Abstract
We present comprehensive maps at single–amino acid resolution of the residues stabilizing the human Gαi1 subunit in nucleotide- and receptor-bound states. We generated these maps by measuring the effects of alanine mutations on the stability of Gαi1 and the rhodopsin–Gαi1 complex. We identified stabilization clusters in the GTPase and helical domains responsible for structural integrity and the conformational changes associated with activation. In activation cluster I, helices α1 and α5 pack against strands β1–β3 to stabilize the nucleotide-bound states. In the receptor-bound state, these interactions are replaced by interactions between α5 and strands β4–β6. Key residues in this cluster are Y320, which is crucial for the stabilization of the receptor-bound state, and F336, which stabilizes nucleotide-bound states. Destabilization of helix α1, caused by rearrangement of this activation cluster, leads to the weakening of the interdomain interface and release of GDP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Gilman, A.G. G proteins: transducers of receptor-generated signals. Annu. Rev. Biochem. 56, 615–649 (1987).
Wall, M.A. et al. The structure of the G protein heterotrimer Gi alpha 1 beta 1 gamma 2. Cell 83, 1047–1058 (1995).
Sprang, S.R. G proteins, effectors and GAPs: structure and mechanism. Curr. Opin. Struct. Biol. 7, 849–856 (1997).
Oldham, W.M., Van Eps, N., Preininger, A.M., Hubbell, W.L. & Hamm, H.E. Mechanism of the receptor-catalyzed activation of heterotrimeric G proteins. Nat. Struct. Mol. Biol. 13, 772–777 (2006).
Oldham, W.M. & Hamm, H.E. Heterotrimeric G protein activation by G-protein-coupled receptors. Nat. Rev. Mol. Cell Biol. 9, 60–71 (2008).
Marin, E.P., Krishna, A.G. & Sakmar, T.P. Rapid activation of transducin by mutations distant from the nucleotide-binding site: evidence for a mechanistic model of receptor-catalyzed nucleotide exchange by G proteins. J. Biol. Chem. 276, 27400–27405 (2001).
Marin, E.P., Krishna, A.G. & Sakmar, T.P. Disruption of the alpha5 helix of transducin impairs rhodopsin-catalyzed nucleotide exchange. Biochemistry 41, 6988–6994 (2002).
Garcia, P.D., Onrust, R., Bell, S.M., Sakmar, T.P. & Bourne, H.R. Transducin-alpha C-terminal mutations prevent activation by rhodopsin: a new assay using recombinant proteins expressed in cultured cells. EMBO J. 14, 4460–4469 (1995).
Onrust, R. et al. Receptor and betagamma binding sites in the alpha subunit of the retinal G protein transducin. Science 275, 381–384 (1997).
Coleman, D.E. et al. Structures of active conformations of Gi alpha 1 and the mechanism of GTP hydrolysis. Science 265, 1405–1412 (1994).
Mixon, M.B. et al. Tertiary and quaternary structural changes in Gi alpha 1 induced by GTP hydrolysis. Science 270, 954–960 (1995).
Chen, Z., Singer, W.D., Danesh, S.M., Sternweis, P.C. & Sprang, S.R. Recognition of the activated states of Galpha13 by the rgRGS domain of PDZRhoGEF. Structure 16, 1532–1543 (2008).
Noel, J.P., Hamm, H.E. & Sigler, P.B. The 2.2 A crystal structure of transducin-α complexed with GTPγS. Nature 366, 654–663 (1993).
Lambright, D.G., Noel, J.P., Hamm, H.E. & Sigler, P.B. Structural determinants for activation of the α-subunit of a heterotrimeric G protein. Nature 369, 621–628 (1994).
Lambright, D.G. et al. The 2.0 A crystal structure of a heterotrimeric G protein. Nature 379, 311–319 (1996).
Rasmussen, S.G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Van Eps, N. et al. Interaction of a G protein with an activated receptor opens the interdomain interface in the alpha subunit. Proc. Natl. Acad. Sci. USA 108, 9420–9424 (2011).
Westfield, G.H. et al. Structural flexibility of the G alpha s alpha-helical domain in the beta2-adrenoceptor Gs complex. Proc. Natl. Acad. Sci. USA 108, 16086–16091 (2011).
Galés, C. et al. Probing the activation-promoted structural rearrangements in preassembled receptor–G protein complexes. Nat. Struct. Mol. Biol. 13, 778–786 (2006).
Chung, K.Y. et al. Conformational changes in the G protein Gs induced by the β2 adrenergic receptor. Nature 477, 611–615 (2011).
Alexander, N.S. et al. Energetic analysis of the rhodopsin–G-protein complex links the α5 helix to GDP release. Nat. Struct. Mol. Biol. 21, 56–63 (2014).
