We describe a 2.1-Å crystal structure of SPF45-UHM in complex with a ULM peptide from SF3b155. Features distinct from those of previously described UHM-ULM structures allowed the design of mutations in the SPF45 UHM that selectively impair binding to individual ULMs. Splicing assays using the ULM-selective SPF45 variants demonstrate that individual UHM-ULM interactions are required for FAS splicing regulation by SPF45 in vivo. Our data suggest that networks of UHM-ULM interactions are involved in regulating alternative splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Black, D.L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Bell, L.R., Horabin, J.I., Schedl, P. & Cline, T.W. Positive autoregulation of sex-lethal by alternative splicing maintains the female determined state in Drosophila. Cell 65, 229–239 (1991).
Lallena, M.J., Chalmers, K.J., Llamazares, S., Lamond, A.I. & Valcarcel, J. Splicing regulation at the second catalytic step by Sex-lethal involves 3′ splice site recognition by SPF45. Cell 109, 285–296 (2002).
Aravind, L. & Koonin, E.V. G-patch: a new conserved domain in eukaryotic RNA-processing proteins and type D retroviral polyproteins. Trends Biochem. Sci. 24, 342–344 (1999).
Silverman, E.J. et al. Interaction between a G-patch protein and a spliceosomal DEXD/H-box ATPase that is critical for splicing. Mol. Cell. Biol. 24, 10101–10110 (2004).
Svec, M., Bauerova, H., Pichova, I., Konvalinka, J. & Strisovsky, K. Proteinases of betaretroviruses bind single-stranded nucleic acids through a novel interaction module, the G-patch. FEBS Lett. 576, 271–276 (2004).
Frenal, K. et al. Structural and functional characterization of the TgDRE multidomain protein, a DNA repair enzyme from Toxoplasma gondii. Biochemistry 45, 4867–4874 (2006).
Chaouki, A.S. & Salz, H.K. Drosophila SPF45: a bifunctional protein with roles in both splicing and DNA repair. PLoS Genet. 2, e178 (2006).
Kielkopf, C.L., Lucke, S. & Green, M.R. U2AF homology motifs: protein recognition in the RRM world. Genes Dev. 18, 1513–1526 (2004).
Maris, C., Dominguez, C. & Allain, F.H. The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression. FEBS J. 272, 2118–2131 (2005).
Kielkopf, C.L., Rodionova, N.A., Green, M.R. & Burley, S.K. A novel peptide recognition mode revealed by the X-ray structure of a core U2AF35/U2AF65 heterodimer. Cell 106, 595–605 (2001).
Selenko, P. et al. Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP. Mol. Cell 11, 965–976 (2003).
Berglund, J.A., Abovich, N. & Rosbash, M. A cooperative interaction between U2AF65 and mBBP/SF1 facilitates branchpoint region recognition. Genes Dev. 12, 858–867 (1998).
Rain, J.C., Rafi, Z., Rhani, Z., Legrain, P. & Krämer, A. Conservation of functional domains involved in RNA binding and protein- protein interactions in human and Saccharomyces cerevisiae pre-mRNA splicing factor SF1. RNA 4, 551–565 (1998).
Rudner, D.Z., Kanaar, R., Breger, K.S. & Rio, D.C. Interaction between subunits of heterodimeric splicing factor U2AF is essential in vivo. Mol. Cell. Biol. 18, 1765–1773 (1998).
Gozani, O., Potashkin, J. & Reed, R. A potential role for U2AF-SAP 155 interactions in recruiting U2 snRNP to the branch site. Mol. Cell. Biol. 18, 4752–4760 (1998).
Das, R., Zhou, Z. & Reed, R. Functional association of U2 snRNP with the ATP-independent spliceosomal complex E. Mol. Cell 5, 779–787 (2000).
Thickman, K.R., Swenson, M.C., Kabogo, J.M., Gryczynski, Z. & Kielkopf, C.L. Multiple U2AF65 binding sites within SF3b155: thermodynamic and spectroscopic characterization of protein-protein interactions among pre-mRNA splicing factors. J. Mol. Biol. 356, 664–683 (2006).
Spadaccini, R. et al. Biochemical and NMR analyses of an SF3b155-p14–U2AF-RNA interaction network involved in branch point definition during pre-mRNA splicing. RNA 12, 410–425 (2006).
Page-McCaw, P.S., Amonlirdviman, K. & Sharp, P.A. PUF60: a novel U2AF65-related splicing activity. RNA 5, 1548–1560 (1999).
Van Buskirk, C. & Schupbach, T. Half pint regulates alternative splice site selection in Drosophila. Dev. Cell 2, 343–353 (2002).
Jung, D.J., Na, S.Y., Na, D.S. & Lee, J.W. Molecular cloning and characterization of CAPER, a novel coactivator of activating protein-1 and estrogen receptors. J. Biol. Chem. 277, 1229–1234 (2002).
Dowhan, D.H. et al. Steroid hormone receptor coactivation and alternative RNA splicing by U2AF65-related proteins CAPERalpha and CAPERbeta. Mol. Cell 17, 429–439 (2005).