Kaya, A.I. et al. A conserved phenylalanine as a relay between the alpha5 helix and the GDP binding region of heterotrimeric Gi protein alpha subunit. J. Biol. Chem. 289, 24475–24487 (2014).
Maeda, S. et al. Crystallization scale preparation of a stable GPCR signaling complex between constitutively active rhodopsin and G-protein. PLoS ONE 9, e98714 (2014).
Matouschek, A., Kellis, J.T. Jr., Serrano, L. & Fersht, A.R. Mapping the transition state and pathway of protein folding by protein engineering. Nature 340, 122–126 (1989).
Flock, T. et al. Universal allosteric mechanism for Gα activation by GPCRs. Nature doi:10.1038/nature14663 (6 July 2015).
Hamm, H.E. et al. Site of G protein binding to rhodopsin mapped with synthetic peptides from the alpha subunit. Science 241, 832–835 (1988).
Osawa, S. & Weiss, E.R. The effect of carboxyl-terminal mutagenesis of Gt alpha on rhodopsin and guanine nucleotide binding. J. Biol. Chem. 270, 31052–31058 (1995).
Standfuss, J. et al. The structural basis of agonist-induced activation in constitutively active rhodopsin. Nature 471, 656–660 (2011).
Dratz, E.A. et al. NMR structure of a receptor-bound G-protein peptide. Nature 363, 276–281 (1993).
Van Eps, N. et al. Interaction of a G protein with an activated receptor opens the interdomain interface in the alpha subunit. Proc. Natl. Acad. Sci. USA 108, 9420–9424 (2011).
Liu, W. & Northup, J.K. The helical domain of a G protein alpha subunit is a regulator of its effector. Proc. Natl. Acad. Sci. USA 95, 12878–12883 (1998).
Jones, J.C. et al. The crystal structure of a self-activating G protein alpha subunit reveals its distinct mechanism of signal initiation. Sci. Signal. 4, ra8 (2011).
Urano, D., Chen, J.G., Botella, J.R. & Jones, A.M. Heterotrimeric G protein signalling in the plant kingdom. Open Biol. 3, 120186 (2013).
Raw, A.S., Coleman, D.E., Gilman, A.G. & Sprang, S.R. Structural and biochemical characterization of the GTPgammaS-, GDP.Pi-, and GDP-bound forms of a GTPase-deficient Gly42 → Val mutant of Gialpha1. Biochemistry 36, 15660–15669 (1997).
Dror, R.O. et al. Signal transduction: structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015).
Sun, D. et al. AAscan, PCRdesign and MutantChecker: a suite of programs for primer design and sequence analysis for high-throughput scanning mutagenesis. PLoS ONE 8, e78878 (2013).
Iwata, S. Methods and Results in Crystallization of Membrane Proteins (International University Line, La Jolla, California, USA, 2003).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 47, 5.6 (2014).
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012).
Mahalingam, M., Martinez-Mayorga, K., Brown, M.F. & Vogel, R. Two protonation switches control rhodopsin activation in membranes. Proc. Natl. Acad. Sci. USA 105, 17795–17800 (2008).
Tajkhorshid, E., Baudry, J., Schulten, K. & Suhai, S. Molecular dynamics study of the nature and origin of retinal's twisted structure in bacteriorhodopsin. Biophys. J. 78, 683–693 (2000).
Nina, M., Roux, B. & Smith, J.C. Functional interactions in bacteriorhodopsin: a theoretical analysis of retinal hydrogen bonding with water. Biophys. J. 68, 25–39 (1995).
Phillips, J.C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
MacKerell, A.D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Acknowledgements
We thank F. Heydenreich, V. Korkhov, R. Kammerer and C. Piscitelli for critical reading of the manuscript and V. Panneels and R. Jaussi for technical support. S. Maeda was supported by Roche postdoctoral fellowship RPF113. T.F. is supported as a Boehringer Ingelheim Fonds PhD fellow. D.M. and S. Mendieta were supported by the Paul Scherrer Institute internship program. This work was supported by Swiss National Science Foundation grants Sinergia 141898 (D.B.V. and G.F.X.S.), 133810 31-135754 (D.B.V.), 31-153145 (G.F.X.S.), 31003A-146520 (X.D.), Swiss National Centre of Competence in Research (NCCR) Structural Biology, NCCR Molecular Systems Engineering, European Cooperation in Science and Technology Action CM1207 GLISTEN: GPCR-Ligand Interactions, Structures, and Transmembrane Signalling: a European Research Network (X.D. and G.F.X.S.), the Marie Curie Initial Training Network NanoMem (D.B.V. and G.F.X.S.), the Medical Research Council (MC_U105185859; M.M.B. and T.F.), and M.M.B. is supported as a Lister Institute Research Prize Fellow. Molecular dynamics simulations were run at the Swiss National Supercomputing Centre (CSCS).
Author information
Authors and Affiliations
Contributions
D.S. collected data and performed all data analysis. T.F. helped with analysis and interpretation. S. Maeda, S. Mendieta and D.M. helped with experiments. X.D. built the molecular models, and M.M. helped with the interpretation of the structural effects of mutations. R.J.P.D., G.F.X.S. and M.M.B. contributed to the discussion and to writing the manuscript. D.S., X.D. and D.B.V. wrote the manuscript. All authors read and provided their comments on the draft. D.S. and D.B.V. conceived research, and D.B.V. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Monitoring the thermal stability of WT Gαi1 by differential scanning fluorimetry (DSF) upon addition of increasing concentrations of nucleotides.
a, b, Melting curves of WT Gαi1 in the presence of GDP (a) and stabilization effect concentration-response curve (b). Stability increase [(Tm (w/ GDP) - Tm (wo/ GDP)] is plotted against the concentration of GDP. Tm of WT Gαi1 in addition of 1mM GDP reflects the thermal stability of WT Gαi1 in the GDP-bound state [Gαi1(WT)-GDP]. c, d, Melting profiles of WT Gγi1 in presence of GTPαS (c) and stability increase concentration-response curve (d). Tm of WT Gαi1 in addition of 100 µM GTPγS was finally chosen to reflect the thermal stability of WT Gαi1 in the GTPγS-bound state. The thermal stability measurement of all Gαi1 alanine mutants in the nucleotide-bound state was performed upon addition of 1mM GDP or 100 µM GTPγS. e, Thermal profile of F336AG.H5.8 in the nucleotide-bond state. The melting curve shows that alanine replacement of F336 G.H5.8 in Gαi1 completely impairs the protein stability and activity of coupling with nucleotides. The details of DSF experiments are described in the methods section.
Supplementary Figure 2 High-throughput (HTP) assay for monitoring the effect of Gαi1 alanine mutants on the Rho*–Gi complex.
The recombinant Gαi1 alanine mutants were prepared by HTP purification as shown in the left panel, and the endogenous rhodopsin and βγ subunit were prepared from bovine retinas. For the formation of Rho*–Gi complex, Gαi1 and βγ subunit were reconstituted to form Gi heterotrimer, followed by mixing with rhodopsin and light activation. The formed Rho*–Gi complex was visualized by native gel electrophoresis and the gel bands of complex were quantified by ImageJ software. In each round, WT Gαi1 was always included as the reference control. The experimental details of HTP assay are described in methods section.
Supplementary Figure 3 Characterization of Gαi1 alanine mutants by native gel electrophoresis.
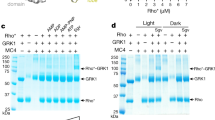
a, Visualization of Rho*–Gi (WT) complex. The Rho*–Gi (WT) complex (+/− GTPγS) was visualized by the native gel electrophoresis, as well as with WT Gαi1, βγ subunit, reconstituted Gi heterotrimer, and the activated rhodopsin as reference markers. The data clearly show that the Rho*–Gi (WT) complex is stable in the absence of nucleotides and that the addition of GTPγS dissociates the complex. b, Gαi1 alanine mutants impairing the formation of Rho*–Gi complex. The Rho*–Gi complexes were formed with WT, Y320AG.S6.2, L348AG.H5.20, G352AG.H5.24 and L353AG.H5.25 of Gαi1 at 4 °C and heated at 36.3 °C, followed by the native gel electrophoresis. The results clearly show that these four Gαi1 alanine mutants are severely impaired in their ability to form the Rho*–Gi complex. c, Monitoring the thermal dissociation (Td) of Rho*–Gi complex. The Rho*–Gi complex formed with WT Gαi1 was heated at the indicated temperature and visualized by the native gel electrophoresis. The gel bands of Rho*–Gi complex were integrated and quantified by ImageJ software. The determined Td50 value of Rho*–Gi(WT) is 36.0 ± 0.1 °C. The same experiment was also performed with Rho*–Gi(R32AG.hns1.3), Rho*–Gi(I55AG.H1.10) and Rho*–Gi(N331AG.H5.3) and the determined Td50 values are 36.7 ± 0.1 °C, 37.4 ± 0.1 °C and 37.8 ± 0.1 °C, respectively, indicating these three alanine mutants stabilize the receptor-bound state. Data points represent mean ± s.d. from three individual experiments.