Cheng, J. et al. Protection from Fas-mediated apoptosis by a soluble form of the Fas molecule. Science 263, 1759–1762 (1994).
Krammer, P.H. CD95's deadly mission in the immune system. Nature 407, 789–795 (2000).
Izquierdo, J.M. et al. Regulation of Fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition. Mol. Cell 19, 475–484 (2005).
Sampath, J. et al. Human SPF45, a splicing factor, has limited expression in normal tissues, is overexpressed in many tumors, and can confer a multidrug-resistant phenotype to cells. Am. J. Pathol. 163, 1781–1790 (2003).
Allain, F.H. et al. Specificity of ribonucleoprotein interaction determined by RNA folding during complex formulation. Nature 380, 646–650 (1996).
Wang, X. et al. Phosphorylation of splicing factor SF1 on Ser20 by cGMP-dependent protein kinase regulates spliceosome assembly. EMBO J. 18, 4549–4559 (1999).
Boudrez, A., Beullens, M., Waelkens, E., Stalmans, W. & Bollen, M. Phosphorylation-dependent interaction between the splicing factors SAP155 and NIPP1. J. Biol. Chem. 277, 31834–31841 (2002).
Wang, C. et al. Phosphorylation of spliceosomal protein SAP 155 coupled with splicing catalysis. Genes Dev. 12, 1409–1414 (1998).
Shi, Y., Reddy, B. & Manley, J.L. PP1/PP2A phosphatases are required for the second step of Pre-mRNA splicing and target specific snRNP proteins. Mol. Cell 23, 819–829 (2006).
Zhang, M., Zamore, P.D., Carmo-Fonseca, M., Lamond, A.I. & Green, M.R. Cloning and intracellular localization of the U2 small nuclear ribonucleoprotein auxiliary factor small subunit. Proc. Natl. Acad. Sci. USA 89, 8769–8773 (1992).
Johnson, J.M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003).
McCoy, A.J., Grosse-Kunstleve, R.W, Storoni, L.C. & Rean, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Sali, A. & Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B.A. & Blevins, R.A. NMRView: a computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Korzhnev, D.M., Skrynnikov, N.R., Millet, O., Torchia, D.A. & Kay, L.E. An NMR experiment for the accurate measurement of heteronuclear spin-lock relaxation rates. J. Am. Chem. Soc. 124, 10743–10753 (2002).
Forch, P. et al. The apoptosis-promoting factor TIA-1 is a regulator of alternative pre-mRNA splicing. Mol. Cell 6, 1089–1098 (2000).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993).
Acknowledgements
We thank the staff at the European Synchrotron Radiation Facility and Swiss Light Source for assistance during data collection, V. Rybin (European Molecular Biology Laboratory, EMBL) for ITC measurements, I. Sinning and I. Tews (University of Heidelberg) for ITC measurement time, G. Stier (EMBL) for expression vectors, and B. Simon (EMBL) for help with NMR measurements. M.H. is a fellow of the Peter and Traudl Engelhorn Foundation. This work was supported by the Deutsche Forschungsgemeinschaft (Sa 823/5) and EU grant 3D Repertoire (LSHG-CT-2005-512028).
Author information
Authors and Affiliations
Contributions
L.C. performed biochemistry, NMR experiments and data analysis; L.C. and J.B. performed crystallization and crystallographic data collection; L.C. and M.H. interpreted the crystallographic data; S.B. carried out molecular biology splicing activity assays; K.S. provided resources; M.S., J.V., L.C. and S.B. conceived the study and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Expression control of SPF54 by western blot with anti-SPF45 (PDF 111 kb)
Supplementary Fig. 2
Isothermal calorimetry titrations of SPF45-UHM with U2AF65-ULM (PDF 117 kb)
Supplementary Fig. 3
In the cocrystals of SPF45-UHM and SF3b155-ULM5, the N terminus of the peptide is involved in lattice formation (PDF 152 kb)
Supplementary Fig. 4
15N T1p data, 1H-15N heteronuclear NOE data and 1H,15N correlation spectra of SPF45 (PDF 318 kb)
Supplementary Fig. 5
Supplementary GST pull-down data (PDF 346 kb)
Rights and permissions
About this article
Cite this article
Corsini, L., Bonnal, S., Basquin, J. et al. U2AF-homology motif interactions are required for alternative splicing regulation by SPF45. Nat Struct Mol Biol 14, 620–629 (2007). https://doi.org/10.1038/nsmb1260
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1260
This article is cited by
-
UHMK1 aids colorectal cancer cell proliferation and chemoresistance through augmenting IL-6/STAT3 signaling
Cell Death & Disease (2022)
-
A 5′-tRNA halve, tiRNA-Gly promotes cell proliferation and migration via binding to RBM17 and inducing alternative splicing in papillary thyroid cancer
Journal of Experimental & Clinical Cancer Research (2021)
-
The role of JMJD6/U2AF65/AR-V7 axis in castration-resistant prostate cancer progression
Cancer Cell International (2021)
-
SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
Nature Communications (2021)
-
Post-transcriptional regulation of BRG1 by FIRΔexon2 in gastric cancer
Oncogenesis (2020)