Supplementary Figure 4 Characterization of Gαi1 alanine mutants in stabilizing the Rho*–Gi complex.
a, Visualization of Rho*–Gi complex by the native gel electrophoresis. The Rho*–Gi complexes were formed with WT, R32AG.hns1.3, K51AG.H1.6, I56AG.H1.11, K54AG.H1.9, I55AG.H1.10, H57AG.H1.12, R176AH.HF.6, N331AG.H5.3 and V332AG.H5.4 of Gαi1 at 4 °C and heated at 36.3 °C, followed by the native gel electrophoresis as described in methods section. b, Gαi1 alanine mutants stabilize the Rho*–Gi complex. The gel bands of Rho*–Gi complexes were integrated and quantified using the ImageJ software. The complex formation efficiency (%) at 4 °C and complex stability (%) at 36.3 °C were defined and determined as described in methods section. The results show that these Gαi1 alanine mutants obviously enhance the thermal stability of Rho*–Gi complex. c, Tm of Gαi1 alanine mutants which stabilize the receptor-bound state. Tm was measured upon addition of 1 mM GDP or 0.1 mM GTPγS using the DSF assay as described in methods section. Data points represent mean ± s.d. from three individual experiments.
Supplementary Figure 5 Effect of alanine substitution of residues involved in the activation and stabilization clusters on formation of the Rho*–Gi complex.
a-d, Effect on Rho*–Gi complex formation of mutation of residues involved in the activation cluster I (a) and stabilization cluster II (b) of the GTPase domain, stabilization cluster III of helical domain (c) and the inter-domain interface (d). The increases in Δcomplex formation efficiency are colored in red and the reductions are colored in blue. The definition of Δcomplex formation efficiency is described in methods section, and the derived numbers are shown in Supplementary Table1.
Supplementary Figure 6 Characterization of heterotrimer (Gi) formation by analytical size-exclusion chromatography (FSEC).
a-f, Characterization of heterotrimer reconstitution of last 48 Gαi1 alanine mutants which are inefficient in formation of Rho*–Gi complex. The reconstitution of Gi and analysis by FSEC were described in methods section. The retention time of WT Gαi1, βγ subunit and reconstituted Gi are 11.15 min, 11.45 min and 10.26 min, respectively. a, b, Retention time of alanine mutants which are efficient in Gi reconstitution. c, d, Retention time of inefficient alanine mutants in formation of heterotrimer. e, f, Retention time of three alanine mutants forming oligomers upon reconstitution.
Supplementary Figure 7 Structural overlay of Gαi1–GDP with G protein α subunit from the plant Arabidopsis thaliana (AtGPA1).
The crystal structure of Gαi1-GDP (PDB 1GDD11) was mapped with the measured ΔTm (in addition of GDP) as spectrum ranging from blue over white to red, as on Fig 1. The structure of AtGPA1-GTPγS (PDB 2XTZ32) is shown in light pink. The enlarged opening of the inter-domain interface suggests that the inter-domain interactions in AtGPA1 are much weaker than those in the Gαi1.
Supplementary Figure 8 Mutations that affect the GTP-bound state cluster around the γ-phosphate of the GTPγS.
a, Correlation of destabilization upon mutation between GDP- and GTP-bound states. The red line shows the linear correlation, blue lines indicate the 95% confidence interval. The most destabilizing residue is N269AG.S5.7, involved in nucleotide binding. b, Alanine substitutions that affect GTP bound state locate near the gamma phosphate of GTPγS. ΔTm values for each single alanine mutant are mapped onto the crystal structure of GTPγS-bound Gαi1 (PDB 1GIA, ref. 11), as a spectrum ranging from blue over white to red. Alanine substitutions that specifically affect the GTP-bound state are deviated from a and the correlated residues are shown as spheres. In addition, alanine mutations in helix α2 also specifically affect the GTP-bound state, consistent with the conformational changes leading to the dissociation of the Gβγ subunit.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note (PDF 1821 kb)
Supplementary Table 1
Summary of alanine-scanning mutagenesis of Gαi1 (XLSX 62 kb)
Supplementary Data Set 1
Multiple sequence alignment of Gα subunit (PDF 431 kb)
Rights and permissions
About this article
Cite this article
Sun, D., Flock, T., Deupi, X. et al. Probing Gαi1 protein activation at single–amino acid resolution. Nat Struct Mol Biol 22, 686–694 (2015). https://doi.org/10.1038/nsmb.3070
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3070
This article is cited by
-
Cryo-EM structure of an activated GPCR–G protein complex in lipid nanodiscs
Nature Structural & Molecular Biology (2021)
-
Structural aspects of rod opsin and their implication in genetic diseases
Pflügers Archiv - European Journal of Physiology (2021)
-
Cryo-EM structure of human rhodopsin bound to an inhibitory G protein
Nature (2018)
-
Structure and dynamics of GPCR signaling complexes
Nature Structural & Molecular Biology (2018)
-
High-throughput mutagenesis using a two-fragment PCR approach
Scientific Reports (2017)